Abstract
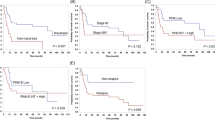
Extra-nodal NK/T-cell lymphoma, nasal type (ENKTCL) is a highly aggressive Epstein-Barr virus associated lymphoma, typically presenting in the nasal and paranasal areas. We assembled a large series of ENKTCL (n = 209) for comprehensive genomic analysis and correlative clinical study. The International Lymphoma Prognostic Index (IPI), site of disease, stage, lymphadenopathy, and hepatomegaly were associated with overall survival. Genetic analysis revealed frequent oncogenic activation of the JAK/STAT3 pathway and alterations in tumor suppressor genes (TSGs) and genes associated with epigenomic regulation. Integrated genomic analysis including recurrent mutations and genomic copy number alterations using consensus clustering identified seven distinct genetic clusters that were associated with different clinical outcomes, thus constituting previously unrecognized risk groups. The genetic profiles of ENTKCLs from Asian and Hispanic ethnic groups showed striking similarity, indicating shared pathogenetic mechanism and tumor evolution. Interestingly, we discovered a novel functional cooperation between activating STAT3 mutations and loss of the TSG, PRDM1, in promoting NK-cell growth and survival. This study provides a genetic roadmap for further analysis and facilitates investigation of actionable therapeutic opportunities in this aggressive lymphoma.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Data availability
The NGS data are deposited in the National Center for Biotechnology Information (NCBI) database of Genotypes and Phenotypes (dbGaP) under the accession number phs002925.v1.p1.
References
Yang Y, Cao JZ, Lan SM, Wu JX, Wu T, Zhu SY, et al. Association of improved locoregional control with prolonged survival in early-stage extranodal nasal-type natural killer/T-cell lymphoma. JAMA Oncol. 2017;3:83–91.
Li X, Cui Y, Sun Z, Zhang L, Li L, Wang X, et al. DDGP versus SMILE in newly diagnosed advanced natural killer/T-cell lymphoma: a randomized controlled, multicenter, open-label study in China. Clin Cancer Res. 2016;22:5223–8.
Vose J, Armitage J, Weisenburger D, International TCLP. International peripheral T-cell and natural killer/T-cell lymphoma study: pathology findings and clinical outcomes. J Clin Oncol. 2008;26:4124–30.
Chiang AK, Tao Q, Srivastava G, Ho FC. Nasal NK- and T-cell lymphomas share the same type of Epstein-Barr virus latency as nasopharyngeal carcinoma and Hodgkin’s disease. Int J Cancer. 1996;68:285–90.
Ho FC, Srivastava G, Loke SL, Fu KH, Leung BP, Liang R, et al. Presence of Epstein-Barr virus DNA in nasal lymphomas of B and ‘T’ cell type. Hematological Oncol. 1990;8:271–81.
Au WY, Weisenburger DD, Intragumtornchai T, Nakamura S, Kim WS, Sng I, et al. Clinical differences between nasal and extranasal natural killer/T-cell lymphoma: a study of 136 cases from the International Peripheral T-Cell Lymphoma Project. Blood. 2009;113:3931–7.
Iqbal J, Weisenburger DD, Chowdhury A, Tsai MY, Srivastava G, Greiner TC, et al. Natural killer cell lymphoma shares strikingly similar molecular features with a group of non-hepatosplenic gammadelta T-cell lymphoma and is highly sensitive to a novel aurora kinase A inhibitor in vitro. Leukemia. 2011;25:348–58.
Huang Y, de Reynies A, de Leval L, Ghazi B, Martin-Garcia N, Travert M, et al. Gene expression profiling identifies emerging oncogenic pathways operating in extranodal NK/T-cell lymphoma, nasal type. Blood. 2010;115:1226–37.
Ko Y, Ree H, Kim W, Choi W, Moon W, Kim SW. Clinicopathologic and genotypic study of extranodal nasal-type natural killer/T-cell lymphoma and natural killer precursor lymphoma among Koreans. Cancer. 2000;89:2106–16.
Jhuang JY, Chang ST, Weng SF, Pan ST, Chu PY, Hsieh PP, et al. Extranodal natural killer/T-cell lymphoma, nasal type in Taiwan: a relatively higher frequency of T-cell lineage and poor survival for extranasal tumors. Hum Pathol. 2015;46:313–21.
Li SY, Feng XL, Li T, Zhang S, Zuo Z, Lin P, et al. Extranodal NK/T-cell lymphoma, nasal type a report of 73 cases at MD Anderson Cancer Center. Am J Surgical Pathol. 2013;37:14–23.
