Abstract
Although squamous cell carcinomas (SCC) are the most frequent human solid tumor at many anatomic sites, the driving molecular alterations underlying their progression from precursor lesions are poorly understood, especially in the context of photodamage. Therefore, we used high-depth, targeted next-generation sequencing (NGS) of RNA and DNA from routine tissue samples to characterize the progression of both well- (cutaneous) and poorly (ocular) studied SCCs. We assessed 56 formalin-fixed paraffin-embedded (FFPE) cutaneous lesions (n = 8 actinic keratosis, n = 30 carcinoma in situ [CIS], n = 18 invasive) and 43 FFPE ocular surface lesions (n = 2 conjunctival/corneal intraepithelial neoplasia, n = 20 CIS, n = 21 invasive), from institutions in the US and Brazil. An additional seven cases of advanced cutaneous SCC were profiled by hybrid capture-based NGS of >1500 genes. The cutaneous and ocular squamous neoplasms displayed a predominance of UV-signature mutations. Precursor lesions had highly similar somatic genomic landscapes to SCCs, including chromosomal gains of 3q involving SOX2, and highly recurrent mutations and/or loss of heterozygosity events affecting tumor suppressors TP53 and CDKN2A. Additionally, we identify a novel molecular subclass of CIS with RB1 mutations. Among TP53 wild-type tumors, human papillomavirus transcript was detected in one matched pair of cutaneous CIS and SCC. Amplicon-based whole-transcriptome sequencing of select 20 cutaneous lesions demonstrated significant upregulation of pro-invasion genes in cutaneous SCCs relative to precursors, including MMP1, MMP3, MMP9, LAMC2, LGALS1, and TNFRSF12A. Together, ocular and cutaneous squamous neoplasms demonstrate similar alterations, supporting a common model for neoplasia in UV-exposed epithelia. Treatment modalities useful for cutaneous SCC may also be effective in ocular SCC given the genetic similarity between these tumor types. Importantly, in both systems, precursor lesions possess the full complement of major genetic changes seen in SCC, supporting non-genetic drivers of invasiveness.
Similar content being viewed by others
Introduction
Squamous cell carcinoma (SCC) is a major cause of cancer mortality, and is the most common form of human solid tumor in many anatomic sites [1]. Although some SCCs are associated with human papillomavirus (HPV), studies suggest that most SCCs are driven by recurrent somatic alterations such as mutations and copy number alterations. Specifically, cutaneous SCCs harbor a high mutation rate caused by UV damage; recurrent mutations in TP53, CDKN2A, and NOTCH1/2; focal or arm-level gains affecting chromosomes 3q26, 5p, 7q21, and 11q22; and CDKN2A loss (on chromosome 9p21) [1,2,3,4,5,6,7,8]. In contrast to these well-characterized tumors, SCC precursors and invasive lesions from other sites, such as the ocular surface (ocular surface squamous neoplasms, referred to as ocular SCCs in this study), are not as well understood [2, 9]. Ocular SCC is the most common cancer of the ocular surface (cornea/conjunctiva) in the US and can be locally destructive and blinding. Though unusual, lethal metastatic ocular SCC cases have been described [10]. Recurrence rates are high despite surgery and adjunctive radiation or chemoradiotherapy. Proposed risk factors for ocular SCCs, such as HPV, remain controversial [11].
SCCs are thought to develop from hyperplastic precursor lesions. Cutaneous and ocular SCCs arise in the setting of environmental factors (UV damage for cutaneous lesions), transforming normal squamous epithelium into actinic keratosis (AK) on the epidermis and intraepithelial neoplasia on the ocular surface (conjunctiva and cornea, CIN). AKs are likely the most prevalent precancerous lesions in humans [12]. Although some AKs undergo regression, a subset progress to carcinoma in situ (CIS), a preinvasive stage characterized by full-thickness atypia, or to cutaneous SCC [13, 14]. Ocular SCC occurs in two forms: preinvasive (i.e., conjunctival/corneal intraepithelial neoplasia (CIN) or CIS) and invasive subtypes. Thus far, the only study that has profiled ocular tissues was limited by analysis of only CIN and CIS lesions, small cohort size, lack of treatment-naïve tumors, and failure to identify actionable targets [9]. To our knowledge, neither treatment naïve, preinvasive ocular lesions (CIN/CIS) nor ocular SCCs have been molecularly profiled previously. The lack of these studies limits our understanding of how these cancers form and hampers our ability to develop molecular therapies against these highly recurrent squamous cancers.
As described in calls to generate a Pre-Cancer Genome Atlas, the molecular progression of precursor SCC lesions to invasive cancer is not completely understood, in part due to technical challenges posed by the profiling of small areas of interest often only available in routinely processed formalin-fixed paraffin-embedded (FFPE) tissues [15]. To date, the genetic alterations underlying epithelial in situ lesions have only been comprehensively profiled in a limited number of cancers, the largest being a recently published study regarding lung cancer [16]. Obstacles to defining stepwise models for tumorigenesis in cutaneous malignancies include high burdens of passenger mutations, variability in driver mutations within a given tumor type, and lack of identifiable precursor lesions for some tumor types such as basal cell carcinoma. Despite these challenges, progression of precursor lesions to malignancy has been correlated with tumor suppressor gene inactivation events for a subset of sweat gland carcinomas and some melanomas; in contrast, oncogene activation events can be observed in both benign and malignant tumors [17,18,19]. Genetic events associated with progression in cutaneous squamous neoplasms are less clear. AKs harbor mutations and methylation profiles similar to cutaneous SCC [6, 20,21,22]. Although AKs display chromosomal aberrations and loss of heterozygosity events [23], these appear to be less numerous than in cutaneous SCC [24, 25]. Some have proposed that tumor suppressor loss of heterozygosity might be a critical step in the transition from AK to cutaneous SCC [21, 26]; however, this hypothesis has not been rigorously addressed. Another area of uncertainty relates to genetic changes in CIS, which is premalignant but has distinct microscopic appearance and clinical management from AK. Finally, although ocular epithelium is a UV-exposed site and displays a similar spectrum of precursor neoplasms and SCC, the genomic changes in treatment naïve or invasive ocular neoplasms remain hitherto uncharacterized, and to our knowledge, mutational signatures in ocular surface lesions have yet to be described.
To better understand genomic changes associated with malignant progression in UV-exposed squamous lesions, we characterized the genomic landscape of cutaneous and ocular neoplasms, and respective precursor lesions at these anatomic sites from routine FFPE tissue samples.
