Abstract
Hairy cell leukemia-variant is rare. Only a small number of cases have been reported in the literature with little cytogenetic or molecular data available. In this study, we describe the clinicopathologic and genetic features of 23 patients with hairy cell leukemia-variant (16 men and 7 women) with a median age of 70 years. Most patients had splenomegaly (90%), leukocytosis (77%), and lymphocytosis (82%); no patients had monocytopenia. Histologically, the bone marrow biopsy specimens showed a mixed pattern of predominantly interstitial and lesser intrasinusoidal infiltration by leukemic cells. In bone marrow aspirate smears most cells had villous cytoplasmic features and a small nucleolus. We describe unusual sites of hairy cell leukemia-variant involvement in 4 patients, including brain, omentum, terminal ileum, and skin at the time of initial presentation. Immunophenotyping showed monotypic B-cells positive for pan B-cell antigens, CD11c, and CD103, and negative for CD25 and annexin A1. Conventional cytogenetic or fluorescence in situ hybridization analysis showed deletions of 17p13/TP53 and 11q22/ATM gene in 5/12 (42%) and 2/9 (22%) cases, respectively. Sequencing of the variable region of IGVH showed mutations (>2% deviation from germline) in 40% of the cases assessed. MAP2K1 mutation (p.C121S) was seen in 1 of 14 (7%) patients tested. No BRAF V600E mutations were detected. The patients were treated in a heterogeneous manner, but most often with therapies designed for classical hairy cell leukemia and the 5-year overall survival was 84%. In summary, hairy cell leukemia-variant exhibits a heterogeneous spectrum of clinical, morphologic, immunophenotypic, and genetic features that may overlap with classic hairy cell leukemia and other hairy cell-like B-cell neoplasms. A subset of patients can have an aggressive clinical course. In our experience MAP2K1 mutations are uncommon in this disease.
Similar content being viewed by others
Introduction
Hairy cell leukemia-variant is a rare neoplasm that was first recognized by Cawley et al. [1] who proposed the designation type II hairy cell leukemia. Almost 40 years later, hairy cell leukemia-variant is included in the current World Health Organization (WHO) classification as a provisional entity [2] and is defined as a B-cell chronic lymphoproliferative disorder that resembles classical hairy cell leukemia, but exhibits a number of variant clinical, laboratory, and morphologic features. Hairy cell leukemia-variant typically involves the spleen, bone marrow and peripheral blood and patients are often resistant to conventional therapy used to treat classical hairy cell leukemia [3].
In part, the provisional status of hairy cell leukemia-variant in the current WHO classification can be attributed to an overall lack of clinicopathologic and genetic studies available in the literature, presumably because the disease is rare. To our knowledge, there are only a limited number of patient cohorts and case reports focusing on hairy cell leukemia-variant in the literature [4,5,6,7,8,9]. These series are, in general, small and they lack comprehensive data on histopathologic features, cytogenetic, or molecular findings, as well as data on therapy and follow-up. There is an obvious need to report additional cases of hairy cell leukemia-variant that have undergone a comprehensive workup including molecular data.
The aim of this study is to better characterize the clinicopathologic and molecular features of hairy cell leukemia-variant. We report our single institution experience of 23 cases and our study includes clinicopathologic features, immunophenotypic data, and the results of conventional cytogenetic and molecular studies.
Materials and methods
Study group
We searched the files of the Department of Hematopathology at The University of Texas MD Anderson Cancer Center from 1 January 2001 through 31 December 2017 for cases diagnosed as “hairy cell leukemia”, either classical hairy cell leukemia or hairy cell leukemia-variant. We identified 24 cases of hairy cell leukemia-variant, of which 23 cases had blood, bone marrow aspirate smears, and bone marrow biopsy sections available for review. One case of hairy cell leukemia-variant was diagnosed based on a splenectomy specimen and not included in this study due to lack of bone marrow material (Fig.1). The diagnosis of hairy cell leukemia-variant was based on criteria described in the current WHO classification [3]. One of these cases has been reported previously [6].
We performed a retrospective review of the medical records for all relevant clinical data, demographic information, therapy, and clinical follow-up. We also reviewed imaging studies for evidence of splenomegaly, hepatomegaly or lymphadenopathy. Splenomegaly was defined as a palpable spleen (≥5 cm below the left costal margin) and/or increased spleen size by magnetic resonance imaging or computerized tomography scans. The study was conducted under an Institutional Review Board–approved protocol.
Histopathologic assessment
Bone marrow specimens obtained at initial diagnosis and/or follow-up were evaluated in all cases. Hematoxylin and eosin–stained slides of core and/or aspirate clot specimens, with corresponding Wright–Giemsa-stained aspirate smears and/or touch imprints, and peripheral blood smears were reviewed. Multiple specimens were reviewed in 14 patients, including splenectomy (n = 4), liver biopsy (n = 3), lymph node biopsy (n = 2), omental mass biopsy (n = 1), and brain biopsy (n = 1) specimens.
Cytochemical and immunohistochemical studies
Cytochemical studies for acid phosphatase and tartrate-resistant acid phosphatase were performed on bone marrow aspirate smears in a subset of cases. Immunohistochemical analysis was performed on formalin-fixed, paraffin-embedded tissue sections of bone marrow aspirate clot and/or biopsy specimens, either at the submitting institution or in our own laboratory. For the latter, heat-induced epitope retrieval with an avidin-biotin complex method was used, and staining was performed using an automated immunostainer (Ventana Medical Systems, Tucson, AZ) as described previously [10]. A variable number of antibodies was used to evaluate these cases over time and included CD3, CD20, PAX5, cyclin D1, annexin A1, DBA.44, VE-1 (BRAF V600E), and p53.