Pongpruttipan T, Sukpanichnant S, Assanasen T, Wannakrairot P, Boonsakan P, Kanoksil W, et al. Extranodal NK/T-cell lymphoma, nasal type, includes cases of natural killer cell and αβ, γδ, and αβ/γδ T-cell origin: a comprehensive clinicopathologic and phenotypic study. Am J Surg Pathol. 2012;36:481–99.
Ng SB, Lai KW, Murugaya S, Lee KM, Loong SL, Fook-Chong S, et al. Nasal-type extranodal natural killer/T-cell lymphomas: a clinicopathologic and genotypic study of 42 cases in Singapore. Mod Pathol. 2004;17:1097–107.
Tse E, Kwong YL. Diagnosis and management of extranodal NK/T cell lymphoma nasal type. Expert Rev Hematol. 2016;9:861–71.
Au WY, Ma SY, Chim CS, Choy C, Loong F, Lie AK, et al. Clinicopathologic features and treatment outcome of mature T-cell and natural killer-cell lymphomas diagnosed according to the World Health Organization classification scheme: a single center experience of 10 years. Ann Oncol. 2005;16:206–14.
Coppo P, Gouilleux-Gruart V, Huang Y, Bouhlal H, Bouamar H, Bouchet S, et al. STAT3 transcription factor is constitutively activated and is oncogenic in nasal-type NK/T-cell lymphoma. Leukemia. 2009;23:1667–78.
Ng SB, Selvarajan V, Huang G, Zhou J, Feldman AL, Law M, et al. Activated oncogenic pathways and therapeutic targets in extranodal nasal-type NK/T cell lymphoma revealed by gene expression profiling. J Pathol. 2011;223:496–510.
Iqbal J, Kucuk C, Deleeuw RJ, Srivastava G, Tam W, Geng H, et al. Genomic analyses reveal global functional alterations that promote tumor growth and novel tumor suppressor genes in natural killer-cell malignancies. Leukemia. 2009;23:1139–51.
Kucuk C, Iqbal J, Hu X, Gaulard P, De Leval L, Srivastava G, et al. PRDM1 is a tumor suppressor gene in natural killer cell malignancies. Proc Natl Acad Sci USA. 2011;108:20119–24.
Kucuk C, Jiang B, Hu X, Zhang W, Chan JK, Xiao W, et al. Activating mutations of STAT5B and STAT3 in lymphomas derived from gammadelta-T or NK cells. Nat Commun. 2015;6:6025.
Jiang L, Gu ZH, Yan ZX, Zhao X, Xie YY, Zhang ZG, et al. Exome sequencing identifies somatic mutations of DDX3X in natural killer/T-cell lymphoma. Nat Genet. 2015;47:1061–6.
Lee S, Park HY, Kang SY, Kim SJ, Hwang J, Lee S, et al. Genetic alterations of JAK/STAT cascade and histone modification in extranodal NK/T-cell lymphoma nasal type. Oncotarget. 2015;6:17764–76.
Song TL, Nairismagi ML, Laurensia Y, Lim JQ, Tan J, Li ZM, et al. Oncogenic activation of the STAT3 pathway drives PD-L1 expression in natural killer/T-cell lymphoma. Blood. 2018;132:1146–58.
Kwong YL, Chan TSY, Tan D, Kim SJ, Poon LM, Mow B, et al. PD1 blockade with pembrolizumab is highly effective in relapsed or refractory NK/T-cell lymphoma failing l-asparaginase. Blood. 2017;129:2437–42.
Iqbal J, Naushad H, Bi C, Yu J, Bouska A, Rohr J, et al. Genomic signatures in B-cell lymphoma: How can these improve precision in diagnosis and inform prognosis? Blood Rev. 2016;30:73–88.
Iqbal J, Wilcox R, Naushad H, Rohr J, Heavican TB, Wang C, et al. Genomic signatures in T-cell lymphoma: How can these improve precision in diagnosis and inform prognosis? Blood Rev. 2016;30:89–100.
Reddy A, Zhang J, Davis NS, Moffitt AB, Love CL, Waldrop A, et al. Genetic and functional drivers of diffuse large B cell lymphoma. Cell 2017;171:481–94.
Dobashi A, Tsuyama N, Asaka R, Togashi Y, Ueda K, Sakata S, et al. Frequent BCOR aberrations in extranodal NK/T-Cell lymphoma, nasal type. Genes Chromosomes Cancer. 2016;55:460–71.
Koo GC, Tan SY, Tang T, Poon SL, Allen GE, Tan L, et al. Janus kinase 3-activating mutations identified in natural killer/T-cell lymphoma. Cancer Discov. 2012;2:591–7.
Kucuk C, Hu X, Jiang B, Klinkebiel D, Geng H, Gong Q, et al. Global promoter methylation analysis reveals novel candidate tumor suppressor genes in natural killer cell lymphoma. Clin Cancer Res. 2015;21:1699–711.
