Abstract
ALK rearranged renal cell carcinoma (ALK-RCC) has recently been included in 2016 WHO classification as a provisional entity. In this study, we describe 12 ALK-RCCs from 8 institutions, with detailed clinical, pathological, immunohistochemical (IHC), fluorescence in situ hybridization (FISH), and next generation sequencing (NGS) analyses. Patients’ age ranged from 25 to 68 years (mean, 46.3 years). Seven patients were females and five were males (M:F = 1:1.4). Tumor size ranged from 17 to 70 mm (mean 31.5, median 25 mm). The pTNM stage included: pT1a (n = 7), pT1b (n = 1), and pT3a (n = 4). Follow-up was available for 9/12 patients (range: 2 to 153 months; mean 37.6 months); 8 patients were alive without disease and one was alive with distant metastases. The tumors demonstrated heterogeneous, ‘difficult to classify’ morphology in 10/12 cases, typically showing diverse architectural and cellular morphologies, including papillary, tubular, tubulocystic, solid, sarcomatoid (spindle cell), rhabdoid, signet-ring cell, and intracytoplasmic vacuoles, often set in a mucinous background. Of the remaining two tumors, one showed morphology resembling mucinous tubular and spindle cell renal cell carcinoma (MTSC RCC-like) and one was indistinguishable from metanephric adenoma. One additional case also showed a focal metanephric adenoma-like area, in an otherwise heterogeneous tumor. By IHC, all tumors were diffusely positive for ALK and PAX8. In both cases with metanephric adenoma-like features, WT1 and ALK were coexpressed. ALK rearrangement was identified in 9/11 tumors by FISH. ALK fusion partners were identified by NGS in all 12 cases, including the previously reported: STRN (n = 3), TPM3 (n = 3), EML4 (n = 2), and PLEKHA7 (n = 1), and also three novel fusion partners: CLIP1 (n = 1), KIF5B (n = 1), and KIAA1217 (n = 1). ALK-RCC represents a genetically distinct entity showing a heterogeneous histomorphology, expanded herein to include unreported metanephric adenoma-like and MTSC RCC-like variants. We advocate a routine ALK IHC screening for “unclassifiable RCCs” with heterogeneous features.
Similar content being viewed by others
Introduction
Anaplastic lymphoma kinase rearrangement-associated renal cell carcinoma (ALK-RCC) is a novel renal entity that is currently considered an “emerging/provisional” type of renal cell carcinoma in the 2016 World Health Organization (WHO) classification [1]. However, ALK gene rearrangements with various partner genes (via translocation), resulting in ALK fusion proteins, as well as ALK point mutations, have been documented in many diverse, nonrenal tumors that range from rare entities to more frequently encountered tumors [2, 3]. These tumors include: anaplastic large cell lymphoma, inflammatory myofibroblastic tumor, non-small-cell lung carcinoma, diffuse large B-cell lymphoma, breast carcinoma, colon carcinoma, serous ovarian carcinoma, squamous esophageal carcinoma, neuroblastoma, anaplastic thyroid carcinoma, and epithelioid inflammatory myofibroblastic sarcoma [2,3,4]. ALK is located at 2p23 and represents a member of the receptor tyrosine kinase family and insulin receptor superfamily [5]. The expression pattern of various ALK fusion/chimeric proteins, ensuing from ALK rearrangement with a corresponding partner gene, is primarily driven by the specific fusion partners, resulting in protein products with variable oncogenic potential [3]. Based on the fusion type, the ALK immunohistochemistry (IHC) may show cytoplasmic, cytoplasmic-nuclear, perinuclear-cytoplasmic, or nuclear membrane staining pattern [2,3,4].
Since the initial description in 2011, to our knowledge, a total of 27 ALK-RCC have been documented in the literature, mostly in case reports and small series; [6,7,8,9,10,11,12,13,14,15,16,17,18,19,20,21,22,23] the accumulated evidence has also been summarized in some recent reviews [24, 25]. The key features of all previously reported ALK-RCC cases are shown in Table 1. The cases were listed only if sufficient data were reported to establish a definitive diagnosis of ALK-RCC. We did not include several additional examples due to lack of pathology data and/or definitive clinical characterization [23, 26, 27]. ALK-RCC has also attracted clinical interest owing to the availability of the first and second line targeted therapies, such as ALK inhibitors crizotinib, alectinib, and entrectinib, with demonstrated clinical effects in various other cancers with ALK rearrangement. [3, 26].
It is evident from previous reports that ALK-RCC exhibited a marked histo-morphologic diversity, which may pose significant diagnostic challenges in establishing ALK-RCC as a distinct morphotype. However, an underlying common feature in all reported ALK-RCCs invariably included ALK rearrangement, documented either by IHC or by fluorescence in situ hybridisation (FISH). Given the limited number of ALK-RCC cases that have been studied to date, we undertook a multi-institutional collaborative study, assembling a cohort of 12 ALK-RCC cases. Despite the limited number of cases, this represents the largest ALK-RCC study to date. Our aim was to further characterize this entity, from a clinicopathologic, IHC, and molecular/genetic aspect, which will hopefully contribute toward its formal recognition and inclusion in the future renal cell carcinoma classifications. In this study, we further delineated and expanded the morphological spectrum of these rare tumors, and we identified three novels, previously unreported fusion partner genes CLIP1, KIF5B, and KIAA1217.
Materials and methods
Case identification
This is a retrospective study of the clinical, histopathologic, immunohistochemical, and molecular features of ALK-RCC. The cases were retrieved from the archives and the consultation files of eight institutions: University of Calgary, Calgary, Alberta, Canada; University of Washington, Seattle, USA; Friedrich-Alexander University Erlangen-Nuremberg, University Hospital of Erlangen, Germany; Cleveland Clinic, Cleveland, OH, USA; University Hospital, Charles University, Pilsen, Czech Republic; Himeji Red Cross Hospital, Kochi, Japan; Cochin Hospital, University of Paris, France; and Taipei Veterans General Hospital, Taipei, Taiwan. The cases were identified by experienced urologic pathologists, who typically see large volumes of renal tumors in their tertiary and quaternary institutions; some of these institutions also serve as regional or national reference centers for renal and genitourinary malignancies. Of note, at the time of the study preparation all cases were unpublished; however, at the time of the submission, one case (#12) has been accepted for publication separately, as a more detailed case report [28].