Flow cytometry immunophenotypic analysis
Multicolor flow cytometric immunophenotypic analysis was performed using bone marrow aspirate specimens, either at the submitting institution or in our laboratory. In all cases, at least 4-color analysis was performed and lymphocytes were gated for analysis using CD45 and side scatter. The antibody panels used in different institutions were highly variable, but most panels included antibodies specific for: CD3, CD4, CD5, CD8, CD10, CD11c, CD19, CD20, CD23, CD25, CD43, CD45, CD56, CD79b, CD103, CD200, and surface immunoglobulin (Ig) light chains. In our institution, the specimens were analyzed using FACSCalibur cytometers (BD Biosciences, San Jose, CA) before 2009, and with FACSCanto II instruments (BD Biosciences, Mountain View, CA) after 2009 as described previously [11].
Cytogenetic studies
Conventional cytogenetic analysis was performed on G-banded metaphase cells prepared from bone marrow aspirate specimens as has been described [12]. Twenty metaphases were assessed, and the results were described using the International System for Human Cytogenetic Nomenclature [13].
Fluorescence in situ hybridization (FISH) was performed on a subset of cases using Abbott Molecular multicolor multiprobe panels designed for simultaneous detection of the 11q22.3 (ATM) region of chromosome 11, the 17p13.1 (TP53) region of chromosome 17, the alpha satellite, centromeric region of chromosome 12 (D12Z3), the D13S319 locus (located between RB1 and D13S25 loci) in the 13q14.3 region of chromosome 13, and the 13q34 region (LAMP1) near the subtelomere of chromosome 13q (Vysis, Abbott Laboratories, Abbott Park, IL). A total of 200 interphases were analyzed for each probe.
Molecular studies
B-cell clonality was determined using a polymerase chain reaction-based methods using V primers derived from the framework 1 (FR1), framework 2 (FR2), and framework 3 (FR3) regions, in combination with a consensus JH primer. PCR product was detected by capillary electrophoresis. The IGH variable region genes were sequenced in 11 cases using RNA extracted from fresh peripheral blood or bone marrow samples. Multiplex PCR amplification of IGH transcripts was performed using consensus V segment primers. Sanger sequencing of PCR products was compared with germline IGH sequence. Somatic hypermutation of IGHV was considered positive when there was ≥2% variation in the sequenced V-segment of clonal IGH sequences compared with the germline sequence. The test sensitivity was 20% [14].
Sequencing of BRAF and MAP2K1
Using genomic DNA extracted from bone marrow aspirate samples, mutations in BRAF and MAP2K1 were assessed either by next-generation sequencing or by PCR-based pyrosequencing (for BRAF: codons 468 to 470 of exon 11 and codons 595 to 600 of exon 15, lower limit of detection 5–10%) or by Sanger sequencing (for MAP2K1: exons 2, codons 28 to 97 and exon 3, codons 98 to 146, lower limit of detection 10%-20%) using standard techniques as described elsewhere [15].
Statistical analysis
Statistical analysis was performed using IBM SPSS Statistics Version 24.0 (SPSS Inc., Chicago, Illinois, USA). For continuous variables, data were reported as median and range. For nominal variables, data were reported as the number of patients unless otherwise specified. Overall survival (OS) was estimated by using Kaplan–Meier curves. Overall survival was defined as the time from the date of diagnosis to the date of death or last follow-up. A P value of less than 0.05 was considered statistically significant.
Results
Clinical data
The study group included 16 men and 7 women with a median age of 70 years (range, 50–89 years) at time of initial diagnosis. The clinical and laboratory features are summarized in Table 1. Clinical symptoms were variable and included left upper quadrant abdominal pain (n = 5) and B-type symptoms (n = 2). Some patients had symptoms attributable to cytopenias. Nine patients were asymptomatic and the diagnostic workup was triggered by an incidental detection of leukocytosis during routine physical examination or during evaluation of other medical conditions.
Splenomegaly was detected either by physical examination or computerized tomography imaging in 19 (90%) patients. Lymphadenopathy was detected by imaging studies in 6 of 17 (33%) patients, often involving abdominal or retroperitoneal nodes. Hepatomegaly was present in 5 of 18 (28%) patients. In four patients, there was involvement of other unusual tissue sites including the brain, omentum, terminal ileum, and skin.
Laboratory findings
A complete blood count was available for 22 patients. Anemia was present in 12 (55%) patients with a median hemoglobin concentration of 11.6 g/dL (range 6.7–16.2 g/dL [reference range 14–18 g/dL]). Seventeen (77%) patients presented with thrombocytopenia with a median platelet count of 104 × 109/L (range, 21–295 × 109/L [normal range, 140–440 × 109/L]). Leukocytosis was present in 17 (77%) patients with a median leukocyte count of 22.0 × 109/L (range, 4.2–74 × 109/L [reference range, 4–11 × 109/L]). The median absolute lymphocyte count was 18.6 × 109/L (range, 1.3–72.0 × 109/L) with median percentage of 75% (range, 31 to 90%). An absolute lymphocytosis was present in 18 of 22 (82%) patients. None of the patients had monocytopenia. The serum lactate dehydrogenase level was within normal limits in 8 (44%) patients and above the upper limit in 10 (56%) patients (median 653 U/L, range 363–1045). The serum total calcium was within normal range in 17 (95%) patients and decreased in 1 (5%) patient (median, 9.2 md/dL; range, 7.3–10.0).
