Abstract
Gastric extranodal marginal zone lymphoma of mucosa-associated lymphoid tissue is a distinct entity in that Helicobacter pylori infection plays the most important causative role in the development of the disease. To investigate the genomic alteration in gastric marginal zone lymphoma that was resistant to the H. pylori eradication therapy, we analyzed 19 cases of the gastric marginal zone lymphoma using fluorescence in situ hybridization for MALT1, BCL10 rearrangement, and targeted sequencing using an Illumina platform. Major genetic alterations affected genes involved in nuclear factor (NF)-κB pathway activation and included MALT1 rearrangement (39%), and somatic mutations of TRAF3 (21%), TNFAIP3 (16%), and NOTCH1 (16%). In the MALT1 rearrangement-negative group, disruptive somatic mutations of TRAF3 were the most common alterations (4/12, 33%), followed by somatic mutations of TNFAIP3 (3/12, 25%), and NOTCH1 (3/12, 25%). The present study confirms that genes involved in activation of NF-κB-signaling pathways are a major driver in oncogenesis of H. pylori eradication-resistant gastric marginal zone lymphoma and revealed that TRAF3 mutation is a major contributor in MALT1 rearrangement-negative gastric marginal zone lymphoma.
Similar content being viewed by others
Introduction
Extranodal marginal zone lymphoma of mucosa-associated lymphoid tissue is a heterogeneous group of low-grade mature B-cell lymphomas that harbor various genetic alterations. The development of this type of tumor is closely associated with chronic microbial infection or autoimmune disease, or both, suggesting a role for antigenic stimulation in their pathogenesis [1]. Helicobacter pylori infection has the most important causative role in the development of gastric extranodal marginal zone lymphoma of mucosa-associated lymphoid tissue [2]. Most tumors are clinically indolent and respond well to therapies to eradicate H. pylori [3]. However, a small proportion of these lymphomas may show a refractory clinical course, which may be associated with acquired genetic alterations [4]. A broad range of common and inherent genetic changes in extranodal marginal zone lymphoma that regulate the nuclear factor (NF)-κB and NOTCH signaling pathways, which are known to be critical to the development and function of marginal zone B cells, have been widely reported [1]. Specific chromosomal translocations including t(1;14)(p22;q32)/BCL10-IGH, t(14;18)(q32;q21)/IGH-MALT1 and t(11;18)(q21;q21)/BIRC3(API2)-MALT1 are major genetic alterations associated with gastric marginal zone lymphoma. These translocations are capable of oncogenic activity by activating both canonical and noncanonical NF-κB pathways [5]. The gene for tumor necrosis factor alpha (TNF α)-inducible protein 3 (TNFAIP3), also known as A20, is a target for the canonical NF-κB pathway, which is expressed by activation of NF-κB, but has negative feedback suppressing various steps of the NF-κB activation pathway. Methylation, deletion, or inactivation mutations of TNFAIP3 result in uncontrolled activation of the NF-κB pathway [6], which shows infrequent or a lack of the chromosome translocation described above [7]. The proliferation of translocation-negative gastric marginal zone lymphoma is largely driven by H. pylori-generated immune responses, which include activation of toll-like receptors and B-cell receptors by pathogen-associated lipopolysaccharides and autoantigens, respectively, in addition to bystander T-cell help via CD40L and B cell activation factor of the tumor necrosis factor family. CD40 activation induces the recruitment of adapter proteins known as tumor necrosis factor receptor-associated factors (TRAFs) to the cytoplasmic domain of CD40 [8]. The TRAF family contains six members designated TRAF1 through TRAF6. In B cells, TRAF3 negatively regulates canonical and noncanonical NF-κB pathways, as well as Jun N-terminal kinase signaling [8]. Deletions and loss of function mutations of TRAF3 frequently occur in human B-cell chronic lymphocytic leukemia, mantle cell lymphoma, splenic marginal zone lymphoma, multiple myeloma, Waldenström’s macroglobulinemia, and Hodgkin lymphoma [9]. However, the TRAF3 mutation has not been reported in gastric marginal zone lymphoma.
NOTCH1/2 is known to recurrently have influence by activating mutations in 10–25% of splenic marginal zone lymphoma and other types of low-grade mature B-cell neoplasms. Genetic alteration of NOTCH2 has been reported mainly associated with splenic marginal zone lymphoma [10], whereas targeted sequencing has identified recurrent mutations of NOTCH1 in 6–8% of ocular adnexal marginal zone lymphoma [11, 12]. To our knowledge, genetic alterations involving NOTCH1/2 in gastric marginal zone lymphoma have not yet been reported or characterized.
H. pylori-positive gastric marginal zone lymphoma seems to have oncogenic cooperation between immunological stimulation and genetic alterations. In this study, we examined the genomic alterations in H. pylori-positive gastric marginal zone lymphoma that were resistant to H. pylori eradication therapy.