The diagnosis of ALK-RCC was either initially established or suspected, based on: (a) testing for ALK IHC and/or FISH in suspected cases; (b) similarities with morphologic features of ALK-RCC described in previous reports; (c) sharing representative images of several index cases among the contributors; and (d) targeted searches and additional work-up of cases signed out as “unclassified RCC with mucinous features or background”. In toto, 10 ALK-RCC cases were identified using these approaches.
Case identification by TMA screening
In an effort to identify additional ALK-RCC cases, we also employed tissue microarray (TMA) IHC screening. We screened 38 TMA blocks with 16 types of renal tumors, containing a total of 456 cases (details provided in Supplemental Table 1). IHC was performed on 2 µm thick sections using Ventana Benchmark Ultra autostainer (Ventana Medical Systems, Tucson, AZ). Primary antibody ALK1 (ALK01, monoclonal, Ventana, ready-to-use) was used as for screening. Using this approach, two additional ALK-RCC cases were identified based on ALK immunoreactivity. One tumor was initially diagnosed on morphology and IHC as “mucinous tubular and spindle cell RCC” (MTSC-RCC) and one additional tumor demonstrated features undistinguishable from “metanephric adenoma”. These two cases were subsequently confirmed to be truly ALK positive by repeating the IHC on whole sections from representative tissue blocks. Both cases were also subsequently confirmed to harbor ALK rearrangements by molecular genetic analysis, with identification of specific partner genes. Prompted by these cases, to further investigate for possible ALK expression in tumors with these types of morphologies, we additionally evaluated 202 MTSC-RCC and 49 MA, using whole sections from representative tissue blocks, and anti-ALK antibody IHC. The cases for additional screening were retrieved from the renal tumor database at the University Hospital, Charles University, Pilsen, Czech Republic. However, no additional cases demonstrated ALK immunoreactivity in these targeted cohorts.
In sum, a total of 12 ALK-RCC were included in the study. The demographic and clinical data for all cases were retrieved from the medical records of the participating institutions. The study was approved by the Ethics Committee of Kochi Red Cross Hospital (no. 144) and by the local institutional ethics committees, as indicated.
Histologic assessment and immunohistochemistry
A 2–3 µm routine sections were cut from formalin-fixed paraffin-embedded (FFPE) tissue blocks and were stained with hematoxylin and eosin. For IHC, the sections were stained using Ventana Benchmark Ultra autostainer (Ventana Medical Systems, Tucson, AZ). Primary antibody against ALK (monoclonal, 5H4, Nichirei, Tokyo, Japan; prediluted), and ALK1 (monoclonal, Ventana Medical Systems, RTU) were used. In addition, we used a core panel that included the following antibodies: cytokeratin 7 (OV-TL12/30, monoclonal, DakoCytomation, 1:200), cytokeratin 20 (M7019, monoclonal, DakoCytomation, 1:100), TTF-1 (SPT24, monoclonal, Novocastra, Newcastle, UK, 1:400), PAX-8 (polyclonal, Cell Marque, Rocklin, CA, 1: 25), TFE3 (polyclonal, Abcam, Cambridge, UK, 1:100), INI-1 (MRQ-27 monoclonal, Ventana, RTU), and GATA 3 (monoclonal, L50-823, Biocare Medical, Pacheco, CA, 1:100). The primary antibodies were visualized using the supersensitive streptavidin-biotin-peroxidase complex (BioGenex), using appropriate positive controls. Of note, additional IHC evaluation was performed in some cases, particularly during the initial case work-up (details are provided in the results and discussion sections).
Fluorescence in situ hybridization and next generation sequencing
FISH for ALK gene rearrangement was performed using a Vysis LSI ALK Dual Color, Break Apart Rearrangement Probe (Abbott Molecular). We typically evaluated 200 cells, and the cut-off value for positivity was considered to be >15% of the neoplastic cells demonstrating ALK gene split signal. Pretreatment using VP-2000 Processor (Abbott Molecular, Tokyo, Japan) was performed according to the manufacturer’s protocol and hybridization was carried out using the ThermoBrite (Abbott Molecular, Tokyo, Japan).
RNA-sequencing utilizing next generation sequencing (NGS) platform was performed in the Bioptical Laboratory, Pilsen, Czech Republic. The CTL Comprehensive Tumor and Lung FusionPlex Kit (ArcherDX Inc., Boulder, CO) containing 203 targets in 36 genes was used for detection of possible ALK rearrangements. Depending on the sample size, up to three FFPE sections (10 µm thick) were macrodissected and total nucleic acid (NA) was extracted using the Agencourt FormaPure Kit (Beckman Coulter, Brea, CA) with modifications recommended by ArcherDX (ArcherDX Inc., Boulder, CO). Total NA was quantified using the Qubit Broad Range RNA Assay Kit (ThermoFisher Scientific, Waltham, MA, USA) and input for each sample’s library preparation was set to 250 ng. Library preparation was performed following the Archer’s Fusion Plex Protocol for Illumina (ArcherDX Inc.). The PreSeq RNA QC Assay using iTaq Universal SYBR Green Supermix (Biorad, Hercules, CA) was performed to assess sample quality. Samples with the cycle threshold value smaller than 30 continued with target enrichment. Final libraries were quantified following the Library Quantification for Illumina Libraries protocol (KAPA, Wilmington, MA) assuming a 200 bp fragment length. Samples were multiplexed and spiked with 20% PhiX control and sequenced on a NextSeq 500 (Illumina, San Diego, CA) to achieve at least 1.5 million reads per sample. The resulting FASTQ files were analyzed using the Archer Analysis software (version 5.1.7; ArcherDX Inc.). Fusion parameters were set to a minimum of five valid fusion reads with a minimum of three unique start sites within the valid fusion reads.
Anchored Multiplex PCR/NGS was also performed to identify the partner gene for one case (#1) in the Clinical Cytogenomics Laboratory of University of Washington using a custom 115-gene FusionPlex solid tumor panel (ArcherDX, Boulder, CO), as described previously [29]. Total nucleic acid (TNA) extracted from unstained sections from the FFPE patient sample was used for the assay.
Case #5 was also examined for fusion using the TruSight RNA Fusion panel (Illumina, Inc., San Diego, CA, USA), as previously described [30].