Histopathologic features
Bone marrow and peripheral blood
Bone marrow core biopsy sections and aspirate smears were examined for all 23 patients and the data are summarized in Table 2. The overall bone marrow cellularity ranged from 30 to 95%, with at least some residual trilineage hematopoiesis observed in all cases. The median bone marrow tumor burden was 50% (range, 10–90%). The tumor showed a predominantly interstitial pattern of infiltration within the bone marrow in 22 (95.7%) of 23 patients (Fig. 2a–c). In 12 (52%) patients, some degree of sinusoidal pattern of infiltration could be appreciated (Fig. 2d–f). The sinusoidal pattern was prominent in 2 patients (#9 and #10), both of which showed a low tumor burden (15 and 20%). Three (13%) patients showed diffuse pattern of bone marrow involvement and 1 patient had an interstitial and nodular pattern of infiltration. Staining for reticulin and trichrome was performed in 8 cases and 6 (75%) showed variable fibrosis; mild (MF-1) in 3 cases and dense (MF-3) in 3 cases.
Bone marrow trephine biopsy showing involvement by hairy cell leukemia-variant with predominant interstitial pattern (a hematoxylin and eosin, ×200), highlighted by CD20 and DBA.44 (b, c immunohistochemistry, ×200). Bone marrow involvement by hairy cell leukemia-variant with sinusoidal pattern which is difficult to appreciate on routine staining (d hematoxylin and eosin, ×200). Lymphoma cells form clusters in the sinusoids (arrowhead) highlighted by PAX5 (e immunohistochemistry, ×200). High-power of sinusoidal involvement highlighted by PAX5 (inset) (f hematoxylin and eosin, ×1000)
The peripheral blood smears showed circulating lymphocytes with “hairy” frayed cytoplasmic edges in all patients. In blood and bone marrow smears the leukemic cells were predominantly intermediate-to-large in size with moderate to abundant lightly basophilic cytoplasm with delicate projections and round to oval nuclei; however the cytomorphology varied within and between individual cases (Fig.3). Distinct nucleoli were present in 14 (61%) cases, occasional inconspicuous nucleoli were observed in 6 (26%) cases, and nucleoli were not identified in 5 (22%) cases. Atypical nuclear features such as irregular nuclear contours, clefted nuclei or blastoid chromatin were seen in 2 (9%) patients (cases #4 and #11). In patient #11, the initial diagnostic bone marrow smears showed predominantly intermediate-sized cells with condensed chromatin, rare cells with prominent nucleoli and occasional intermediate-to-large-sized nuclei with occasional nuclear clefts. At the time of relapse (8 years later) bone marrow smears showed predominantly large leukemic cells with irregular nuclear contours, frequent forms with nuclear clefts, open chromatin with prominent single large nucleolus (Fig. 3c–e). Tartrate treated acid phosphatase was performed on bone marrow smears in 15 cases; 8 cases were weakly positive and the remaining 7 cases were negative.
Peripheral blood smear in case #12 showing neoplastic lymphocytes with light basophilic cytoplasm and delicate villous/“hairy” projections (a hematoxylin and eosin, ×400; inset, ×1000). Bone marrow aspirate smear in patient #15 showing leukemic cells with oval nuclei, dispersed chromatin and small nucleoli (b hematoxylin and eosin, ×400; inset, ×1000). Leukemic cells with binucleation, convoluted nuclei (c, d, case #11) and blastic chromatin (e case #11 and f case#4) (c–f Wright–Giemsa, ×1000)
Immunophenotypic findings
The results of immunophenotypic studies are summarized in Table 3. Immunohistochemical studies showed that the neoplastic cells were positive for CD20 and PAX-5 in all cases assessed. DBA.44 (CD72) was strongly expressed in 14 of 15 (93%) cases. The neoplastic cells were negative for annexin A-1 (n = 18), cyclin D1 (n = 10), and BRAF V600E (n = 7). Flow cytometry immunophenotyping showed that the leukemia cells had a mature B-cell immunophenotype, positive for CD19, CD20, CD22, and CD79b in all cases. The leukemic cells were also positive for CD11c (100%; bright), CD52 (100%), CD103 (95%; usually bright), FMC7 (91%; bright), and CD200 (58%; dim). The neoplastic cells expressed monotypic surface immunoglobulin light chains in 20 of 22 cases (10 kappa and 10 lambda) with bright expression in 12 of 19 (63%) of cases. CD25 expression was uncommon, observed in only 3 (14%) cases and was usually dim/partial (Fig. 4). CD123 was performed on only 1 case and it was positive. The neoplastic cells were negative for CD5, CD10, CD23, CD38, and CD43. In 13 patients, flow cytometry immunophenotyping was performed at multiple time points during the course of therapy and the immunophenotype was similar to the diagnostic specimen, with the exception of CD20 being negative in patients receiving rituximab.
Histopathologic findings in other sites
Five patients underwent splenectomy and spleen specimens were available for review in four patients. Grossly, the spleen was enlarged in all four patients with a median weight of 2465 g (range, 1740–4500). Histologic sections showed extensive infiltration of red pulp cords and sinusoids by a monomorphous infiltrate of lymphoid cells with variably prominent nucleoli and moderately abundant cytoplasm (Fig. 5a–d). Red blood cell lakes were noted in 1 patient (case #4).
