Abstract
Hydatidiform mole is an aberrant human pregnancy characterized by early embryonic arrest and excessive trophoblastic proliferation. Recurrent hydatidiform moles are defined by the occurrence of at least two hydatidiform moles in the same patient. Fifty to eighty percent of patients with recurrent hydatidiform moles have biallelic pathogenic variants in NLRP7 or KHDC3L. However, in the remaining patients, the genotypic types of the moles are unknown. We characterized 80 new hydatidiform mole tissues, 57 of which were from patients with no mutations in the known genes, and we reviewed the genotypes of a total of 123 molar tissues. We also reviewed mutation analysis in 113 patients with recurrent hydatidiform moles. While all hydatidiform moles from patients with biallelic NLRP7 or KHDC3L mutations are diploid biparental, we demonstrate that those from patients without mutations are highly heterogeneous and only a small minority of them are diploid biparental (8%). The other mechanisms that were found to recur in patients without mutations are diploid androgenetic monospermic (24%) and triploid dispermic (32%); the remaining hydatidiform moles were misdiagnosed as moles due to errors in the analyses and/or their unusual mechanisms. We compared three parameters of genetic susceptibility in patients with and without mutations and show that patients without mutations are mostly from non-familial cases, have fewer reproductive losses, and more live births. Our data demonstrate that patients with recurrent hydatidiform moles and no mutations in the known genes are, in general, different from those with mutations; they have a milder genetic susceptibility and/or a multifactorial etiology underlying their recurrent hydatidiform moles. Categorizing these patients according to the genotypic types of their recurrent hydatidiform moles may facilitate the identification of novel genes for this entity.
Similar content being viewed by others
Introduction
Hydatidiform mole is an aberrant human pregnancy characterized by abnormal embryonic development and excessive proliferation of the trophoblast. It occurs once in every 600–1000 pregnancies in Western countries but at higher frequencies in Latin America, Africa, the Middle East, and the Far East [1, 2]. Among women with one hydatidiform mole (sporadic hydatidiform mole), about 1–9% develops a second mole (recurrent hydatidiform moles), depending on populations and studies [3,4,5,6,7,8].
At the histopathological level, hydatidiform mole is classified as complete hydatidiform mole or partial hydatidiform mole. Complete hydatidiform moles have marked circumferential trophoblastic proliferation and, in general, lack embryonic tissues, with the exception of extremely rare cases where inner cell derivatives were observed [9, 10]. Partial hydatidiform moles have moderate focal trophoblastic proliferation and may contain embryonic tissues [11]. At the genotypic level, complete hydatidiform moles are mostly diploid androgenetic (both chromosome complements are of paternal origin), while partial hydatidiform moles are mostly triploid dispermic (one maternal chromosome and two paternal chromosome complements). Non-molar miscarriages lack significant trophoblastic proliferation and are mostly diploid biparental (one maternal and one paternal chromosome complements) with or without aneuploidies. As microscopic evaluation of products of conception is mainly descriptive and not always sufficient to classify subgroups of hydatidiform moles and distinguish them from non-molar miscarriages, various methods have been developed to determine the parental contribution to such tissues and establish a more reliable diagnosis that is based on histopathological examination and parental contribution to the product of conception. Among these methods, short tandem repeat genotyping is used to identify the ploidy of the products of conception and the parental contribution to their genomes; immunohistochemistry of p57KIP2 (a paternally imprinted, maternally expressed gene) is used to distinguish diploid androgenetic hydatidiform mole from most other genotypic types due to the lack of p57KIP2 expression in the cytotrophoblast and villous stroma of androgenetic hydatidiform mole; and flow cytometry is used to determine the ploidy of the tissues. Fluorescent in situ hybridization (FISH) is also used for determining the ploidy of the tissues, as well as for s investigating the presence of mosaicisms. Previous studies have shown an important improvement in the diagnosis of hydatidiform mole by using different methods and integrating their results [12, 13].
By studying familial cases of recurrent hydatidiform moles, NLRP7 and KHDC3L were found to be responsible for recurrent hydatidiform moles [14, 15]. To date (including this study), ~70 and 6 pathogenic variants, observed in a recessive state, have been described in NLRP7 (http://fmf.igh.cnrs.fr/ISSAID/infevers/) and KHDC3L, respectively. Recently, we demonstrated a high frequency of Alu-mediated deletions and rearrangements in NLRP7 [16]. All recurrent hydatidiform moles from patients with mutations in these two genes are diploid biparental with the exception of three cases, one reported to be triploid dispermic [17] and two reported to be triploid digynic [18].
In this study, we report ten novel pathogenic variants in NLRP7, two of which mediated by Alu repeats, and review our mutation analysis in NLRP7 and KHDC3L on 113 unrelated patients with at least two hydatidiform moles. We used different approaches to comprehensively characterize the parental contribution to 23 and 57 products of conception from patients with and without recessive mutations in the known genes, respectively. We show that all products of conception from patients with recessive mutations in NLRP7 or KHDC3L are diploid biparental, while those from patients without mutations are highly heterogeneous and only a minority of them are diploid biparental. We compared various parameters of genetic susceptibility between patients with and without mutations and found that a family history of moles is very rare among patients without mutations in the known genes. In addition, these patients have less reproductive losses and more live births. Our data suggest that patients with recurrent hydatidiform moles and no mutations in the known genes have a milder genetic susceptibility to reproductive losses and some of them may have polygenic and/or multifactorial aetiologies underlying their recurrent hydatidiform moles.
Materials and methods
Patients
The study was approved by the McGill Institutional Review Board (IRB# A01-M07-98 03A). Patients with at least two hydatidiform moles (all forms combined) were referred to our laboratory between 2002 and 2017 from various collaborators or recruited at the Quebec Trophoblastic Disease Registry (http://www.rmtq.ca/en/). All patients provided written consent to participate in our study, gave blood samples for mutation analysis, and agreed for us to retrieve their products of conception from various histopathology laboratories for research purposes.
Mutation analysis
NLRP7 and KHDC3L mutation analyses were performed on genomic DNA by PCR amplification of all their exons and Sanger sequencing in both directions, as previously described [19, 20]. PCR conditions and the sequences of the primers used to amplify the junction fragments are provided in Supplementary Table 1. Variant nomenclature for NLRP7 is given according to the following references, NM_001127255.1 (cDNA), NG_008056.1 (genomic), and Q8WX94 (protein). Exon numbering is as in NG_008056.1.