Cases #2, #3, and #4 were additionally evaluated by the Variant Plex Solid Tumor kit (Archer DX, Boulder, CO, USA) for copy number variations (CNV) and for mutational analysis of a 67 gene panel (including ABL1, AKT1, ALK, APC, ATM, AURKA, BRAF, CCNE1, CDH1, CDK4, CDKN2A, CSF1R, CTNNB1, DDR2, EGFR, ERBB2, ERBB3, ERBB4, ESR1, EZH2, FBXW7, FGFR1, FGFR2, FGFR3, FLT3, FOXL2, GNA11, GNAQ, GNAS, H3F3A, HNF1A, HRAS, IDH1, IDH2, JAK2, JAK3, KDR, KIT, KRAS, MAP2K1, MET, MLH1, MPL, NOTCH1, NPM1, NRAS, PDGFRA, PIK3CA, PIK3R1, PTEN, PTPN11, RB1, RET, RHOA, ROS1, SMAD4, SMARCB1, SMO, SRC, STK11, TERT, TP53, VHL). The library was constructed using the manufacturer’s protocol. Successfully prepared libraries were sequenced on Nextseq 500 (Illumina, San Diego, CA, USA) with at least 2.5 million reads per sample. Data were analysed in proprietary software Archer Analysis version 5.1.7. Analytical sensitivity was set up to 5% of the mutated alleles.
Results
Clinical features and follow-up
The clinical and demographic features of the 12 patients are summarized in Table 2. All patients had single tumors, and no association was found with any known clinical syndromes or other nonrenal tumors harboring ALK rearrangements. The age of patients ranged from 25 to 68 years, with a mean age of 46.3 years (median 45.5 years). Seven patients were females and five were males (M:F = 1:1.4). Tumor size (largest dimension) ranged from 17 to 70 mm (mean 31.5, median 25 mm). The pTNM stage included: pT1a (n = 7), pT1b (n = 1), and pT3a (n = 4) (AJCC/UACC 8th Edition, 2017) [31]. Follow-up data ranging from 2 to 153 months (mean 37.6, median 20 months) were available for nine patients (follow-up information was not available for three patients). Of the nine patients with clinical follow-up, eight were alive with no evidence of disease and one patient was alive with metastatic disease to cervical and mediastinal lymph nodes, and brain. None of the patients died of disease during the available follow-up.
Pathology—gross and microscopic features
The tumors were located mostly in the renal cortex (8/9). One tumor (#1) involved both the cortex and the medulla and one other tumor (#8) was located only in the medulla. In 2/8 cortical cases, an involvement of the renal sinus was also found, and one case (#7) showed extension into the pelvicalyceal system (all stage pT3a). 11/12 tumors appeared circumscribed and one tumor (#5) showed poorly defined borders with microscopic infiltration into the adjacent renal parenchyma. Some cases showed either a complete or incomplete fibrous pseudocapsule at the periphery, which varied in thickness. The cut surface color was variegated in some cases, but in some it was more homogeneous gray/white, white, or tan (Fig. 1). Cystic changes were present in 3/12 ALK-RCC cases. Necrosis was documented grossly in 3/12 cases, and hemorrhage was noted in 2/12 tumors. The macroscopic and gross findings are summarized in Supplementary Table 2.
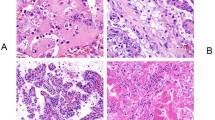
The microscopic findings are summarized in Table 3. One of the key observations was that the morphology was highly variable and heterogeneous, not only between the cases, but also within individual cases, as shown in Table 3 and illustrated in Figs. 2–4 (and Suppl. Fig. 1). The majority of cases showed mixed architectural patterns, including papillary, tubular, trabecular, tubulocystic, and solid. The cells showed eosinophilic cytoplasm, typically without areas of cytoplasmic clearing (“clear cell” areas). In 9/12 cases, at least focal mucinous deposition was found in the background; in some cases the mucinous deposition was easily recognizable at low-power (Suppl. Fig. 1a). In 6/12 cases, there were also cells showing rhabdoid morphology (Figs. 2e and 3d). Prominent intracytoplasmic vacuoles were found in 4/12 tumors, and signet ring-cell morphology was documented in 5/12 cases (Suppl. Fig. 1d). Focal sarcomatoid (spindle cell) differentiation was present in 2/12 cases (illustrated in Fig. 2d).
a Lower power magnification showed admixed solid (center), papillary and trabecular (right), and sarcomatoid (spindle cell) areas (lower-left) (×100). b Exuberant papillary formations and abundant psammoma bodies were present focally, resembling papillary RCC, type 1; note the mucinous background (×200). c Other areas showed small tubules and glands with low-grade nuclei, set in a fibrous stroma (×200). d There were focal sarcomatoid (spindle cell) areas, with scattered individual epithelioid cells (×200). e Foci of atypical rhabdoid and signet-ring cells were also present (×600). f ALK immunostaining highlighted the infiltrating neoplastic cells, while the stromal and inflammatory cells were negative (×200).
a Low power demonstrated a circumscribed tumor. Scattered psammoma bodies were present (top), as well as extensive areas of coagulative necrosis (bottom) (×100). b At higher power, the neoplastic cells focally exhibited irregular tubular, trabecular, and cord-like growth (×200). c Other neoplastic areas demonstrated solid sheath growth, shown here next to an area of coagulative necrosis (upper left) (×200). d In other areas, there were individual epithelioid and rhabdoid cells set in a background of mononuclear inflammation (×200). e The neoplastic cells showed focal nuclear atypia and multinucleation (×400). f In contrast, a very small neoplastic area showed low-grade morphology, resembling “mucinous tubular and spindle cell RCC” (×200).
a At low power, a circumscribed, but non-encapsulated tumor was seen, showing tubular and papillary/trabecular growth (center) (×100). b Areas of papillary growth are shown at higher power (×200). c Most of the tumor was composed of small tubules and acini (×200). d Focally, the tubules showed eosinophilic luminal content resembling thyroid follicles (×400). e The neoplastic cells were diffusely and strongly immunoreactive for TTF1 (thyroglobulin was negative, not shown) (×200). f ALK immunostaining was also positive in the neoplastic cells (×200).