Spleen, lymph node and brain involvement by hairy cell leukemia-variant. a–d Hairy cell leukemia-variant diffusely infiltrates the red pulp of the spleen. Neoplastic cells display a fried egg appearance (a hematoxylin and eosin, ×200; b hematoxylin and eosin, ×1000). CD20 immunostain shows a diffuse replacement of the red pulp with scattered CD3-positive T-cells (c, d immunohistochemistry with hematoxylin counter stain, ×200). e–h Lymph node diffusely infiltrated by hairy cell leukemia-variant. The infiltrate is characterized by intermediate-sized lymphocytes with oval nuclei and moderately abundant clear cytoplasm, some of the leukemic cells show prominent nucleoli (arrowhead) (e hematoxylin and eosin, ×200; f hematoxylin and eosin, ×1000). Hairy cell leukemia-variant cells highlighted by immunohistochemical B-cell marker CD20 admixed with background small CD3-positive T-cells. (g, h immunohistochemistry with hematoxylin counter stain, ×200). i–k Involvement of the brain by hairy cell leukemia-variant (case #3): Multiple brain lesions by MRI (i). Brain biopsy specimen showing brain parenchyma with hemorrhages resembling blood lakes containing atypical lymphocytes (j hematoxylin and eosin, ×100). High-power showing blastoid morphology of hairy cell leukemia-variant cells highlighted by CD20 (inset) (k hematoxylin and eosin, ×400)
In addition, six patients had biopsy proven hairy cell leukemia-variant involving unusual sites including the liver, lymph node, brain, omentum, terminal ileum, and skin. The three patients with liver biopsies showed leukemic infiltration of the portal tracts and leukemic cells within dilated sinusoids. Two patients showed lymph node involvement (cases #4 and #22). The nodal architecture was effaced by sheets of monotonous, intermediate-to-large in size lymphoid cells with abundant cytoplasm and irregular nuclear contours, many of which showed prominent nucleoli. Infrequent mitotic activity and increased large cells were noted both patients (Fig.5e–h). The infiltrate expanded the interfollicular areas with rare residual follicles (best appreciated by CD21 immunostain). The biopsy specimen of an omental nodule showed fibro-adipose tissue with areas of hemorrhage containing atypical lymphocytes. The brain biopsy specimen (case #3) showed clusters of intermediately sized lymphoid cells with variably open chromatin, distinct nucleoli, and a moderate amount of cytoplasm involving brain parenchyma associated with areas of hemorrhage (Fig.5j, k). The immunophenotype of leukemic cells in the unusual sites was similar to that described in the bone marrow infiltrate.
Cytogenetic findings
Conventional karyotypes performed on bone marrow aspirates were available in 17 patients (Table 4). Twelve (71%) patients had an abnormal karyotype. A complex karyotype, defined here as three or more chromosomal aberrations, was present in five (29%) patients. Recurrent aberrations included loss of Y (n = 4), 17p abnormalities (n = 3) and del[18q] (n = 3).
FISH analysis showed TP53 deletion or monosomy 17 in 5 of 12 (42%) cases with a median percentage of positive nuclei of 47% (range, 41.5%-96%); of note, only 1 of these cases was also detected by conventional cytogenetics as part of a complex karyotype. ATM deletion or monosomy 11 was detected in 2 of 9 (22.2%) cases analyzed by FISH of which one was also detected by karyotype. One case showed del[7q] by both conventional karyotype and FISH analysis.
Molecular results
Thirteen of fourteen (93%) patients showed monoclonal IGH rearrangements. Somatic hypermutation analysis was performed in 10 patients; 4 (40%) patients showed mutated IGHV. The IGHV4-34 variable region was utilized in 2 (20%) patients. No patients had BRAF mutations (n = 9). In 1 of 14 (~7%) patients, a MAP2K1 mutation was detected in codon 121, exon 3 (TGC to AGC) that would change the encoded amino acid from cystine to serine (p.C121S).
Therapy and clinical follow-up
Treatment information and clinical follow-up were available in 17 patients (Table 5). Combined chemoimmunotherapy with cladribine (2-chlorodeoxyadenosine, 2-CDA) plus rituximab was used in 11 patients. Multiple therapeutic regimens were explored in 4 patients, all of whom showed resistance to initial treatment with purine analogues as a single agent or with rituximab. One patient received rituximab alone and 1 patient did not receive treatment for hairy cell leukemia-variant. Therapeutic splenectomy was performed in 5 patients, either as a part of initial treatment or during the course of the disease.
The median follow-up duration from initial diagnosis to last follow-up or patient death was 37.3 months (range, 14–254 months). During the follow-up interval, 12 (70%) patients had at least one relapse and 6 patients had 3 or more relapses, involving bone marrow or other extramedullary sites (brain, liver and lymph node). One patient, developed ataxia and aphasia with altered mental status at 15 months while on therapy with cladribine. A computerized tomography scan showed multiple brain lesions. Craniotomy and brain biopsy confirmed involvement of the brain parenchyma by hairy cell leukemia-variant.
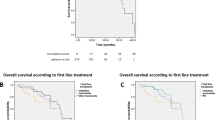
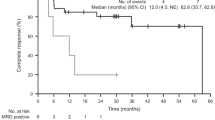
The five-year overall survival for the entire cohort was 85% and the median overall survival time was 89.2 months (95% CI: 86.0–92.5) (Fig.6). At the end of the follow-up interval, 11 (64.7%) patients were alive; 4 were without disease activity (negative bone marrow studies and/or negative flow cytometry of blood) and 7 had persistent/relapsed hairy cell leukemia-variant disease. The remaining six patients died: three died from hairy cell leukemia-variant or its complications, and three died from complications from other malignancies (two lung carcinoma, one primary myelofibrosis). The patient with lung carcinoma had minimal residual hairy cell leukemia-variant in bone marrow (1.9%) detected by flow cytometry.