Histopathological review
Morphological evaluation of the products of conception was performed on hematoxylin and eosin-stained tissue sections independently by at least two experts in the histology of molar pregnancies (JA, RK, and RS) according to standard criteria [11].
Parental contribution to the molar tissues
p57KIP2 immunohistochemistry
p57KIP2 immunohistochemistry was performed on 4-µm sections of formalin-fixed paraffin-embedded tissues, as previously described [21]. For each product of conception, the p57KIP2 immunostaining result was interpreted as negative when maternal decidua and/or extravillous trophoblastic cells, which serve as internal positive control, exhibited nuclear p57KIP2 staining but villous stromal and cytotrophoblast cells did not exhibit staining. The result was interpreted as positive when villous stromal and cytotrophoblast cells showed nuclear staining of p57KIP2.
Flow cytometry
Flow cytometry was performed on formalin fixed paraffin embedded tissues that were prepared according to standard methods. Cellular preparation for flow cytometry was performed according to a Hedley’s protocol [22], as previously described [23]. Briefly, two sections of 60 µm were cut from each formalin fixed paraffin embedded block, deparaffinized with xylene, and gradually rehydrated. The proteins were digested in 1 ml of 5 mg/ml pepsin (Sigma-Aldrich, St Louis, USA) in 0.9% NaCl (adjusted to pH 1.5 with HCl). The cellular suspension was then suspended in propidium iodide solution (0.1 mg/µl, Sigma-Aldrich) and 50 µl RNase (1 mg/ml) and incubated at 37 °C for 30 min. Finally, they were filtered through a 48 µm mesh nylon filter and analyzed using a BD FACS Canto II at the Immunophenotyping Core Facility of the McGill University Health Centre Research Institute. Data files were analyzed using FCSalyzer (Wien, Austria).
Microsatellite DNA genotyping
On the basis of the quantity of chorionic villi in the formalin fixed paraffin embedded blocks, 5–12 serial 10 µm sections were prepared from the blocks with the largest amount of chorionic villi that are separated from maternal tissues. The sections were mounted on slides and stained with hematoxylin and eosin (H&E). Under a stereomicroscope, chorionic villi were collected from the slides using Kimwipes and forceps and used for DNA extraction using the QIAamp DNA formalin fixed paraffin embedded Tissue Kit (Qiagen, Hilden, Germany). Extracted DNA was quantified using a Nanodrop and loaded on an agarose gel to evaluate its quality and the required amount for multiplex fluorescent microsatellite genotyping with the PowerPlex 16 HS System (Promega, Corporation, Fitchburg, Wisconsin, USA). The reaction consists of short tandem repeat multiplex PCR assay that amplifies DNA at 15 different short tandem repeat loci and a fragment from the Amelogenin gene. DNA from the product of conception and their available parents was amplified, and the PCR products were resolved by capillary electrophoresis using an Applied Biosystems 3730xl DNA Analyzer (Applied Biosystems, Foster City, CA, USA) at the Centre for Applied Genomics (http://www.tcag.ca). The data were analyzed with PeakScanner, version 1.0 (Applied Biosystems, Foster City, CA, USA) and the product of conception alleles were compared to the parental alleles to determine their origin.
Fluorescent in situ hybridization
Fluorescent in situ hybridisation (FISH) was performed on 4 μm sections, which were hybridised systematically with centromeric probes from chromosomes, X, Y and 18, as previously described [24]. On some tissues, other probes were also used. At least 100 cells for each product of conception, from different microscopic fields, were scored with each probe.
Microarrays analysis
Microarray analysis was performed on blood DNA from patients with no identified mutations in either NLRP7 or KHDC3L to search for large deletions or rearrangements using Cytoscan HD (Affymetrix, Santa Carla, CA, USA). We note that the Cytoscan HD microarray contains 32 markers in the genomic sequence of NLRP7, of which 6 are single nucleotide polymorphisms (SNP); most of these markers are located in the region spanning from intron 4 to exon 11. Consequently, this microarray is not sensitive to detect NLRP7 deletion upstream of intron 4. For KHDC3L, because of its small genomic size (1495-bp, from the first nucleotide of exon 1 to the last nucleotide of exon 3), there are only two markers from its genomic sequence on the Cytoscan HD microarray and only one of them is a SNP marker. Consequently, the Cytoscan HD does not allow reaching a conclusion on the presence of deletions or rearrangements in KHDC3L. Another SNP microarray platform was performed at Combimatrix to search for aneuploidies in formalin fixed paraffin embedded tissues from one product of conception, as previously described [25].
Search for deletions by quantitative real time PCR (qPCR) and multiplex ligation-dependent probe amplification
qPCR on genomic DNA from patient 1566 was performed using Quantifast SYBR-green PCR kit (Qiagen, Toronto, ON, Canada). Each sample was checked in duplicates using the Bio-Rad Miniopticon Real Time PCR system (Bio-Rad Laboratories, Mississauga, ON, Canada) and analyzed by the Opticon Monitor software (Bio-Rad Laboratories). The comparative CT method (ΔΔCT method) was used for relative quantification, and data was normalized against an endogenous control primer that amplifies exon 11 of NLRP7, for which the two alleles are amplified based on the presence of a heterozygous SNP. For the other samples, patient 1 and her family members and patient 6, multiplex ligation-dependent probe amplification was performed, as previously described [26].
Results
Ten novel pathogenic variants in NLRP7
To date, our laboratory has performed NLRP7 mutation analysis on a total of 113 unrelated patients with at least 2 hydatidiform moles. This analysis revealed NLRP7 pathogenic variants observed in a recessive state in 62 out of 113 unrelated patients (55%). Many of these variants were previously reported by our group [14, 16, 27,28,29,30,31] and others [32,33,34] and ten are novel. The novel variants are a missense, three stop codon gain, four small deletions, an insertion of Alu Yb element, and an Alu-mediated large deletion removing 9287-bp from the promoter region. All these variants are listed in Table 1 and their chromatograms are shown in Fig. 1 and Supplementary Figures S1A and S1B. The missense variant affects a conserved amino acid and is predicted to be pathogenic (polyphen score = 0.998); eight variants lead to protein truncations and one removes part of the promoter region and is predicted to affect NLRP7 transcription. Consequently, these 10 mutations are most likely pathogenic.
