Abstract
Human papillomavirus (HPV)-independent vulvar squamous cell carcinoma (VSCC) is an aggressive clinical entity. Current diagnostic guidelines for premalignant lesions are ambiguous, and their molecular profile and progression events are still unclear. We selected 75 samples, from 40 patients, including 33 VSCC, 8 verrucous carcinomas (VC), 13 differentiated-type vulvar intraepithelial neoplasia (dVIN), 11 suspicious for dVIN (?dVIN), 6 differentiated exophytic vulvar intraepithelial lesions (DE-VIL), 2 vulvar acanthosis with altered differentiation (VAAD), and 2 usual-type vulvar intraepithelial neoplasia (uVIN/HSIL). Invasive and precursor lesions were matched in 29 cases. Clinical information, p16 immunohistochemistry, and mutation analysis were performed on all lesions. All dVIN, ?dVIN, DE-VIL, and VAAD were p16 negative, all uVIN/HSIL were p16 positive. In the HPV-independent group, mutations were identified in 6 genes: TP53 (n = 40), PIK3CA (n = 20), HRAS (n = 12), MET (n = 5), PTEN (n = 4), and BRAF (n = 1). TP53 mutations occurred in 73% (22/30) VSCC, 85% (11/13) dVIN, 70% (7/10) ?dVIN and no VC (0/8), DE-VIL (0/6) nor VAAD (0/2). Basal atypia was the only reliable feature of TP53 mutations. ?dVIN lesions that were non-acanthotic and atypical but obscured by inflammation, all harbored TP53 mutations. In lesions without TP53 mutations, PIK3CA (50% VC, 33% DE-VIL, 100% VAAD, 40% VSCC) and HRAS (63% VC, 33% DE-VIL, 0% VAAD, 20% VSCC) mutations were found. Mutational progression from in situ to invasive was seen (7/26, 27%) and usually involved TP53 (4/26, 15%). Cases with TP53 and PIK3CA co-mutations had the worse clinical outcomes (p < 0.001). We recommend testing for p53 in all HPV-independent lesions suspicious for dVIN, even in the presence of marked inflammation or non-acanthotic skin, particularly when close to a margin. VC, VAAD, and DE-VIL, were almost never mutated for TP53, but instead often harbored PIK3CA and HRAS mutations. In VSCC, combined TP53 and PIK3CA mutations may inform prognosis.
Similar content being viewed by others
Introduction
Vulvar carcinoma represents 3–5% of all gynecologic malignancies with an annual incidence of 1–2 per 100,000, accounting for 6070 cases per year and 1280 deaths in the United States in 2019 [1, 2]. Squamous cell carcinoma (VSCC) is the most common, constituting 80–90% of malignancies at this site [1]. The 5-year overall survival (OS) varies from 86.3% when disease is localized, to 52.6% when regional lymph nodes are involved, and 22.7% for distant metastases [2]. The current mainstays of treatment are surgical resection, radiation, and/or chemotherapy [3,4,5]. The high-risk of severe morbidity with treatment escalation is a major challenge in managing vulvar carcinomas, which is magnified when patients undergo pelvic exenteration for treatment refractory or recurrent disease [6,7,8,9]. There is a need for better prognostic stratification tools and treatment strategies in this patient group to help optimize outcomes and limit complications.
There has been an effort to separate VSCC into two main categories based on their pathophysiology: human papillomavirus (HPV)-associated and HPV-independent [10,11,12,13]. HPV-associated VSCC typically occur in younger women and show basaloid and warty histomorphology. HPV-independent VSCC are seen in older women, often in association with lichen sclerosus and tend to exhibit keratinizing histomorphology [12, 14]. The HPV-independent:HPV-associated VSCC ratio varies between 0.60 and 0.83, and is expected to rise further as we see rates of HPV-associated premalignant genital lesions decrease following the widespread adoption of HPV vaccination in developed countries [15,16,17,18]. Most studies have revealed worse outcomes in HPV-independent VSCC [14, 15, 17, 19,20,21], similar to what is observed in squamous cell carcinomas of the head and neck region [22].
The premalignant (in situ) squamous lesions preceding VSCC are a rapidly evolving field. VSCC precursors include high-grade squamous intraepithelial lesions, otherwise known as usual-type vulvar intraepithelial neoplasia (uVIN/HSIL), which leads to HPV-associated VSCC, and differentiated vulvar intraepithelial lesion (dVIN), which leads to HPV-independent VSCC. In recent years, two new lesions termed vulvar acanthosis with altered differentiation (VAAD) and differentiated exophytic vulvar intraepithelial lesion (DE-VIL), have been raised as alternate precursor lesions to HPV-independent VSCC [23,24,25], with a new category of HPV-independent VIN proposed to encompass all three of these precursors i.e., dVIN, VAAD, and DE-VIL [26]. The absence of TP53 mutations in VAAD and DE-VIL, and frequent association with verrucous carcinoma, a more indolent neoplasm, have led investigators to believe they constitute a third pathway to VSCC (p53-independent/HPV-independent), which is separate from dVIN [23,24,25].