Second malignancies
Four patients in this study had a second malignancy. Two patients developed lung adenocarcinoma, 17 and 92 months after hairy cell leukemia-variant diagnosis and treatment with cladribine. Two other patients had other concomitant hematologic neoplasms: One patient (case 2) had chronic myelomonocytic leukemia and hairy cell leukemia-variant (~10% bone marrow involvement) diagnosed simultaneously. The hematopoietic cells in this case showed moderate trilineage dysplasia. Treatment of hairy cell leukemia-variant with cladribine and rituximab resulted in remission within three months after which treatment for chronic myelomonocytic leukemia was initiated. Another patient (case 8) had primary myelofibrosis associated with dense reticulin (MF-3) fibrosis first. This patient was treated with a JAK2 inhibitor and later developed hairy cell leukemia-variant. No data regarding the details of hairy cell leukemia-variant therapy were available, but follow-up bone marrow evaluation three months after hairy cell leukemia-variant diagnosis was negative for residual hairy cell leukemia-variant and showed persistent primary myelofibrosis.
Discussion
We compared the clinical and pathologic characteristics of this patient cohort to previously reported studies (Table 6). Most patients with hairy cell leukemia-variant were elderly men. The male-to-female ratio in this cohort was 2:1 and the median age was 70 years. However, the male predominance in hairy cell leukemia-variant is less pronounced than in patients with classical hairy cell leukemia (reported male predominance of 5-to-1) [4]. The most common clinical manifestations of hairy cell leukemia-variant, such as abdominal discomfort/pain and cytopenias, can be attributed to splenomegaly, hypersplenism, and/or bone marrow infiltration [3, 16]. However, the advent of more sensitive methods for detection of hairy cell leukemia-variant is leading to diagnosis at an early stage with lower disease burden. Thrombocytopenia and anemia were the most common initial presentations seen in about half of the patients in this cohort, followed by incidental detection of leukocytosis and/or neoplastic lymphocytes in the peripheral blood. Overall, B-symptoms were uncommon in patients with hairy cell leukemia-variant, in about 10%. In most patients with leukocytosis, the elevation was moderate, commonly 20–40 × 109/L. An absolute lymphocytosis was present in about 80% of patients and the absolute number and proportion of monocytes were normal or even slightly elevated. These findings are in contrast to patients with classical hairy cell leukemia who often present with pancytopenia and monocytopenia.
Physical examination and imaging studies showed splenomegaly, often massive, in over 90% of the patients in this study. Hepatomegaly was documented in about 25% of patients, in accord with a frequency of 14–30% reported by others [5, 16, 17]. In the literature, intra-abdominal lymphadenopathy has been reported in 5–15% of patients [16]. However, imaging studies in this cohort showed a higher frequency of lymphadenopathy in about one-third of patients involving abdominal, retroperitoneal, diaphragmatic, and axillary lymph nodes. Therefore, the imaging studies are helpful in the diagnostic workup to detect lymphadenopathy and estimate the extent of the disease.
Involvement of sites other than spleen, peripheral blood, and bone marrow by hairy cell leukemia-variant is rarely documented, as isolated case reports of cutaneous lesions [16, 18] or a periarticular joint infiltration [19]. In this study, we describe the pathologic features of unusual sites of involvement including the brain, omentum, terminal ileum, and skin developing during the course of disease. The patient with brain involvement is of particular interest. During treatment with cladribine he developed ataxia and aphasia with altered mental status 15 months after initial diagnosis. Multiple bilateral brain lesions were seen on imaging and biopsy showed hairy cell leukemia-variant involving the brain parenchyma. To our knowledge, this is the first documented case of leukemic involvement of the brain by hairy cell leukemia-variant. Brain involvement is also rare in patients with classical hairy cell leukemia [20, 21].
Although the diagnosis of hairy cell leukemia-variant can be made on peripheral blood, bone marrow biopsy, and aspirate examination is crucial at the time of initial presentation. Bone marrow evaluation allowed an accurate estimation of the extent of involvement at baseline to assess future treatment response and provided sufficient material for ancillary studies. Unlike in classical hairy cell leukemia, in patients with hairy cell leukemia-variant, the bone marrow aspirate is often cellular with variable degree of tumor involvement [3, 16]. This fibrosis was not associated with the degree of lymphomatous infiltration. In one case, this finding was most likely due to the underlying primary myelofibrosis.
Hairy cell leukemia-variant cells on peripheral blood and bone marrow aspirate smears are usually monomorphic, small to intermediate in size with abundant cytoplasm, cytoplasmic projections, and a single nucleolus. However, none of these features is specific or constant in all cases. Prominent nucleoli have been reported as a typical characteristic of hairy cell leukemia-variant, they are not a constant feature in all hairy cell leukemia-variant [7]. As shown in our study, some hairy cell leukemia-variant cases exhibited inconspicuous nucleoli (27%) or nucleoli were not identified (23%). Therefore, the description of hairy cell leukemia-variant as “prolymphocytoid” can be misleading. Cases of hairy cell leukemia-variant usually display regular nuclear contours [16]. Rare cases, however, are composed of leukemic cells with convoluted nuclear contours, binucleation or immature chromatin (blastoid variant) [16]. Nuclear atypia was seen in 2 (9%) cases in this cohort. In one of those cases, the nuclear atypia developed over eight years. A second case presented to our institution with blastoid morphology, extensive bone marrow involvement, and brain lesions after being treated with cladribine. Both cases with nuclear atypia suggested disease progression or aggressive behavior.