Location of the Alu insertion in exon 4 of NLPR7. a PCR amplification of genomic DNA from exon 4 followed by gel electrophoresis analysis revealed a second larger fragment by approximately 350-bp in patient 1590 and her father but not in her mother who carries a missense variant p.(Arg693Trp) in a heterozygous state. b A schematic of the region of exon 4 where the Alu Yb8 element is inserted between nucleotides c.1548 and c.1565 and results in a 18-bp duplication (blue box) at the site of the insertion. The red arrow indicates the 5′ to 3′ orientation of the inserted Alu Yb8 element that begins at genomic position g.106 and ends at g.419 in the poly (A) tract (poly (T) in the reverse complementary strand) of reference sequence AF15169.2. c Chromatograms showing the 5′ and 3′ junctions of the 18-bp duplication flanking the inserted Alu Yb8 element. d Long-range PCR amplification showing the amplification of an abnormal genomic DNA fragment of 7135-bp overlapping the deletion in patient 1566, her mother, and maternal grandmother, but not in three control subjects. The promoter region predicted by the “Eukaryotic Promoter Database” (http://epd.vital-it.ch/) is indicated above exon 1 and starts 499-bp upstream of exon 1 and ends 63-bp downstream of exon 1. “GM” stands for Grandmother. e. Schematic of the deletion that is mediated by recombination between two Alu Y elements and a microhomology of 23-bp shown in capital letters
Mutation analysis of patient 1590 was initially performed by Whole Exome Sequencing by the referring laboratory and only revealed a previously reported missense pathogenic variant, c.2077 C > T, p.(Arg693Trp), in a heterozygous state. Exome sequencing analysis had shown a significant low coverage in exon 4 with high number of recurrent soft-clipped reads (sequences that did not align with the reference sequence) that displayed significant similarities with Alu sequences when compared to databases. Because the patient had three recurrent hydatidiform moles and such phenotype is considered to be severe in our judgment and is associated with recessive mutations, we repeated the search for NLRP7 mutations by PCR amplification of genomic DNA followed by Sanger sequencing. Analysis of the amplified fragments by gel electrophoresis revealed an abnormal amplicon with primers located in exon 4 that is approximately 350-bp larger than the normal fragment in the patient and in her father (Fig. 1). Gel extraction and sequencing of the abnormal fragment showed that it contains an Alu Yb8 element inserted in exon 4 at position c.1548 and identified a duplication of 18-bp at the site of the Alu Yb8 insertion, c. [1548_1566dup;1566_1567insAF15169.2: g.106_419] (Fig. 1).
In patient 1566, only a stop gain variant c.2227 G > T, p.(Glu743*), was found in a heterozygous state in exon 6. Because the patient had five recurrent hydatidiform moles and again such phenotype is in our judgment severe, we suspected the presence of a deletion in the other allele. Cytoscan HD microarray did not reveal any deletion (Supplementary Figure S2) but since this microarray does not cover the region upstream of intron 4 of NLRP7, we performed quantitative PCR (qPCR) with three amplicons located in the region of the suspected deletion. The result of this analysis with two amplicons suggested the presence of hemizygosity (Supplementary Figure S3). We next sequenced NLRP7 amplicons containing common SNP, intronic, and promoter regions not covered by our standard NLRP7 mutation analysis in this patient and in her mother and grandmother. We also used long range PCR to amplify various large fragments covering the suspected deleted region. The results of these analyses led to the identification of a large deletion of 9287-bp that starts 6950-bp upstream of exon 1 and ends 2300-bp downstream of exon 1, that we define at the nucleotide level to c.-7026_-40 + 2300del (Fig. 1 and Supplementary table 1). This deletion removes the entire NLRP7 promoter that is predicted by the Eukaryote Promoter Database (http://epd.vital-it.ch/), which spans from 499-bp upstream of exon 1 to 63-bp downstream of exon 1. This variant is therefore most likely pathogenic and is the first regulatory mutation to be described in NLRP7.
From the aforementioned analysis, 51 patients were found negative for recessive NLRP7 pathogenic variants and were then screened for KHDC3L mutations. This analysis revealed recessive KHDC3L pathogenic variants in 6 out of the 51 patients (12%), all of which were previously reported [16, 35]. In conclusion, among the 113 unrelated patients with at least 2 hydatidiform moles, recessive KHDC3L pathogenic variants accounted for 5% of patients with at least 2 hydatidiform moles, which is in agreement with previous findings [15, 18]
Comprehensive analysis of the parental contribution to the molar tissues
Strategy of the analysis
To better understand the pathology of recurrent hydatidiform moles in patients with no mutations in the known genes, we made extensive efforts to retrieve archived formalin fixed paraffin embedded tissues from their products of conception from various hospitals. From most products of conception, formalin fixed paraffin embedded blocks were retrieved, which gave us the possibility to use various methods to determine the parental contribution to their genomes. From others, only sections on microscopic slides were available, which limited our investigation. All products of conception referred to us as molar pregnancies were systematically assessed by morphology by two experts in the histopathology of hydatidiform mole, and by flow cytometry, p57KIP2 immunohistochemistry, and microsatellite DNA genotyping. The results of the various methods were compared and in cases of discrepancies, the experiments were repeated and/or FISH was used to clarify discrepancies. On several triploid dispermic partial hydatidiform moles, FISH was also used to investigate the presence of mosaicisms. On most tissues, probes that detect chromosomes X, Y, and 18 were used. Additional probes from other chromosomes were also used on some products of conception to clarify discrepancies between our results and those sent to us from the referring laboratories or to clarify distortions between the heights of alleles observed with the microsatellite genotyping.
