Abstract
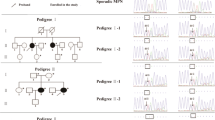
Myeloperoxidase (MPO) gene alterations with variable clinical penetrance have been found in hereditary MPO deficiency, but their leukemia association in patients and carriers has not been established. Germline MPO alterations were found to be significantly enriched in myeloid neoplasms: 28 pathogenic/likely pathogenic variants were identified in 100 patients. The most common alterations were c.2031-2 A > C, R569W, M519fs* and Y173C accounting for about half of the cases. While functional experiments showed that the marrow stem cell pool of Mpo−/− mice was not increased, using competitive repopulation demonstrated that Mpo−/− grafts gained growth advantage over MPO wild type cells. This finding also correlated with increased clonogenic potential after serial replating in the setting of H2O2-induced oxidative stress. Furthermore, we demonstrated that H2O2-induced DNA damage and activation of error-prone DNA repair may result in secondary genetic damage potentially predisposing to leukemia leukemic evolution. In conclusion, our study for the first time demonstrates that germline MPO variants may constitute risk alleles for MN evolution.

This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Horwitz MS, Corey SJ, Grimes HL, Tidwell T. ELANE mutations in cyclic and severe congenital neutropenia: genetics and pathophysiology. Hematol Oncol Clin North Am. 2013;27:19–41.
Li ST, Wang J, Wei R, Shi R, Adema V, Nagata Y, et al. Rare germline variant contributions to myeloid malignancy susceptibility. Leukemia. 2020;34:1675–8.
Morishita K, Kubota N, Asano S, Kaziro Y, Nagata S. Molecular cloning and characterization of cDNA for human myeloperoxidase. J Biol Chem. 1987;262:3844–51.
Klebanoff SJ. Myeloperoxidase: friend and foe. J Leukoc Biol. 2005;77:598–625.
Nauseef WM. Myeloperoxidase in human neutrophil host defence. Cell Microbiol. 2014;16:1146–55.
Mantovani A, Cassatella MA, Costantini C, Jaillon S. Neutrophils in the activation and regulation of innate and adaptive immunity. Nat Rev Immunol. 2011;11:519–31.
Pahwa R, Modi P, Jialal I Myeloperoxidase Deficiency. StatPearls. StatPearls Publishing Copyright © 2021, StatPearls Publishing LLC.: Treasure Island (FL), 2021.
Marchetti C, Patriarca P, Solero GP, Baralle FE, Romano M. Genetic studies on myeloperoxidase deficiency in Italy. Jpn J Infect Dis. 2004;57:S10–12.
Endo D, Saito T, Umeki Y, Suzuki K, Aratani Y. Myeloperoxidase negatively regulates the expression of proinflammatory cytokines and chemokines by zymosan-induced mouse neutrophils. Inflamm Res. 2016;65:151–9.
Milla C, Yang S, Cornfield DN, Brennan ML, Hazen SL, Panoskaltsis-Mortari A, et al. Myeloperoxidase deficiency enhances inflammation after allogeneic marrow transplantation. Am J Physiol Lung Cell Mol Physiol. 2004;287:L706–714.
Kremserova S, Perecko T, Soucek K, Klinke A, Baldus S, Eiserich JP, et al. Lung Neutrophilia in Myeloperoxidase Deficient Mice during the Course of Acute Pulmonary Inflammation. Oxid Med Cell Longev. 2016;2016:5219056.
Nicholls SJ, Hazen SL. Myeloperoxidase and cardiovascular disease. Arterioscler Thromb Vasc Biol. 2005;25:1102–11.
Strzepa A, Pritchard KA, Dittel BN. Myeloperoxidase: A new player in autoimmunity. Cell Immunol. 2017;317:1–8.
Malle E, Buch T, Grone HJ. Myeloperoxidase in kidney disease. Kidney Int. 2003;64:1956–67.
Kutter D, Devaquet P, Vanderstocken G, Paulus JM, Marchal V, Gothot A. Consequences of total and subtotal myeloperoxidase deficiency: risk or benefit? Acta Haematol. 2000;104:10–15.
Pfeilstöcker M, Tuechler H, Sanz G, Schanz J, Garcia-Manero G, Solé F, et al. Time-dependent changes in mortality and transformation risk in MDS. Blood. 2016;128:902–10.