Materials and methods
Cohort
The study was conducted with local IRB approval. Our cohort was composed of 56 cutaneous tissues (n = 8 AK, n = 30 CIS, n = 18 SCC), and 43 ocular tissues (n = 2 CIN, n = 20 CIS, n = 21 SCC), from institutions in the US and Brazil with available archived FFPE tissues suitable for next-generation sequencing (NGS) analysis (Tables 1, S1 and S2). Additional cutaneous SCC cases from the Mi-Oncoseq program (n = 7), described below, were also included (Table S3). For each case, regions of interest were identified on hematoxylin and eosin stained slides and classified as AK or CIN, CIS, or SCC (Fig. 1) and given a histology-based tumor content by board certified anatomic pathologists. FFPE blocks were cut to make 4-8 10-µm sections. Although most areas with high tumor purity were macrodissected using a scalpel, lesions classified as AK or CIN were mainly dissected under the microscope. DNA and RNA from each sample were co-isolated as described (Supplementary methods).
a Actinic keratosis displaying atypia of the basal layer of the epidermis, with maturation in the upper layers. Copy number profiling demonstrated CDKN2A loss. Nonsynonymous mutations include truncating TP53 and CDKN2A mutations. b Squamous cell carcinoma in situ, demonstrating full-thickness squamous atypia without invasion. Molecular features include CDKN2A loss and truncating mutation, accompanied by TP53 mutations. c Invasive squamous cell carcinoma adjacent to the in situ lesion in the panel above, displaying malignant squamous cells infiltrating collagen. Molecular findings include CDKN2A loss and distinct TP53 mutations.
Multiplexed PCR-based DNA next-generation sequencing
We performed DNA-based NGS using a highly scalable approach optimized for routine FFPE material. To identify oncogenic and tumor suppressive somatic mutations and copy number aberrations, we performed multiplexed PCR-based DNA NGS (mxDNAseq) on spatially defined, minute cell populations using the Oncomine Cancer Panel, which targets 134 cancer-related genes, including nearly all genes known to be recurrently mutated or amplified/deleted in SCCs. Targeted mxDNAseq NGS was performed using 20 ng of DNA from each sample. DNA library preparation, sequencing, and analysis was done as described in Supplementary methods. All samples underwent variant analysis, with the exception of eight which only underwent copy number analysis due to mutation signatures indicative of over-fixation/low library complexity (Table S2). Sample-level variant allele frequencies were used to determine tumor content. Variants were then classified as homozygous, heterozygous, or germline according to estimated tumor content. Most germline variants had a variant allele frequency between 40 and 60% or >90%; however, if the sample had a tumor content of ~50 or >80%, these thresholds were not applicable. Variants classified as germline and present in population databases were excluded unless occurring at a well-supported somatic mutation hotspot in COSMIC (https://cancer.sanger.ac.uk/cosmic). Validation of selected variants was conducted through Sanger Sequencing with custom designed primers (Supplementary methods; Table S4; Fig. S1).
For most cases, estimated tumor contents based upon variant allele frequency were used to correct copy number estimates to account for variability between samples. The exceptions were 10 ocular samples for which copy number estimates were corrected by histology-based tumor content. Specifically, variant analysis was not conducted on eight samples (as described above) and there were no driver mutations to provide variant allele frequency data for two additional samples (Table S2). After correction of copy number estimates, the following copy number thresholds were used: loss (1 copy loss), deep deletion (2 copy loss), gain (1 or 2 copy gain), amplification (>2 copy gain). Further details are in Supplementary methods.
Mi-Oncoseq
Seven cases of advanced cutaneous SCC were identified from the Mi-Oncoseq program, which performs comprehensive somatic and germline sequencing for patients with rare or advanced cancers to guide clinical trial enrollment and precision medicine approaches. Clinical grade, hybrid capture-based exome, or targeted sequencing of >1500 cancer-related genes in tumor and normal tissue were performed to identify somatic mutations, fusions, copy number aberrations, and viral transcripts, as previously described [27].
cBioPortal
Selected prioritized variants for all samples were visualized using the public OncoPrinter tool available from the cBioPortal for Cancer Genomics. In addition, the MutationMapper tool was used to map TP53 (NM_000546), RB1 (NM_000321), and CDKN2A (NM_000077) mutations across all samples [28, 29].
Amplicon-based whole-transcriptome sequencing and analysis
We performed amplicon-based whole-transcriptome sequencing of cutaneous samples in singlicate, as previously described [30]. The linear models used to fit the contrasts for AK versus SCC, CIS versus SCC, and RB1 Mut vs Wild-Type (WT) CIS samples did not have an intercept term and followed the model “~0 + factor.” Overlapping differentially expressed genes of AK versus SCC and CIS versus SCC were used for heatmap visualization. Functional analysis of differentially expressed transcripts between RB1 Mut vs. WT populations was performed using Gene Set Enrichment Analysis version 3.0, developed by the Broad Institute (Cambridge, MA) [31, 32]. The enrichment was done using a pre-ranked list with the ranking metric being the corrected p value divided by the sign of the fold change. The expression data were tested against the hallmark gene set.
RNAscope HPV
To determine HPV status of ocular SCCs and TP53 WT cutaneous cases with adequate remaining tumor material, we used the RNAscope 2.5 HD Red Reagent Kit and target probes HPV-HR18 (pool probe of 18 high-risk HPV strains, 16, 18, 26, 31, 33, 35, 39, 45, 51, 52, 53, 56, 58, 59, 66, 68, 73, 82), and HPV-LR10 (pool probe of 10 low-risk HPV strains, 6, 11, 40, 43, 44, 54, 70; 69, 71, 74) (Advanced Cell Diagnostics Inc., Newark, CA), according to manufacturer’s instructions. Cervical SCC and cutaneous verrucae were used as positive control samples for HR-HPV and LR-HPV, respectively. Positive (Hs-PPIB) and negative (DapB) control probes were also used as sample quality control and assay background control. DapB was uniformly negative. Samples without Hs-PPIB staining were excluded. FFPE tissue blocks were cut into 5-μm sections. After deparaffinization and pretreatments, tissue sections were hybridized with target probes, followed by a series of signal amplification steps and chromogenic staining with Fast Red dye. Stained slides were then evaluated for HR- and LR-HPV infection according to the staining results.