Other unusual and previously unreported morphologies were also found in several ALK-RCCs. In one case (#3), the morphology and the IHC features were indistinguishable from metanephric adenoma, as illustrated in Fig. 5. One other case (#4), in addition to the marked morphologic heterogeneity that is illustrated in Fig. 6, also showed a focal metanephric adenoma-like area (5 mm in size) that was separated from the remaining tumor by sclerosis that contained numerous psammoma bodies, as shown in Fig. 7. One other case (#12) demonstrated tubulopapillary growth, with relatively small basophilic cells, vaguely resembling metanephric adenoma or papillary RCC (for details, see Hang et al. [28]). Another case (#2) demonstrated virtually identical morphology and IHC profile to a typical MTSC-RCC, as illustrated in Fig. 8.
a The tumor was well-circumscribed, but capsule was not present (×20). b The tumor cells were arranged in small tubules and occasional tubules were elongated (×40). c The cells showed scant cytoplasm and basophilic nuclei (×100). d ALK immunostaining was diffusely positive in the neoplastic cells (×200).
a Low power showed a circumscribed tumor with thick peripheral pseudocapsule. The tumor exhibited a tubulocystic and papillary/trabecular growth (right), with extensive adjacent necrosis (left) (×100). b Tubulocystic areas with marked mucinous background are shown at higher power (×400). Other areas showed papillary and trabecular growth (c), solid sheaths (d) and signet-ring cell areas (e) (c, d ×200; e ×400). f ALK immunostaining was positive in the neoplastic cells (×200).
a–b The area resembling metanephric adenoma was found separated from the remaining neoplasm by a sclerotic area containing numerous psammoma bodies; rare mucin-containing tubules and cysts can be seen in the sclerotic area (a, bottom) (a, ×100; b, ×200). c ALK immunostaining was positive in the neoplastic cells (×200). d WT1 was also coexpressed in the neoplastic cells of the metanephric adenoma area (×200).
a On low-power view, fascicles of neoplastic cells are clearly visible (×20). b Neoplastic cells are arranged in pseudotubular structures that showed focal spindle cells proliferation (×100). c On high-power, the spindle cells were low-grade and lacked sarcomatoid features (×200). d ALK staining was variably positive in the neoplastic cells (×100).
Immunohistochemistry
IHC results are summarized in Table 4. ALK protein was diffusely expressed in all tumors. All tumors also showed uniform reactivity for PAX8 and 9/11 cases were positive for CK7. Other positive stains included INI1 (10/11) and vimentin (9/10). TTF1 reactivity was documented in 3/11 cases (all without thyroglobulin expression; not shown in Table 4). TFE3 was weakly positive in only 1/10 cases, but TFE3 rearrangement by FISH was not identified in this tumor (#11). SDHB and FH were retained (both 4/4) and CD117 was uniformly negative (4/4) (not shown in Table 4).
The ALK-RCC case morphologically resembling MTSC-RCC (#2) showed immunoreactivity compatible with MTSC-RCC (positive stains for PAX8, CK7, CD10, AMACR, and vimentin).
Two ALK-RCC cases demonstrated morphology resembling metanephric adenoma, one with exclusive morphology (#3) and one with focal areas (#4). The IHC evaluation of the case with an exclusive morphology (#3) showed a typical IHC profile for metanephric adenoma (WT1 positive, CD57 positive, CK7 focally positive, vimentin negative), along with a strong coexpression for ALK1 in the neoplastic cells (Fig. 5; IHC results not shown in Table 4). In the case with a focal metanephric adenoma-like area (#4), WT1 was positive in the metanephric adenoma part of the tumor, but was completely negative in the other parts; ALK1 was coexpressed in both areas with and without metanephric adenoma morphology (Fig. 7).
Molecular genetic evaluation
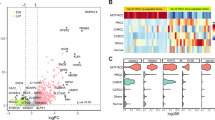
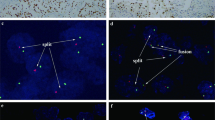
Genetic results are summarized in Table 5. FISH showed significant split signals of ALK gene in 9/11 examined tumors. One of 11 tumors was negative by FISH (#7), and one additional tumor showed an equivocal result (#8); both however demonstrated convincing IHC reactivity for ALK. Using NGS, fusion partner gens were identified in all 12 cases, including those with negative or equivocal FISH result. The fusion partner genes included: STRN (3 cases), TPM3 (3 cases), EML4 (2 cases), and PLEKHA7, CLIP1, KIF5B and KIAA1217, each in one case, respectively. Of these, STRN, TPM3, PLEKHA7, and EML4 have previously been reported in ALK-RCC, while CLIP1, KIF5B, and KIAA1217 have not been previously identified in ALK-RCC (Supplementary Fig. 2a, b). The cases showing an exclusive metanephric adenoma morphology (#3), and focal metanephric adenoma-like area (#4), were additionally tested using the 67 gene panel. No specific mutations, including BRAF V600 mutations, were found in both cases. Case #3 revealed only a polysomy of chromosome 17. In the case with focal metanephric adenoma-like area (#4), no CNV abnormalities were found.
ALK-RCC with novel gene partners CLIP1, KIF5B, and KIAA1217
In this study, we identified three novel and previously undocumented ALK fusion partners, each identified in one case, respectively: CLIP1 (#1), KIF5B (#2), and KIAA1217 (#9). In this section, we provide additional description of the morphology and the IHC findings in these three cases.
ALK-RCC with gene partner CLIP1
ALK-RCC with CLIP1 gene fusion partner (#1) was initially signed-out as “unclassified RCC” due to extreme morphologic heterogeneity, as shown in Fig. 2. At low power, it showed different admixed growth patterns including solid, papillary, trabecular, tubular, and sarcomatoid. There were focal papillary formations with abundant psammoma bodies and a mucinous background, resembling papillary RCC, type 1. Other areas showed small tubules, glands, and nests with low-grade nuclei, set in a fibrous stroma. Sarcomatoid (spindle cell) areas were also present, while other areas showed rhabdoid and signet-ring cells. These cells had eosinophilic cytoplasm and exhibited slightly pleomorphic and hyperchromatic nuclei with frequent intranuclear vacuoles. On IHC, the neoplastic cells were immunoreactive for: ALK1, pan-keratin, PAX8, CK7, INI1, GATA3 (rare positive cells); negative stains included CK20, OCT3/4, TTF1, thyroglobulin, and TFE3.
ALK-RCC with gene partner KIF5B
The ALK-RCC that demonstrated KIF5B-ALK gene fusion (#2) was initially signed out as “MTSC-RCC” because it demonstrated morphology that was essentially indistinguishable from this tumor type, as illustrated in Fig. 8. This case was identified on TMA screening. The IHC profile of this case was also identical to MTSC-RCC, showing reactivity for PAX8, CK7, CD10, AMACR, and vimentin. Unfortunately, it was not possible to additionally evaluate this case for CNV because the residual specimen had low DNA concentration.