Makishima H, Yoshida K, Nguyen N, Przychodzen B, Sanada M, Okuno Y, et al. Somatic SETBP1 mutations in myeloid malignancies. Nat Genet. 2013;45:942–6.
Nagata Y, Makishima H, Kerr CM, Przychodzen BP, Aly M, Goyal A, et al. Invariant patterns of clonal succession determine specific clinical features of myelodysplastic syndromes. Nat Commun. 2019;10:5386.
Hirsch CM, Nazha A, Kneen K, Abazeed ME, Meggendorfer M, Przychodzen BP, et al. Consequences of mutant TET2 on clonality and subclonal hierarchy. Leukemia. 2018;32:1751–61.
Makishima H, Yoshizato T, Yoshida K, Sekeres MA, Radivoyevitch T, Suzuki H, et al. Dynamics of clonal evolution in myelodysplastic syndromes. Nat Genet. 2017;49:204–12.
Wang K, Li M, Hakonarson H. ANNOVAR: functional annotation of genetic variants from high-throughput sequencing data. Nucleic Acids Res. 2010;38:e164.
Sanger F, Nicklen S, Coulson AR. DNA sequencing with chain-terminating inhibitors. Proc Natl Acad Sci USA. 1977;74:5463–7.
Nauseef WM, Cogley M, Bock S, Petrides PE. Pattern of inheritance in hereditary myeloperoxidase deficiency associated with the R569W missense mutation. J Leukoc Biol. 1998;63:264–9.
DeLeo FR, Goedken M, McCormick SJ, Nauseef WM. A novel form of hereditary myeloperoxidase deficiency linked to endoplasmic reticulum/proteasome degradation. J Clin Invest. 1998;101:2900–9.
Romano M, Dri P, Da Dalt L, Patriarca P, Baralle FE. Biochemical and molecular characterization of hereditary myeloperoxidase deficiency. Blood. 1997;90:4126–34.
Lanza F, Fietta A, Spisani S, Castoldi GL, Traniello S. Does a relationship exist between neutrophil myeloperoxidase deficiency and the occurrence of neoplasms? J Clin Lab Immunol. 1987;22:175–80.
Cappelletti P, Lippi U. Hereditary myeloperoxidase deficiency: a rare condition? Diagnostic possibilities of a differential white cell autoanalyzer (Hemalog-D). Haematologica. 1983;68:736–41.
Lanza F. Clinical manifestation of myeloperoxidase deficiency. J Mol Med (Berl). 1998;76:676–81.
Calado RT, Regal JA, Hills M, Yewdell WT, Dalmazzo LF, Zago MA, et al. Constitutional hypomorphic telomerase mutations in patients with acute myeloid leukemia. Proc Natl Acad Sci USA. 2009;106:1187–92.
Awan A, Malcolm Taylor G, Gokhale DA, Dearden SP. Increased Frequency of Fanconi Anemia Group C Genetic Variants in Children With Sporadic Acute Myeloid Leukemia. Blood. 1998;91:4813–4.
Berwick M, Satagopan JM, Ben-Porat L, Carlson A, Mah K, Henry R, et al. Genetic heterogeneity among Fanconi anemia heterozygotes and risk of cancer. Cancer Res. 2007;67:9591–6.
Rao S, Yao Y, Soares de Brito J, Yao Q, Shen AH, Watkinson RE, et al. Dissecting ELANE neutropenia pathogenicity by human HSC gene editing. Cell Stem Cell. 2021;28:833–45. 2021/05/06/e835
Germeshausen M, Deerberg S, Peter Y, Reimer C, Kratz CP, Ballmaier M. The spectrum of ELANE mutations and their implications in severe congenital and cyclic neutropenia. Hum Mutat. 2013;34:905–14.
Rydzynska Z, Pawlik B, Krzyzanowski D, Mlynarski W, Madzio J. Neutrophil Elastase Defects in Congenital Neutropenia. Front. Immunol. 2021;12:1–12.
Skokowa J, Steinemann D, Katsman-Kuipers JE, Zeidler C, Klimenkova O, Klimiankou M, et al. Cooperativity of RUNX1 and CSF3R mutations in severe congenital neutropenia: a unique pathway in myeloid leukemogenesis. Blood. 2014;123:2229–37.