Materials and methods
Patients and samples
A total of 534 cases of gastric marginal zone lymphoma from 2007 to 2015 were retrieved retrospectively from the computerized records system at Samsung Medical Center, Seoul, Korea. Among them, 325 cases were H. pylori-positive gastric marginal zone lymphoma with a history of eradication therapy, 205 cases were H. pylori-negative gastric marginal zone lymphoma, and there were four cases of consultation from another hospital that could not be reviewed for H. pylori infection status because there were no slides or samples from which to review the histologic findings. Among cases of H. pylori-positive gastric marginal zone lymphoma, there were 29 cases (29/325, 9%) resistant to H. pylori eradication therapy alone and ultimately 19 cases with available formalin-fixed and paraffin-embedded tumor tissue. Two pathologists (Hyeon and Ko) reviewed the pathological findings of all cases to reaffirm a diagnosis as a gastric marginal zone lymphoma according to the recent WHO classification criteria [13] and to determine a pathologic grade after post-eradication therapy of each case according to the Groupe d’Etude des Lymphomes de l’Adulte histological scoring system [14]. Post-treatment response in each case was defined based on the European Gastro-Intestinal Lymphoma Study consensus report [2].
The study was approved by the Institutional Review Board of Samsung Medical Center (IRB File No. 2013-12-076-005) and all 19 patients had provided written informed consent for their tissue to be used in the study. All investigations were conducted according to the principles expressed in the Declaration of Helsinki and its contemporary amendments.
Fluorescence in situ hybridization analysis for MALT1 and BCL10
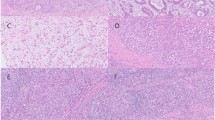
Interphase fluorescence in situ hybridization (FISH) analysis was performed in all cases using a Vysis LSI MALT1 Dual Color Break Apart Rearrangement Probe (Abbott/Vysis, Des Plaines, IL, USA) for detection of MALT1 rearrangement and Empire Genomics BCL10 Dual-Color Break Apart Rearrangement Probe (Empire Genomics, Buffalo, NY, USA) for detection of BCL10 rearrangement. At least 250 nuclei per sample were evaluated under a fluorescence microscope (Zeiss Axioskop, Oberkochen, Germany) using filter sets recommended by Vysis (DAPI/Spectrum Orange dual bandpass, DAPI/Spectrum Green dual bandpass). All overlapping nuclei were excluded. In normal cells, two fused or colocalized hybridization signals of LSI MALT1 are seen, whereas in the case of MALT1-associated translocation, one fused, one red, and one green signal are observed. The determined cutoff value for the detection of a rearrangement of MALT1 and BCL10 was 5%. After analyzing the FISH results, cases were divided into subgroups according to MALT1 and BCL10 rearrangement.
DNA extraction from formalin-fixed and paraffin-embedded tumor samples
All available hematoxylin and eosin-stained slides from formalin-fixed and paraffin-embedded H. pylori eradication-resistant gastric marginal zone lymphoma were reviewed. Tumor DNA (n = 19) was extracted using a QIAamp DNA Mini kit (Qiagen, Valencia, CA, USA), according to the manufacturer’s protocol. DNA quality and quantity were analyzed using a Nanodrop 8000 UV–Vis spectrometer (NanoDrop Technologies), Qubit 2.0 Fluorometer (Life Technologies), and 2200 TapeStation Instrument (Agilent Technologies, Santa Clara, CA, USA) (Supplementary Table 1).
Somatic variant detection in targeted sequencing data
Targeted sequencing was performed using HemaSCAN. This panel contains the whole exome of 425 genes related to hematological malignancies (Supplementary Table 2). Genomic DNA was sheared using a Covaris S220 (Covaris, Woburn, MA, USA). Target capture was performed using the SureSelect XT Reagent Kit, HSQ (Agilent Technologies) and a paired-end sequencing library was constructed with a barcode. Sequencing was performed on a HiSeq 2500 with 100-bp reads (Illumina, San Diego, CA, USA). The paired-end reads were aligned to the human reference genome (hg19) using BWA-MEM v0.7.5, Samtools v0.1.18, GATK v3.1-1, and Picard v1.93. The mean coverage across all samples was 703 after deduplication, ranging from 350.1 to 906.2 (Supplementary Table 3). We called single-nucleotide variants using MuTect version 1.1.4, Lowfreq version 0.6.1, and VarDict version 1.06 software with a variant allele frequency of ≥1% or the number of variant supporting reads of > 4. We filtered out sequencing errors using a machine learning algorithm with features extracted from SAM files. By this algorithm, we could increase specificity of the results [15]. We manually reviewed variants with supporting reads <20 using an Integrative Genomics Viewer browser and filtered out sequencing errors. We identified small insertions and deletions using Pindel version 0.2.5a4 with the number of variant supporting reads of > 9. We further filtered out variants present with a minor allele frequency of ≥1% in the 1000 Genomes Project database (http://www.internationalgenome.org/), the Exome Aggregation Consortium database (http://exac.broadinstitute.org/), the National Heart, Lung, and Blood Institute’s Exome Sequencing Project database (https://esp.gs.washington.edu/drupal/), the Korean Reference Genome Database (http://152.99.75.168/KRGDB/), the Korean Variant Archive (https://kobic.re.kr/kova/), or an in-house database from 192 Korean individuals. We selected missense variants predicted to have a functional consequence by Mendelian Clinically Applicable Pathogenicity score [16] with the author’s recommended threshold. Copy number alterations and tumor purity were called using a custom-built in-house algorithm. Genes with a copy number >6 were selected as amplification, and genes with a copy number <1.2 were selected as deletion. Structure variants and large indels were identified using a custom-built algorithm with >20 variant supporting reads.