While the utilization of molecular information for patient diagnosis, prognostication, and treatment planning has been fervently adopted in breast cancer, malignant gliomas, colorectal cancer, and more recently, endometrial carcinoma [11,12,13, 27], very few studies have explored the application of molecular information in VSCC. In this study, we examine the mutational profile of VSCC, focusing particularly on the more aggressive HPV-independent subgroup, in order to uncover any biologic, prognostic, and therapeutic insights in this understudied disease. We have also included a variety of paired squamous precursor lesions, which have not yet been clearly defined on a molecular level.
Materials and methods
Patient selection
Cases were selected from the archives of Vancouver General Hospital from 1998 to 2019. Vulvectomy specimens were reviewed and preference was given to cases with features suggestive of HPV-independent VSCC with an associated in situ lesion [13, 14]. The invasive and in situ components had to be distinct enough to be separated by macrodissection (coring). In some cases, where one of the components could not be separated, only one component was included for subsequent mutational analysis. The cohort was also enriched for DE-VIL, VAAD, and VC.
Histomorphologic review
Hematoxylin and eosin (H&E) stained slides for each case was re-reviewed and the lesions were re-classified into the following categories: conventional squamous cell carcinoma (VSCC: invasive carcinoma comprising of malignant squamous cells with variable keratinization and definite stromal invasion), verrucous carcinoma (VC: well differentiated squamous tumors with minimal cytologic atypia, bullous epithelial pegs and a broad pushing front into the stroma), differentiated-type vulvar intraepithelial neoplasia (dVIN: in situ squamous precursor lesion usually located adjacent to VSCC exhibiting moderate to marked basal nuclear atypia as well as variable degrees of hyperchromasia, karyomegaly, enlarged nucleoli, atypical mitoses, dyskeratosis, elongated and anastomosing rete ridges), usual-type vulvar intraepithelial neoplasia/high-grade squamous intraepithelial lesion (uVIN/HSIL: in situ squamous lesion exhibiting basaloid morphology, hyperchromasia, crowding, anisonucleosis), differentiated exophytic vulvar intraepithelial lesion (DE-VIL: as defined by Watkins et al. [23], in situ squamous precursor lesion exhibiting prominent acanthosis or verruciform morphology, absence of conspicuous basal atypia and abnormalities in keratinocyte differentiation such as hypogranulosis, hyperkeratosis, parakeratosis and dyskeratosis), and vulvar acanthosis with altered differentiation (VAAD: as defined by Nascimento et al. [24], triad of marked acanthosis with variable verruciform architecture, loss of the granular cell layer with superficial epithelial cell pallor and multilayered parakeratosis). Cases that morphologically appeared as DE-VIL or VC, but exhibited moderate to severe basal nuclear atypia, were upgraded to dVIN and SCC, respectively. We also included a category of in situ lesions that were suspicious for dVIN but did not display all the classic morphologic features for a confident diagnosis. These lesions were designated under the “?dVIN” category.
Immunohistochemistry (IHC)
IHC for p16 was used to discriminate between HPV-associated and HPV-independent neoplasms, as previously described [28]. p16 IHC was scored as positive if there was diffuse block-like cytoplasmic and nuclear staining and as negative for any lesser staining (such as patchy staining or absence of staining), in accordance with the LAST (Lower Anogenital Squamous Terminology) recommendations [29].
Sequencing
Mutational analysis was performed on macrodissected formalin fixed paraffin embedded tissue using a commercial next-generation sequencing (NGS) panel that targeted 123 hotspot mutations and 17 exons in 33 known cancer-related genes (AKT, ALK, AR, BRAF, CTNNB1, DDR2, EGFR, ERBB2, ESR1, FGFR1, FGFR2, GNA11, GNAQ, GNAS, HRAS, IDH1, IDH2, JAK, KIT, KRAS, MEK1, MAP2K1, MAP2K2, MET, NRAS, PDGFRA, PIK3CA, PTCH1, PTEN, RET, ROS1, SMO, STK11, TP53), as previously described [30]. The TP53 status for many of these tumors have been reported previously [28]. Only variants with a minimum read depth of 500, an allelic ratio ≥5%, a base quality score ≥30, and a probability score ≥0.90 for single nucleotide changes or a quality score of ≥1000 for insertion/deletion events were reported. To ensure the areas of interest were captured, post-core hematoxylin and eosin (H&E) stained slides were reviewed.
Statistics and survival analysis
The statistical significance, when comparing frequencies of events across two different groups, was assessed using a comparison of proportions analysis [31].
We performed univariate regression analyses of overall survival (OS) and progression-free survival (PFS) using TP53 and PIK3CA mutational status, age, tumor size, tumor depth, focality, presence of lichen sclerosus, surgical margins, lymphovascular invasion, perineural invasion, tumor grade, nodal status, tumor stage, and post-surgical treatment as covariates. In our multivariate analysis the model included TP53 and PIK3CA mutational status, age, tumor focality, surgical margins, tumor depth, tumor grade, and nodal status.
The Kaplan-Meier analyses for OS and PFS compared four molecular conditions: TP53 and PIK3CA mutated, TP53 mutated only, PIK3CA mutated only, no TP53 or PIK3CA mutation.
