Abstract
The unbalanced translocation dic(1;7)(q10;p10) in myelodysplastic syndromes (MDS) is originated by centromeric juxtaposition resulting into 1q trisomy and 7q monosomy. More than half of cases arise after chemo/radio-therapy. To date, given the absence of genes within the centromeric regions, no specific molecular events have been identified in this cytogenetic subgroup. We performed the first comprehensive genetic and epigenetic analysis of MDS with dic(1;7)(q10;p10) compared to normal controls and therapy-related myeloid neoplasms (t-MNs). RNA-seq showed a unique downregulated signature in dic(1;7) cases, affecting more than 80% of differentially expressed genes. As revealed by pathway and gene ontology analyses, downregulation of ATP-binding cassette (ABC) transporters and lipid-related genes and upregulation of p53 signaling were the most relevant biological features of dic(1;7). Epigenetic supervised analysis revealed hypermethylation at intronic enhancers in the dicentric subgroup, in which low expression levels of enhancer putative target genes accounted for around 35% of the downregulated signature. Enrichment of Krüppel-like transcription factor binding sites emerged at enhancers. Furthermore, a specific hypermethylated pattern on 1q was found to underlie the hypo-expression of more than 50% of 1q-deregulated genes, despite trisomy. In summary, dic(1;7) in MDS establishes a specific transcriptional program driven by a unique epigenomic signature.
Similar content being viewed by others
Introduction
The dic(1;7)(q10;p10) is a rare cytogenetic alteration found in 1.5–6% of myelodysplastic syndromes (MDS), and less frequently in acute myeloid leukemia (AML) and myeloproliferative neoplasms [1]. Of note, dic(1;7)-associated MDS/AML are frequent after chemo/radio-therapy exposure. Interestingly, in both de novo and treatment-induced disorders, this abnormality usually occurs as isolated or associated with one additional cytogenetic change [2].
Dic(1;7) differs from other typical chromosome translocations that act as leukemia drivers by originating fusion genes and chimeric transcripts, as it derives from the translocation of whole arms of chromosomes 1 and 7 with juxtaposition at centromeric regions, without production of abnormal fusion genes.
Human centromeres are made by chromosome-specific no-gene DNA encompassing transposable elements and repetitive alpha-satellite (α-sat) sequences. Their function is under the control of epigenetic marks that promote binding of protein complexes directing segregation during cellular divisions (Fig. 1a). Aberrant recombinations of chromosomes 1, 9 and 16 at centromeric level, characterize a rare autosomal recessive disease with a constitutional methylation defect, immunodeficiency and facial anomalies (Online Mendelian Inheritance in Man (OMIM) #614069, OMIM #616910, OMIM #616911) [3]. In multiple myeloma, cytogenetic instability of 1q originating jumping translocations prevalently involves pericentromeric heterochromatin decondensation rather than α-sat [4].
Cytogenetics, fluorescent in situ hybridization (FISH) and single-nucleotide polymorphism array (SNPa) result summary. a Human centromeric regions. Top: representative chromosome structure with position of repetitive sequences. Upper middle: small different colored circles represent monomers of alpha-satellite DNA at centromeres; large red arrows indicate high-order repeat (HOR) units that form centromeric chromatin. Lower middle: specific histone modifications promote binding of centromeric (CENP-A, CENPB, CENP-C) and pericentromeric (HP-1) proteins. Bottom: the constitutive 16 proteins, centromere-associated network (CCAN), interact with centromeric proteins and promote microtubule attachment to the centromere. b Schematic representation with relative orientation of genomic FISH probes at centromere 1 (D1Z5 SpectrumOrange) and centromere 7 (D7Z1 SpectrumGreen). c Metaphase dual-color FISH with D1Z5 (orange) and D7Z1 (green) showing a fusion signal on dic(1;7), indicated by a yellow arrow (DIC 4 case). d Copy number alterations (CNAs) detected by SNP-arrays in all five dicentric cases. CN copy number, chr chromosome, cytostart cytogenetic band start, cytoend cytogenetic band end
Due to an absence of genes within the centromeric regions, specific molecular consequences have not been identified in dic(1;7) cases and detailed genomic information is still lacking. Using a comprehensive genetic and epigenetic approach we delineated the specific (epi)genomic features underlying the typical 1;7 centromere–centromere recombination in MDS.
Materials and methods
Samples
Cases were retrospectively collected from the files of the Laboratory of Cytogenetics and Molecular Genetics at the Hematology Department of the University of Perugia, Italy. All patients gave their written informed consent to sample collection and biological analyses in accordance with the Declaration of Helsinki. The study was approved by the Institutional Bioethics Committee (Prot.1.X.2011). Hematological and cytogenetic features of patients are listed in Table 1. dic(1;7)(q10;p10) cases were compared with a series of controls and therapy-related myeloid neoplasms (t-MNs). As controls we used bone marrow samples from non-neoplastic cytopenias, normal bone marrow morphology and normal karyotype. Mutational diagnostic screening of 14 MDS-related genes was performed on both controls and t-MNs (Supplementary Table 1).