Results
Ocular and cutaneous SCCs: clinical features
Our final cohort included 106 samples from 87 distinct clinical lesions (Tables 1, S1, and S3). Patients had a mean age at diagnosis of 72.4 years (for cutaneous lesions) or 65.2 years (for ocular lesions), with no significant differences among subgroups. Cutaneous lesions in our cohort were relatively evenly divided between men and women, whereas ocular lesions were strongly skewed toward men. Altered immune status (related to iatrogenic immunosuppression, lymphoma, or human immunodeficiency virus) was present in 27% of patients with cutaneous lesions and 38% of patients with ocular lesions. Of patients with ocular lesions, four had a known history of human immunodeficiency virus (Table S1).
Comparison of chromosomal aberrations present in ocular and cutaneous SCCs
In this study, we observed a combination of alterations involving those previously reported in non UV-driven SCCs and UV-driven SCCs. Invasive cutaneous and ocular SCCs harbored CDKN2A copy number loss (n = 12/25 [48%] cutaneous, n = 17/21 [81%] ocular SCCs) and 3q gain (n = 5/25 [20%] cutaneous, n = 4/21 [19%] ocular SCCs), with additional focal SOX2 gains. Furthermore, we observed CCND1, MYC, and EGFR gains in a minority of invasive ocular and cutaneous SCCs at similar frequencies (Figs. 2, 3, S2 and S3). We also observed copy number gains in chromosomes 7, and 11q as well as copy number loss in chromosome 11q in both cutaneous (Figs. 2a and S3) and ocular (Figs. 2b and S2b) SCCs. Hence, cutaneous and ocular SCCs display striking similarity in patterns of chromosomal aberrations, including genomic loss of CDKN2A, 3q gains, and amplification of other oncogenes.
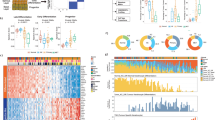
Somatic, autosomal copy number profiles generated by targeted next-generation sequencing (NGS) are presented for (a) 56 cutaneous and (b) 43 ocular tissues. Each copy number profile was GC and tumor content corrected. Normalized read counts per amplicon were divided by those from composite normal tissue, yielding a copy number ratio for each gene (cancer/composite normal), with red and blue indicating gain and loss, respectively, according to the log2 color scale (right). Unsupervised clustering was used on all log2 copy number ratios within lesion groups. Copy number ratios between the range of −1 and 1 were not visualized. Genes part of low arm-level gains and losses are shown with a different shade and border. Columns represent individual targeted genes in genome order (from chromosome 1 to 22). Clinicopathologic features are indicated in the figure legend. Mi-Oncoseq cases are not shown due to differences in normalization. ND: Not Determined, NA: Not Available.
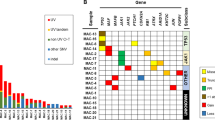
Integrated table of prioritized nonsynonymous mutations and copy number aberrations from (a) 63 cutaneous and (b) 43 ocular tissues. Rows represent genes and columns represent individual samples. Clinicopathologic features are indicated in the figure legend. Copy number aberrations and prioritized mutation types are indicated below the table. A total of eight ocular tissues samples were only analyzed for copy number aberrations and not for mutations, as labeled. Thresholds used: Loss (1 copy loss), Deep deletion (2 copy loss), Gain (1 or 2 copy gain), and Amplification (>2 copy gain).
Copy number aberrations found in SCC lesions are also recurrent in CIS lesions
After correcting for tumor content, we observed similar, recurrent copy number aberrations within cutaneous and ocular lesions that were present in all three types of lesions (AK, CIS, and SCC), indicating that these precursor and invasive lesions were essentially indistinguishable at the genomic level (Figs. 2 and S2). MYC, CCND1, and EGFR gains were also present in some precursor lesions. In addition, CDKN2A loss was observed in 8/8 (100%) AK and 4/30 (13%) CIS lesions. Notably, pair 9, composed of in situ (ID51) and invasive (ID50) regions from a single tumor, harbored CDKN2A copy number loss in both components accompanied by distinct mutations, suggesting that CDKN2A loss in this case was an early event (Fig. 1; Table S5), although we could not exclude the possibility that this might represent collision of unrelated squamous malignancies. Similarly, shared CDKN2A loss was observed in pair 16 composed of ocular lesions IE140A and 140B (Fig. 2b; Table S5). In addition, as shown in Figs. 2b and S2b, both CIS and ocular SCCs can harbor 3q gains, an arm-level gain characteristic of SCCs. Therefore, here we show that many of the copy number aberrations presumed to be characteristic of SCC are also found prior to invasion and overt malignant cytology.
Prioritized somatic variants recurrent in SCCs
After sequencing, we used filtering criteria as specified in the methods section to identify and prioritize somatic variants (Tables S6 and S7). Across all cutaneous and ocular tumor types with the exception of CIN (considering each group in aggregate), we observed a predominance of C > T transitions at dipyrimidine sites among single nucleotide variants, as well as a predominance of CC > TT tandem substitutions among dinucleotide substitutions, consistent with UV-mediated DNA damage. Despite a reported association with tobacco smoking, we found that C > A substitutions characteristic of tobacco signature mutations [33] are rare in ocular neoplasms. Prioritized mutations were highly conserved in multiple samples collected from the same clinical lesion (Table S5), supporting the clonal nature of driver mutations, with the exception of one SCCIS-SCC pair from separate blocks (ID50, ID51) as noted above. As previously reported in SCC, we observed recurrent mutations in genes such as TP53, CDKN2A, NOTCH1, PIK3CA, and EGFR (Fig. 3). TP53 is among the most frequently mutated genes in non-HPV-driven SCCs from all anatomic sites, including 83% of lung SCC, 71% of head and neck SCC, and 48% of penile SCC. However, second TP53 mutations are infrequent to absent at other sites, identified in 0% of lung SCC, 16% of head and neck SCC, and 8% of penile SCC (data downloaded from TCGA PanCancer Atlas, accessed on March 31, 2019 [28, 29]). Interestingly, both of our SCC cohorts had a high frequency of TP53 mutations but also had a significant frequency of a second or even third mutation across all types of lesions (Fig. 3). Upon mapping the location of the mutations, we confirmed that TP53 mutations in precursor and invasive lesions are present in similar hotspot regions (e.g., p.R248, p.P278) across all types of tissues (Fig. S4).
TP53 copy-neutral loss of heterozygosity in SCC progression
Studies on photodamaged skin have reported the presence of multiple TP53 mutations in the absence of clinical lesions, consistent with multiple small, clonally independent cell populations largely defined by these mutations [34]. Other studies have used the ratio of heterozygous to homozygous mutations within a region of copy-neutral loss of heterozygosity to identify the temporal order of genomic events [35], demonstrating that in cutaneous SCC, TP53 loss of heterozygosity is an early event and may gate most of the remaining mutations present in invasive tumors.