Results
Study cohort and pathology review
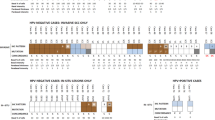
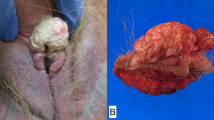
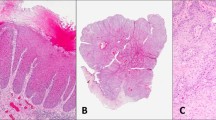
A total of 75 tissue samples, from 40 patients were included for analysis. Histology review of the cohort confirmed the presence of 33 VSCC, 8 VC, 13 dVIN, 11 ?dVIN, 6 DE-VIL, 2 VAAD, and 2 uVIN/HSIL samples (Fig. 1). Two cases originally diagnosed as well-differentiated VSCC were revised to VC after H&E review. One case comprised of areas of mixed VC and VSCC, both components were sampled and sequenced separately. Two cases had a prior history of possible VC. One case appeared to represent VAAD but there was moderate nuclear atypia worrisome for dVIN, therefore it was categorized as ?dVIN.
In 29 patients, at least one sample from both the invasive and in situ components could be matched from a single case. We also included 7 VSCC and 4 VC where the corresponding invasive or in situ samples could not be sequenced. In two cases (1 VSCC and 1 VAAD), the corresponding lesion (1?dVIN and 1 VSCC respectively) failed to be sequenced because the core missed the lesion on post-core H&E review. p16 IHC testing was negative in all but three samples, where morphologic features were also consistent with an HPV-associated neoplasm (two uVIN/HSIL and one VSCC). In one of these uVIN/HSIL cases, the associated VSCC was actually negative for p16 and the VSCC showed morphologic features of HPV-independence, such as keratinization and infiltrative growth, suggesting that the uVIN/HSIL in this case was an incidental/unrelated finding.
Mutational analysis
The mutational findings are summarized in Fig. 2 and Table 1.
HPV-independent (p16-negative) squamous lesions
Mutational analysis was successfully performed on 69 HPV-independent squamous lesions, including 25 cases with matched invasive and in situ samples. The NGS analysis identified significant mutations in 6 different genes: TP53 (n = 40), PIK3CA (n = 20), HRAS (n = 12), MET (n = 5), PTEN (n = 4), and BRAF (n = 1). As shown in Fig. 2, there appeared to be two major molecular groups, one group with TP53 mutations and a group without TP53 mutations.
The first major group of lesions exhibited TP53 mutations (61% [23/38] of patients; 58% of samples), with co-occurring mutations in PIK3CA (17%), HRAS (13%), PTEN (4%) and MET (4%). TP53 was the most frequent mutation overall, occurring in 73% (22/30) VSCC, 85% (11/13) dVIN, and 70% (7/10) ?dVIN. No TP53 mutations were found in VC (0/8), DE-VIL (0/6), or VAAD (0/2). Thus, evidence of TP53 mutation was substantially more frequent in dVIN and ?dVIN compared to DE-VIL and VAAD (p < 0.0001). Similarly, invasive squamous carcinomas arising in association from dVIN and ?dVIN lesions were much more likely to be driven by TP53 compared to other precursors (p < 0.0001). The presence of lichen sclerosus was not associated with TP53 mutation (p = 0.7676). Multiple TP53 mutations were found in one dVIN (two missense mutations) and four VSCC (one case had two missense mutations, one case had three missense mutations, one case a missense and splice site mutation, one case had a missense and frameshift mutation).
The second group without TP53 mutations (37% [15/38] of patients), was more molecularly heterogeneous. In this group, PIK3CA and HRAS mutations were both frequent; PIK3CA mutations were found in 29% (11/38) of patients (29% of samples) and HRAS mutations in 26% (10/38) patients (17% of samples). HRAS and PIK3CA mutations co-occurred in five patients (13%). PIK3CA and HRAS mutations were more frequent in cases without evidence of TP53 mutations (n = 7 for both) than in lesions with TP53 mutations (n = 4 and 3, respectively). All (100%) of the VC, DE-VIL and VAAD lesions resided within this group, compared to only 8/30 (27%) VSCC, 2/13 (15%) dVIN and 3/10 (30%) ?dVIN. VC, DE-VIL, VAAD and associated VSCC were more commonly mutated for PIK3CA, HRAS, MET, and PTEN compared to dVIN and ?dVIN and associated VSCC (p = 0.0001). One case showed morphologic features of both VSCC as well as VC and mutational analysis of both components were mutated for PIK3CA and MET, just like its associated DE-VIL. One VC had a BRAF mutation.
Two VSCC arising in association with ?dVIN, one unpaired VSCC, one VC, two dVIN, and three DE-VIL (13% [9/69] of all samples), had no mutations detected by the targeted NGS panel. PTEN mutations were not specific to any molecular group or any particular histologic diagnosis.
In HPV-associated (p16-positive) squamous lesions
The representation of HPV-associated squamous lesions in our cohort was very small with only four samples, including one uVIN/HSIL associated with an VSCC, one VSCC, and one uVIN/HSIL that was an incidental finding adjacent to a p16-negative VSCC. A PIK3CA mutation was identified in one uVIN/HSIL and a TP53 mutation (splice site mutation) was found in one unrelated VSCC.
Molecular progression
In 26 (25 p16-negative, 1 p16-positive) patients the invasive lesion was paired to at least one precursor lesion. Overall, 7/26 (27%) cases showed evidence of molecular progression. The invasive component often showed additional mutations (five gained TP53, one gained HRAS, one gained BRAF mutations) and in one case a different (n = 1, TP53 c.817 C > G to c.817 C > T) mutation compared to the in situ component. Three cases had mutations in the in situ lesion (one TP53, one HRAS, one PIK3CA) that were not found in the paired VSCC.