Karyotype, fluorescent in situ hybridization (FISH), denaturing high-performance liquid chromatography (DHPLC) and Sanger
Cytogenetic analysis was performed following standard procedure and G-banded karyotypes. FISH experiments on metaphase chromosomes were performed using the following probes: Vysis CEP 1 (D1Z5, SpectrumOrange Probe, Abbott) for alphoid sequences of chromosome 1 and Vysis CEP 7 (D7Z1, SpectrumGreen Probe, Abbott) for chromosome 7 α-sat DNA (Fig. 1b). Analyses were carried out using a fluorescence microscope Olympus BX61 equipped with a highly sensitive camera JAI and driven by CytoVision 4.5.4 software. At least seven abnormal metaphases were analyzed in each experiment.
Nucleic acids were extracted from unsorted bone marrow cells of patients using All Prep DNA/RNA Mini Kit (Qiagen), quantified with Qubit fluorimeter using Quant-i-T dsDNA HS Assay Kit and RNA HS Assay Kit (Invitrogen) respectively and samples quality was evaluated using Tapestation visualization (Agilent 2100 Bioanalyzer). Mutational diagnostic screening of 14 MDS-related genes was performed using DHPLC (Wavemaker software, Wave System, MD Transgenomic Inc., USA) and Sanger sequencing (3500 Genetic Analyzer, Applied Biosystems) (Table 1; Supplementary Table 1).
Single-nucleotide polymorphism array (SNPa)
SNPa was performed on CytoScan HD Affymetrix platform (Affymetrix, Santa Clara, CA, USA) following the manufacturer’s instructions. A total of 250 ng of high-quality genomic DNA obtained from bone marrow cells was used to study copy number alterations (CNAs) and copy neutral loss of heterozygosity (cnLOH). The protocol was supported by Affymetrix GeneChip Command Console (AGCC) software and analyzed through Affymetrix Chromosome Analysis 3.1 (ChAS) software with filters set at 100 Kb dimension for CNAs and 10 Mb dimensions for cnLOH. Our data refer to NetAffx Build 32.3 (hg19) database. Polymorphic copy number variations were excluded from the analysis.
Whole exome sequencing (WES)
WES was performed in dic(1;7) (DIC) 2 and DIC 5 (Table 1) using germline DNA extracted from bone marrow fibroblasts with normal karyotype. All variations identified by WES were confirmed as somatic on matched tumor/normal samples and tested on the remaining three DIC cases by Sanger sequencing. Illumina paired-end libraries were generated according to the manufacturer’s protocol (Illumina, San Diego, CA). Image processing and basecall were performed using the Illumina Real Time Analysis Software. Paired whole-exome fastq data were aligned to the human reference genome (GRCh38/hg38) with the BWA-MEM algorithm [5]. Duplicates were marked using Samblaster. Quality of the aligned reads, somatic variant calling and copy number analysis were performed through CEQer2 tool, as previously described [6]. Variants were annotated using dbSNP142. Variants with minor allele frequency <0.01 or carrying a ‘Clinical’ single-nucleotide polymorphism database (dbSNP) flag were further processed; the other variants were discarded from subsequent analyses. Filtered variants were exported as vcf files and used as input for Annovar [7] analysis/annotation.
RNA-seq and quantitative real-time PCR (qRT-PCR)
All samples used for RNA-sequencing (RNA-seq) had RNA integrity number of 6 or above. Ribosomal RNA depletion and library preparation were obtained following the manufacturer’s instructions using RiboGone Mammalian Kit (Clontech Laboratories Inc.) and NEBNext® Ultra™ RNA Library Prep Kit for Illumina (New England Biolabs, NEB), respectively. Reads were aligned against hg19 build of the human genome using TopHat [8], counts were generated using HTseq [9] and DESeq2 was used for differential expression analysis (false discovery rate (FDR) <0.1 and log2 fold change ≥I1I) [10]. To validate RNA-seq results, 1 μg RNA was retrotranscribed using 100 U of Superscript II and esarandom primers (Invitrogen, Carlsbad, CA, USA) and qRT-PCR was performed on ZMAT3 and NR1H2 using TaqMan assay probes Hs00536976_m1 and Hs01027215_g1 (Applied Biosystems, Life Technologies, Carlsbad, USA) respectively with endogenous ABL1 (Hs00245445 m1, Applied Biosystems, Life Technologies, Carlsbad, USA) and Universal Human Reference RNA (Stratagene, Cedar Creek, TX, USA) as reference controls. Real-time reactions were performed in triplicate using the Roche Light Cycler 480; fluorescence data were analyzed with the software version 1.5 and second derivative maximum method; gene expression was expressed as Cp (crossing point) values. Statistical significance was tested by Mann–Whitney test (*p < 0.05).