Of cases that were evaluable for mutation events, our analysis identified homozygous TP53 mutations in 40% (n = 17/42) of invasive lesions and 23% (n = 13/56) of precursor lesions. An additional second or even third mutation at a heterozygous variant allele frequency was found in 8 out of the 17 invasive lesions and 4 out of the 13 precursor lesions (Fig. S4). In cutaneous lesions, this event was more frequently observed in SCC than precursor lesions; however, a similar trend was not observed for ocular lesions. Most samples lacked TP53 copy number loss, consistent with TP53 copy-neutral loss of heterozygosity through a duplication event following the initial loss of the TP53 wild-type allele. Our data also support either (1) continued acquisition of TP53 mutations after the duplication event or (2) the presence of multiple histologically indistinguishable clonal populations, as heterozygous TP53 mutations were found in both precursor and invasive samples with homozygous TP53 mutations. Taken together, these results support TP53 copy-neutral loss of heterozygosity as an early event in SCC development, frequently occurring before invasion.
CDKN2A loss of heterozygosity in SCC progression
As described above, CDKN2A copy number loss is a recurrent event in cutaneous and ocular SCCs (n = 12/25 [48%] cutaneous, n = 1/7 [14%] copy-neutral loss of heterozygosity reported in Mi-Oncoseq cutaneous SCCs, n = 17/21 [81%] ocular). Specifically, CDKN2A deep deletions (two copy if diploid) were observed in cutaneous SCCs (n = 3/25) and ocular SCCs (n = 3/21). In addition, we report that CDKN2A copy number loss is also present in cutaneous and ocular precursor lesions (Figs. 3 and 4). Interestingly, cutaneous AKs had the highest frequency (n = 8/8, 100%) of copy number loss (Fig. 3a) and the highest frequency (n = 2/8, 20%) of homozygous CDKN2A mutations (Fig. 4a). In fact, we found that most samples with a CDKN2A copy number loss harbor a CDKN2A mutation at a homozygous variant allele frequency. Similarly, this observation was also seen in the ocular lesions, with both CIN and invasive samples displaying CDKN2A homozygous mutation (Fig. 4b) and copy number loss (Fig. 3b). Therefore, like TP53, our analysis supports CDKN2A loss of heterozygosity as occurring at the earliest stages of cutaneous and ocular squamous neoplasia; however, CDKN2A loss of heterozygosity more frequently occurs through copy loss. Furthermore, upon mapping the location of CDKN2A mutations, we confirmed that mutations in precursor and invasive lesions are found in similar regions (e.g., p.R58) (Fig. 4).
Two-level concentric pie charts show zygosity (each level) and co-occurrence (overlapping regions of the two levels) of CDKN2A mutations and CDKN2A variant mapping across (a) AK, CIS, cutaneous SCC and (b) CIS and ocular SCC. Outside circle of the pie chart gives the number of samples with homozygous (Homo) mutations. Inside circle gives the number of samples with heterozygous (Het) mutations. The number of heterozygous and homozygous CDKN2A mutations in each section is denoted by shading. Overlapping regions of the pie chart indicate samples with co-ocurrence of homozygous and heterozygous mutations. Lollipop plots show CDKN2A (NM_000077) mutations in cutaneous and ocular lesions arranged by amino acid location. Histological classification is noted by the color of each line. Mutation type is noted by the colored dot.
RB1 nonsense and splice mutations enriched in cutaneous CIS lesions
Unexpectedly, one of the differences between cutaneous CIS and SCC lesions was the frequency of RB1 mutations (Fig. 3a), which is oftentimes mutually exclusive with CDKN2A alterations in many cancers [36, 37]. Not only did we observe the same mutual exclusivity, but RB1 homozygous/heterozygous nonsense and splice site mutations were enriched in cutaneous CIS lesions (n = 0/8 [0%] AK, n = 8/30 [27%] CIS, n = 2/25 [8%] SCC) (Figs. 3a and S5). Of cutaneous SCCs, RB1 mutations in our cohort were exclusively found in recurrent or metastatic tumors. Comparison to two TCGA studies of cutaneous SCC confirmed that driving RB1 mutations were infrequently found in either cohort (6/68 samples with deleterious RB1 mutations) (Fig. S5) [3, 4], and were associated with significantly increased bone invasion and shorter overall survival (Fig. S6) [3]. Microscopically, RB1-mutated CIS were morphologically heterogeneous from tumor to tumor, with a tendency to display increased inflammatory infiltrate. The findings suggest that RB1 mutations may characterize a distinct subclass of cutaneous SCC that is enriched for CIS, but may display aggressive behavior when present in SCC.
HPV in SCC
Ocular lesions in our cohort with adequate quantity and quality of tumor material for RNA-ISH were uniformly negative (n = 21/21) for HPV, regardless of TP53 status. Of cutaneous squamous neoplasms that lacked detectable TP53 mutation, three matched samples from one clinical lesion (ID05, ID06, and ID27) demonstrated HPV-associated viropathic changes and were positive for high-risk HPV transcript expression. The remaining TP53 wild-type lesions were negative for HPV and lacked characteristic HPV morphologic changes.
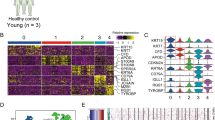
Transcriptome profiles distinguish precursor and invasive cutaneous squamous cell neoplasia, and correlate with mutation events
As our mutation and copy number-based comparison of precursor and invasive lesions in cutaneous/ocular SCCs did not identify alterations likely driving invasion, we pursued RNA-seq on 20 cutaneous samples selected based on appropriate tumor content (n = 4 AK, n = 8 CIS, n = 8 SCC) to determine whether differences were present at the transcriptome level (Table S8). We found that 129 genes were differentially expressed when comparing cutaneous SCC against AK and CIS. These included genes previously associated with invasiveness in SCC or other cancer types, including MMP1, MMP3, MMP9, LAMC2, LGALS1, and TNFRSF12A (Fig. 5). CDKN2A mutations correlated with loss of transcript expression. Transcriptome profiling and gene ontology analysis of RB1-mutated versus wild-type CIS lesions revealed enrichment for interferon gamma/alpha, inflammatory response, and allograft rejection signatures (Fig. S5).