Hydatidiform moles from patients with recessive pathogenic variants in NLRP7 or KHDC3L are all diploid biparental
We previously reported the parental contribution to the genotypes of 41 molar tissues, 36 from patients with recessive pathogenic variants in NLRP7 [29] and 5 from patients with recessive pathogenic variants in KHDC3L [20] and found them all diploid biparental. Other groups obtained similar results on most of their patients with only three exceptions reported to date [17, 18]. In an effort to better understand the mechanisms leading to mole formation in patients with mutations in either gene and better understand the functions of these genes, we extended our genotypic analysis to 23 additional hydatidiform mole tissues, 22 from patients with recessive pathogenic variants in NLRP7 and one from a patient with a recessive pathogenic variant in KHDC3L. Our analysis demonstrated that all 23 products of conception are diploid biparental.
In conclusion, all 64 products of conception from patients with recessive mutations in the two known genes analyzed by our laboratory to date were found diploid biparental.
Hydatidiform moles from patients with no recessive mutations in NLRP7 or KHDC3L are highly heterogeneous and a minority of them are diploid biparental
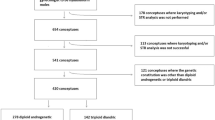
From the aforementioned analysis, a total of 45 unrelated patients did not have any variant in a recessive state in NLRP7 or KHDC3L that are believed to be pathogenic. To better understand the aetiologies of their recurrent hydatidiform moles, extensive efforts were done and allowed for the retrieval of 57 “hydatidiform mole” tissues from 25 patients that we comprehensively genotyped using various approaches. A summary of the results is recapitulated in Fig. 2 and the detailed results with the various approaches are provided in Supplementary Table 2. To our surprise, only a minority of patients (8%) were found to have diploid biparental hydatidiform moles. Six other patients (24%) had recurrent diploid androgenetic monospermic complete hydatidiform moles, 8 patients (32%) had recurrent triploid dispermic partial hydatidiform moles, and 3 patients had, each, only one hydatidiform moles available for analysis. The remaining 6 patients did not have recurrent hydatidiform moles because one or two of their “hydatidiform mole” were revised to non-molar miscarriages. Below is a summary of our analyses and results.
Summary of the comprehensive characterization of 57 hydatidiform mole tissues from 25 patients with no recessive pathogenic variants in NLRP7 or KHDC3L. Recapitulation of the characterization of the referred hydatidiform mole tissues with the various results obtained by morphological analysis, flow cytometry, karyotype analysis, and fluorescent in situ hybridization, p57KIP2 immunohistochemistry, microsatellite genotyping, and the conclusion of the analyses. Unrelated patients are numbered from 1 to 25, and only one of them has a half-sister with recurrent hydatidiform moles. The number of analyzed tissues for each patient is provided under the “Histopathology” column. More than one tissue of the same type are indicated by their numbers; lack of a number indicates one such tissue or item. In the other columns, results on the different tissues are listed by their chronological order. POC stands for product of conception; NM, for non-molar; MC, miscarriage; CHM, complete hydatidiform mole; PHM, partial hydatidiform mole; n.a., indicates result not available and is used to indicate an inconclusive result or not available samples; Dip, diploid; Bip, biparental; Dis, dispermic; Tri, triploid.
Patients with recurrent hydatidiform moles
Two unrelated patients with diploid biparental complete hydatidiform moles
Patient 1 has a half-sister, who also had recurrent hydatidiform moles. Six products of conceptions from these two sisters were re-examined by us and five fulfilled the morphological criteria of complete hydatidiform mole; all were evaluated for p57KIP2 expression, two were found positive, two negative, and two with negative and positive cells; three complete hydatidiform moles, from patient 1’s sister, were genotyped and found diploid biparental. The full description and the characterization of these tissues are described by Scurry et al (in preparation). From patient 2, three products of conception were analyzed, all fulfilled the criteria of complete hydatidiform mole; two were evaluated for p57KIP2 expression, one was found positive and one was found negative; and two were genotyped and found diploid biparental. These are the only patients out of 25 (without mutations in the two known genes) and with recurrent diploid biparental hydatidiform moles (8%).
Because (1) these patients were found to have diploid biparental complete hydatidiform moles, (2) most patients with diploid biparental complete hydatidiform moles are found to have NLRP7 recessive mutations, and (3) NLRP7 has a high frequency of Alu-mediated deletions [16], we performed microarray analysis using Cytoscan HD (Supplementary Figure 2) and multiplex ligation-dependent probe amplification for NLRP7 (Supplementary Figure S3) on blood DNA from patient 1, her half-sister and father, and on patient 2. However, we did not detect any deletion or rearrangement in NLRP7 or KHDC3L. These three patients are the only ones in our cohort of 113 patients with recurrent hydatidiform moles, for which we found evidence for diploid biparental complete hydatidiform moles but without mutations in the known genes. Another atypical finding in these two patients is that both of them had complete hydatidiform moles that expressed p57KIP2 in cytotrophoblast and villous stroma cells, which we have never seen in any other sample.
Six patients with diploid androgenetic complete hydatidiform moles
Six patients, 3–8, each had 2–4 available hydatidiform mole tissues and their analysis revealed their diploid androgenetic monospermic origin. The morphological evaluation of all these tissues fulfilled the criteria of complete hydatidiform mole and all analyzed tissues were negative for p57KIP2 expression.
Eight patients with triploid dispermic partial hydatidiform moles
Eight patients, 9–16, each had two triploid dispermic conceptions. Morphological evaluation confirmed the diagnosis of partial hydatidiform mole with the exception of one product of conception (from patient 15) that did not fulfill the morphological criteria for a partial hydatidiform mole and was diagnosed as a non-molar miscarriage. Indeed, the triploidy of this case was discovered incidentally by flow cytometry while analyzing the partial hydatidiform mole from this patient. Patient 10 had, in addition to her two partial hydatidiform moles, one complete hydatidiform mole that was confirmed by microscopic morphological evaluation and found to be diploid androgenetic monospermic by microsatellite DNA genotyping.
Three patients with various molar genotypes and only one available tissue
Patients 17–19 each had two hydatidiform moles but we were able to access or reach a conclusive result on only one of the two hydatidiform mole. This group consisted of patient 17 with one triploid dispermic partial hydatidiform mole and one complete hydatidiform mole that is diploid by karyotype analysis but no tissues were available for p57KIP2 and genotyping from the complete hydatidiform mole; patient 18 had one diploid dispermic complete hydatidiform mole, and patient 19 with one androgenetic monospermic complete hydatidiform mole.