Multiplex enhanced reduced representation bisulfite sequencing (mERRBS)
mERRBS libraries were prepared with modified size selection fragments for multiplex application, as previously described [11]. Only genomic regions with coverage ranging from 10 to 450 times were used for the downstream analysis. Reads were aligned against a bisulfite-converted human genome (hg19) using Bismark with Bowtie2 [12]. Downstream analysis was performed with the MethylKit [13] and MethylSig [14] packages. We investigated cytosine methylation using beta-binomial regression (FDR ≤0.1) with a minimum cut-off of 25% methylation difference to identify differential methylated cytosines. Unsupervised clustering analysis with principal component analysis was conducted using only high-variance CpG tiles across all samples. Unsupervised analysis of DNA methylation using hierarchical clustering (distance = 1-Pearson Correlation, Ward’s agglomeration method) was performed using MethylKit. Differentially methylated regions (DMRs) were identified by first summarizing the methylation status of the genomic regions into 25 bp tiles and then identifying regions with an absolute methylation difference ≥25% and an FDR ≤0.1. DMRs were annotated to the RefSeq genes (National Center for Biotechnology Information (NCBI)) using the following criteria: (a) DMRs overlapping with a gene were annotated to that gene; (b) intergenic DMRs were annotated to all neighboring genes within a 50 kb window; and (c) if no gene was detected within a 50 kb window, the DMR was annotated to the nearest transcription start site [15]. For Chip-Enrich annotation we used the option locus.def = “nearest_tss” [16]. Enhancer annotation was performed using chromatin immunoprecipitation sequencing (ChIP-seq) data sets generated in human-mobilized CD34+ cells. Enhancers were defined as regions with H3K4me1 and H3K27ac, with absence of the promoter marker H3K4me3 [17]. Moreover, we used data sets generated in human-adult CD34+ cells based on histone repressive marks (H3K27me3 and H3K9me3) and histone active marks (H3K36me3, H3K4me1 and H3K4me3) to investigate if aberrant DNA methylation in dic(1;7) was associated with a particular chromatin and/or structural microenvironment modification [17].
Sequencing
All amplified libraries for RNA-seq and mERRBS underwent quality control steps as described for DNA and RNA extraction and were sequenced on Illumina HiSeq2500 using the manufacturer’s recommendations (Illumina San Diego, CA). The data were deposited in NCBI Gene Expression Omnibus (GEO) under the accession number GSE118648.
Integrative functional analysis
To analyze the functional pathway and Gene Ontology (GO) enrichment of the identified gene list, we used the Database for Annotation, Visualization, and Integrated Discovery 6.8 (DAVID) [18] and RNA-Enrich [19]. Known and de novo DNA motif enrichment were identified with Hypergeometric Optimization of Motif EnRichment (HOMER) [20] using all tiled CpGs from our sequencing as background in the case of mERRBS analysis. In order to analyze the potential recruitment of transcriptional repressors that could be responsible for gene silencing, we intersected DMRs and extracted promoter position from the identified downregulated genes between dic(1;7) and controls with ChIP-seq data generated in CD34+ cells for CTCF [21], one of the most well-characterized transcriptional repressor, and for seven well-known hematopoietic transcription factors [22].
Results
The dic(1;7) breakpoints fell within α-sat DNA
Double-color FISH with Vysis D1Z5 and D7Z1 probes generated a fusion signal between centromeric regions in all our five dic(1;7) cases (Fig. 1c). Three cases bore one or two additional events as seen at both conventional karyotypes and SNPa, in particular, trisomy 8, trisomy 9 and del(13q) (Table 1 and Fig. 1d). In DIC 3, trisomy 8 characterized a subclone of a main line including dic(1;7) and del(13q) (Fig. 1d). No CNAs or cnLOH, with reduplication of a chromosomal allele alongside with loss of its respective homologous region, were detected by SNPa.
dic(1;7) bore no common somatic mutations
WES on DIC 2 (Table 1) identified a mutation in the ankyrin 7 domain of ANKS1B/EB1 gene (12q23.1), a promiscuous gene originating multiple fusion transcripts in solid tumors [23] and overexpressed in a subset of B-cell acute lymphoblastic leukemia (B-ALL) [24]. The same case harbored a variation in the polyprenyl synthetase domain of GGPS1 (1q42.3), a gene transcribing for a long non-coding RNA which is overexpressed in lung cancer and mutated in adult T-cell lymphoma cases [25, 26]. In DIC 5 we identified two mutations, the first one within the alanine-rich domain of CCDC8 gene (19q13), involved in maintaining microtubule integrity and mutated in the genetic 3 M growth retardation syndrome (OMIM #614205) [27]. The second mutation occurred in the proline-rich domain of PSMF1 gene (20p13), a proteasome regulator previously described as mutated in a case of ALL [28]. In silico analysis with three different software predicted the GGPS1 mutation to be disease causing, while the other three mutations showed conflicting results (Supplementary Figure 1). None of these variations were identified by Sanger sequencing in the other three patients (DIC 1, DIC 3 and DIC 4, Table 1).