Sanger sequencing for TRAF3, NOTCH1, CARD11, and TNFAIP3 mutations in tumor and matched normal samples
We performed Sanger sequencing of matched normal tissue to demonstrate whether TRAF3, NOTCH1, CARD11, or TNFAIP3 mutations found in targeted sequencing were somatic alterations. We used paired bone marrow specimens from patients or biopsy specimens obtained from another organ with no histopathological alteration as matched normal samples. Amplification was performed in a total volume of 25 μL containing 1 μL of DNA from formalin-fixed and paraffin-embedded tissue using AmpliTaq Gold Master mix (Thermo Fisher Scientific, Waltham, MA, USA). Samples were denatured for 10 min at 95 °C, followed by 35 cycles of 30 s at 95 °C, 30 s at 56 or 59 °C, and 1 min at 72 °C, with a final elongation of 7 min at 72 °C. The primer sequences and annealing temperature are shown in Supplementary Table 4. For Sanger sequencing, the PCR products were purified and analyzed using an ABI3730XL DNA analyzer to validate the presence of each mutation.
Mutation detection by droplet digital polymerase chain reaction
We performed droplet digital polymerase chain reaction of tumor that harbors TRAF3, NOTCH1, or TNFAIP3 mutation detected by targeted sequencing, but with negative results by Sanger sequencing because of low variant allele frequency. Simultaneously, paired normal tissue from the patients was also evaluated using droplet digital polymerase chain reaction. Droplet digital polymerase chain reaction assays were performed on a QX200 Droplet Digital PCR System (Bio-Rad, Hercules, CA, USA), consisting of a T100 Thermal Cycler, a QX200 Droplet Generator, and a QX200 Droplet Reader. The PCR reaction mixture (20 μL) contained 10 μL of droplet digital polymerase chain reaction Supermix (no dUTP) for probes, 2.4 μL of each primer/probe mix (target and reference, labeled with FAM and HEX fluorophores, respectively), 1 μL of HindIII (4 U/μL) restriction enzymes, 3 μL of DNA from formalin-fixed and paraffin-embedded tissue, and 3.6 μL of water. Droplet generation oil (70 μL) was then loaded and the cartridge was placed into the droplet generator. Under vacuum, sample and oil were mixed, generating monodisperse droplets. Thereafter, 40 μL of packed droplets was transferred into a 96-well PCR plate for thermal cycling amplification. The thermal cycling started with 10 min at 95 °C, followed by 45 cycles of 94 °C for 30 s, 58 °C for 30 s, and 98 °C for 10 min. The droplet reader (Bio-Rad) was used for fluorescence signal quantification. Results were analyzed using Quantasoft v.1.7.4 software (Bio-Rad).
Results
Prevalence of the MALT1 and BCL10 rearrangement
The general clinicopathological characteristics of 19 gastric marginal zone lymphomas are presented in Table 1. The cases with MALT1 rearrangement comprised 7 of the 19 (37%) (Fig. 1a), which is consistent with the literature [4, 17]. Because the Break Apart probe was used, the partner gene that rearranges with MALT1 cannot be determined through the FISH test results. From the targeted sequencing data, although the intronic region of MALT1 was not included in the sequencing panel, three BIRC3–MALT1 fusions (Fig. 1b) were detected. These three cases also showed MALT1 translocation on FISH results. The breakpoints of these fusions were located at or near the MALT1 exon. In the remaining four cases, which revealed MALT1 translocation on FISH results, fusions were not observed on targeted sequencing data, presumably because the translocation site was not included in the targeted sequencing region. No BCL10 rearrangement, which only occurs in ~ 4% of patients with gastric marginal zone lymphoma [18], was observed in any case.
MALT1 gene rearrangement. a Fluorescence in situ hybridization result showing rearrangement of MALT1 region. Separate green dot and red dot represent the rearrangement (arrow). Yellow dot represents normal allele. b BIRC3 intron 6 (chr11:102203657) was fused to MALT1 exon 6 (chr18:56377246) in case 17. c BIRC3 intron 6 (chr11:102202770) was fused to MALT1 intron 8 (chr18:56381355) in case 5
Prevalence of the TRAF3 and TNFAIP3 mutations
We called somatic single-nucleotide variants and identified an average of 14 nonsilent substitutions per tumor sample (median, 12; range, 0–26) (Supplementary Table 5). To increase the specificity of excluding mutations that were predicted to be present in normal populations or at low risk of protein structural changes, we again selected single-nucleotide variants with a minor allele frequency of ≥0.1% in normal population data (described under Methods) and that were predicted to have a functional consequence by Mendelian Clinically Applicable Pathogenicity score >0.025 (average of 14 nonsilent substitutions per tumor sample, median, 5; range 0–18) (Supplementary Table 6). To confirm that each mutation was a somatic alteration, we examined the presence of protein-altering variants detected in each tumor in matched normal tissues using conventional Sanger sequencing or droplet digital polymerase chain reaction.