Patients with “hydatidiform mole” revised to nonmolar miscarriages
Four patients with diploid biparental non-molar miscarriages
Patients 20–23 were each referred with a history of two hydatidiform mole conceptions; however, after morphological evaluation, the hydatidiform moles from these patients were revised to non-molar miscarriages (Fig. 2). These products of conception did have some, but not all the morphological features of hydatidiform mole and mainly lacked significant trophoblastic proliferation. p57KIP2 immunohistochemistry on these products of conception revealed that three of them are positive for p57KIP2 expression, which is in agreement with their diagnosis as miscarriages; two products of conception were not available for p57KIP2 expression analysis; one product of conception was inconclusive, and 2 products of conception from patient 23 were negative for p57KIP2 expression.
The absence of p57KIP2 expression in two products of conception from patient 23 was in contradiction with their histopathological diagnosis as a non-molar miscarriage mainly due to the lack of trophoblastic proliferation (Fig. 3a–d). Microsatellite genotyping on the first product of conception showed the presence of a non-maternal, 173-bp, and the lack of the maternal allele, 176-bp, at marker TH01 located on 11p15. However, this product of conception had the maternal allele, 232-bp, on a proximal marker, D11S1983, located on 11q12–13 suggesting a partial deletion of chromosome 11 (Fig. 3c). Microarray analysis at Combimatrix on DNA from formalin fixed paraffin embedded tissues from this product of conception revealed a terminal microduplication of 3.9-Mb on 4p16.3 and a microdeletion of 3.6-Mb on 11p15.5-p15.4 (NCBI build GRCh37/hg, Feb. 2009) (Fig. 3f) that removes CDKN1C, the gene coding for p57KIP2. The microarray data in combination with those of microsatellite genotyping demonstrate that the 3.6-Mb deletion is on the maternal chromosome 11 and explains the lack of p57KIP2 expression in this product of conception. The second product of conception from this patient, which was also diploid biparental by microsatellite genotyping, was uninformative for marker TH01, which showed a single allele of 176-bp that could be in a hemi- or homozygous state. Since the partner's DNA was not available, no conclusion could be reached on marker TH01 in the second product of conception and there were not enough tissues from this product of conception for microarray analysis. Consequently, we could not further investigate why this product of conception is negative for p57KIP2 expression.
Histopathology and genotyping data of the two non-molar miscarriages from patient 4. Microphotograhs of hematoxilin and eosin-stained sections of the 1st (a) and 2nd (b) product of conception (POC) showing that chorionic villi (some of which are indicated by “CV”) lack trophoblastic proliferation. c and d Microphotographs of p57KIP2 immunostaining on the 1st (c) and 2nd (d) product of conception showing negative staining of the cytotrophoblast (arrows) and villous mesenchyme (asterisks) nuclei, in contrast to the positive staining of the endometrial nuclei (arrowheads), which serve as an internal positive control. e Deletion of the distal end of chromosome 11 is shown by the loss of the maternal allele (176-bp) in the 1st product of conception at TH01 marker located on 11p15. Marker D11S1983, located on 11q12.1, shows the presence of one maternal (232-bp) and one non-maternal (225-bp) alleles, demonstrating the diploid biparental origin of alleles at this locus in the 1st product of conception. f Combimatrix microarray results on the 1st product of conception demonstrating a duplication of 3.9-Mb on chromosome 4p16.3 and a deletion of 3.6-Mb on 11p15.5-p15.4
Two patients with tetraploid non-molar miscarriages
Two patients, 24 and 25, were referred to us because they each had 2 hydatidiform moles, but after comprehensive analysis, only one product of conception from each patient was found triploid dispermic and fulfilled morphological criteria of partial hydatidiform mole. The two other products of conception did not fulfill morphological criteria of hydatidiform mole and were therefore diagnosed as non-molar miscarriage. In patient 24, the tetraploid genome was not detected by microsatellite genotyping since the tissues had two identical maternal and two identical paternal genomes (XXYY) and displayed a normal diploid biparental genotype. Because the microsatellite genotyping result was in contradiction with a medical report indicating a trisomy for chromosome 17, obtained by FISH on tissue sections by the referring laboratory, we performed FISH on tissue sections with probes from chromosomes X, Y, 10, 11, and 17. Our results showed a tetraploid genotype with XXYY in more than 100 analyzed cells. In patient 25, microsatellite genotyping did not lead to a conclusive result on her first product of conception because of the presence of several non-maternal alleles and the absence of DNA from the first partner of the patient with whom she had the tetraploid conception. FISH on this tissue failed to reach a conclusion because of a technical reason (the provided slides by the referring laboratory were not positively charged). However, karyotype analysis from the referring laboratory documented a tetraploid karyotype, 92, XXXY, which most likely does not originate from contamination with maternal cells. We therefore concluded that this tissue is tetraploid.
Comparison of the genetic susceptibility for recurrent hydatidiform moles between patients with and without recessive mutations in NLRP7 or KHDC3L
Less familial cases of recurrent hydatidiform moles among patients without recessive mutations in the known genes
Our laboratory has been analyzing patients for mutations in NLRP7 and KHDC3L genes since their identification in 2006 [14] and 2011 [15], respectively. Our consistent observation over these years has always been that most patients from familial cases of recurrent hydatidiform moles are found to have recessive mutations mostly in NLRP7 and the remaining in KHDC3L. Among the 113 unrelated patients with recurrent hydatidiform moles that we analyzed, 68 had recessive mutations in either gene and of these, 19 patients were from familial cases and had other relatives with recurrent hydatidiform moles. However, among a total of 45 unrelated patients with no recessive mutations in either gene, only one patient has a half-sister with recurrent hydatidiform moles. The difference in the number of familial cases between patients with and without mutations is statistically significant p = 0.00027 (Fisher exact test, 95% confidence) (Fig. 4a).
Comparison of the genetic susceptibility for recurrent hydatidiform moles between patients with and without recessive mutations in NLRP7 or KHDC3L. a A significantly lower number of familial cases was found in patients without NLRP7 or KHDC3L mutations (p = 0.00029). b A history of <3 pregnancy losses was more frequent in patients without mutations in either gene while a history of 4–6 pregnancy losses or at least 7 pregnancy losses was more frequent in patients with NLRP7 or KHDC3L mutations. c Patients without mutations in either gene had a significantly higher number of live births than patients with mutations (p = 0).