ALK-RCC with gene partner KIAA1217
A third ALK-RCC (#9) with a novel gene partner KIAA1217 also showed an unusual morphology with variable admixed growth patterns, including solid, papillary, irregular tubular, trabecular, cord-like, and discohesive single cells growth, as illustrated in Fig. 3. A very small area mimicking MTSC-RCC morphology was also noted in this case (less than 2 mm in size), as shown in Fig. 3f. There were geographic areas of coagulative necrosis with hemorrhage; some neoplastic areas showed marked mononuclear inflammation. The neoplastic cells had voluminous eosinophilic cytoplasm with round to oval, focally pleomorphic, and multinucleated nuclei, with prominent nucleoli. Rhabdoid morphology, intracytoplasmic vacuoles, psammomatous calcifications, and extracellular mucin were focally present. IHC evaluation showed that neoplastic cells were diffusely positive for ALK1, pan-keratin, Cam 5.2, EMA, PAX-8, vimentin, CK7, and AMACR; there was focal reactivity for CD10 and CK5/6. INI-1, SDHB, and FH were all retained. Negative IHC stains included: CA9, CD117, CK20, cathepsin K, smooth muscle actin, desmin, TFE3, HMB45, and melan A.
Discussion
ALK-RCC is a novel renal tumor that has been recently described and it is currently considered an “emerging/provisional” category in the 2016 WHO classification [1]. The current study contributes 12 ALK-RCC cases to the growing body of literature on this entity that will hopefully further solidify the existing data and collected evidence, and will also provide new insights into this entity.
Based on the limited number of cases that have been described to date, and the ones included in this study, ALK-RCC present as solitary tumors, not associated with any known clinical syndrome or other extra-renal tumors harboring ALK rearrangement. In the previously reported ALK-RCCs, the patients showed a diverse racial background, including African-American, Caucasian, and Asian; in this study we also found that Caucasian and Asian patients can present with this type of tumor. Of note, a recent study of 1019 renal tumors in adult Polish patients did not identify a single ALK-RCC [32], highlighting the rarity of this entity in some ethnic groups.
In the current study, ALK-RCC occurred slightly more often in females (M:F = 1:1.4), which is somewhat different from the gender distribution documented in the previously reported cases that were slightly more common in males with a M:F = 1.5:1 ratio (based on recent review by Siadat and Trpkov [25]). ALK-RCCs have been reported in patients showing a wide age range, including pediatric and adolescent patients with sickle cell trait, as well as adult patients, who typically did not harbor a sickle cell trait (see also Table 1). The ALK-RCCs reported in the pediatric/adolescent group typically demonstrated a higher degree of histomorphologic coherence, resembling medullary renal carcinoma and showed either VCL or TPM3 as ALK gene partners [6, 7, 11, 13, 15]. On the other hand, a more heterogeneous and diverse morphologic spectrum has been documented in adult and older patients harboring ALK-RCC, accompanied by several different ALK fusion partners. The current study included exclusively adult patients and confirmed the general theme of a broad morphologic diversity observed in adult ALK-RCC, as well as the involvement of several known ALK partner genes, such as STRN, TPM3, and EML4.
Although the clinical behavior of ALK-RCC has not been fully established, it is evident from the reported cases with available follow-up that they may exhibit aggressive behavior, including metastatic disease in 27% (6/22) and death in 14% (3/22) of patients (based on previously published cases listed in Table 1) [9, 15, 16, 22, 23]. In this study, only 1/9 (11%) patients with available follow-up developed a metastatic disease, but no deaths of disease were documented, which further confirms their malignant potential, even when they are of relatively small size. However, the available patient follow-up of all published ALK-RCC cases so far (including those from the current study), was relatively limited: 77% (24/31) had a follow-up of ≤24 months.
In the previous reports, ALK-RCC were generally circumscribed tumors, showing solid or solid and focally cystic appearances, with either tan-gray or variegated cut surface, ranging in size from 3 to 7 cm. We observed similar gross features and 11/12 tumors were well-circumscribed, often with a peripheral pseudocapsule that varied in thickness. In the current study, the tumor size was below 5 cm in 11/12 cases. The ALK-RCC cases reported herein were typically centered in the cortex, in contrast to the previously reported pediatric cases with VCL-ALK fusion that were typically centered in the medulla or in the renal pelvis.
On histology, we found that ALK-RCCs indeed demonstrated variable and diverse architectural and cellular morphologies, which may pose diagnostic difficulties resulting in labeling these cases as “unclassified RCCs” at the time of the initial sign-out. As documented previously, multiple growth patterns were typically found in a single case, such as papillary (or pseudopapillary), solid, tubular, tubulo-cystic, cribriform, trabecular, signet-ring, and individual cell growth. The presence of interstitial mucin or intracytoplasmic mucin was a frequent feature on microscopy, confirming the previous observations that this may represent a helpful clue for the diagnosis [8, 16, 19]. The cells typically demonstrated an eosinophilic cytoplasm and variable morphologies, often including rhabdoid, vacuolated, pleomorphic, and even small cell with basophilic scanty cytoplasm, as seen in metanephric adenoma. Scattered psammoma bodies were frequently seen, along with coagulative necrosis, as previously reported [9, 16]. The previously reported pediatric ALK-RCC cases with VCL–ALK and TPM3–ALK fusions showed some morphologic similarities to adult renal medullary carcinoma and collecting duct carcinoma, but we did not have any such cases in this study [6, 7, 11, 13, 15].
All patients with ALK-RCC in this study showed diffuse cytoplasmic and membranous ALK protein expression by IHC, which was essential to identify these cases. Of note, using FISH, 9/11 cases showed an ALK split signal in more than 50% of the evaluated tumor cells; this technique, however, failed to identify a signal in one ALK-RCC (#7), while in another one (#8), the result was equivocal. The discrepancy between the IHC and FISH highlights the necessity to pursue further molecular testing in cases where high degree of suspicion persists, when the FISH result is negative or equivocal.
We have also confirmed that ALK-RCC, in addition to the ALK expression by IHC, usually shows a nonspecific immunoprofile that includes reactivity for PAX8, CK7, vimentin, and INI1; other studies have also documented reactivity for 34βE12 and AMACR [11]. Negative markers in the current study included CK20 and GATA3. Other studies showed lack of expression for melan A, S100, HMB45, and cathepsin K in ALK-RCC [16]. Immunoreactivity for TFE3 has been reported previously in some pediatric cases, but without a genuine TFE3-rearrangement by FISH [11, 13, 15, 18], and it has been recently reported in one adult patient with ALK-RCC showing TPM3-ALK fusion [19]. In this study, we also found a TFE3 IHC reactivity in 1/10 cases (#11), in a 38 year-old male, with TPM3-ALK fusion, but TFE3 gene rearrangement by FISH was also not identified. We also identified TTF1 reactivity in 3/11 cases, 2 of which demonstrated ALK-STRN gene fusion (#4 and #7), and one (#10) showed ALK-EML4 fusion. The presence of thyroid primary has been ruled out in the cases with TTF1 reactivity, to completely rule out the possibility of metastatic disease. Interestingly, TTF1 reactivity by IHC has also been previously reported in 2 cases with ALK-STRN fusion [16].