Bone marrow biopsy examination showed a mixed pattern of interstitial and intrasinusoidal infiltration with the interstitial component being the most prevalent component. Half of the cases showed some degree of intrasinusoidal involvement best visualized by immunohistochemical assessment, but in most cases it was subtle compared with a more extensive interstitial infiltration. Interestingly, in two of our cases the predominant pattern was intrasinusoidal, which was difficult to appreciate on routine H&E stain due to low tumor burden and background preserved hematopoiesis. However, intrasinusoidal infiltration in hairy cell leukemia-variant is rarely seen as an exclusive and “single” pattern [22, 23]. Intrasinusoidal bone marrow infiltration has been considered a specific histologic finding in splenic marginal zone lymphoma but it can occasionally occur in other B-cell lymphomas with splenomegaly, particularly classical hairy cell leukemia, hairy cell leukemia-variant, and splenic diffuse red pulp small B-cell lymphoma, as well as other nodal small B-cell lymphoma [24]. A nodular pattern of involvement, frequently described in some of the earlier studies on hairy cell leukemia-variant, was seen in only one of our cases in which the nodules were distributed in interstitial and juxtatrabecular fashion, without paratrabecular localization. In this cohort nearly 75% of cases had some degree of reticulin fibrosis.
Due to the overlapping morphologic features, immunophenotypic analysis is essential to reliably distinguish hairy cell leukemia-variant from classic hairy cell leukemia and other B-cell lymphomas. Consistent with data in the literature, the neoplastic cells in hairy cell leukemia-variant show bright expression of pan-B-cell markers, such as CD19, CD20, CD22, CD79b, and also brightly positive for surface light chains [8, 25]. Two cases (<10%) did not express surface immunoglobulins as reported in earlier studies [26]. We did not observe λ-light chain predominance in this cohort, in contrast to the previously reported significantly higher prevalence of λ-expression that is seen in classical hairy cell leukemia [26]. Cases of hairy cell leukemia-variant shared many immunophenotypic features with classical hairy cell leukemia, such as expression of CD11c and CD103. However, hairy cell leukemia-variant cells lacked CD25, CD123, tartrate-resistant alkaline phosphatase, and annexin A1 [3]. CD103, a cell surface glycoprotein of the integrin β-7 family, is regarded as a useful diagnostic criterion for diagnosis. CD103 expression in hairy cell leukemia-variant greatly varies in the literature from 36–100%; [8, 25] in our study, it was expressed in 21 of 22 cases (~95%) with predominantly bright intensity. CD200 is a type I immunoglobulin superfamily membrane protein and is another marker helpful in distinguishing hairy cell leukemia-variant from classical hairy cell leukemia, which is bright in the latter. In this study, 64% of hairy cell leukemia-variant cases showed positivity for CD200 and in all of them the expression was consistently dim. All hairy cell leukemia-variant cases were negative for annexin-A1 and cyclin D1 [27, 28], unlike classical hairy cell leukemia.
Cytogenetic data in hairy cell leukemia-variant are very limited. Generally, hairy cell leukemia-variant cells show few chromosomal abnormalities and no recurrent genetic abnormalities have been described. In our cohort, a complex karyotype was noted in about 30% of patients. Three of these included abnormalities in 17p. FISH analysis showed TP53 deletion in 5 cases. None of the tested patients showed abnormalities in 8q24 or chromosome 5 gains, as previously described [16, 29]. One case showed loss of 7q, a characteristic feature of splenic marginal zone lymphoma [30]. However, the morphologic findings of large blastoid cells with prominent nucleolus, bright expression of CD11c and CD103, weak tartrate-resistant alkaline phosphatase staining, as well as aggressive behavior with brain involvement, helped differentiating this case from splenic marginal zone lymphoma. Although abnormalities involving 17p are rare, studies had shown that hairy cell leukemia-variant patients with monoallelic deletions of TP53 and the patients with higher numbers of cells with deleted p53 tend to progress and/or undergo transformation [23, 31]. In this study, patients with TP53 abnormalities tended to have a more aggressive clinical course with abdominal lymphadenopathy and/or liver involvement (n = 4), and unresponsive to treatments with multiple relapses (n = 2).
In the era of precision medicine, identification of therapeutic targets offers additional options for patients who do not respond to conventional chemotherapy. Waterfall et al. [32] assessed 24 patients with hairy cell leukemia-variant and showed 10 (42%) with activating MAP2K1 (MEK1) mutations. A recent study on 8 relapsed refractory hairy cell leukemia-variant patients showed MAP2K1 mutations in 3 (38%) [33]. However, Mason et al. [7] found only 1 of 11 (9%) hairy cell leukemia-variant patients mutated for MAP2K1. In the current study, we identified only 1 of 14 (~7%) hairy cell leukemia-variant patients with a MAP2K1 mutation, in accord with Mason et al. [7]. The reason for the discrepancy between the studies is unclear. In our study, we performed Sanger sequencing (targeting exons 2 and 3) on patient samples with >40% tumor burden in the bone marrow aspirate to ensure adequate representation, given that the lower limit of detection of our assay was between 10–20%. The study by Waterfall et al. [32] employed whole exome sequencing on seven cases (range of median depth of coverage reported between 24 × and 89 × ); the remaining 17 cases (7 of which were mutated, 41%) underwent Sanger sequencing for exons 2 and 3, similar to our study. This suggests that testing methodology is likely not a reason for the discrepancy. Furthermore, Mason et al. [7] also reported a low mutation frequency despite using a targeted NGS-based sequencing that interrogated all of the coding exons of MAP2K1. Based on this, we believe that the heterogeneity in the genetic findings reported between different groups is most likely related to the patient selection. Another possibility, so far unexplored, includes potential selection and expansion of MAP2K1 mutant clones over the course of the disease and likely modified by therapy. Notably, the C121S mutation identified in our patient is the most common MAP2K1 alteration in hairy cell leukemia-variant and is known to confer resistance to current MEK inhibitors [32]. However, certain MAP2K1 mutations that do not disrupt drug binding may be amenable for treatment with MEK inhibitors, as recently demonstrated in a case report [18].
