Abstract
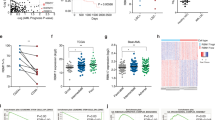
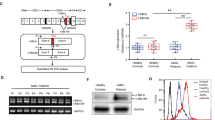
Patients with chronic myeloid leukemia (CML) who are treated with tyrosine kinase inhibitors (TKIs) experience significant heterogeneity regarding depth and speed of responses. Factors intrinsic and extrinsic to CML cells contribute to response heterogeneity and TKI resistance. Among extrinsic factors, cytokine-mediated TKI resistance has been demonstrated in CML progenitors, but the underlying mechanisms remain obscure. Using RNA-sequencing, we identified differentially expressed splicing factors in primary CD34+ chronic phase (CP) CML progenitors and controls. We found SRSF1 expression to be increased as a result of both BCR-ABL1- and cytokine-mediated signaling. SRSF1 overexpression conferred cytokine independence to untransformed hematopoietic cells and impaired imatinib sensitivity in CML cells, while SRSF1 depletion in CD34+ CP CML cells prevented the ability of extrinsic cytokines to decrease imatinib sensitivity. Mechanistically, PRKCH and PLCH1 were upregulated by elevated SRSF1 levels, and contributed to impaired imatinib sensitivity. Importantly, very high SRSF1 levels in the bone marrow of CML patients at presentation correlated with poorer clinical TKI responses. In summary, we find SRSF1 levels to be maintained in CD34+ CP CML progenitors by cytokines despite effective BCR-ABL1 inhibition, and that elevated levels promote impaired imatinib responses. Together, our data support an SRSF1/PRKCH/PLCH1 axis in contributing to cytokine-induced impaired imatinib sensitivity in CML.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout








Similar content being viewed by others
References
Arrigoni E, Del ReM, Galimberti S, Restante G, Rofi E, Crucitta S, et al. Concise review: chronic myeloid leukemia: stem cell niche and response to pharmacologic treatment. Stem Cells Transl Med. 2018;7:305–14.
Mahon F-X, Réa D, Guilhot J, Guilhot F, Huguet F, Nicolini F, et al. Discontinuation of imatinib in patients with chronic myeloid leukaemia who have maintained complete molecular remission for at least 2 years: the prospective, multicentre Stop Imatinib (STIM) trial. Lancet Oncol. 2010;11:1029–35.
Rea D, Nicolini FE, Tulliez M, Guilhot F, Guilhot J, Guerci-Bresler A, et al. Discontinuation of dasatinib or nilotinib in chronic myeloid leukemia: interim analysis of the STOP 2G-TKI study. Blood. 2017;129:846–54.
Copland M, Hamilton A, Elrick LJ, Baird JW, Allan EK, Jordanides N, et al. Dasatinib (BMS-354825) targets an earlier progenitor population than imatinib in primary CML but does not eliminate the quiescent fraction. Blood. 2006;107:4532–9.
Graham SM, Jørgensen HG, Allan E, Pearson C, Alcorn MJ, Richmond L, et al. Primitive, quiescent, Philadelphia-positive stem cells from patients with chronic myeloid leukemia are insensitive to STI571 in vitro. Blood. 2002;99:319–25.
Corbin AS, Agarwal A, Loriaux M, Cortes J, Deininger MW, Druker BJ. Human chronic myeloid leukemia stem cells are insensitive to imatinib despite inhibition of BCR-ABL activity. J Clin Investig. 2011;121:396–409.
Graf L, Iwata M, Torok-Storb B. Gene expression profiling of the functionally distinct human bone marrow stromal cell lines HS-5 and HS-27a. Blood. 2002;100:1509–11.
Ng KP, Manjeri A, Lee KL, Huang W, Tan SY, Chuah CT, et al. Physiologic hypoxia promotes maintenance of CML stem cells despite effective BCR-ABL1 inhibition. Blood, J Am Soc Hematol. 2014;123:3316–26.
Jiang X, Lopez A, Holyoake T, Eaves A, Eaves C. Autocrine production and action of IL-3 and granulocyte colony-stimulating factor in chronic myeloid leukemia. Proc Natl Acad Sci. 1999;96:12804–9.
Traer E, MacKenzie R, Snead J, Agarwal A, Eiring AM, O’Hare T, et al. Blockade of JAK2-mediated extrinsic survival signals restores sensitivity of CML cells to ABL inhibitors. Leukemia. 2012;26:1140.