Less reproductive losses and more live births among patients without recessive mutations in the known genes
Another consistent observation is that most patients with a high number of recurrent hydatidiform moles, even those who are not from familial cases, turn out to have recessive mutations mainly in NLRP7 or in KHDC3L. To investigate the relevance of this observation, we compared the number of pregnancy losses in patients with (mutations positive) and without recessive pathogenic variants (mutations negative) (Fig. 4b). In this analysis, we included the patients and their affected relatives (a total of 137 patients) and counted all their reproductive losses. We also combined all reproductive losses (hydatidiform moles and miscarriages) to avoid variations due to histopathological misdiagnosis of hydatidiform mole since not all the reproductive losses were available to us for re-evaluation. Pregnancy losses were divided into three groups based on their frequency, as follows: at least 7 ( ≥ 7) pregnancy losses, 4–6 pregnancy losses, and less or equal to 3 ( ≤ 3) pregnancy losses and their analysis showed that the highest numbers of pregnancy losses, ≥ 7 and 4–6, are more frequent in patients with recessive pathogenic variants in either NLRP7 or KHDC3L, while the lowest number of pregnancy losses ( ≤ 3) was more frequent among patients with no mutations in either gene and this difference was statistically significant (p = 0.01088). In addition, patients with identified recessive pathogenic variants had less live births than patients with no recessive pathogenic variants (8 live births out of a total of 429 pregnancies in patients with mutations versus 28 live births out of 196 in patients with no mutations, respectively) (p-value = 0).
Discussion
In this study, we report the identification of seven novel variants in NLRP7, two of which are mediated by Alu elements, an insertion in exon 4 and a deletion in the promoter regulatory region. Both Alu-mediated variants, the insertion in the coding region and the deletion that only affects the regulatory non-coding region, are described here for the first time in NLRP7. Our data confirm our previous observation on the high frequency of Alu-mediated deletions and rearrangements in NLRP7 [16]. Among the 113 unrelated patients referred with at least two hydatidiform moles analyzed to date by our laboratory, 68 had pathogenic variants in a recessive state, 62 (55%) in NLRP7 and 6 in KHDC3L (5%); the remaining 45 patients did not have any detectable pathogenic variants in a recessive state by the methods utilised (Fig. 5).
A recapitulation of the mutation screening in 113 patients and comprehensive characterization of their hydatidiform mole tissues. Among 113 patients, 55% (62 patients) have NLRP7 and 5% have KHDC3L pathogenic variants in recessive state; the remaining 40% (45) do not have mutations in either gene. In patients with NLRP7 and KHDC3L recessive mutations, all analyzed products of conception (60 and 6, respectively) were found diploid biparental. In patients without mutations in these two genes, the genotypes were highly heterogeneous and diploid biparental moles were found in only a minority of cases (8%). The sizes of the pies are proportional to the number of patients. HM stands for hydatidiform mole; CHM, complete HM; PHM, partial HM; RHM, recurrent hydatidiform mole; NM MC, non-molar miscarriages; POC, product of conception.
We previously reported a comprehensive analysis of the parental contribution to 41 products of conceptions from patients with two defective alleles in NLRP7 [29] or KHDC3L [20] and demonstrated that they are all diploid biparental. In this study, we extended our comprehensive analysis to 23 additional products of conception and again found all of them diploid biparental. Other groups have also reported that molar tissues from such patients are mostly diploid biparental [36]. Deviation from this genotype has thus far been reported only in three hydatidiform mole tissues [17, 18]. However, these three tissues were not comprehensively analyzed with various methods. Based on our data and our extensive experience in genotyping formalin fixed paraffin embedded archived tissues, we believe that no single genotyping method is perfect and all methods have their own limitations, especially when working with formalin fixed paraffin embedded tissues. It was the integration and reconciliation of the results from various methods that revealed several mistakes and inaccuracies of results obtained by some methods. We therefore recommend that any deviation from the diploid biparental genotype, in patients with recessive pathogenic variants in NLRP7 or KHDC3L, be investigated and documented by various methods. This will improve our understanding of the consequences of mutations in the two genes on oogenesis and normal fertilization.
Because recurrent hydatidiform moles may have different genotypes, in an effort to homogenize the category of patients with no mutations and facilitate the identification of novel genes responsible for this entity, we retrieved 57 hydatidiform mole conceptions (as per referral) from 25 patients without recessive pathogenic variants in the known genes. Genotypic analysis of these 57 hydatidiform moles using various approaches demonstrated that recurrent hydatidiform moles from these patients are highly heterogeneous. The three mechanisms that were found to recur in our cohort of 25 patients are diploid biparental in 8%, diploid androgenetic monospermic in 24%, and triploid dispermic in 32%. In 6 patients, one or two of their tissues were misdiagnosed as hydatidiform mole because of two main reasons, (i) errors in the analyses (at the level of histopathology and/or genotyping) and (ii) the presence of very rare mechanisms such as the 3.6-Mb deletion on 11p15 and the tetraploid non-molar miscarriages. In such unusual cases, the use of additional characterization methods helped reaching a correct diagnosis. Interestingly, the breakpoint on 11p15 in our patient is very close to a previously reported deletion in two patients with a molar like-phenotype and deletions of 3.2-Mb and 3.6-Mb from the short arm telomere of chromosome 11 [37], which suggests the presence of repetitive elements in this region that could be mediating these different deletions that remain to be investigated in the future. The presence of a small deletion on 11p15 and duplication on 4p16.3 suggests that the patient may be a carrier for a cryptic balanced reciprocal translocation involving 11p15 and 4p16.3 that may be responsible for her four miscarriages.
In our cohort, the number of triploid dispermic partial hydatidiform moles did not exceed two in any of the eight patients, which is in agreement with a previous observation [38] and suggests a mild genetic susceptibility underlying the genotypic entity of triploid dispermic partial hydatidiform mole. However, androgenetic monospermic complete hydatidiform mole recurred 3–4 times in three out of six patients. In general, recurrent hydatidiform moles in the same patients tended to have the same genotypes with only three exceptions. One patient had 2 triploid dispermic partial hydatidiform moles and 1 complete hydatidiform mole by histopathology and two others had each one tetraploid and one triploid dispermic product of conception.