Morphologic spectrum of dVIN lesions
We included 11 ?dVIN samples that had some but not all features of dVIN, all of which were adjacent to a VSCC. Morphologically, we identified two scenarios: (i) inflamed atrophic (or normal thickness) skin with mild to moderate basal atypia (n = 8 but only seven successfully cored), where the nuclear atypia was difficult to assess due to the confounding inflammation, and (ii) acanthotic lesions with mild (bordering on moderate) basal atypia (n = 3) (Fig. 3). While all cases with inflamed non-acanthotic skin/mucosa showed evidence of a TP53 mutation, no TP53 mutations were identified in the other group. Of those three cases with prominent acanthosis, one had an activating PIK3CA (c.3140 A > T) and PTEN (c.389 G > C) mutation, but no mutations were detected in the other two. Although this lesion was speculated to be dVIN, its mutational profile fits better under DE-VIL.
There were two cases called dVIN that did not harbor TP53 mutations, although TP53 mutations (one with a missense mutation, and the other with an in-frame deletion) were found in the adjacent VSCCs. On re-review both cases were still regarded as morphologically consistent with dVIN.
Survival analysis based on molecular characteristics
In a univariate analysis of the HPV-independent cases (n = 36), tumor depth of invasion (p = 0.0430) and perineural invasion (p = 0.0025) correlated with OS. The presence of a TP53 status alone trended towards worse outcome but did not reach statistical significance (p = 0.2160). Kaplan-Meier analysis of TP53 and PIK3CA mutations significantly showed that the worse OS was in tumors mutated for both TP53 and PIK3CA versus no TP53 or PIK3CA mutations (p < 0.001) (Fig. 4). Amongst the TP53 mutated tumors, those with concurrent PIK3CA mutations had worse OS and PFS than those without PIK3CA mutations (p = 0.0017 and p = 0.0337 respectively).
The multivariate analysis also showed that the combination of TP53 and PIK3CA mutations had a worse PFS compared to TP53 or PIK3CA mutations alone, or neither mutations (p = 0.0184). TP53 mutation status alone was not an independent predictor of OS or PFS. The other significant variables in our multivariate analysis were margin status (OS and PFS, p = 0.0067 and 0.0230), tumor focality (PFS, p = 0.0184), and nodal status (PFS, p = 0.0009).
Discussion
In this study we assessed the molecular profile of invasive and in situ squamous lesions of the vulva using targeted sequencing and correlated it with clinicopathologic features. Our analysis showed that, among HPV-independent cases, TP53 mutations were present in almost all dVIN, ?dVIN, and associated VSCC, and absent in VC, DE-VIL, and VAAD, which were enriched in PIK3CA and HRAS mutations. dVIN was morphologically heterogenous, with basal atypia being the only consistent histologic feature. Finally, tumors with both TP53 and PIK3CA had the worse survival profile.
The presence of TP53 mutations in HPV-independent VSCC has ranged widely, between 17 and 44% using single-strand conformation polymorphism testing [32,33,34] and higher rates, 56–77%, with more recent Sanger and NGS technologies [35,36,37,38,39]. In our series, 76% of HPV-independent VSCC harbored TP53 mutations, although this rate would probably be higher if we included a consecutive population-based series, because we had selectively enriched for VC, DE-VIL, and VAAD, entities which did not harbor TP53 mutations. The majority of studies addressing the prognostic relevance of p53 in VSCC have largely used immunohistochemistry, with highly varied approaches to interpretation, leading to muddled results, as previously discussed by our group [28]. In the few studies that have evaluated TP53 mutation status as a prognosticator, Nooij et al. have examined the largest series to date (n = 118) [38]. The authors found worse recurrence-free (p = 0.044) and disease-specific survival (p = 0.049) when comparing 3 groups of VC (HPV+, HPV−/p53 abn, HPV-/p53wt). However, when comparing the two groups directly, HPV-/p53abn vs HPV-/p53wt, no clinical or tumor characteristics were statistically different [38]. Kashofer et al. (n = 72) [37, 38] observed worse OS (69% vs 100% at 5 years) and Regauer et al. (n = 24) [40] noticed shorter disease-free intervals (33 vs 65 months) in TP53 mutated tumors compared to TP53 wild tumors, although no formal statistical analyses were made. In contrast, Trietch et al. (n = 107) and Choschzick et al. (n = 25) did not find that TP53 was an independent prognosticator [35, 39]. In our small study, we found that TP53 mutations alone were not an independent predictor of outcome, however, the combination of a TP53 and PIK3CA mutations was a strong predictor of worse outcome. A much more sizeable series will be needed to clarify the prognostic significance of the molecular profile of vulvar squamous lesions.
TP53 was important in tumor progression, where 5/26 (19%) tumors showed additional TP53 mutations in the invasive component compared to the adjacent in situ lesion. This acquisition of TP53 mutations in tumor progression (in situ vs invasive lesions, primary vs recurrent VSCC) has been reported by other groups [37, 40,41,42]. Regauer et al. reported that 60% of patients with wild-type TP53 in the primary VSCC, had a recurrence harboring a TP53 mutation [40]. It is likely that the development multiple structural variations, associated with p53 dysfunction, over time increases malignant potential, but the details surrounding this process are still unknown [43, 44].