Bewry NN, Nair RR, Emmons MF, Boulware D, Pinilla-Ibarz J, Hazlehurst LA. Stat3 contributes to resistance toward BCR-ABL inhibitors in a bone marrow microenvironment model of drug resistance. Mol Cancer Ther. 2008;7:3169–75.
Wang Y, Cai D, Brendel C, Barett C, Erben P, Manley PW, et al. Adaptive secretion of granulocyte-macrophage colony-stimulating factor (GM-CSF) mediates imatinib and nilotinib resistance in BCR/ABL+ progenitors via JAK-2/STAT-5 pathway activation. Blood. 2007;109:2147–55.
Chang JS, Santhanam R, Trotta R, Neviani P, Eiring AM, Briercheck E, et al. High levels of the BCR/ABL oncoprotein are required for the MAPK-hnRNP-E2–dependent suppression of C/EBPα-driven myeloid differentiation. Blood. 2007;110:994–1003.
Holm F, Hellqvist E, Mason CN, Ali SA, Delos-Santos N, Barrett CL, et al. Reversion to an embryonic alternative splicing program enhances leukemia stem cell self-renewal. Proc Natl Acad Sci. 2015;112:15444–9.
Salesse S, Dylla SJ, Verfaillie CM. p210BCR/ABL-induced alteration of pre-mRNA splicing in primary human CD34+ hematopoietic progenitor cells. Leukemia. 2004;18:727–33.
Iervolino A, Santilli G, Trotta R, Guerzoni C, Cesi V, Bergamaschi A, et al. hnRNP A1 nucleocytoplasmic shuttling activity is required for normal myelopoiesis and BCR/ABL leukemogenesis. Mol Cell Biol. 2002;22:2255–66.
Das S, Krainer AR. Emerging functions of SRSF1, splicing factor and oncoprotein, in RNA metabolism and cancer. Mol Cancer Res. 2014;12:1195–204.
de Miguel FJ, Sharma RD, Pajares MJ, Montuenga LM, Rubio A, Pio R. Identification of alternative splicing events regulated by the oncogenic factor SRSF1 in lung cancer. Cancer Res. 2014;74:1105–15.
Anczukow O, Rosenberg AZ, Akerman M, Das S, Zhan L, Karni R, et al. The splicing factor SRSF1 regulates apoptosis and proliferation to promote mammary epithelial cell transformation. Nat Struct Mol Biol. 2012;19:220–8.
Sheng J, Zhao J, Xu Q, Wang L, Zhang W, Zhang Y. Bioinformatics analysis of SRSF1‑controlled gene networks in colorectal cancer. Oncol Lett. 2017;14:5393–9.
Karni R, Hippo Y, Lowe SW, Krainer AR. The splicing-factor oncoprotein SF2/ASF activates mTORC1. Proc Natl Acad Sci USA. 2008;105:15323–7.
Shultz JC, Goehe RW, Murudkar CS, Wijesinghe DS, Mayton EK, Massiello A, et al. SRSF1 regulates the alternative splicing of caspase 9 via a novel intronic splicing enhancer affecting the chemotherapeutic sensitivity of non-small cell lung cancer cells. Mol Cancer Res. 2011;9:889–900.
Adesso L, Calabretta S, Barbagallo F, Capurso G, Pilozzi E, Geremia R, et al. Gemcitabine triggers a pro-survival response in pancreatic cancer cells through activation of the MNK2/eIF4E pathway. Oncogene. 2013;32:2848–57.
Gout S, Brambilla E, Boudria A, Drissi R, Lantuejoul S, Gazzeri S, et al. Abnormal expression of the pre-mRNA splicing regulators SRSF1, SRSF2, SRPK1 and SRPK2 in non small cell lung carcinoma. PloS One. 2012;7:e46539.
van der Werf I, Cloos J. Involvement of SRSF1 in alternative splicing of FPGS and methotrexate resistance in children with acute lymphoblastic leukemia. Student Undergrad Res E-J 2015;1.
Zou L, Zhang H, Du C, Liu X, Zhu S, Zhang W, et al. Correlation of SRSF1 and PRMT1 expression with clinical status of pediatric acute lymphoblastic leukemia. J Hematol Oncol. 2012;5:42.