Discussion
Our study defines the genetic landscape of ocular SCC and describes molecular alterations in precursor lesions at cutaneous and ocular sites. We report that squamous epithelium at both sites undergoes similar pathways of tumorigenesis characterized by UV-signature mutations, an accumulation of driver mutations in precursor lesions, and frequent detection of multiple TP53 mutations in a single lesion.
TP53 mutations and clonality in UV-associated squamous neoplasia
Although TP53 alterations are a hallmark of non-HPV-driven SCCs, we report a high percentage of samples harboring a second or even third TP53 mutation in our cutaneous and ocular SCCs. Multiple TP53 mutations, described in normal skin and vulvar intraepithelial neoplasia [34, 38, 39], have been interpreted as multiple intermingled clonal populations with distinct TP53 mutations. While we acknowledge this possibility, our data suggest there may instead be a single clonal population harboring multiple TP53 mutations. In fact, studies report that clinically normal sun-exposed skin already has a high mutation burden, and that mutations known to drive cutaneous SCC, such as NOTCH1 and TP53, are already under strong positive selection. Therefore, a clone must acquire the proper combination of somatic alterations to outcompete all the other clones present in the skin for malignant transformation to begin [34]. Reeves et al., drawing upon observations from transgenic mouse models of squamous neoplasia, suggest that while a terminally benign papilloma has a small number of subclones driving growth, a malignant tumor develops after a clonal sweep followed by the development of additional subclones originating from the progressing clone [40]. Since we identify TP53 mutations at homozygous and heterozygous variant allele frequencies at all stages of squamous neoplasia, our analysis suggests that a similar selection process in photodamaged human epithelia occurs prior to the formation of microscopically identifiable neoplastic lesions. Furthermore, our findings predict that photodamage results in subclinical proliferations of mutated keratinocytes in ocular epithelium, similar to those described in sun-damaged epidermis.
As precursors likely already harbor the full complement of highly recurrent genomic aberrations associated with invasive disease, other epigenetic or non-genomic events may trigger a transition to invasive disease. In support of this, we find gene expression profile differences between precursor and invasive squamous lesions. However, the mechanism for the shift in transcriptional profile remains unclear.
RB1 mutations in cutaneous squamous cell carcinoma in situ
RB1 mutations have been associated with highly metastatic forms of cutaneous carcinoma such as porocarcinoma and virus-negative Merkel cell carcinoma [18, 41, 42], but have not been recognized as a major driver in cutaneous SCC. We identify RB1 mutations in a substantial subset of cutaneous CIS, as well as a small fraction of cutaneous SCC. RB1 mutations were mutually exclusive with CDKN2A mutations and were associated with inflammatory gene signatures suggestive of distinct changes in the tumor microenvironment. When present in cutaneous SCC, RB1 mutations were uniformly associated with recurrent or metastatic disease in our cohort. Although the sample size of RB1-mutated cutaneous SCC is small, precluding rigorous outcomes analysis, our findings suggest that RB1 mutations characterize a distinct molecular subclass of SCC with a propensity to form in situ lesions, with a decreased rate of progression to invasiveness, but paradoxically increased aggressiveness in the setting of invasive carcinoma.
Molecular alterations in UV-associated precursor lesions: previous reports
Similar to SCCs, AKs have been shown to harbor mutations in tumor suppressors such as TP53 and CDKN2A [6, 21], as well as amplifications of EGFR and MYC [43, 44]. In contrast to a previous report describing intratumoral heterogeneity in preinvasive squamoproliferative lesions [45], we found that major genomic events were relatively consistent across multiple areas of a given lesion. Despite a previous hypothesis to the contrary [21, 26], we found that CDKN2A mutation and loss of heterozygosity are frequent events in AKs, and thus do not represent a likely candidate driver for transition to invasiveness. Similarly, despite studies suggesting p53 inactivation to be a late event [46, 47], we observed that the p53 inactivation is already being selected for at early stages of cancer development in cutaneous and ocular squamous neoplasia, as predicted by a whole genome-based study of cutaneous SCC [35].
Published reports comparing expression patterns in AKs and SCCs have had mixed results [6]. This may be due in part to the relatively low power of some studies. The largest such study identified significant differences in gene expression between AKs and SCCs [48]. Our cohort is of similar size to Lambert et al., and corroborates findings from that report. Although we cannot exclude the possibility that a subset of the gene expression differences detected between AK and cutaneous SCC may be related to differences in background tissue rather than neoplastic cells, the similarity of our results with results obtained using microdissection [48], and the established role for many differentially expressed genes in promoting tumor invasiveness, supports these expression changes to be occurring in neoplastic cells.
Molecular profiles of ocular surface squamous neoplasia
Previous mutational studies in ocular neoplasms have been limited, with a disputed role for UV-associated TP53 mutations [9, 49]. A recent exome sequencing study did not comment on mutational profiles within their cohort [9]. To our knowledge, this is the first NGS study to profile ocular lesions at preinvasive, invasive, and treatment-naïve stages. Our results demonstrate the utility of an NGS-based approach, using small ocular surface biopsies and excisions, to nominate precision therapeutic approaches for ocular squamous neoplasms. The current therapies, surgical excision and topical chemotherapies (i.e., interferon-α2b, mitomycin C, 5-fluorouracil), are not genetically tailored and are variably effective inasmuch as ocular squamous neoplasms have an unusually high relapse rate, even when surgical margins are negative. These treatments can also be associated with ocular pain, limbal stem cell loss, conjunctivitis, and other ocular surface toxicities. Such features create an unmet need that could potentially be addressed by currently available oral or systemic therapies, or potential future topical adjunctive therapies, that target aberrant pathways related to genetic alterations in EGFR, FGFR, and PIK3CA genes present in ocular squamous surface neoplasms that we identify for the first time in this study [50,51,52]. Furthermore, the molecular similarity between ocular and cutaneous SCC, including a likely hypermutated UV-signature profile in many cases, suggests that therapeutic approaches with promise in cutaneous SCC, such as immunotherapy [53, 54], may also be effective in advanced ocular SCC.
In agreement with several previous studies, we did not find evidence of a role for HPV in ocular SCC [55,56,57,58]. The discrepancy between this result, and other studies reporting significant rates of HPV detection in these lesions, might be related to differences in clinical cohorts such as the rate of atopy [59]. However, the balance of evidence suggests a less prominent role for HPV in ocular SCC than oropharyngeal SCC.