Sim SH, Kim S, Kim TM, Jeon YK, Nam SJ, Ahn YO, et al. Novel JAK3-activating mutations in extranodal NK/T-cell lymphoma, nasal type. Am J Pathol. 2017;187:980–6.
Xiong J, Cui BW, Wang N, Dai YT, Zhang H, Wang CF, et al. Genomic and transcriptomic characterization of natural killer T cell lymphoma. Cancer Cell. 2020;37:403–19.
Kuilman T, Velds A, Kemper K, Ranzani M, Bombardelli L, Hoogstraat M, et al. CopywriteR: DNA copy number detection from off-target sequence data. Genome Biol. 2015;16:49.
Chapuy B, Stewart C, Dunford AJ, Kim J, Kamburov A, Redd RA, et al. Molecular subtypes of diffuse large B cell lymphoma are associated with distinct pathogenic mechanisms and outcomes. Nat Med. 2018;24:679–90.
Brunet JP, Tamayo P, Golub TR, Mesirov JP. Metagenes and molecular pattern discovery using matrix factorization. Proc Natl Acad Sci USA. 2004;101:4164–9.
Lee J, Suh C, Park YH, Ko YH, Bang SM, Lee JH, et al. Extranodal natural killer T-cell lymphoma, nasal-type: a prognostic model from a retrospective multicenter study. J Clin Oncol. 2006;24:612–8.
Montes-Mojarro IA, Chen BJ, Ramirez-Ibarguen AF, Quezada-Fiallos CM, Perez-Baez WB, Duenas D, et al. Mutational profile and EBV strains of extranodal NK/T-cell lymphoma, nasal type in Latin America. Mod Pathol. 2020;33:781–91.
Wen Z, Zhong Z, Darnell JE Jr. Maximal activation of transcription by Stat1 and Stat3 requires both tyrosine and serine phosphorylation. Cell. 1995;82:241–50.
Kataoka H, Miura Y, Joh T, Seno K, Tada T, Tamaoki T, et al. Alpha-fetoprotein producing gastric cancer lacks transcription factor ATBF1. Oncogene. 2001;20:869–73.
Hurlin PJ, Steingrìmsson E, Copeland NG, Jenkins NA, Eisenman RN. Mga, a dual-specificity transcription factor that interacts with Max and contains a T-domain DNA-binding motif. EMBO J. 1999;18:7019–28.
Davidsson J, Puschmann A, Tedgard U, Bryder D, Nilsson L, Cammenga J. SAMD9 and SAMD9L in inherited predisposition to ataxia, pancytopenia, and myeloid malignancies. Leukemia. 2018;32:1106–15.
Jiang Y, Janku F, Subbiah V, Angelo LS, Naing A, Anderson PM, et al. Germline PTPRD mutations in Ewing sarcoma: biologic and clinical implications. Oncotarget. 2013;4:884–9.
Spina V, Khiabanian H, Messina M, Monti S, Cascione L, Bruscaggin A, et al. The genetics of nodal marginal zone lymphoma. Blood. 2016;128:1362–73.
Faronato M, Nguyen VT, Patten DK, Lombardo Y, Steel JH, Patel N, et al. DMXL2 drives epithelial to mesenchymal transition in hormonal therapy resistant breast cancer through Notch hyper-activation. Oncotarget. 2015;6:22467–79.
Aster JC, Pear WS, Blacklow SC. The varied roles of notch in cancer. Annu Rev Pathol. 2017;12:245–75.
Yan J, Li B, Lin B, Lee PT, Chung TH, Tan J, et al. EZH2 phosphorylation by JAK3 mediates a switch to noncanonical function in natural killer/T-cell lymphoma. Blood. 2016;128:948–58.
Soto-Rifo R, Ohlmann T. The role of the DEAD-box RNA helicase DDX3 in mRNA metabolism. Wiley Interdiscip Rev RNA. 2013;4:369–85.
Yin Y, Stephen CW, Luciani MG, Fahraeus R. p53 Stability and activity is regulated by Mdm2-mediated induction of alternative p53 translation products. Nat Cell Biol. 2002;4:462–7.
Takahara M, Kishibe K, Bandoh N, Nonaka S, Harabuchi Y. P53, N- and K-Ras, and beta-catenin gene mutations and prognostic factors in nasal NK/T-cell lymphoma from Hokkaido, Japan. Hum Pathol. 2004;35:86–95.
Hongyo T, Hoshida Y, Nakatsuka S, Syaifudin M, Kojya S, Yang WI, et al. p53, K-ras, c-kit and beta-catenin gene mutations in sinonasal NK/T-cell lymphoma in Korea and Japan. Oncol Rep. 2005;13:265–71.