We compared various parameters and indicators of genetic susceptibility between patients with and without recessive mutations in the two known genes. Our analysis demonstrated that a family history of recurrent hydatidiform moles is very rare among patients without pathogenic variants in the two genes and is significantly associated with the presence of pathogenic variants in NLRP7 or KHDC3L (p = 0.00027). In addition, low numbers of pregnancy losses (hydatidiform moles or miscarriages) and high numbers of live births were significantly associated with patients without recessive pathogenic variants in the known genes (p = 0.01088 and p = 0.0000, respectively).
Altogether, our data demonstrate that recurrent hydatidiform moles from patients without mutations in the known genes are highly heterogeneous and a minority of them (8%) have diploid biparental genomes. Moreover, our data suggest that patients without recessive pathogenic variants in the known genes have a milder genetic susceptibility for recurrent hydatidiform moles. Consequently, a genetic defect segregating in their families may not always manifest as molar pregnancies. One explanation for this observation is that recurrent hydatidiform moles in some of these patients may have polygenic and/or multifactorial aetiologies that may decrease their manifestation in other siblings since only those who inherited variants in several genes may manifest the defect.
In conclusion, patients with no recessive mutations in the known genes are different from those with mutations and have other mechanisms and molecular bases at the origin of their recurrent hydatidiform moles that remain to be elucidated in future studies.
References
Grimes DA. Epidemiology of gestational trophoblastic disease. Am J Obstet Gynecol. 1984;150:309–18.
Savage P, Williams J, Wong SL, et al. The demographics of molar pregnancies in England and Wales from 2000–2009. J Reprod Med. 2010;55:341–5.
Berkowitz RS, Im SS, Bernstein MR, et al. Gestational trophoblastic disease. Subsequent pregnancy outcome, including repeat molar pregnancy. J Reprod Med. 1998;43:81–6.
Boufettal H, Coullin P, Mahdaoui S, et al. Complete hydatiforme mole in Morocco: epidemiological and clinical study. J Gynecol Obstet Biol Reprod. 2011;40:419–29.
Horn LC, Kowalzik J, Bilek K, et al. Clinicopathologic characteristics and subsequent pregnancy outcome in 139 complete hydatidiform moles. Eur J Obstet Gynecol Reprod Biol. 2006;128:10–4.
Kim JH, Park DC, Bae SN, et al. Subsequent reproductive experience after treatment for gestational trophoblastic disease. Gynecol Oncol. 1998;71:108–12.
Kronfol NM, Iliya FA, Hajj SN. Recurrent hydatidiform mole: a report of five cases with review of the literature. J Med Liban. 1969;22:507–20.
Sebire NJ, Fisher RA, Foskett M, et al. Risk of recurrent hydatidiform mole and subsequent pregnancy outcome following complete or partial hydatidiform molar pregnancy. BJOG. 2003;110:22–6.
Baergen RN, Kelly T, McGinniss MJ, et al. Complete hydatidiform mole with a coexistent embryo. Hum Pathol. 1996;27:731–4.
Zaragoza MV, Keep D, Genest DR, et al. Early complete hydatidiform moles contain inner cell mass derivatives. Am J Med Genet. 1997;70:273–7.
Szulman AE, Surti U. The syndromes of hydatidiform mole. II. Morphologic evolution of the complete and partial mole. Am J Obstet Gynecol. 1978;132:20–7.
Gupta M, Vang R, Yemelyanova AV, et al. Diagnostic reproducibility of hydatidiform moles: ancillary techniques (p57 immunohistochemistry and molecular genotyping) improve morphologic diagnosis for both recently trained and experienced gynecologic pathologists. Am J Surg Pathol. 2012;36:1747–60.
Murphy KM, McConnell TG, Hafez MJ, et al. Molecular genotyping of hydatidiform moles: analytic validation of a multiplex short tandem repeat assay. J Mol Diagn. 2009;11:598–605.
Murdoch S, Djuric U, Mazhar B, et al. Mutations in NALP7 cause recurrent hydatidiform moles and reproductive wastage in humans. Nat Genet. 2006;38:300–2.
Parry DA, Logan CV, Hayward BE, et al. Mutations causing familial biparental hydatidiform mole implicate c6orf221 as a possible regulator of genomic imprinting in the human oocyte. Am J Hum Genet. 2011;89:451–8.
Reddy R, Nguyen NM, Sarrabay G, et al. The genomic architecture of NLRP7 is Alu rich and predisposes to disease-associated large deletions. Eur J Hum Genet. 2016;24:1516.
Ulker V, Gurkan H, Tozkir H, et al. Novel NLRP7 mutations in familial recurrent hydatidiform mole: are NLRP7 mutations a risk for recurrent reproductive wastage? Eur J Obstet Gynecol Reprod Biol. 2013;170:188–92.
Fallahian M, Sebire NJ, Savage PM, et al. Mutations in NLRP7 and KHDC3L confer a complete hydatidiform mole phenotype on digynic triploid conceptions. Hum Mutat. 2013;34:301–8.
Qian J, Deveault C, Bagga R, et al. Women heterozygous for NALP7/NLRP7 mutations are at risk for reproductive wastage: report of two novel mutations. Hum Mutat. 2007;28:741.
Reddy R, Akoury E, Phuong Nguyen NM, et al. Report of four new patients with protein-truncating mutations in C6orf221/KHDC3L and colocalization with NLRP7. Eur J Hum Genet. 2013;21:957–64.
Castrillon DH, Sun D, Weremowicz S, et al. Discrimination of complete hydatidiform mole from its mimics by immunohistochemistry of the paternally imprinted gene product p57KIP2. Am J Surg Pathol. 2001;25:1225–30.
Hedley DW. Flow cytometry using paraffin-embedded tissue: five years on. Cytometry. 1989;10:229–41.
Khawajkie Y, Buckett W, Nguyen NMP, et al. Recurrent triploid digynic conceptions and mature ovarian teratomas: Are they different manifestations of the same genetic defect? Genes Chromosomes Cancer. 2017;56:832–40.