Study limitations
Our study has several limitations. Most ocular CIS samples were from the United States (Ann Arbor, MI; Nashville, TN; Baltimore, MD) and the invasive, from Sao Paulo, Brazil. One practical reason for this is that ocular SCCs are relatively rare in the United States, but are more common in regions with increased UV radiation exposure, such as Sao Paulo, Brazil. The dearth of invasive samples from the United States precluded meaningful comparison of histopathologic differences, such as aggressiveness, by nation of origin. We acknowledge that the possibility that differences related to region of origin might influence comparisons between CIS and ocular SCCs in our study; however, this factor is unlikely to be a significant confounder, as we report fundamental similarities between these groups rather than differences.
Furthermore, there was limited follow-up for many patients and few episodes of recurrence, precluding robust associations between genomic profiles and outcome. Our targeted approach does not detect events that affect genes not included in our cancer panel, and therefore exome-wide sequencing might provide more definitive analysis of mutational spectra and chromosomal copy number aberrations in these lesions. However, findings from previous exome-wide sequencing studies indicate that our panel encompasses the highly recurrent drivers of cutaneous SCC. Of note, our panel does not include KMT2 and FAT family genes reported to display recurrent (but not universal) mutations in SCC. Another potential limitation is lack of comparison to normal germline DNA for many samples, which may impact individual mutation calls; however, we compared our findings with multiple large germline sequencing databases (see “Materials and methods”) to minimize potential inclusion of germline variants. As noted above, we cannot exclude the possibility that background tissue may influence expression profiling results. Finally, our approach does not address tumor mutation burden or epigenetic alterations.
In conclusion, ocular and cutaneous squamous neoplasms demonstrate a similar spectrum of genetic changes and hence represent parallel models for squamous neoplasia on UV-exposed epithelia. We profiled invasive and treatment naïve preinvasive ocular lesions for the first time. In both ocular and cutaneous settings, precursor lesions already possess the full complement of major genetic changes that are seen in SCC. By contrast, cutaneous precursor lesions demonstrate a distinct transcriptome profile from SCC. In contrast to the stepwise accumulation of mutations proposed for some other malignancies, our findings support the hypothesis that transition to invasiveness in cutaneous SCC may be driven by changes in the transcriptional program or other epigenetic features rather than acquisition of additional genomic insults. Finally, the alterations we identify here are targetable and provide crucial insights toward novel precision therapies for ocular surface lesions, which frequently recur despite current treatment modalities of surgical resection and topical chemotherapy.
References
Dotto GP, Rustgi AK. Squamous cell cancers: a unified perspective on biology and genetics. Cancer Cell. 2016;29:622–37.
Campbell JD, Yau C, Bowlby R, Liu Y, Brennan K, Fan H, et al. Genomic, pathway network, and immunologic features distinguishing squamous carcinomas. Cell Rep. 2018;23:194–212.
Pickering CR, Zhou JH, Lee JJ, Drummond JA, Peng SA, Saade RE, et al. Mutational landscape of aggressive cutaneous squamous cell carcinoma. Clin Cancer Res. 2014;20:6582–92.
Li YY, Hanna GJ, Laga AC, Haddad RI, Lorch JH, Hammerman PS. Genomic analysis of metastatic cutaneous squamous cell carcinoma. Clin Cancer Res. 2015;21:1447–56.
Inman GJ, Wang J, Nagano A, Alexandrov LB, Purdie KJ, Taylor RG, et al. The genomic landscape of cutaneous SCC reveals drivers and a novel azathioprine associated mutational signature. Nat Commun. 2018;9:3667.
Chitsazzadeh V, Coarfa C, Drummond JA, Nguyen T, Joseph A, Chilukuri S, et al. Cross-species identification of genomic drivers of squamous cell carcinoma development across preneoplastic intermediates. Nat Commun. 2016;7:12601.
Mueller SA, Gauthier MA, Ashford B, Gupta R, Gayevskiy V, Ch’ng S, et al. Mutational patterns in metastatic cutaneous squamous cell carcinoma. J Invest Dermatol. 2019;139:1449.
Zilberg C, Lee MW, Yu B, Ashford B, Kraitsek S, Ranson M, et al. Analysis of clinically relevant somatic mutations in high-risk head and neck cutaneous squamous cell carcinoma. Mod Pathol. 2018;31:275–87.
Galor A, Karp CL, Sant D, Joag M, Shalabi N, Gustafson CB, et al. Whole exome profiling of ocular surface squamous neoplasia. Ophthalmology. 2016;123:216–7.
Tabbara KF, Kersten R, Daouk N, Blodi FC. Metastatic squamous cell carcinoma of the conjunctiva. Ophthalmology. 1988;95:318–21.
Sayed-Ahmed IO, Palioura S, Galor A, Karp CL. Diagnosis and medical management of ocular surface squamous neoplasia. Expert Rev Ophthalmol. 2017;12:11–19.
Rosen T, Lebwohl MG. Prevalence and awareness of actinic keratosis: barriers and opportunities. J Am Acad Dermatol. 2013;68:S2–9.
Rowert-Huber J, Patel MJ, Forschner T, Ulrich C, Eberle J, Kerl H, et al. Actinic keratosis is an early in situ squamous cell carcinoma: a proposal for reclassification. Br J Dermatol. 2007;156(Suppl 3):8–12.
Boukamp P. Non-melanoma skin cancer: what drives tumor development and progression? Carcinogenesis. 2005;26:1657–67.
Campbell JD, Mazzilli SA, Reid ME, Dhillon SS, Platero S, Beane J, et al. The case for a Pre-Cancer Genome Atlas (PCGA). Cancer Prev Res. 2016;9:119–24.
Teixeira VH, Pipinikas CP, Pennycuick A, Lee-Six H, Chandrasekharan D, Beane J, et al. Deciphering the genomic, epigenomic, and transcriptomic landscapes of pre-invasive lung cancer lesions. Nat Med. 2019;25:517–25.
Bosic M, Kirchner M, Brasanac D, Leichsenring J, Lier A, Volckmar AL, et al. Targeted molecular profiling reveals genetic heterogeneity of poromas and porocarcinomas. Pathology. 2018;50:327–32.
Harms PW, Hovelson DH, Cani AK, Omata K, Haller MJ, Wang ML, et al. Porocarcinomas harbor recurrent HRAS-activating mutations and tumor suppressor inactivating mutations. Hum Pathol. 2016;51:25–31.
Shain AH, Yeh I, Kovalyshyn I, Sriharan A, Talevich E, Gagnon A, et al. The genetic evolution of melanoma from precursor lesions. N Engl J Med. 2015;373:1926–36.