In our cohort, the dVIN lesions, notorious for their rapid progression to invasive cancer, almost always showed identical mutations to their invasive counterpart and almost all harbored a TP53 mutation [45]. This suggests that TP53 is an early genetic event in the development of HPV-independent VSCC. Similar to Pinto et al., we also found multiple TP53 mutations in dVIN. Pinto et al. further sequenced multiple foci of dVIN from four patients, and identified TP53 mutations which overlapped amongst samples [46]. No two areas of dVIN were genetically distinct. The authors suggest that there may be multiple independent foci of in situ disease originating from different neoplastic clones. This concept of “field effect”, akin to the phenomenon seen in colon cancers arising from inflammatory bowel disease and bladder cancer in smokers, makes intuitive sense given the frequent VSCC association with chronic vulvar inflammation. However, unless genetically distinct (non-overlapping) foci have been identified, we cannot confirm this theory, and it is possible that the mutations occur sequentially in tumors with high mutation burden, with the first being the main driver mutation and the second being a passenger.
The term dVIN was introduced by the International Society for the Study of Vulvovaginal Disease in 1986, based on the description of the entity by Abell in 1961, and is now a well-recognized morphologic entity in vulvar pathology [47, 48]. Its diagnosis however is challenging, especially at the lower end of the morphologic atypia spectrum [49]. There is tremendous variation across pathologists on how to approach the diagnosis of these lesions and what, if any, ancillary test should be used [50, 51]. Previous research has shown much higher proportions of dVIN without TP53 alterations, reflecting its diagnostic irreproducibility. Our study showed that dVIN has a wide range of morphologies, and that current diagnostic criteria can lead to missing a large proportion of precursor lesions with clonal association to VSCC. The results from our ?dVIN group suggest that the diagnosis of dVIN should still be considered even in the absence of acanthosis, parakeratosis, hyperkeratosis, and even with moderate/marked atypia as the only finding. The presence of basal atypia was the only sensitive morphologic feature for dVIN, as previously suggested by others [52]. in situ lesions with prominent acanthosis, parakeratosis and hyperkeratosis without nuclear atypia often harbored PIK3CA and HRAS mutations, and are better classified under DE-VIL/VAAD. We also noted that the inflammatory reaction surrounding the squamous lesions often complicated the assessment of the basal atypia, especially in cases with an atrophic/non-acanthotic appearance. Pathologists should not confidently exclude the diagnosis of dVIN in a background of inflammation, in the presence of atypia.
The entities VAAD and DE-VIL can be enigmatic to the practicing pathologist. Although the concept of VAAD was published over 15 years ago, only 4 studies have since been published on this topic to date [38, 41, 53, 54]. Despite small numbers in our series, DE-VIL and VAAD were not associated with TP53 mutations, instead they showed PIK3CA and/or HRAS mutations or sometimes no mutations. This is unlike what was shown in a previous small series of VAAD where all 7 VAAD lesions lacked PIK3CA mutations; Nooij et al. thereby proposed that PIK3CA status may be used to differentiate VAAD from DE-VIL [38], because PIK3CA mutations were reported in >60% of DE-VIL by Watkins et al. [23]. However, 5/7 (71%) of VAAD by Nooij et al. did harbor HRAS mutations [38], a mutation that is also frequently found in DE-VIL (22–38%) [23, 55]. Certainly, there is significant morphologic and molecular overlap between DE-VIL and VAAD. The authors who initially described VAAD [24], in a subsequent study [23], combined VAAD, atypical verruciform hyperplasia, verruciform lichen simplex chronicus and verruciform dVIN, under an umbrella term “atypical verruciform lesions”, and ultimately proposed a new name DE-VIL, which would encompass this spectrum of lesions. More importantly, the clinical outcomes for these p53 wild-type in situ lesions has not been reported. If the clinical differences between p53 mutated VIN (dVIN) and p53 wild-type lesions (DE-VIL, VAAD) can be confirmed, it seems prudent to adopt simpler nomenclature, such as p53-abn VIN and p53-wt VIN, as has been suggested avoiding many issues with overlapping morphologic features [26].
We recommend the use of p53 IHC or TP53 sequencing in any HPV-independent lesions suspicious for dVIN, particularly when the lesion is close to a resection margin. Superimposed inflammation and normal thickness epithelium should not dissuade the pathologist. In our center, HPV status is first determined using p16 IHC. If the lesion is p16 negative and shows evidence of a p53 mutational staining pattern, dVIN can be diagnosed. The abnormal p53 pattern should match that seen in the adjacent squamous cell carcinoma. Comparison to wild-type p53 staining in hair follicles or normal skin is helpful. If there is no evidence of a p53 mutational pattern in the in situ lesion in question and the adjacent VSCC shows a p53 mutational pattern, then the lesion is likely reactive. If there is a question about DE-VIL or VAAD, p53 IHC will not help. The presence of HRAS or PIK3CA mutations support the diagnosis or DE-VIL/VAAD, but we understand most institutions will not have ready access to sequencing technologies. Currently, no reliable immunohistochemical biomarker for DE-VIL/VAAD exists. Lesions which raise the possibility of DE-VIL or VAAD, warrants close clinical follow-up.