Jiang L, Huang J, Higgs BW, Hu Z, Xiao Z, Yao X, et al. Genomic landscape survey identifies SRSF1 as a key oncodriver in small cell lung cancer. PLoS Genet. 2016;12:e1005895.
Bhatia R, McGlave PB, Dewald GW, Blazar B, Verfaillie C. Abnormal function of the bone marrow microenvironment in chronic myelogenous leukemia: role of malignant stromal macrophages. Blood. 1995;85:3636–45.
Chu S, Holtz M, Gupta M, Bhatia R. BCR/ABL kinase inhibition by imatinib mesylate enhances MAP kinase activity in chronic myelogenous leukemia CD34+ cells. Blood. 2004;103:3167–74.
Anders S, Pyl PT, Huber W. HTSeq—a Python framework to work with high-throughput sequencing data. Bioinformatics. 2015;31:166–9.
Love MI, Huber W, Anders S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 2014;15:550.
Ritchie ME, Phipson B, Wu D, Hu Y, Law CW, Shi W, et al. limma powers differential expression analyses for RNA-sequencing and microarray studies. Nucleic acids Res. 2015;43:e47–e.
Dobin A, Davis CA, Schlesinger F, Drenkow J, Zaleski C, Jha S, et al. STAR: ultrafast universal RNA-seq aligner. Bioinformatics. 2013;29:15–21.
Baccarani M, Deininger MW, Rosti G, Hochhaus A, Soverini S, Apperley JF, et al. European LeukemiaNet recommendations for the management of chronic myeloid leukemia: 2013. Blood. 2013;122:872–84.
Giulietti M, Piva F, D’Antonio M, D’Onorio De Meo P, Paoletti D, Castrignano T, et al. SpliceAid-F: a database of human splicing factors and their RNA-binding sites. Nucleic Acids Res. 2012;41(D1):D125–31.
Skorski T, Kanakaraj P, Nieborowska-Skorska M, Ratajczak M, Wen S-C, Zon G, et al. Phosphatidylinositol-3 kinase activity is regulated by BCR/ABL and is required for the growth of Philadelphia chromosome-positive cells. Blood 1995;86:726–36.
Cortez D, Reuther G, Pendergast AM. The Bcr-Abl tyrosine kinase activates mitogenic signaling pathways and stimulates G1-to-S phase transition in hematopoietic cells. Oncogene. 1997;15:2333.
Tago K, Kaziro Y, Satoh T. Functional involvement of mSos in interleukin-3 and thrombin stimulation of the Ras, mitogen-activated protein kinase pathway in BaF3 murine hematopoietic cells. J Biochem. 1998;123:659–67.
Valacca C, Bonomi S, Buratti E, Pedrotti S, Baralle FE, Sette C, et al. Sam68 regulates EMT through alternative splicing–activated nonsense-mediated mRNA decay of the SF2/ASF proto-oncogene. J Cell Biol. 2010;191:87–99.
Lee CR, Kang JA, Kim HE, Choi Y, Yang T, Park SG. Secretion of IL‐1β from imatinib‐resistant chronic myeloid leukemia cells contributes to BCR–ABL mutation‐independent imatinib resistance. FEBS Lett. 2016;590:358–68.
Levescot A, Flamant S, Basbous S, Jacomet F, Féraud O, Bourgeois EA, et al. BCR-ABL–induced deregulation of the IL-33/ST2 pathway in CD34 (+) progenitors from chronic myeloid leukemia patients. Cancer Res. 2014;74:2669–76.
Zhang B, Ho YW, Huang Q, Maeda T, Lin A, Lee SU, et al. Altered microenvironmental regulation of leukemic and normal stem cells in chronic myelogenous leukemia. Cancer Cell. 2012;21:577–92.
Quentmeier H, Eberth S, Romani J, Zaborski M, Drexler HG. BCR-ABL1-independent PI3Kinase activation causing imatinib-resistance. J Hematol Oncol. 2011;4:6.
Wöhrle FU, Halbach S, Aumann K, Schwemmers S, Braun S, Auberger P, et al. Gab2 signaling in chronic myeloid leukemia cells confers resistance to multiple Bcr-Abl inhibitors. Leukemia. 2013;27:118.
Wagle M, Eiring A, Wongchenko M, Lu S, Guan Y, Wang Y, et al. A role for FOXO1 in BCR–ABL1-independent tyrosine kinase inhibitor resistance in chronic myeloid leukemia. Leukemia. 2016;30:1493.