Among the 19 tumor samples, four (21%) carried disruptive alterations in TRAF3 including frameshift deletion, nonsynonymous single nucleotide variants, splicing mutation, and frameshift insertion. One of the four cases with TRAF3 mutations had both a splicing mutation and a frameshift insertion (Table 2). Of the 19 samples, three (16%) harbored somatic TNFAIP3 alterations, which were a frameshift insertion, a frameshift deletion, and a splicing mutation (Table 2). Inactivation of these genes is implicated in the pathogenesis of B cell lymphoma via the activation of the NF-κB pathway [6, 7, 19, 20]. Because we applied targeted sequencing using an NGS platform, subclonal mutations with low variant allele frequency were identified, which might have been missed in the results of conventional Sanger sequencing reported previously. Reflecting this limitation, one frameshift deletion (case 15) and one nonsynonymous single nucleotide variants of TRAF3 (case 12) were identified in the tumor sample with conventional Sanger sequencing, but the other three mutations (case 1 and case 10) with low variant allele frequency ( <5%) were not detected in Sanger sequencing (Fig. 2a and Supplementary Figure 1). We performed droplet digital polymerase chain reaction to detect TRAF3 mutations with low variant allele frequency. One mutation (case 1, splicing mutation in exon3 of TRAF, variant allele frequency: 3.42%) was detected by droplet digital polymerase chain reaction (Fig. 2b); however, droplet digital polymerase chain reaction could not be performed for the other two mutations involving exon10 of TRAF3 (case 1 and case 10) because of failure to design the probe. Each variant of TRAF3 was absent in the matched normal DNA. The Integrative Genomics Viewer images of each mutation in exon10 of TRAF3 are depicted in Supplementary Figure 3A–B.
Results of Sanger sequencing and droplet digital polymerase chain reaction. a Results of Sanger sequencing for TRAF3, TNFAIP3, and CARD11. Two of 5 TRAF3 (case 12, variant allele fraction: 25.41%; case 15, variant allele fraction: 13.15%), two of three TNFAIP3 (case 6, variant allele fraction: 9.62%; case 7, variant allele fraction: 26.71%) and one of two CARD11 (case 6, variant allele fraction: 9.62%) somatic mutations were identified in tumor tissue, but not in normal tissue. Altered reference sequence is marked as a box. b Results of droplet digital polymerase chain reaction for TRAF3 with low variant allele fraction. The two graphs at the top show droplets (event number) versus fluorescence amplitude (blue dots: designation of the droplet populations as positives (mutant type); green dots: designation of the droplet populations as positives (wild type); gray dots: designation of the droplet populations as negatives). The last graph shows sample concentrations (blue bar: designation of the concentration (copies/μL) of mutant-type DNA; green bars: designation of the concentration of wild-type DNA) and fractional abundance. One of three TRAF3 (case 1, variant allele fraction: 3.42% in targeted sequencing data) somatic mutations with low variant allele fraction identified in tumor tissue with 2.0% of fractional abundance, but not in matched normal tissue from small intestine. c Results of droplet digital polymerase chain reaction for TNFAIP3. One TNFAIP3 (case 11, variant allele fraction: 6.78% in targeted sequencing data) somatic mutations with low variant allele fraction identified in tumor tissue with 5.4% of fractional abundance, but not in matched normal tissue from bone marrow. DW: distilled water used for negative control
In the case of TNFAIP3 mutation, frameshift insertion (case 6) and frameshift deletion (case 7) were detected in the tumor sample with Sanger sequencing, but splicing mutation (case 11) with 6.78% of variant allele frequency was not identified by Sanger sequencing (Fig. 2a and Supplementary Figure 1). Then we performed droplet digital polymerase chain reaction of case 11 and confirmed the same TNFAIP3 splicing mutation (Fig. 2c). Another somatic mutation activating the NF-κB pathway was the nonframeshift deletion and insertion of CARD11 as found in one case (5%). Alterations in CARD11 and TNFAIP3 would enhance the activation of the canonical NF-κB pathway, whereas those in BIRC3 and TRAF3 would likely augment the signaling of the noncanonical NF-κB pathway. Taken together, seven of the 19 samples (37%) demonstrated somatic single nucleotide variants or frameshift indels involving activation of the NF-κB pathway.
Mutual exclusivity test results between MALT1 rearrangements and genetic alterations including TRAF3 and TNFAIP3 mutations
Seven MALT1 rearrangements, four TRAF3 mutations, and three TNFAIP3 somatic mutations were mutually exclusive in 19 cases of refractory gastric marginal zone lymphoma (Fig. 3). None of the alterations in these three genes coexisted in the same case. This was significant in a CoMEt (Combinations of Mutually Exclusive Alterations) mutual exclusivity test [21] (p = 0.004).
Heatmap of somatic alterations in gastric marginal zone lymphoma. MALT1, TRAF3, and TNFAIP3 are mutually exclusive. Sixteen out of 19 samples have one or more alterations on the genes related to the NF-κB pathway. Top bar plot and right-side bar plot show total number of mutations in each case and for each gene. Truncation: truncation mutation including frameshift indel, stop gain single nucleotide variants, and splice site mutation. Nontruncation: nontruncation mutation including missense single nucleotide variants and in-frame indel
Prevalence of the NOTCH1 mutation
Somatic alterations in NOTCH1 were found in three of the 19 tumor samples (16%) and included nonsynonymous single nucleotide variants (n = 1) and frameshift deletions (n = 2) (Table 2). Each protein-altering NOTCH1 mutation was confirmed using tumor tissue and matched normal tissue by conventional Sanger sequencing or droplet digital polymerase chain reaction (Fig. 4 and Supplementary Figure 2), except one mutation in exon5 of NOTCH1 (case 10) with very low variant allele frequency of 1.02% (Supplementary Figure 2). An Integrative Genomics Viewer image of a mutation in exon5 of NOTCH1 is depicted in Supplementary Figure 3D.