Our NGS panel offers possibilities for targeted molecular therapies in VSCC. PIK3CA, PTEN and HRAS mutations may be amendable to mTOR and MEK inhibitors. We also identified MET and BRAF mutations, which may be susceptible to MET and BRAF inhibitors. However, we acknowledge that the efficacies of some of these novel therapies can be quite variable [56].
There are several limitations to our study. First the targeted mutation panel we used did not include certain genes that have been previously reported in squamous lesions of the vulva such as CDKN2A, NOTCH1 or ARID1A, BRCA2, or FBXW7 [23, 36, 38, 39]. As discussed above our NGS assay did not cover the entire TP53 gene, and could have missed uncommon TP53 alterations and large deletions [30]. Our sampling of HPV-associated lesions only included two uVIN/HSIL and two VSCC, which limits our ability to detect molecular overlap between HPV-associated and HPV-independent vulvar squamous lesions.
The diagnosis and stratification of HPV-independent vulvar squamous lesions are one of the most challenging areas in anatomical pathology, morphologic features are subtle and outcome data is rare at best. p53 IHC or TP53 sequencing should be used to support the diagnosis of HPV-independent squamous lesions with basal atypia suggestive of dVIN. Our results support that TP53 and PIK3CA mutation status can help to inform prognostication. We hope this study can lay the ground for more rigorous clinical and molecular studies to address the large gap in our knowledge, in this under-studied yet clinically important disease.
References
Weinberg D, Gomez-Martinez RA. Vulvar cancer. Obstet Gynecol Clin North Am. 2019;46:125–35.
SEER cancer Stat Facts: Vulvar Cancer. https://seer.cancer.gov/statfacts/html/vulva.html. 2020.
Hopkins MP, Reid GC, Morley GW. The surgical management of recurrent squamous cell carcinoma of the vulva. Obstet Gynecol. 1990;75:1001–5.
Parthasarathy A, Cheung MK, Osann K, Husain A, Teng NN, Berek JS, et al. The benefit of adjuvant radiation therapy in single-node-positive squamous cell vulvar carcinoma. Gynecol Oncol. 2006;103:1095–9.
Gould N, Kamelle S, Tillmanns T, Scribner D, Gold M, Walker J, et al. Predictors of complications after inguinal lymphadenectomy. Gynecol Oncol. 2001;82:329–32.
Figge DC, Gaudenz R. Invasive carcinoma of the vulva. Am J Obstet Gynecol. 1974;119:382–95.
Rutledge F, Smith JP, Franklin EW. Carcinoma of the vulva. Am J Obstet Gynecol. 1970;106:1117–30.
De Hullu JA, Hollema H, Lolkema S, Boezen M, Boonstra H, Burger MP, et al. Vulvar carcinoma. price less Radic Surg Cancer. 2002;95:2331–8.
Stehman FB, Bundy BN, Dvoretsky PM, Creasman WT. Early stage I carcinoma of the vulva treated with ipsilateral superficial inguinal lymphadenectomy and modified radical hemivulvectomy: a prospective study of the Gynecologic Oncology Group. Obstet Gynecol. 1992;79:490–7.
Bloss JD, Liao SY, Wilczynski SP, Macri C, Walker J, Peake M, et al. Clinical and histologic features of vulvar carcinomas analyzed for human papillomavirus status: evidence that squamous cell carcinoma of the vulva has more than one etiology. Hum Pathol. 1991;22:711–8.
Hording U, Junge J, Daugaard S, Lundvall F, Poulsen H, Bock JE. Vulvar squamous cell carcinoma and papillomaviruses: indications for two different etiologies. Gynecol Oncol. 1994;52:241–6.
Kurman RJ, Toki T, Schiffman MH. Basaloid and warty carcinomas of the vulva. Distinctive types of squamous cell carcinoma frequently associated with human papillomaviruses. Am J Surg Pathol. 1993;17:133–45.
Toki T, Kurman RJ, Park JS, Kessis T, Daniel RW, Shah KV. Probable nonpapillomavirus etiology of squamous cell carcinoma of the vulva in older women: a clinicopathologic study using in situ hybridization and polymerase chain reaction. Int J Gynecol Pathol. 1991;10:107–25.
Dong F, Kojiro S, Borger DR, Growdon WB, Oliva E. Squamous cell carcinoma of the vulva: a subclassification of 97 cases by clinicopathologic, Immunohistochemical, and molecular features (p16, p53, and EGFR). Am J Surg Pathol. 2015;39:1045–53.
McAlpine JN, Leung SCY, Cheng A, Miller D, Talhouk A, Gilks CB, et al. Human papillomavirus (HPV)-independent vulvar squamous cell carcinoma has a worse prognosis than HPV-associated disease: a retrospective cohort study. Histopathology. 2017;71:238–46.
Santos M, Landolfi S, Olivella A, Lloveras B, Klaustermeier J, Suárez H, et al. p16 overexpression identifies HPV-positive vulvar squamous cell carcinomas. Am J Surg Pathol. 2006;30:1347–56.
van de Nieuwenhof HP, van Kempen LC, de Hullu JA, Bekkers RL, Bulten J, Melchers WJ, et al. The etiologic role of HPV in vulvar squamous cell carcinoma fine tuned. Cancer Epidemiol Biomark Prev. 2009;18:2061–7.