Published data regarding the outcome of hairy cell leukemia-variant are very limited due to disease rarity. Hairy cell leukemia-variant patients respond poorly to interferon alpha or single agent purine analogs, with a median overall survival of ~9 years [23, 25]. In this cohort, we report a 5-year overall survival of 84%, higher than reported in the literature, although the median follow-up duration was relatively short (37 months). The higher survival rate in this cohort may be attributable to differences in therapy compared with those used in the past [23]. The current recommended therapy is cladrabine followed by rituximab, with a reported 65% 5-year FFS rate [5, 34]. This was the most frequently used regimen in our cohort (11 patients). A study from Memorial Sloan Kettering reported that hairy cell leukemia-variant patients had shorter time to next treatment than patients with classic hairy cell leukemia, but had a similar overall survival (median follow-up was 47 months) [35]. In addition, recent report suggests that treatment with anti-CD22 immunotoxins, anti-CD52 antibody (alemtuzumab), fludarabine, or ibrutinib is effective in hairy cell leukemia-variant, and these agents need further investigation [16, 34]. The absence of BRAF V600E mutations and rarity of MAP2K1 mutation preclude currently available treatment modalities with MEK inhibitors in many patients.
In summary, patients with hairy cell leukemia-variant can exhibit a heterogeneous spectrum of clinical, morphologic, immunophenotypic, and genetic features. The diagnosis of hairy cell leukemia-variant is frequently a challenge, due to lack of specific histological features and immunophenotypic markers, and the close resemblance to other poorly characterized lymphomas. Evaluation of immunophenotype by flow cytometry and immunohistochemistry is a useful tool in distinguishing hairy cell leukemia-variant from other splenic B-cell neoplasms. In this study, MAP2K1 mutations were infrequent.
References
Cawley JC, Burns GF, Hayhoe FGJ. A chronic lymphoproliferative disorder with distinctive features: a distinct variant of hairy-cell leukaemia. Leuk Res. 1980;4:547–59.
Foucar K, Catovsky D. Hairy cell leukaemia. In: Jaffe E, Harris N, Stein H, (eds.) WHO Classification Tumours of Haematopoietic and Lymphoid Tissues. Lyon, France: IARC Press, 2001. p. 138-41.
Piris M, Foucar K, Mollejo M. et al. In: Swerdlow S, Campo E, Harris N, (eds.) Splenic B-cell lymphoma/leukemia, unclassifiable. WHO Classification of Tumours of Haematopoietic and Lymphoid Tissues. Revised 4th Edition Lyon, France: IARC; 2017. p. 229–31. .
Cessna MH, Hartung L, Tripp S, et al. Hairy cell leukemia-variant: fact or fiction. Am J Clin Pathol. 2005;123:132–8.
Hockley SL, Else M, Morilla A, et al. The prognostic impact of clinical and molecular features in hairy cell leukaemia variant and splenic marginal zone lymphoma. Br J Haematol. 2012;158:347–54.
Hu Z, Sun Y, Wang W, et al. Refractory hairy cell leukemia-variant. Am J Hematol. 2016;92:1398–9.
Mason EF, Brown RD, Szeto DP, et al. Detection of activating MAP2K1 mutations in atypical hairy cell leukemia and hairy cell leukemia variant. Leuk Lymphoma. 2017;58:233–6.
Shao H, Calvo KR, Grönborg M, et al. Distinguishing hairy cell leukemia variant from hairy cell leukemia: development and validation of diagnostic criteria. Leuk Res. 2013;37:401–9.
Zinzani PL, Lauria F, Buzzi M, et al. Hairy cell leukemia variant: a morphologic, immunologic and clinical study of 7 cases. Haematologica. 1990;75:54–7.
Khoury JD, Wang WL, Prieto VG, et al. Validation of immunohistochemical assays for integral biomarkers in the NCI-MATCH EAY131 clinical trial. Clin Cancer Res. 2018;24:521–31.
Hu Z, Sun Y, Schlette EJ, et al. CD200 expression in mantle cell lymphoma identifies a unique subgroup of patients with frequent IGHV mutations, absence of SOX11 expression, and an indolent clinical course. Mod Pathol. 2018;31:327–36.
Boddu PC, Wang SA, Pemmaraju N, et al. 8q24/MYC rearrangement is a recurrent cytogenetic abnormality in blastic plasmacytoid dendritic cell neoplasms. Leuk Res. 2018;66:73–8.
ISCN 2016: An International System for Human Cytogenomic Nomenclature. In: McGowan-Jordan JAS Schmid M, (eds.) 1st ed. Switzerland: Karger Publishers; 2016;149:1-140.