Results of Sanger sequencing for NOTCH1. Two of 3 NOTCH1 somatic mutations (case 7, variant allele fraction: 9.26%; case 11, variant allele fraction: 12.21%) were identified in tumor tissue, but not in normal tissue. Altered reference sequence is marked as a box. Tumor tissue of case 11 is sequenced on the sense strand. Normal tissue of case 11 and both tissues of case 7 are sequenced on the anti-sense strand
Alterations of other genes
Genes involved in epigenetic modification, such as chromatin modification and transcriptional regulation, were also frequently mutated in gastric marginal zone lymphoma. TBL1XR1, which is essential for transcriptional repression and is known as a mutation often accompanied by ocular adnexal marginal zone lymphoma [11], was a commonly mutated gene in gastric marginal zone lymphoma (n = 3, 16%). In addition, nonsynonymous single nucleotide variants of CIITA, which are involved in immune regulation or evasion and frequently altered in primary mediastinal large B-cell lymphoma, were identified in three cases. The other genes mutated in this epigenetic modifier group included WHSC1 (n = 2), EWSR1 (n = 2), JARID2 (n = 2), KDM5A (n = 2), TET2 (n = 1), and ARID1A (n = 1). In total, eleven of the 19 samples (58%) carried mutations in this group, suggesting that alterations to epigenetic modifiers may also contribute to the mutational characteristics of gastric marginal zone lymphoma (Fig. 3). Several genes involving DNA repair and integrity were mutated. These included a frameshift deletion of TP53 (n = 1) and nonsynonymous single nucleotide variants of ATM (n = 1). Although one case (case 15) was accompanied by a TP53 mutation, it showed the persistence of intractable gastric marginal zone lymphoma without transformation into a diffuse large B-cell lymphoma. Other tumor suppressor genes were mutated as follows: NF1 (n = 3), MSH3 (n = 2), MLH1 (n = 2), BRCA1 (n = 2), BRCA2 (n = 2), BRIP1 (n = 2), APC (n = 2), and RB1 (n = 1). Overall, twelve of the 19 samples (63%) harbored mutations in the tumor suppressor groups (Fig. 3). The specific impact of some of these mutations on lymphomagenesis remains to be investigated.
Discussion
We showed recurrent genetic alterations in multiple genes of gastric marginal zone lymphomas that were resistant to H. pylori eradication therapy. Commonly mutated genes are involved in the NF-κB and NOTCH signaling pathways, and epigenetic modification. The frequency of mutation in each gene varies, and some have been reported previously [5]. The frequency of MALT1 rearrangement (37%) was consistent with that reported in the literature [4, 17, 22]. Although the TRAF3 mutation in gastric marginal zone lymphoma has not been clarified, in the present study, we found that the frequency of TRAF3 mutation was high and mutually exclusive with MALT1 rearrangement.
H. pylori-positive gastric marginal zone lymphoma is causatively linked to chronic inflammation associated with H. pylori infection and seems to involve oncogenic cooperation between immunological stimulation and genetic alterations. It may depend on the interaction between malignant B cells and a microenvironment including infection-associated T cells, B cell helper signals such as CD40L and B-cell activation factor of the tumor necrosis factor family, those from T cells, dendritic cells, and innate lymphoid cells [5, 18]. Several recent studies suggest that the B-cell activation factor of the tumor necrosis factor family signal transduction pathway, which is affected by both immunological stimulation and genetic changes contributes to gastric marginal zone lymphoma without chromosomal translocation [23, 24].
NF-κB is a positive mediator for the development, proliferation, and survival of T cells and B cells [25]. Constitutive activation of NF-κB signaling is a common characteristic of many lymphoid malignancies and has an important role in the neoplastic transformation of mature B cells and the progression of marginal zone lymphoma [18]. In the case of extranodal marginal zone lymphoma of mucosa-associated lymphoid tissue derived from other sites, particularly the ocular adnexa and spleen, somatic mutation appears to be the major genetic event that activates NF-κB, whereas in gastric marginal zone lymphoma, the major event is thought to be chromosomal translocation of NF-κB signaling molecules that leads to antigen-independent lymphoma growth [1, 5, 10, 18, 26].
Kuo et al. [27] published studies of the pathogenesis of H. pylori-independent gastric marginal zone lymphoma [4] and H. pylori-dependent gastric marginal zone lymphoma. In H. pylori-dependent gastric marginal zone lymphoma, expression of cytotoxin-associated gene A, which is known as the most important H. pylori virulence factor [28], was higher than in the H. pylori-independent group. Patients with cytotoxin-associated gene A expression responded to eradication therapy more quickly than those without the expression. In the H. pylori-independent gastric marginal zone lymphoma group, B-cell activation factor of the tumor necrosis factor family expression was significantly higher than in the H. pylori-dependent group. They found that B cell activation factor of the tumor necrosis factor family activated the canonical NF-kB pathway and AKT pathway as well as the noncanonical NF-kB pathway through TRAF3 degradation.