Previous studies of ALK-RCC have identified five different ALK fusion gene partners, including VCL, TPM3, EML4, STRN, and HOOK1 [6,7,8, 11, 13,14,15,16, 19,20,21,22,23]. In the current study, we also found three of these genes to partner with ALK in 8/11 cases: STRN (3), TPM3 (3), and EML4 (2). A novel ALK gene partner PLEKHA7 has also been most recently documented in a case report (#12) [28]. PLEKHA7 represents a pleckstrin homology domain containing A7 gene that is located on chromosome 11p15 and encodes for a specific adherens junction protein [33]. In the current study, we also expand the list of ALK partner genes by identifying three novels, previously undocumented fusion partners, each identified in one case, respectively: CLIP1 (#1), KIF5B (#2), and KIAA1217 (#9), all representing protein coding genes. CLIP1 promotes microtubule growth and microtubule bundling and links cytoplasmic vesicles to microtubules and thereby plays an important role in intracellular vesicle trafficking [34]. KIF5B gene is so-called ‘microtubule-dependent motor’ required for normal distribution of mitochondria and lysosomes [35]. KIAA1217 is required for normal development of intervertebral disks [36]. Of note, CLIP1-ALK fusion has been previously identified in Spitz tumor and lung carcinoma [37, 38]. KIF5B-ALK fusion has been described in lung carcinoma and inflammatory myofibroblastic tumor [39]. To our knowledge, KIAA1217-ALK fusion has not been reported previously, but KIAA1217-RET fusion has been identified in lung adenocarcinoma [40].
Targeted therapies, such as ALK inhibitors alectinib and crizotinib have been used in other, nonrenal tumors demonstrating ALK rearrangement, raising the possibility of their effectiveness in patients with ALK-RCC. Pal et al. recently reported on three patients with “metastatic ALK-rearranged papillary RCC” demonstrating ALK fusion (all with EML4-ALK fusion), treated with alectinib, all showing a profound short term clinical and radiographic response to the treatment [23]. In another 22 year-old patient with ALK-RCC with VCL-ALK fusion, a treatment with Entrectinib resulted in a measurable response over a 19 month period [26].
A novel aspect of this study also represents identification of 2 ALK-RCCs with features virtually identical to metanephric adenoma, either exclusively (#3; illustrated in Fig. 5) or focally (#4; illustrated in Fig. 7). Both tumors were ALK positive by IHC and FISH, and both showed an ALK-STRN rearrangement confirmed by NGS. We were able to also analyze for CNV in cases #3 and #4. In case #3 only a polysomy of chromosome 17 was found, but chromosome 7 was diploid; case #4 showed no abnormal CNV. Another previously unreported finding represents an identification of ALK-RCC with morphology resembling MTSC RCC (#2; illustrated in Fig. 8). This case showed a previously unreported gene partner KIF5B. Interestingly, another ALK-RCC case also showed a very limited MTSC RCC-like area, in addition to the heterogeneous morphology in the remaining tumor, and with a different partner gene KIAA1217 (#9, illustrated in Fig. 3f). Because metanephric adenoma-like and MTSC RCC-like morphologies were not previously reported in ALK-RCC, it is important to be aware of the expanding morphologic spectrum of ALK-RCC, particularly when working-up these types of tumors that are encountered more commonly in practice. We have undertaken an additional systematic evaluation for ALK by IHC of 49 metanephric adenomas and 202 MTSC RCCs from our files and have not identified any additional ALK-RCC cases with these types of morphologies. Therefore, we can confirm that these ALK-RCC morphotypes are indeed very rare. The clinical behavior of such renal tumors with previously unreported morphologies and confirmed ALK rearrangement is uncertain, but given their rarity, we would still maintain the designation “RCC” till more cases are identified. Although a frequent BRAF mutation has been recently reported in metanephric adenoma [41, 42], we did not detect a BRAF V600E or other mutations using NGS in cases #3 and #4, that resembled metanephric adenoma either completely or focally.
From a practical diagnostic perspective, the presence of variable and heterogeneous histologic patterns (typically, papillary + other patterns) encountered in a single case, is somewhat reminiscent of the “pattern multiplicity” seen, for example in FH deficient RCC, clear cell RCC, or MiTF RCC [43, 44]. Therefore, we would recommend a diagnostic consideration of ALK-RCC with IHC screening for ALK in all “difficult to classify” renal tumors showing variable admixed patterns, unusual morphologies, and a mucinous component (intracellular or interstitial). Of note, the presence of mucin has also been described in other ALK rearranged (ALK positive) tumors in different anatomic locations, for example in lung carcinomas, demonstrating either mucinous cribriform pattern or signet-ring cell pattern. [45, 46].
The differential diagnosis of ALK-RCC is broad, because the heterogeneous and diverse morphologies seen in ALK-RCC may mimic other renal entities, such as renal medullary carcinoma (in children and adolescents), collecting duct carcinoma, papillary RCC, MiTF RCC (TFE3 and TFEB), “rhabdoid” RCC (or clear cell with rhabdoid features), metanephric adenoma, MTSC RCC, and thyroid-like follicular RCC.
In conclusion, in this retrospective, multi-institutional study, the largest reported to date, we described a cohort of 12 adults ALK-RCC. We confirmed that ALK-RCC represents a genetically distinct, but histologically heterogeneous group of renal neoplasms, characterized by diverse growth patterns and varied morphologies, set in a mucinous background. We also expand the morphologic spectrum of ALK-RCC by identifying novel morphologies resembling metanephric adenoma and MTSC RCC, previously not reported in this entity. ALK positivity, documented by IHC, and ALK rearrangement established by FISH, are the key elements in establishing the diagnosis. We were able to identify specific gene partners in all ALK-RCCs in this study, and in addition to the previously reported genes STRN, TPM3, EML4, and PLEKHA7, we also identified three previously unreported fusion partners CLIP1, KIF5B, and KIAA1217. We hope that further recognition of additional examples of these rare renal neoplasms will contribute toward their full characterization, and hopefully to their formal recognition in the future classifications of kidney tumors. It is also of utmost importance to establish their long-term clinical behavior, to potentially identify patients with ALK-RCC who may benefit from targeted therapies.
Change history
05 June 2020
An amendment to this article has been published and can be accessed via a link at the top of the article.