Rodriguez-Paredes M, Bormann F, Raddatz G, Gutekunst J, Lucena-Porcel C, Kohler F, et al. Methylation profiling identifies two subclasses of squamous cell carcinoma related to distinct cells of origin. Nat Commun. 2018;9:577.
Kanellou P, Zaravinos A, Zioga M, Stratigos A, Baritaki S, Soufla G, et al. Genomic instability, mutations and expression analysis of the tumour suppressor genes p14(ARF), p15(INK4b), p16(INK4a) and p53 in actinic keratosis. Cancer Lett. 2008;264:145–61.
Nelson MA, Einspahr JG, Alberts DS, Balfour CA, Wymer JA, Welch KL, et al. Analysis of the p53 gene in human precancerous actinic keratosis lesions and squamous cell cancers. Cancer Lett. 1994;85:23–9.
Rehman I, Takata M, Wu YY, Rees JL. Genetic change in actinic keratoses. Oncogene. 1996;12:2483–90.
Jin Y, Jin C, Salemark L, Wennerberg J, Persson B, Jonsson N. Clonal chromosome abnormalities in premalignant lesions of the skin. Cancer Genet Cytogenet. 2002;136:48–52.
Garcia-Diez I, Hernandez-Munoz I, Hernandez-Ruiz E, Nonell L, Puigdecanet E, Bodalo-Torruella M, et al. Transcriptome and cytogenetic profiling analysis of matched in situ/invasive cutaneous squamous cell carcinomas from immunocompetent patients. Genes Chromosomes Cancer. 2019;58:164–74.
Mortier L, Marchetti P, Delaporte E, Martin de Lassalle E, Thomas P, Piette F, et al. Progression of actinic keratosis to squamous cell carcinoma of the skin correlates with deletion of the 9p21 region encoding the p16(INK4a) tumor suppressor. Cancer Lett. 2002;176:205–14.
Robinson DR, Wu YM, Vats P, Su F, Lonigro RJ, Cao X, et al. Activating ESR1 mutations in hormone-resistant metastatic breast cancer. Nat Genet. 2013;45:1446–51.
Gao J, Aksoy BA, Dogrusoz U, Dresdner G, Gross B, Sumer SO, et al. Integrative analysis of complex cancer genomics and clinical profiles using the cBioPortal. Sci Signal. 2013;6:pl1.
Cerami E, Gao J, Dogrusoz U, Gross BE, Sumer SO, Aksoy BA, et al. The cBio cancer genomics portal: an open platform for exploring multidimensional cancer genomics data. Cancer Disco. 2012;2:401–4.
Lazo de la Vega L, Samaha MC, Hu K, Bick NR, Siddiqui J, Hovelson DH, et al. Multiclonality and marked branched evolution of low-grade endometrioid endometrial carcinoma. Mol Cancer Res. 2019;17:731–40.
Mootha VK, Lindgren CM, Eriksson KF, Subramanian A, Sihag S, Lehar J, et al. PGC-1alpha-responsive genes involved in oxidative phosphorylation are coordinately downregulated in human diabetes. Nat Genet. 2003;34:267–73.
Subramanian A, Tamayo P, Mootha VK, Mukherjee S, Ebert BL, Gillette MA, et al. Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc Natl Acad Sci USA. 2005;102:15545–50.
Alexandrov LB, Ju YS, Haase K, Van Loo P, Martincorena I, Nik-Zainal S, et al. Mutational signatures associated with tobacco smoking in human cancer. Science. 2016;354:618–22.
Martincorena I, Roshan A, Gerstung M, Ellis P, Van Loo P, McLaren S. et al. Tumor evolution. High burden and pervasive positive selection of somatic mutations in normal human skin. Science. 2015;348:880–6.
Durinck S, Ho C, Wang NJ, Liao W, Jakkula LR, Collisson EA, et al. Temporal dissection of tumorigenesis in primary cancers. Cancer Discov. 2011;1:137–43.
Kim N, Song M, Kim S, Seo Y, Kim Y, Yoon S. Differential regulation and synthetic lethality of exclusive RB1 and CDKN2A mutations in lung cancer. Int J Oncol. 2016;48:367–75.
Knudsen ES, Knudsen KE. Tailoring to RB: tumour suppressor status and therapeutic response. Nat Rev Cancer. 2008;8:714–24.
Pinto AP, Miron A, Yassin Y, Monte N, Woo TY, Mehra KK, et al. Differentiated vulvar intraepithelial neoplasia contains Tp53 mutations and is genetically linked to vulvar squamous cell carcinoma. Mod Pathol. 2010;23:404–12.
Yizhak K, Aguet F, Kim J, Hess JM, Kubler K, Grimsby J, et al. RNA sequence analysis reveals macroscopic somatic clonal expansion across normal tissues. Science. 2019;364:eaaw0726.
Reeves MQ, Kandyba E, Harris S, Del Rosario R, Balmain A. Multicolour lineage tracing reveals clonal dynamics of squamous carcinoma evolution from initiation to metastasis. Nat Cell Biol. 2018;20:699–709.
Cimino PJ, Robirds DH, Tripp SR, Pfeifer JD, Abel HJ, Duncavage EJ. Retinoblastoma gene mutations detected by whole exome sequencing of Merkel cell carcinoma. Mod Pathol. 2014;27:1073–87.
Harms PW, Harms KL, Moore PS, DeCaprio JA, Nghiem P, Wong MKK, et al. The biology and treatment of Merkel cell carcinoma: current understanding and research priorities. Nat Rev Clin Oncol. 2018;15:763–76.
Toll A, Salgado R, Yebenes M, Martin-Ezquerra G, Gilaberte M, Baro T, et al. MYC gene numerical aberrations in actinic keratosis and cutaneous squamous cell carcinoma. Br J Dermatol. 2009;161:1112–8.
Toll A, Salgado R, Yebenes M, Martin-Ezquerra G, Gilaberte M, Baro T, et al. Epidermal growth factor receptor gene numerical aberrations are frequent events in actinic keratoses and invasive cutaneous squamous cell carcinomas. Exp Dermatol. 2010;19:151–3.
South AP, Purdie KJ, Watt SA, Haldenby S, den Breems N, Dimon M, et al. NOTCH1 mutations occur early during cutaneous squamous cell carcinogenesis. J Invest Dermatol. 2014;134:2630–8.
Hosoda W, Chianchiano P, Griffin JF, Pittman ME, Brosens LA, Noe M, et al. Genetic analyses of isolated high-grade pancreatic intraepithelial neoplasia (HG-PanIN) reveal paucity of alterations in TP53 and SMAD4. J Pathol. 2017;242:16–23.