Our results showed that somatic mutations in many genes belonging to different functional groups leading to the activation of NF-κB are also important factors in the lymphomagenesis of refractory gastric marginal zone lymphoma. In particular, TRAF3 and TNFAIP3 appear to be the major driver genes in terms of frequency and known functional impact. TRAF3 is a member of a large and diverse group of receptors used for signaling by receptors of the tumor necrosis factor receptor superfamily, as well as some viral mimics and pattern recognition receptors including Toll-like receptors [9, 29]. TRAF3 is also known to be a tumor suppressor gene and functions as a negative regulator of the noncanonical NF-κB pathway [9]. TRAF3 targets NF-κB-inducing kinase (NIK) for constant ubiquitination and degradation, thus maintaining a low basal level of NIK. Downregulation of TRAF3 causes NIK stabilization and constitutive NF-κB activity through the noncanonical pathway. There is an experimental study that reveals that deletion of TRAF3 in B lymphocytes results in marked peripheral B cell hyperplasia and eventually leads to development of splenic marginal zone lymphoma, at least in mice [30]. In addition, there is a preponderance of mutations affecting TRAF3 in canine and human B-cell lymphoma [20]. In extranodal marginal zone lymphoma, studies to date showed rare somatic mutations of TRAF3 as well as CD79A, CD79B, CARD11, BIRC3, and TNFRSF11A [1], whereas our data showed a relatively high frequency of TRAF3 loss of function mutations or frameshift indels in gastric marginal zone lymphoma. This result is remarkable in that, to our knowledge, it is the first that reveals the frequent mutation of TRAF3 in gastric marginal zone lymphoma, and mutation of TRAF3 is mutually exclusive to other genes involved in NF-κB pathway activation.
TNFAIP3, also known as A20, exerts dual ubiquitin editing functions to downregulate NF-κB signaling and TNF-mediated programmed cell death [31, 32]. Because it can inactivate a number of NF-κB-signaling molecules, under physiological conditions, TNFAIP3/A20 negatively regulates the signaling of several surface receptors, which are related to the activation of the canonical NF-κB activation pathway, including B cell receptors, tumor necrosis factor receptors, Toll-like receptors, and interleukin 1 receptor [33]. TNFAIP3-deleted mice with uncontrolled NF-κB activation show an overproduction of proinflammatory cytokines, multiorgan inflammation, enhanced proliferation and survival, and ultimately development of autoimmune disease [34, 35]. In extranodal marginal zone lymphoma, TNFAIP3/A20 deletion and inactivating mutation are largely seen in various organs including ocular adnexa, salivary glands, and thyroid [6, 7, 11, 26]. Our data suggest that relatively frequent TNFAIP3/A20 mutations are also identified in gastric marginal zone lymphoma that has no MALT1 translocation. Alteration of NOTCH1, which is a well-known oncogenic factor in chronic lymphocytic leukemia, mantle cell lymphoma, and splenic marginal zone lymphoma [1], may play a potential role in the oncogenesis of gastric marginal zone lymphoma when considering the frequency of mutations in the results.
TBL1XR1 overexpression has been observed in various solid tumors [36, 37] and hematopoietic neoplasms [11, 38,39,40,41]. In some types of hematological malignancies, recurrent genetic alteration of TBL1XR1 through chromosome rearrangement and point mutations is reported. TBL1XR1 interferes with glucocorticoid receptor mobilization to chromatin and causes glucocorticoid resistance [42]. We found that TBL1XR1 mutation is also frequently present in gastric marginal zone lymphoma.
In summary, the major genetic alterations associated with gastric marginal zone lymphoma include destructive changes of genes that play a role in the NF-κB signaling pathway, with alterations in genes regulating B-cell differentiation, chromatin modification, and transcriptional regulation. The major driver genes other than MALT1 seem to be TRAF3 and TNFAIP3, which were inactivated by recurrent disruptive somatic mutations and are mutually exclusive to MALT1 rearrangement, along with NOTCH1 and TBL1XR1. In addition to these potential major driver genes, our results expand the list of mutated genes in gastric marginal zone lymphoma with potential contributions to the disease. The functions of these minor mutated genes and their roles in the initiation and progression of gastric marginal zone lymphoma remain to be explored in future studies.
Our study focused on identifying genetic changes that predispose to treatment response in those who harbor H. pylori, but who are refractory to eradication therapy. Therefore, this study only included patients who were positive for H. pylori and did not respond to eradication treatment. We did not evaluate the genetic alteration of H. pylori-eradication-sensitive gastric marginal zone lymphoma or H. pylori-negative gastric marginal zone lymphoma. It would be meaningful to compare the results of this study with those of H. pylori-negative gastric marginal zone lymphoma or H. pylori-eradication-sensitive gastric marginal zone lymphoma in the future.
Although with some limitations, the genomic characterization shown in our data provides new information regarding refractory gastric marginal zone lymphoma biology that may improve our understanding of the underlying pathogenesis, provide better prediction of prognosis with therapeutic effect, and identify adequate therapeutic plans earlier for the few refractory cases of gastric marginal zone lymphoma.