Germeshausen M, Ballmaier M, Welte K. Incidence of CSF3R mutations in severe congenital neutropenia and relevance for leukemogenesis: Results of a long-term survey. Blood. 2007;109:93–99.
Kettle AJ, Winterbourn CC. Myeloperoxidase: a key regulator of neutrophil oxidant production. Redox Rep. 1997;3:3–15.
Rosales C, Lowell CA, Schnoor M, Uribe-Querol E. Neutrophils: Their role in innate and adaptive immunity 2017. J Immunol Res. 2017;2017:9748345.
Burdon RH. Superoxide and hydrogen peroxide in relation to mammalian cell proliferation. Free Radic Biol Med. 1995;18:775–94.
Bae YS, Kang SW, Seo MS, Baines IC, Tekle E, Chock PB, et al. Epidermal growth factor (EGF)-induced generation of hydrogen peroxide. Role in EGF receptor-mediated tyrosine phosphorylation. J Biol Chem. 1997;272:217–21.
Hansberg W, Aguirre J. Hyperoxidant states cause microbial cell differentiation by cell isolation from dioxygen. J Theor Biol. 1990;142:201–21.
Arnold RS, Shi J, Murad E, Whalen AM, Sun CQ, Polavarapu R, et al. Hydrogen peroxide mediates the cell growth and transformation caused by the mitogenic oxidase Nox1. Proc Natl Acad Sci USA. 2001;98:5550–5.
Buttke TM, Sandstrom PA. Oxidative stress as a mediator of apoptosis. Immunol Today. 1994;15:7–10.
Forman HJ, Bernardo A, Davies KJ. What is the concentration of hydrogen peroxide in blood and plasma? Arch Biochem Biophys. 2016;603:48–53.
Sies H. Hydrogen peroxide as a central redox signaling molecule in physiological oxidative stress: Oxidative eustress. Redox Biol. 2017;11:613–9.
Valverde M, Lozano-Salgado J, Fortini P, Rodriguez-Sastre MA, Rojas E, Dogliotti E. Hydrogen Peroxide-Induced DNA Damage and Repair through the Differentiation of Human Adipose-Derived Mesenchymal Stem Cells. Stem Cells Int. 2018;2018:1615497.
Driessens N, Versteyhe S, Ghaddhab C, Burniat A, De Deken X, Van Sande J, et al. Hydrogen peroxide induces DNA single- and double-strand breaks in thyroid cells and is therefore a potential mutagen for this organ. Endocr Relat Cancer. 2009;16:845–56.
McDonald RJ, Pan LC, St George JA, Hyde DM, Ducore JM. Hydrogen peroxide induces DNA single strand breaks in respiratory epithelial cells. Inflammation. 1993;17:715–22.
Acknowledgements
This work was supported in parts by grants from NIH (R35HL135795 and RO1 HL132071 to JPM and R01 CA257544-01A1 to BKJ). We thank the Vera and Joseph Dresner Foundation (to VV) and the American Italian Cancer Foundation (to CG).
Author information
Authors and Affiliations
Contributions
SK, designed research studies, performed experiments, acquired data, analyzed data, and wrote the draft manuscript. LT., MC performed flow cytometry analysis, collected clinical specimens, and performed Sanger sequencing; VA analyzed RNA sequencing; WW processed raw data of RNA-sequencing; ST analyzed raw data of genomic sequencing; HA collected clinical data and interpreted sequencing data; CG provided and interpreted clinical data and wrote the manuscript; SP provided insights on statistical analysis and data interpretation and wrote the manuscript. YG, MH, TD, contributed with reagents and discussed the design. VV, helped, designed and supervised mouse experiments, data analysis, discussed the results, and wrote manuscript. HJR, reviewed bone marrow histopathology. TL help and perform data analysis. DL and YP helped with mouse experiments and edited the manuscript. MM, TH provided genomic data. BKJ conceived and conceptualize the idea, designed, and supervised the research, acquired, and discussed the results. JPM conceived and conceptualization, read and edited the manuscript and generated resources. All authors read and edited the manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
About this article
Cite this article
Kongkiatkamon, S., Terkawi, L., Guan, Y. et al. Rare germline alterations of myeloperoxidase predispose to myeloid neoplasms. Leukemia 36, 2086–2096 (2022). https://doi.org/10.1038/s41375-022-01630-0
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41375-022-01630-0