Baker SJ, Fearon ER, Nigro JM, Hamilton SR, Preisinger AC, Jessup JM, et al. Chromosome 17 deletions and p53 gene mutations in colorectal carcinomas. Science. 1989;244:217–21.
Lambert SR, Mladkova N, Gulati A, Hamoudi R, Purdie K, Cerio R, et al. Key differences identified between actinic keratosis and cutaneous squamous cell carcinoma by transcriptome profiling. Br J Cancer. 2014;110:520–9.
Ateenyi-Agaba C, Dai M, Le Calvez F, Katongole-Mbidde E, Smet A, Tommasino M, et al. TP53 mutations in squamous-cell carcinomas of the conjunctiva: evidence for UV-induced mutagenesis. Mutagenesis. 2004;19:399–401.
Seshacharyulu P, Ponnusamy MP, Haridas D, Jain M, Ganti AK, Batra SK. Targeting the EGFR signaling pathway in cancer therapy. Expert Opin Ther Targets. 2012;16:15–31.
Chae YK, Ranganath K, Hammerman PS, Vaklavas C, Mohindra N, Kalyan A, et al. Inhibition of the fibroblast growth factor receptor (FGFR) pathway: the current landscape and barriers to clinical application. Oncotarget. 2017;8:16052–74.
Mizrachi A, Shamay Y, Shah J, Brook S, Soong J, Rajasekhar VK, et al. Tumour-specific PI3K inhibition via nanoparticle-targeted delivery in head and neck squamous cell carcinoma. Nat Commun. 2017;8:14292.
Fitzgerald K, Tsai KK. Systemic therapy for advanced cutaneous squamous cell carcinoma. Semin Cutan Med Surg. 2019;38:E67–74.
Migden MR, Rischin D, Schmults CD, Guminski A, Hauschild A, Lewis KD, et al. PD-1 blockade with cemiplimab in advanced cutaneous squamous-cell carcinoma. N Engl J Med. 2018;379:341–51.
Guthoff R, Marx A, Stroebel P. No evidence for a pathogenic role of human papillomavirus infection in ocular surface squamous neoplasia in Germany. Curr Eye Res. 2009;34:666–71.
Manderwad GP, Kannabiran C, Honavar SG, Vemuganti GK. Lack of association of high-risk human papillomavirus in ocular surface squamous neoplasia in India. Arch Pathol Lab Med. 2009;133:1246–50.
Carreira H, Coutinho F, Carrilho C, Lunet N. HIV and HPV infections and ocular surface squamous neoplasia: systematic review and meta-analysis. Br J Cancer. 2013;109:1981–8.
Shrestha T, Choi W, Kim GE, Yang JM, Yoon KC. Human papilloma virus identification in ocular surface squamous neoplasia by p16 immunohistochemistry and DNA chip test: a strobe-compliant article. Medicine. 2019;98:e13944.
Griffin H, Mudhar HS, Rundle P, Shiraz A, Mahmood R, Egawa N, et al. Human papillomavirus type 16 causes a defined subset of conjunctival in situ squamous cell carcinomas. Mod Pathol. 2020;33:74–90.
Acknowledgements
This research was supported in part by NIH K08EY026654 (to RCR), P30CA046592 (to the University of Michigan Comprehensive Cancer Center); the Research to Prevent Blindness (to the University of Michigan Kellogg Eye Center and RCR), A. Alfred Taubman Medical Research Institute Leslie and Abigail Wexner Emerging Scholar Program (to RCR), A. Alfred Taubman Medical Research Institute A. Alfred Taubman Emerging Scholar Program (to SAT), Grossman Research Fund (to RCR), Leonard G. Miller Professorship and Ophthalmic Research Fund at the Kellogg Eye Center (to RCR), Barbara Dunn Research Fund (to RCR), Roz Greenspon Research Fund (to RCR), Beatrice & Reymont Paul Foundation (to RCR), and March Hoops to Beat Blindness (to RCR). NIH/NEI 5K08EY027464-02 (to ABD), Research to Prevent Blindness Career Development Award (to ABD), AMC is an NCI Outstanding Investigator (R35CA231996), Howard Hughes Medical Institute Investigator, A. Alfred Taubman Scholar, and American Cancer Society Professor.
Author information
Authors and Affiliations
Contributions
LLV, NB, SAT, RCR, and PWH substantially contributed to conception or design of the work. LLV, NB, KH, SER, CDS, SM, PP, XW, AS, HKS, SIM, DRR, AMC, HD, ABD, FW, CGE, SAT, RCR, and PWH contributed to acquisition, analysis, or interpretation of data. LLV, NB, SAT, RCR, and PWH drafted the work and significantly revised it. All authors have approved the submitted version, and have agreed both to be personally accountable for the authors’ own contributions and to ensure that questions related to the accuracy or integrity of any part of the work, even ones in which the author was not personally involved, are appropriately investigated, resolved, and the resolution documented in the literature.
Corresponding authors
Ethics declarations
Conflict of interest
SAT has had a prior sponsored research agreement with ThermoFisher Scientific that provided access to the OCP. SAT is a co-founder of, prior consultant to, equity holder in, and current employee of Strata Oncology. AMC is a consultant and SAB member of Tempus.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
About this article
Cite this article
Lazo de la Vega, L., Bick, N., Hu, K. et al. Invasive squamous cell carcinomas and precursor lesions on UV-exposed epithelia demonstrate concordant genomic complexity in driver genes. Mod Pathol 33, 2280–2294 (2020). https://doi.org/10.1038/s41379-020-0571-7
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41379-020-0571-7
This article is cited by
-
Emerging precision diagnostics in advanced cutaneous squamous cell carcinoma
npj Precision Oncology (2022)
-
Genomic evidence suggests that cutaneous neuroendocrine carcinomas can arise from squamous dysplastic precursors
Modern Pathology (2022)
-
RB1-deficient squamous cell carcinoma: the proposed source of combined Merkel cell carcinoma
Modern Pathology (2022)
-
MicroRNA31 and MMP-1 contribute to the differentiated pathway of invasion -with enhanced epithelial-to-mesenchymal transition- in squamous cell carcinoma of the skin
Archives of Dermatological Research (2021)
-
MACE RNA sequencing analysis of conjunctival squamous cell carcinoma and papilloma using formalin-fixed paraffin-embedded tumor tissue
Scientific Reports (2020)