References
Rosenquist R, Bea S, Du MQ, et al. Genetic landscape and deregulated pathways in B-cell lymphoid malignancies. J Intern Med. 2017;282:371–94.
Ruskone-Fourmestraux A, Fischbach W, Aleman BM, et al. EGILS consensus report. Gastric extranodal marginal zone B-cell lymphoma of MALT. Gut. 2011;60:747–58.
Fischbach W, Goebeler-Kolve ME, Dragosics B, et al. Long term outcome of patients with gastric marginal zone B cell lymphoma of mucosa associated lymphoid tissue (MALT) following exclusive Helicobacter pylori eradication therapy: experience from a large prospective series. Gut. 2004;53:34–7.
Liu H, Ye H, Ruskone-Fourmestraux A, et al. T(11;18) is a marker for all stage gastric MALT lymphomas that will not respond to H. pylori eradication. Gastroenterology. 2002;122:1286–94.
Du MQ. MALT lymphoma: Genetic abnormalities, immunological stimulation and molecular mechanism. Best Pract Res Clin Haematol. 2017;30:13–23.
Chanudet E, Huang Y, Ichimura K, et al. A20 is targeted by promoter methylation, deletion and inactivating mutation in MALT lymphoma. Leukemia. 2010;24:483–7.
Chanudet E, Ye H, Ferry J, et al. A20 deletion is associated with copy number gain at the TNFA/B/C locus and occurs preferentially in translocation-negative MALT lymphoma of the ocular adnexa and salivary glands. J Pathol. 2009;217:420–30.
Elgueta R, Benson MJ, de Vries VC, et al. Molecular mechanism and function of CD40/CD40L engagement in the immune system. Immunol Rev. 2009;229:152–72.
Moore CR, Edwards SK, Xie P. Targeting TRAF3 downstream signaling pathways in B cell neoplasms. J Cancer Sci Ther. 2015;7:67–74.
Spina V, Rossi D. Molecular pathogenesis of splenic and nodal marginal zone lymphoma. Best Pract Res Clin Haematol. 2017;30:5–12.
Jung H, Yoo HY, Lee SH, et al. The mutational landscape of ocular marginal zone lymphoma identifies frequent alterations in TNFAIP3 followed by mutations in TBL1XR1 and CREBBP. Oncotarget. 2017;8:17038–49.
Johansson P, Klein-Hitpass L, Grabellus F, et al. Recurrent mutations in NF-κB pathway components, KMT2D, and NOTCH1/2 in ocular adnexal MALT-type marginal zone lymphomas. Oncotarget. 2016;7:62627–39.
Cook JR, Isaacson PG, Chott A, Nakamura S, Müller-Hermelink HK, Harris NL, Swerdlow SH. Extranodal marginal zone lymphoma of mucosa-associated lymphoid tissue (MALT lymphoma). In: WHO Classification of Tumours of Haematopoietic and Lymphoid Tissues. Swerdlow SH, Campo E, Harris NL, et al., Editors. Revised 4th edn. Lyon, International Agency for Research on Cancer, 2017. pp. 259-62.
Copie-Bergman C, Gaulard P, Lavergne-Slove A, et al. Proposal for a new histological grading system for post-treatment evaluation of gastric MALT lymphoma. Gut. 2003;52:1656.
Shin H-T, Choi Y-L, Yun JW, et al. Prevalence and detection of low-allele-fraction variants in clinical cancer samples. Nat Commun. 2017;8:1377.
Jagadeesh KA, Wenger AM, Berger MJ, et al. M-CAP eliminates a majority of variants of uncertain significance in clinical exomes at high sensitivity. Nat Genet. 2016;48:1581–6.
Du MQ, Isaccson PG. Gastric MALT lymphoma: from aetiology to treatment. Lancet Oncol. 2002;3:97–104.
Du MQ. MALT lymphoma: a paradigm of NF-κB dysregulation. Semin Cancer Biol. 2016;39:49–60.
Tusche MW, Ward LA, Vu F, et al. Differential requirement of MALT1 for BAFF-induced outcomes in B cell subsets. J Exp Med. 2009;206:2671–83.
Bushell KR, Kim Y, Chan FC, et al. Genetic inactivation of TRAF3 in canine and human B-cell lymphoma. Blood. 2015;125:999–1005.
Leiserson MD, Wu HT, Vandin F, et al. CoMEt: a statistical approach to identify combinations of mutually exclusive alterations in cancer. Genome Biol. 2015;16:160.
Liu H, Ruskone-Fourmestraux A, Lavergne-Slove A, et al. Resistance of t(11;18) positive gastric mucosa-associated lymphoid tissue lymphoma to Helicobacter pylori eradication therapy. Lancet. 2001;357:39–40.
Craig VJ, Cogliatti SB, Arnold I, et al. B-cell receptor signaling and CD40 ligand-independent T cell help cooperate in Helicobacter-induced MALT lymphomagenesis. Leukemia. 2010;24:1186–96.
Kuo SH, Tsai HJ, Lin CW, et al. The B-cell-activating factor signalling pathway is associated with Helicobacter pylori independence in gastric mucosa-associated lymphoid tissue lymphoma without t(11;18)(q21;q21). J Pathol. 2017;241:420–33.