Hoshida Y, Hongyo T, Jia X, He Y, Hasui K, Dong Z, et al. Analysis of p53, K-ras, c-kit, and beta-catenin gene mutations in sinonasal NK/T cell lymphoma in northeast district of China. Cancer Sci. 2003;94:297–301.
Kurniawan AN, Hongyo T, Hardjolukito ES, Ham MF, Takakuwa T, Kodariah R, et al. Gene mutation analysis of sinonasal lymphomas in Indonesia. Oncol Rep. 2006;15:1257–63.
Shilatifard A. The COMPASS family of histone H3K4 methylases: mechanisms of regulation in development and disease pathogenesis. Annu Rev Biochem. 2012;81:65–95.
Lohr JG, Stojanov P, Lawrence MS, Auclair D, Chapuy B, Sougnez C, et al. Discovery and prioritization of somatic mutations in diffuse large B-cell lymphoma (DLBCL) by whole-exome sequencing. Proc Natl Acad Sci USA. 2012;109:3879–84.
Okosun J, Bodor C, Wang J, Araf S, Yang CY, Pan C, et al. Integrated genomic analysis identifies recurrent mutations and evolution patterns driving the initiation and progression of follicular lymphoma. Nat Genet. 2014;46:176–81.
Chen BJ, Chapuy B, Jing OY, Sun HH, Roemer MGM, Xu ML, et al. PD-L1 expression is characteristic of a subset of aggressive B-cell lymphomas and virus-associated malignancies. Clin Cancer Res. 2013;19:3462–73.
Kataoka K, Shiraishi Y, Takeda Y, Sakata S, Matsumoto M, Nagano S, et al. Aberrant PD-L1 expression through 3’-UTR disruption in multiple cancers. Nature. 2016;534:402–6.
Dong G, Li Y, Lee L, Liu X, Shi Y, Liu X, et al. Genetic manipulation of primary human natural killer cells to investigate the functional and oncogenic roles of PRDM1. Haematologica. 2021;106:2427–38.
Acknowledgements
Research reported in this publication included work performed in the Flow Cytometry core, the Integrative Genomics Core, and the Gene Editing and Viral Vector Core supported by the National Cancer Institute of the National Institutes of Health under grant number P30CA033572. It was also partly supported by Start-up funds from the City of Hope National Medical Center, the Dr. Norman and Melinda Payson Professorship in Hematologic Cancers and the Toni Stephenson Lymphoma Center; Beijing Natural Science Funding (No. 7132062) and Beijing Golden Bridge Engineer Seed Funding (No. 2014-37) awarded to GD. We thank Drs. Stephen Forman and Jiing Kuan Yee for kindly providing the vector epHIV7. The in vitro experiments in this study utilized cytokines from BRB Preclinical Biologics Repository at NCI.
Author information
Authors and Affiliations
Contributions
GD, XL, and WCC conceived and designed the project and wrote the manuscript; GD, LW, WY, JW, HL, LK, WS, WZ, and WL contributed the Asian ENKTCL cases and collected the clinical information. GD performed the DNA extraction and coordinate sequencing studies. QG, AB, SS, JZ, TM, LW and JI performed data management data extraction and analysis; LC performed statistical analysis. JW and XW did the sequencing experiment. CLM, LQM, and JYS contributed ENKTCL case specimens and clinical information from their institutions. XL and KS performed the in vitro functional study. YS, LK, and YL performed experiments and prepared figures and assisted in manuscript preparation. JI and WCC finalized the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Ethics approval and consent to participate
Written informed consent was obtained from all patients. The institutional review boards (IRBs) of the respective institutions approved this study.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
About this article
Cite this article
Dong, G., Liu, X., Wang, L. et al. Genomic profiling identifies distinct genetic subtypes in extra-nodal natural killer/T-cell lymphoma. Leukemia 36, 2064–2075 (2022). https://doi.org/10.1038/s41375-022-01623-z
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41375-022-01623-z
This article is cited by
-
Full-spectral genome analysis of natural killer/T cell lymphoma highlights impacts of genome instability in driving its progression
Genome Medicine (2024)
-
Genomic features reveal potential benefit of adding anti-PD-1 immunotherapy to treat non-upper aerodigestive tract natural killer/T-cell lymphoma
Leukemia (2024)
-
Updates in the Classification of T-cell Lymphomas and Lymphoproliferative Disorders
Current Hematologic Malignancy Reports (2023)
-
New concepts in EBV-associated B, T, and NK cell lymphoproliferative disorders
Virchows Archiv (2023)
-
Clinical and prognostic role of sarcopenia based on masticatory muscle index on MR images in patients with extranodal natural killer/T cell lymphoma, nasal type
Annals of Hematology (2023)