Mitchell R, Hopcroft LE, Baquero P, Allan EK, Hewit K, James D, et al. Targeting BCR-ABL-independent TKI resistance in chronic myeloid leukemia by mTOR and autophagy inhibition. J Natl Cancer Inst. 2017;110:467–78.
Ly C, Arechiga AF, Melo JV, Walsh CM, Ong ST. Bcr-Abl kinase modulates the translation regulators ribosomal protein S6 and 4E-BP1 in chronic myelogenous leukemia cells via the mammalian target of rapamycin. Cancer Res. 2003;63:5716–22.
Nakamura Y, Fukami K. Regulation and physiological functions of mammalian phospholipase C. J Biochem. 2017;161:315–21.
Deka SJ, Trivedi V. Potentials of PKC in cancer progression and anticancer drug development. Curr Drug Discov Technol. 2019;16:135–47.
Holyoake TL, Vetrie D. The chronic myeloid leukemia stem cell: stemming the tide of persistence. Blood. 2017;129:1595–606.
Shah M, Bhatia R. Preservation of quiescent chronic myelogenous leukemia stem cells by the bone marrow microenvironment. In: Biological mechanisms of minimal residual disease and systemic cancer. Springer International Publishing, 2018. p. 97–110.
Ma L, Shan Y, Bai R, Xue L, Eide CA, Ou J, et al. A therapeutically targetable mechanism of BCR-ABL–independent imatinib resistance in chronic myeloid leukemia. Sci Transl Med. 2014;6:252ra121–252ra121.
Wilkinson SE, Parker P, Nixon J. Isoenzyme specificity of bisindolylmaleimides, selective inhibitors of protein kinase C. Biochemical J. 1993;294:335–7.
Gschwendt M, Muller H, Kielbassa K, Zang R, Kittstein W, Rincke G, et al. Rottlerin, a novel protein kinase inhibitor. Biochem Biophys Res Commun. 1994;199:93–8.
Acknowledgements
This work was supported by grants from the National Medical Council Singapore (NMRC/CSA/0051/2013 and NMRC/CIRG/1404/2014).
Author information
Authors and Affiliations
Contributions
JRS designed and performed experiments for cloning, western blots, quantitative PCR, and colony formation. MGY and SPT performed experiments for cloning, western blots, and qPCR. KLL performed the Fluidigm experiments for validation of alternative splicing isoforms. SC, JL, KLL, SR, and HY analyzed the RNA-sequencing data for gene expression and ASEs. CC and TH provided the primary CML samples. CO and JI performed the immunohistochemistry (IHC) staining and scoring. JH performed statistical analysis on the IHC scoring by CO and JI. OA-C, XR, and ARK provided experimental advice and essential reagents. STO supervised the project, and JRS prepared figures and wrote the manuscript which was approved by all authors.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
41375_2020_732_MOESM6_ESM.xlsx
Supp Table 6: List of detected alternative splicing events from K562 cells with SRSF1 overexpression vs vector in DMSO and IM conditions
41375_2020_732_MOESM7_ESM.xlsx
Supp Table 7: Differential alternative splicing event analysis of K562 cells with SRSF1 overexpression vs vector control in DMSO
41375_2020_732_MOESM8_ESM.xlsx
Supp Table 8: Differential alternative splicing event analysis of K562 cells with SRSF1 overexpression vs vector control in IM
41375_2020_732_MOESM11_ESM.xlsx
Supp Table 11: List of significant and differential genes from analysis of K562 cells with SRSF1 overexpression vs vector control in DMSO
41375_2020_732_MOESM12_ESM.xlsx
Supp Table 12: List of significant and differential genes from analysis of K562 cells with SRSF1 overexpression vs vector control in IM
Rights and permissions
About this article
Cite this article
Sinnakannu, J.R., Lee, K.L., Cheng, S. et al. SRSF1 mediates cytokine-induced impaired imatinib sensitivity in chronic myeloid leukemia. Leukemia 34, 1787–1798 (2020). https://doi.org/10.1038/s41375-020-0732-1
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41375-020-0732-1
This article is cited by
-
Ppm1d truncating mutations promote the development of genotoxic stress-induced AML
Leukemia (2023)
-
EVI1 upregulates PTGS1 (COX1) and decreases the action of tyrosine kinase inhibitors (TKIs) in chronic myeloid leukemia cells
International Journal of Hematology (2023)