References
Moch H, Humphrey PA, Ulbright TM, Reuter VE. WHO classification of tumours of the urinary system and male genital organs. 4th ed. Lyon, France: International Agency for Research on Cancer; 2016.
Mano H. ALKoma: a cancer subtype with a shared target. Cancer Disco. 2012;2:495–502.
Hallberg B, Palmer RH. Mechanistic insight into ALK receptor tyrosine kinase in human cancer biology. Nat Rev Cancer. 2013;13:685–700.
Marino-Enriquez A, Wang WL, Roy A, Lopez-Terrada D, Lazar AJ, Fletcher CD, et al. Epithelioid inflammatory myofibroblastic sarcoma: An aggressive intra-abdominal variant of inflammatory myofibroblastic tumor with nuclear membrane or perinuclear ALK. Am J Surg Pathol. 2011;35:135–44.
Pulford K, Lamant L, Morris S, Butler L, Wood KM, Stroud D, et al. Detection of Anaplastic Lymphoma Kinase (ALK) and Nucleolar Protein Nucleophosmin (NPM)-ALK Proteins in Normal and Neoplastic Cells With the Monoclonal Antibody ALK1. Blood. 1997;89:1394–404.
Marino-Enriquez A, Ou WB, Weldon CB, Fletcher JA, Perez-Atayde AR. ALK rearrangement in sickle cell trait-associated renal medullary carcinoma. Genes Chromosomes Cancer. 2011;50:146–53.
Debelenko LV, Raimondi SC, Daw N, Shivakumar BR, Huang D, Nelson M, et al. Renal cell carcinoma with novel VCL-ALK fusion: new representative of ALK-associated tumor spectrum. Mod Pathol. 2011;24:430–42.
Sugawara E, Togashi Y, Kuroda N, Sakata S, Hatano S, Asaka R, et al. Identification of anaplastic lymphoma kinase fusions in renal cancer: large-scale immunohistochemical screening by the intercalated antibody-enhanced polymer method. Cancer. 2012;118:4427–36.
Sukov WR, Hodge JC, Lohse CM, Akre MK, Leibovich BC, Thompson RH, et al. ALK alterations in adult renal cell carcinoma: frequency, clinicopathologic features and outcome in a large series of consecutively treated patients. Mod Pathol. 2012;25:1516–25.
Lee C, Park JW, Suh JH, Nam KH, Moon KC. ALK-Positive Renal Cell Carcinoma in a Large Series of Consecutively Resected Korean Renal Cell Carcinoma Patients. Korean J Pathol. 2013;47:452–7.
Smith NE, Deyrup AT, Marino-Enriquez A, Fletcher JA, Bridge JA, Illei PB, et al. VCL-ALK renal cell carcinoma in children with sickle-cell trait: the eighth sickle-cell nephropathy? Am J Surg Pathol. 2014;38:858–63.
Ryan C, Mayer N, Cunningham J, Hislop G, Pratt N, Fleming S. Increased ALK1 copy number and renal cell carcinoma-a case report. Virchows Arch. 2014;464:241–5.
Cajaiba MM, Jennings LJ, Rohan SM, Perez-Atayde AR, Marino-Enriquez A, Fletcher JA, et al. ALK-rearranged renal cell carcinomas in children. Genes Chromosomes Cancer. 2016;55:442–51.
Cajaiba MM, Jennings LJ, George D, Perlman EJ. Expanding the spectrum of ALK-rearranged renal cell carcinomas in children: identification of a novel HOOK1-ALK fusion transcript. Genes Chromosomes Cancer. 2016;55:814–7.
Thorner PS, Shago M, Marrano P, Shaikh F, Somers GR. TFE3-positive renal cell carcinomas are not always Xp11 translocation carcinomas: Report of a case with a TPM3-ALK translocation. Pathol Res Pract. 2016;212:937–42.
Kusano H, Togashi Y, Akiba J, Moriya F, Baba K, Matsuzaki N, et al. Two Cases of Renal Cell Carcinoma Harboring a Novel STRN-ALK Fusion Gene. Am J Surg Pathol. 2016;40:761–9.
Jeanneau M, Gregoire V, Desplechain C, Escande F, Tica DP, Aubert S, et al. ALK rearrangements-associated renal cell carcinoma (RCC) with unique pathological features in an adult. Pathol Res Pract. 2016;212:1064–6.
Oyama Y, Nishida H, Kusaba T, Kadowaki H, Arakane M, Daa T, et al. A case of anaplastic lymphoma kinase-positive renal cell carcinoma coincident with Hodgkin lymphoma. Pathol Int. 2017;67:626–31.
Yu W, Wang Y, Jiang Y, Zhang W, Li Y. Genetic analysis and clinicopathological features of ALK-rearranged renal cell carcinoma in a large series of resected Chinese renal cell carcinoma patients and literature review. Histopathology. 2017;71:53–62.
Bodokh Y, Ambrosetti D, Kubiniek V, Tibi B, Durand M, Amiel J, et al. ALK-TPM3 rearrangement in adult renal cell carcinoma: report of a new case showing loss of chromosome 3 and literature review. Cancer Genet. 2018;221:31–7.
Yang J, Dong L, Du H, Li XB, Liang YX, Liu GR. ALK-TPM3 rearrangement in adult renal cell carcinoma: a case report and literature review. Diagn Pathol. 2019;14:112.
Wang XT, Fang R, Ye SB, Zhang RS, Li R, Wang X, et al. Targeted next-generation sequencing revealed distinct clinicopathologic and molecular features of VCL-ALK RCC: a unique case from an older patient without clinical evidence of sickle cell trait. Pathol Res Pract. 2019;215:152651.
Pal SK, Bergerot P, Dizman N, Bergerot C, Adashek J, Madison R, et al. Responses to alectinib in ALK-rearranged papillary renal cell carcinoma. Eur Urol. 2018;74:124–8.
Trpkov K, Hes O. New and emerging renal entities: a perspective post-WHO 2016 classification. Histopathology. 2019;74:31–59.
Siadat F, Trpkov K. ESC, ALK, HOT and LOT: Three letter acronyms of emerging renal entities knocking on the door of the WHO classification. Cancers (Basel) 2020;12:168.
Tao J, Wei G, Patel R, Fagan P, Hao X, Bridge J, et al. ALK fusions in renal cell carcinoma: response to entrectinib. JCO Precis Oncol. 2018;2:1–8.
Chen YB, Xu J, Skanderup AJ, Dong Y, Brannon AR, Wang L, et al. Molecular analysis of aggressive renal cell carcinoma with unclassified histology reveals distinct subsets. Nat Commun. 2016;7:13131.