Jost PJ, Ruland J. Aberrant NF-κB signaling in lymphoma: mechanisms, consequences, and therapeutic implications. Blood. 2007;109:2700–7.
Streubel B, Simonitsch-Klupp I, Mullauer L, et al. Variable frequencies of MALT lymphoma-associated genetic aberrations in MALT lymphomas of different sites. Leukemia. 2004;18:1722–6.
Kuo SH, Chen LT, Lin CW, et al. Expressions of the CagA protein and CagA-signaling molecules predict Helicobacter pylori dependence of early-stage gastric DLBCL. Blood. 2017;129:188–98.
Hatakeyama M. Oncogenic mechanisms of the Helicobacter pylori CagA protein. Nat Rev Cancer. 2004;4:688–94.
Hildebrand JM, Yi Z, Buchta CM, et al. Roles of tumor necrosis factor receptor associated factor 3 (TRAF3) and TRAF5 in immune cell functions. Immunol Rev. 2011;244:55–74.
Moore CR, Liu Y, Shao C, et al. Specific deletion of TRAF3 in B lymphocytes leads to B-lymphoma development in mice. Leukemia. 2012;26:1122–7.
Wertz IE, O’Rourke KM, Zhou H, et al. De-ubiquitination and ubiquitin ligase domains of A20 downregulate NF-κB signalling. Nature. 2004;430:694–9.
Shembade N, Ma A, Harhaj EW. Inhibition of NF-κB signaling by A20 through disruption of ubiquitin enzyme complexes. Science. 2010;327:1135–9.
Catrysse L, Vereecke L, Beyaert R, et al. A20 in inflammation and autoimmunity. Trends Immunol. 2014;35:22–31.
Lee EG, Boone DL, Chai S, et al. Failure to regulate TNF-induced NF-κB and cell death responses in A20-deficient mice. Science. 2000;289:2350–4.
Chu Y, Vahl JC, Kumar D, et al. B cells lacking the tumor suppressor TNFAIP3/A20 display impaired differentiation and hyperactivation and cause inflammation and autoimmunity in aged mice. Blood. 2011;117:2227–36.
McAllister-Lucas LM, Baens M, Lucas PC. MALT1 protease: a new therapeutic target in B lymphoma and beyond? Clin Cancer Res. 2011;17:6623–31.
Chen SP, Yang Q, Wang CJ, et al. Transducin β-like 1 X-linked receptor 1 suppresses cisplatin sensitivity in nasopharyngeal carcinoma via activation of NF-κB pathway. Mol Cancer. 2014;13:195.
Gonzalez-Aguilar A, Idbaih A, Boisselier B, et al. Recurrent mutations of MYD88 and TBL1XR1 in primary central nervous system lymphomas. Clin Cancer Res. 2012;18:5203–11.
Scott DW, Mungall KL, Ben-Neriah S, et al. TBL1XR1/TP63: a novel recurrent gene fusion in B-cell non-Hodgkin lymphoma. Blood. 2012;119:4949–52.
Andersson E, Eldfors S, Edgren H, et al. Novel TBL1XR1, EPHA7 and SLFN12 mutations in a Sezary syndrome patient discovered by whole exome sequencing. Exp Dermatol. 2014;23:366–8.
Kiel MJ, Sahasrabuddhe AA, Rolland DC, et al. Genomic analyses reveal recurrent mutations in epigenetic modifiers and the JAK-STAT pathway in Sezary syndrome. Nat Commun. 2015;6:8470.
Jones CL, Bhatla T, Blum R, et al. Loss of TBL1XR1 disrupts glucocorticoid receptor recruitment to chromatin and results in glucocorticoid resistance in a B-lymphoblastic leukemia model. J Biol Chem. 2014;289:20502–15.
Acknowledgements
This study was supported by a grant from the National Research Foundation of Korea (NRF-2017R1A2B1010739).
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Electronic supplementary material
Rights and permissions
About this article
Cite this article
Hyeon, J., Lee, B., Shin, SH. et al. Targeted deep sequencing of gastric marginal zone lymphoma identified alterations of TRAF3 and TNFAIP3 that were mutually exclusive for MALT1 rearrangement. Mod Pathol 31, 1418–1428 (2018). https://doi.org/10.1038/s41379-018-0064-0
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41379-018-0064-0
This article is cited by
-
Role of upfront autologous stem cell transplantation in patients newly diagnosed with primary CNS lymphoma treated with R-MVP: real-world data from a retrospective single-center analysis
Bone Marrow Transplantation (2022)
-
Genomic landscape of Epstein–Barr virus-positive extranodal marginal zone lymphomas of mucosa-associated lymphoid tissue
Modern Pathology (2022)
-
Mutational landscape of marginal zone B-cell lymphomas of various origin: organotypic alterations and diagnostic potential for assignment of organ origin
Virchows Archiv (2022)
-
Characterization of molecular genetics and clinicopathology in thymic MALT lymphoma
Annals of Hematology (2022)
-
Comprehensive analysis of clinical, pathological, and genomic characteristics of follicular helper T-cell derived lymphomas
Experimental Hematology & Oncology (2021)