Drolet M, Benard E, Perez N, Brisson M, Group HPVVIS. Population-level impact and herd effects following the introduction of human papillomavirus vaccination programmes: updated systematic review and meta-analysis. Lancet. 2019;394:497–509.
Allo G, Yap ML, Cuartero J, Milosevic M, Ferguson S, Mackay H, et al. HPV-independent vulvar squamous cell carcinoma is associated with significantly worse prognosis compared with HPV-associated tumors. Int J Gynecol Pathol. 2019;39:391–9.
Hay CM, Lachance JA, Lucas FL, Smith KA, Jones MA. Biomarkers p16, human papillomavirus and p53 predict recurrence and survival in early stage squamous cell carcinoma of the vulva. J Low Genit Trac Dis. 2016;20:252–6.
Lee LJ, Howitt B, Catalano P, Tanaka C, Murphy R, Cimbak N, et al. Prognostic importance of human papillomavirus (HPV) and p16 positivity in squamous cell carcinoma of the vulva treated with radiotherapy. Gynecol Oncol. 2016;142:293–8.
Ang KK, Harris J, Wheeler R, Weber R, Rosenthal DI, Nguyen-Tân PF, et al. Human papillomavirus and survival of patients with oropharyngeal cancer. N Engl J Med. 2010;363:24–35.
Watkins JC, Howitt BE, Horowitz NS, Ritterhouse LL, Dong F, MacConaill LE, et al. Differentiated exophytic vulvar intraepithelial lesions are genetically distinct from keratinizing squamous cell carcinomas and contain mutations in PIK3CA. Mod Pathol. 2017;30:448–58.
Nascimento AF, Granter SR, Cviko A, Yuan L, Hecht JL, Crum CP. Vulvar acanthosis with altered differentiation: a precursor to verrucous carcinoma? Am J Surg Pathol. 2004;28:638–43.
Yang B, Hart WR. Vulvar intraepithelial neoplasia of the simplex (differentiated) type: a clinicopathologic study including analysis of HPV and p53 expression. Am J Surg Pathol. 2000;24:429–41.
Singh N, Gilks CB. Vulval squamous cell carcinoma and its precursors. Histopathology. 2020;76:128–38.
McAlpine J, Leon-Castillo A, Bosse T. The rise of a novel classification system for endometrial carcinoma; integration of molecular subclasses. J Pathol. 2018;244:538–49.
Tessier-Cloutier B, Kortekaas KE, Thompson E, Pors J, Chen J, Ho J, et al. Major p53 immunohistochemical patterns in in situ and invasive squamous cell carcinomas of the vulva and correlation with TP53 mutation status. Mod Pathol. 2020;33:1595–1605.
Darragh TM, Colgan TJ, Cox JT, Heller DS, Henry MR, Luff RD, et al. The lower anogenital squamous terminology standardization project for HPV-Associated Lesions: background and consensus recommendations from the College of American Pathologists and the American Society for Colposcopy and Cervical Pathology. Arch Pathol Lab Med. 2012;136:1266–97.
Prentice LM, Miller RR, Knaggs J, Mazloomian A, Aguirre Hernandez R, Franchini P, et al. Formalin fixation increases deamination mutation signature but should not lead to false positive mutations in clinical practice. PLoS One. 2018;13:e0196434.
Richardson JT. The analysis of 2 x 2 contingency tables-yet again. Stat Med. 2011;30:890. author reply 1-2
Lee YY, Wilczynski SP, Chumakov A, Chih D, Koeffler HP. Carcinoma of the vulva: HPV and p53 mutations. Oncogene. 1994;9:1655–9.
Pilotti S, D’Amato L, Della Torre G, Donghi R, Longoni A, Giarola M, et al. Papillomavirus, p53 alteration, and primary carcinoma of the vulva. Diagn Mol Pathol. 1995;4:239–48.
Aulmann S, Schleibaum J, Penzel R, Schirmacher P, Gebauer G, Sinn HP. Gains of chromosome region 3q26 in intraepithelial neoplasia and invasive squamous cell carcinoma of the vulva are frequent and independent of HPV status. J Clin Pathol. 2008;61:1034–7.
Choschzick M, Hantaredja W, Tennstedt P, Gieseking F, Wölber L, Simon R. Role of TP53 mutations in vulvar carcinomas. Int J Gynecol Pathol. 2011;30:497–504.
Han MR, Shin S, Park HC, Kim MS, Lee SH, Jung SH, et al. Mutational signatures and chromosome alteration profiles of squamous cell carcinomas of the vulva. Exp Mol Med. 2018;50:e442.
Kashofer K, Regauer S. Analysis of full coding sequence of the TP53 gene in invasive vulvar cancers: Implications for therapy. Gynecol Oncol. 2017;146:314–8.
Nooij LS, Ter Haar NT, Ruano D, Rakislova N, van Wezel T, Smit VTHBM, et al. Genomic characterization of vulvar (pre)cancers identifies distinct molecular subtypes with prognostic significance. Clin Cancer Res. 2017;23:6781–9.