Hang JF, Chung HJ, Pan CC. ALK-rearranged renal cell carcinoma with a novel PLEKHA7-ALK translocation and metanephric adenoma-like morphology. Vinchows Arch. 2020. [Online ahead of print].
Tretiakova MS, Wang W, Wu Y, Tykodi SS, True L, Liu YJ. Gene fusion analysis in renal cell carcinoma by FusionPlex RNA-sequencing and correlations of molecular findings with clinicopathological features. Genes Chromosomes Cancer 2019, [Online ahead of print].
Agaimy A, Moskalev EA, Weisser W, Bach T, Haller F, Hartmann A. Low-grade endometrioid stromal sarcoma of the paratestis: a novel report with molecular confirmation of JAZF1/SUZ12 translocation. Am J Surg Pathol. 2018;42:695–700.
Sobin LH, Gospodarowicz M, Wittekind C. International union against cancer. TNM classification of malignant tumours. 7th ed. New York, NY: Wiley-Liss; 2009.
Gorczynski A, Czapiewski P, Korwat A, Budynko L, Prelowska M, Okon K, et al. ALK-rearranged renal cell carcinomas in Polish population. Pathol Res Pract. 2019;215:152669.
Pulimeno P, Bauer C, Stutz J, Citi S. PLEKHA7 is an adherens junction protein with a tissue distribution and subcellular localization distinct from ZO-1 and E-cadherin. PLoS One. 2010;5:e12207.
Weisbrich A, Honnappa S, Jaussi R, Okhrimenko O, Frey D, Jelesarov I, et al. Structure-function relationship of CAP-Gly domains. Nat Struct Mol Biol. 2007;14:959–67.
Navone F, Niclas J, Hom-Booher N, Sparks L, Bernstein HD, McCaffrey G, et al. Cloning and expression of a human kinesin heavy chain gene: interaction of the COOH-terminal domain with cytoplasmic microtubules in transfected CV-1 cells. J Cell Biol. 1992;117:1263–75.
Nagase T, Ishikawa K, Kikuno R, Hirosawa M, Nomura N, Ohara O. Prediction of the coding sequences of unidentified human genes. XV. The complete sequences of 100 new cDNA clones from brain which code for large proteins in vitro. DNA Res. 1999;6:337–45.
Yeh I, de la Fouchardiere A, Pissaloux D, Mully TW, Garrido MC, Vemula SS, et al. Clinical, histopathologic, and genomic features of Spitz tumors with ALK fusions. Am J Surg Pathol. 2015;39:581–91.
Vendrell JA, Taviaux S, Beganton B, Godreuil S, Audran P, Grand D, et al. Detection of known and novel ALK fusion transcripts in lung cancer patients using next-generation sequencing approaches. Sci Rep. 2017;7:12510.
Maruggi M, Malicki DM, Levy ML, Crawford JR. A novel KIF5B-ALK fusion in a child with an atypical central nervous system inflammatory myofibroblastic tumour. BMJ Case Rep. 2018;2018:bcr2018226431.
Lee MS, Kim RN, I H, Oh DY, Song JY, Noh KW, et al. Identification of a novel partner gene, KIAA1217, fused to RET: functional characterization and inhibitor sensitivity of two isoforms in lung adenocarcinoma. Oncotarget. 2016;7:36101–14.
Choueiri TK, Cheville J, Palescandolo E, Fay AP, Kantoff PW, Atkins MB, et al. BRAF mutations in metanephric adenoma of the kidney. Eur Urol. 2012;62:917–22.
Udager AM, Pan J, Magers MJ, Palapattu GS, Morgan TM, Montgomery JS, et al. Molecular and immunohistochemical characterization reveals novel BRAF mutations in metanephric adenoma. Am J Surg Pathol. 2015;39:549–57.
Trpkov K, Hes O, Agaimy A, Bonert M, Martinek P, Magi-Galluzzi C, et al. Fumarate hydratase-deficient renal cell carcinoma is strongly correlated with fumarate hydratase mutation and hereditary leiomyomatosis and renal cell carcinoma syndrome. Am J Surg Pathol. 2016;40:865–75.
Muller M, Guillaud-Bataille M, Salleron J, Genestie C, Deveaux S, Slama A, et al. Pattern multiplicity and fumarate hydratase (FH)/S-(2-succino)-cysteine (2SC) staining but not eosinophilic nucleoli with perinucleolar halos differentiate hereditary leiomyomatosis and renal cell carcinoma-associated renal cell carcinomas from kidney tumors without FH gene alteration. Modern Pathol. 2018, [Online ahead of print].
Takeuchi K, Soda M, Togashi Y, Suzuki R, Sakata S, Hatano S, et al. RET, ROS1 and ALK fusions in lung cancer. Nat Med. 2012;18:378–81.
Kuroda N, Ohara M, Wada Y, Yasuoka K, Mizuno K, Yorita K, et al. Cytological features in eight patients with ALK-rearranged lung cancer. Diagnostic Cytopathol. 2018;46:516–9.
Acknowledgements
The study was supported in part by the Charles University Research Fund (project number Q39) and by the project Institutional Research Fund of University Hospital Plzen (Faculty Hospital in Plzen- FNPl 00669806). Dr. Gao’ s fellowship in uropathology was supported by the Alberta Precision Laboratories (formerly Calgary Laboratory Services), Calgary, AB, Canada.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
About this article
Cite this article
Kuroda, N., Trpkov, K., Gao, Y. et al. ALK rearranged renal cell carcinoma (ALK-RCC): a multi-institutional study of twelve cases with identification of novel partner genes CLIP1, KIF5B and KIAA1217. Mod Pathol 33, 2564–2579 (2020). https://doi.org/10.1038/s41379-020-0578-0
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41379-020-0578-0
This article is cited by
-
Genomic alterations and diagnosis of renal cancer
Virchows Archiv (2024)
-
ALK-rearranged renal cell carcinoma with TPM3::ALK gene fusion and review of the literature
Virchows Archiv (2023)
-
Papillary Renal Cell Carcinoma: A Review of Prospective Clinical Trials
Current Treatment Options in Oncology (2023)
-
Concomitant novel ALK-SSH2, EML4-ALK and ARID2-ALK, EML4-ALK double-fusion variants and confer sensitivity to crizotinib in two lung adenocarcinoma patients, respectively
Diagnostic Pathology (2022)
-
ALK rearrangement-associated renal cell carcinoma morphologically mimicking mucinous tubular and spindle cell carcinoma: a case report
Diagnostic Pathology (2022)