Jain P, Nogueras Gonzalez GM, Kanagal-Shamanna R, et al. The absolute percent deviation of IGHV mutation rather than a 98% cut-off predicts survival of chronic lymphocytic leukaemia patients treated with fludarabine, cyclophosphamide and rituximab. Br J Haematol. 2018;180:33–40.
Alayed K, Medeiros LJ, Patel KP, et al. BRAF and MAP2K1 mutations in Langerhans cell histiocytosis: a study of 50 cases. Hum Pathol. 2016;52:61–7.
Matutes E, Martinez-Trillos A, Campo E. Hairy cell leukaemia-variant: disease features and treatment. Best Pract Res Clin Haematol. 2015;28:253–63.
Sainati L, Matutes E, Mulligan S, et al. A variant form of hairy cell leukemia resistant to alpha-interferon: clinical and phenotypic characteristics of 17 patients. Blood. 1990;76:157–62.
Andritsos LA, Grieselhuber NR, Anghelina M, et al. Trametinib for the treatment of IGHV4-34, MAP2K1-mutant variant hairy cell leukemia. Leuk Lymphoma. 2018;59:1008–11.
McKay MJ, Rady KL, McKay TA. Hairy cell leukaemia variant with periarticular joint infiltration and excellent radiotherapy response. Ann Transl Med. 2017;5:162.
Chandana SR, Kotecha R, Al-Janadi A, et al. Rare case of hairy cell leukemia with brain parenchymal involvement: a diagnostic dilemma. J Clin Oncol. 2013;31:e186–8.
Tadmor T, Polliack A. Hairy cell leukemia: Uncommon clinical features, unusual sites of involvement and some rare associations. Best Pract Res Clin Haematol. 2015;28:193–9.
Ya-In C, Brandwein J, Pantalony D, et al. Hairy cell leukemia variant with features of intrasinusoidal bone marrow involvement. Arch Pathol Lab Med. 2005;129:395–8.
Matutes E, Wotherspoon A, Brito-Babapulle V, et al. The natural history and clinico-pathological features of the variant form of hairy cell leukemia. Leukemia. 2001;15:184–6.
Ponzoni M, Kanellis G, Pouliou E, et al. Bone marrow histopathology in the diagnostic evaluation of splenic marginal-zone and splenic diffuse red pulp small B-cell lymphoma: a reliable substitute for spleen histopathology? Am J Surg Pathol. 2012;36:1609–18.
Arons E, Suntum T, Stetler-Stevenson M, et al. VH4-34 + hairy cell leukemia, a new variant with poor prognosis despite standard therapy. Blood. 2009;114:4687–95.
Matutes E. Immunophenotyping and differential diagnosis of hairy cell leukemia. Hematol Oncol Clin North Am. 2006;20:1051–63.
Dong HY, Weisberger J, Liu Z, et al. Immunophenotypic analysis of CD103 + B-lymphoproliferative disorders: hairy cell leukemia and its mimics. Am J Clin Pathol. 2009;131:586–95.
Falini B, Tiacci E, Liso A, et al. Simple diagnostic assay for hairy cell leukaemia by immunocytochemical detection of annexin A1 (ANXA1). Lancet. 2004;363:1869–70.
Hockley SL, Morgan GJ, Leone PE, et al. High-resolution genomic profiling in hairy cell leukemia-variant compared with typical hairy cell leukemia. Leukemia. 2011;25:1189–92.
Watkins AJ, Huang Y, Ye H, et al. Splenic marginal zone lymphoma: characterization of 7q deletion and its value in diagnosis. J Pathol. 2010;220:461–74.
Vallianatou K, Brito-Babapulle V, Matutes E, et al. p53 gene deletion and trisomy 12 in hairy cell leukemia and its variant. Leuk Res. 1999;23:1041–5.
Waterfall JJ, Arons E, Walker RL, et al. High prevalence of MAP2K1 mutations in variant and IGHV4-34-expressing hairy-cell leukemias. Nat Genet. 2014;46:8–10.
Durham BH, Getta B, Dietrich S, et al. Genomic analysis of hairy cell leukemia identifies novel recurrent genetic alterations. Blood. 2017;130:1644–8.
Chihara D, Kantarjian H, O’Brien S, et al. Long-term durable remission by cladribine followed by rituximab in patients with hairy cell leukaemia: update of a phase II trial. Br J Haematol. 2016;174:760–6.
Getta BM, Woo KM, Devlin S, et al. Treatment outcomes and secondary cancer incidence in young patients with hairy cell leukaemia. Br J Haematol. 2016;175:402–9.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Rights and permissions
About this article
Cite this article
Angelova, E.A., Medeiros, L.J., Wang, W. et al. Clinicopathologic and molecular features in hairy cell leukemia-variant: single institutional experience. Mod Pathol 31, 1717–1732 (2018). https://doi.org/10.1038/s41379-018-0093-8
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41379-018-0093-8
This article is cited by
-
A comparison of the International Consensus and 5th World Health Organization classifications of mature B-cell lymphomas
Leukemia (2023)
-
Hairy Cell Leukemia (HCL) and HCL Variant: Updates and Spotlights on Therapeutic Advances
Current Oncology Reports (2022)
-
Advances in the Treatment of Hairy Cell Leukemia Variant
Current Treatment Options in Oncology (2022)
-
Purine nucleoside analogs plus rituximab are an effective treatment choice for hairy cell leukemia-variant
Annals of Hematology (2022)
-
The 5th edition of the World Health Organization Classification of Haematolymphoid Tumours: Lymphoid Neoplasms
Leukemia (2022)