Trietsch MD, Spaans VM, ter Haar NT, Osse EM, Peters AA, Gaarenstroom KN, et al. CDKN2A(p16) and HRAS are frequently mutated in vulvar squamous cell carcinoma. Gynecol Oncol. 2014;135:149–55.
Regauer S, Kashofer K, Reich O. Time series analysis of TP53 gene mutations in recurrent HPV-negative vulvar squamous cell carcinoma. Mod Pathol. 2019;32:415–22.
Watkins JC, Yang E, Crum CP, Herfs M, Gheit T, Tommasino M, et al. Classic vulvar intraepithelial neoplasia with superimposed lichen simplex chronicus: a unique variant mimicking differentiated vulvar intraepithelial neoplasia. Int J Gynecol Pathol. 2019;38:175–82.
Xing D, Liu Y, Park HJ, Baek I, Tran H, Cheang G, et al. Recurrent genetic alterations and biomarker expression in primary and metastatic squamous cell carcinomas of the vulva. Hum Pathol. 2019;92:67–80.
Donehower LA, Soussi T, Korkut A, Liu Y, Schultz A, Cardenas M, et al. Integrated analysis of TP53 gene and pathway alterations in the cancer genome atlas. Cell Rep. 2019;28:1370–84 e5.
Lane DP. Cancer. p53, guardian of the genome. Nature. 1992;358:15–6.
van de Nieuwenhof HP, Massuger LF, van der Avoort IA, Bekkers RL, Casparie M, Abma W, et al. Vulvar squamous cell carcinoma development after diagnosis of VIN increases with age. Eur J Cancer. 2009;45:851–6.
Pinto AP, Miron A, Yassin Y, Monte N, Woo TY, Mehra KK, et al. Differentiated vulvar intraepithelial neoplasia contains Tp53 mutations and is genetically linked to vulvar squamous cell carcinoma. Mod Pathol. 2010;23:404–12.
Wilkinson EJ, Kneale B, Lynch PJ. Report of the ISSVD terminology committee. J Reprod Med Obstetrician Gynecol. 1986;31:973–4.
Abell MR, Gosling JR. Intraepithelial and infiltrative carcinoma of vulva: Bowen’s type. Cancer. 1961;14:318–29.
Hoang LN, Park KJ, Soslow RA, Murali R. Squamous precursor lesions of the vulva: current classification and diagnostic challenges. Pathology. 2016;48:291–302.
van den Einden LC, de Hullu JA, Massuger LF, Grefte JM, Bult P, Wiersma A, et al. Interobserver variability and the effect of education in the histopathological diagnosis of differentiated vulvar intraepithelial neoplasia. Mod Pathol. 2013;26:874–80.
Kokka F, Singh N, Faruqi A, Gibbon K, Rosenthal AN. Is differentiated vulval intraepithelial neoplasia the precursor lesion of human papillomavirus-negative vulval squamous cell carcinoma? Int J Gynecol Cancer. 2011;21:1297–305.
Reutter JC, Walters RA, Selim MA. Differentiated vulvar intraepithelial neoplasia: what criteria do we use in practice? J Low Genit Trac Dis. 2016;20:261–6.
Al-Bannai R, Miller D, Sadownik L, Blake Gilks C. Vulvar acanthosis with altered differentiation (VAAD): report of a case with progression to poorly differentiated carcinoma over a 5-yr period. Int J Gynecol Pathol. 2015;34:385–9.
Walton DB, Stearns L, Fillman EP, Banks N, Dalton SR. Vulvar acanthosis with altered differentiation: is this entity a variant of hypertrophic lichen sclerosus? J Cutan Pathol. 2015;42:1038–42.
Akbari A, Pinto A, Amemiya Y, Seth A, Mirkovic J, Parra-Herran C. Differentiated exophytic vulvar intraepithelial lesion: clinicopathologic and molecular analysis documenting its relationship with verrucous carcinoma of the vulva. Mod Pathol. 2020.
Pilotto S, Gkountakos A, Carbognin L, Scarpa A, Tortora G, Bria E. MET exon 14 juxtamembrane splicing mutations: clinical and therapeutical perspectives for cancer therapy. Ann Transl Med. 2017;5:2.
Acknowledgements
This study has supported by the Carraresi Foundation, Sumiko Kobayashi Marks Memorial Fund, OVCare and the UBC & VGH Hospital Foundations.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Tessier-Cloutier, B., Pors, J., Thompson, E. et al. Molecular characterization of invasive and in situ squamous neoplasia of the vulva and implications for morphologic diagnosis and outcome. Mod Pathol 34, 508–518 (2021). https://doi.org/10.1038/s41379-020-00651-3
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41379-020-00651-3
This article is cited by
-
HPV-independent, p53-wild-type vulvar intraepithelial neoplasia: a review of nomenclature and the journey to characterize verruciform and acanthotic precursor lesions of the vulva
Modern Pathology (2022)
-
Molecular landscape of vulvovaginal squamous cell carcinoma: new insights into molecular mechanisms of HPV-associated and HPV-independent squamous cell carcinoma
Modern Pathology (2022)
-
Histological interpretation of differentiated vulvar intraepithelial neoplasia (dVIN) remains challenging—observations from a bi-national ring-study
Virchows Archiv (2021)