Surti U, Hoffner L, Kolthoff M, et al. Persistent gestational trophoblastic disease after an androgenetic/biparental fetal chimera: a case report and review. Int J Gynecol Pathol. 2006;25:366–72.
Sahoo T, Dzidic N, Strecker MN, et al. Comprehensive genetic analysis of pregnancy loss by chromosomal microarrays: outcomes, benefits, and challenges. Genet Med. 2017;19:83–9.
Pallares-Ruiz N, Philibert L, Dumont B, et al. Combined mutation and rearrangement screening by quantitative PCR high-resolution melting: is it relevant for hereditary recurrent Fever genes? PLoS ONE. 2010;5:e14096.
Deveault C, Qian JH, Chebaro W, et al. NLRP7 mutations in women with diploid androgenetic and triploid moles: a proposed mechanism for mole formation. Hum Mol Genet. 2009;18:888–97.
Messaed C, Chebaro W, Di Roberto RB, et al. NLRP7 in the spectrum of reproductive wastage: rare non-synonymous variants confer genetic susceptibility to recurrent reproductive wastage. J Med Genet. 2011;48:540–8.
Nguyen NM, Zhang L, Reddy R, et al. Comprehensive genotype-phenotype correlations between NLRP7 mutations and the balance between embryonic tissue differentiation and trophoblastic proliferation. J Med Genet. 2014;51:623–34.
Qian J, Cheng Q, Murdoch S, et al. The genetics of recurrent hydatidiform moles in China: correlations between NLRP7 mutations, molar genotypes, and reproductive outcomes. Mol Hum Reprod. 2011;17:612–9.
Slim R, Bagga R, Chebaro W, et al. A strong founder effect for two NLRP7 mutations in the Indian population: an intriguing observation. Clin Genet. 2009;76:292–5.
Hayward BE, De Vos M, Talati N, et al. Genetic and epigenetic analysis of recurrent hydatidiform mole. Hum Mutat. 2009;30:E629–39.
Wang M, Dixon PH, Decordova S, et al. Identification of 13 novel NLRP7 mutations in 20 families with recurrent hydatidiform mole; missense mutations cluster in the leucine rich region. J Med Genet. 2009;46:569–55.
Dixon PH, Trongwongsa P, Abu-Hayyah S, et al. Mutations in NLRP7 are associated with diploid biparental hydatidiform moles, but not androgenetic complete moles. J Med Genet. 2012;49:206–11.
Rezaei M, Nguyen NM, Foroughinia L, et al. Two novel mutations in the KHDC3L gene in Asian patients with recurrent hydatidiform mole. Hum Genome Var. 2016;3:16027.
Sebire NJ, Savage PM, Seckl MJ, et al. Histopathological features of biparental complete hydatidiform moles in women with NLRP7 mutations. Placenta. 2013;34:50–6.
Sunde L, Lund H, J Sebire N, et al. Paternal hemizygosity in 11p15 in mole-like conceptuses: two case reports. Medicine. 2015;94:e1776.
Eagles N, Sebire NJ, Short D, et al. Risk of recurrent molar pregnancies following complete and partial hydatidiform moles. Hum Reprod. 2015;30:2055–63.
Acknowledgements
We thank the patients and their families for their cooperation. We thank Sam Dougaparsad from Affymetrix for his kind technical assistance in the analysis of Cytoscan HD; Urvashi Surti and Lori Hoffner for performing FISH on tissue sections; and Judith St-Onge for her help in the primer design for long-range PCR. We acknowledge the use of the McGill University and Génome Québec Innovation Centre for Sanger sequencing. We also thank the Departments of Pathology of the following hospitals and medical centers, Greater Nashville Perinatology (TN, USA), Nashville Fertility Center (TN, USA), INTEGRIS Baptist Medical Center (OK, USA), Wellington Hospital-Genetics Health Service (New Zealand), Essentia Health Duluth (MN, USA), Falls Memorial Hospital (MN, USA), Maisonneuve Rosemont Hospital (QC, Canada), Notre Dame Hospital (QC, Canada), Pathology Queensland Laboratory (Australia), Toowooba Hospital (Australia), Auckland City Hospital (New Zealand), Touro Laboratory (NO, USA), Saint Justine Hospital (QC, Canada), Vaudois University Hospital (Switzerland), Evergreen Hospital Medical Center (Kirkland, USA), Providence Regional Laboratories (Portland, USA), St Mary’s Hospital Center (QC, Canada), Johnston Memorial Hospital (NC, USA), Erlanger Medical Center (TN, USA), Pole Hospitalo-Universitaire Naissance Et Pathologie De La Femme (Montpellier, France), East Tallinn Central Hospital (Estonia), IWK Health Centre (Halifax, Canada), Victoria General Hospital (Nova Scotia, Canada), for providing archived molar tissues.
Funding
NMPN was supported by fellowships from Réseau Québécois en Reproduction, McGill Faculty of Medicine, RI-MUHC Desjardins Studentship in Child Health Research, and CRRD. Yassemine Khawajkie was supported by a CRRD trainee fellowship. RS is supported by the Canadian Institute of Health Research (MOP-86546, POP-122897, and MOP-130364).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Electronic supplementary material
Rights and permissions
About this article
Cite this article
Nguyen, N.M.P., Khawajkie, Y., Mechtouf, N. et al. The genetics of recurrent hydatidiform moles: new insights and lessons from a comprehensive analysis of 113 patients. Mod Pathol 31, 1116–1130 (2018). https://doi.org/10.1038/s41379-018-0031-9
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41379-018-0031-9
This article is cited by
-
Imprinting disorders
Nature Reviews Disease Primers (2023)
-
Practical guidelines of the EOTTD for pathological and genetic diagnosis of hydatidiform moles
Virchows Archiv (2023)
-
Trans-acting genetic variants causing multilocus imprinting disturbance (MLID): common mechanisms and consequences
Clinical Epigenetics (2022)
-
Recurrent Androgenetic Complete Hydatidiform Moles with p57KIP2-Positive in a Chinese Family
Reproductive Sciences (2022)
-
Association of extended culture to blastocyst and pre-malignant gestational trophoblastic disease risk following IVF/ICSI-assisted reproduction cycles: an analysis of large UK national database
Journal of Assisted Reproduction and Genetics (2022)