dic(1;7) has a distinct expression signature
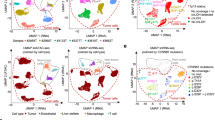
To characterize the transcriptomic differences between dic(1;7) cases and the other two groups, controls and t-MNs, we analyzed their gene expression profile. Unsupervised analysis using principal component analysis of transformed RNA-Seq count data separated cases bearing dic(1;7) from controls and t-MNs along the first principal component (Fig. 2a, b). We identified a total of 4860 differentially expressed genes (DEGs) between dic(1;7) and controls, and 4317 between dic(1;7) and t-MNs (Fig. 2c). Gene expression analysis showed a prevalent downregulation within the signature of dic(1;7) cases, affecting more than 80% of DEGs (Fig. 2c, d; Supplementary Figure 2). By analyzing promoters of downregulated genes for the presence of hematopoietic transcription factors binding sites, we found a low percentage of overlap, ranging from 0.58% of LYL to 10.8% of FLI (Supplementary Figure 3). Interestingly, the CTCF analysis at the level of promoters of silenced genes revealed that 920 out of 3971 downregulated genes between dic(1;7) and controls (23%) were CTCF targets, suggesting a potential contribution of transcriptional repressors in mediating the observed downregulated gene expression profile (Supplementary Table 2 and Supplementary Figure 3).
Summary of RNA-sequencing (RNA-seq) results. a Scatter plot showing results of a principal component analysis (PCA) of transformed count data from RNA-seq using DESeq2 (varianceStabilizingtransformation). Samples are colored according to their disease status, observe that the DIC are separated from all other samples along dimension 1 (the x-axis). b The scree plot line indicates the amount of the total variance preserved by a principal component. c Table summary of RNA-seq result analysis. Up refers to upregulated genes, down refers to downregulated genes. d Heatmap representing the top 50 differentially expressed genes (DEGs) across all three groups of samples. The heatmap colors reflect values representing the degree of expression from low to high as blue to red, respectively, as shown on the scale at the top-left hand side of the figure. Red indicates control (CTL), blue therapy-related myeloid neoplasm (t-MN) and green dic(1;7). e Significance for ZMAT3 and NR1H2 expression were tested by Mann–Whitney test (*p < 0.05); values are expressed as means ± SEM
To capture biological differences, we analyzed functional pathways within DEGs. Dicentric cases showed 18 and 30 significantly enriched pathways when analyzed against controls and t-MNs, respectively (FDR ≤0.1) (Supplementary Table 2 and 3). The t-MN cohort compared to controls showed 102 enriched pathways (Supplementary Table 4). When compared to controls, the two neoplastic subgroups included in this study, i.e., dic(1;7) and t-MNs, showed aberrant downregulation of biologically relevant pathways, specifically: mitogen-activated protein kinase (MAPK) signaling, spliceosome, ribosome, gonadotropin-releasing hormone (GnRH) signaling, IL12/Stat4 (interleukin-12/signal transducer and activator of transcription 4)-dependent signaling and tumor necrosis factor (TNF)/stress-related signaling (Supplementary Tables 2-4). In addition, six pathways emerged as specifically deregulated (4 downregulated and 2 upregulated) in dic(1;7) cases (Supplementary Tables 2-4). Notably, the signaling of ATP-binding cassette (ABC) transporters, a superfamily of proteins involved in the transport of metabolic products and drugs across biological membranes, emerged as specifically downregulated (Supplementary Tables 2-5). Of the ABC genes detected as enriched by our RNA pathway analysis, seven were found to be significantly deregulated in dic(1;7) compared to the other two subgroups (Supplementary Table 5). In particular, 5/7 were downregulated: ABCA6 and ABCA10, known as phospholipid and cholesterol transporters; ABCB4 and ABCG2 involved in phosphatidylcholine and drug transport; and ABCB6, which plays a crucial role in heme synthesis by mediating porphyrin uptake into mitochondria [29]. Two other ABC genes, the phospholipid and drug transporters ABCA3 and ABCC3, were upregulated (Supplementary Table 5). Moreover, the cholesterol transporter ABCA9 was significantly downregulated in dic(1;7) only when compared to t-MNs, while the peptide transporter ABCB2 was significantly upregulated only against controls (Supplementary Table 5).
GO analysis of differentially expressed genes showed significantly enrichment in lipid metabolism terms involving 12 genes, 10 of which were downregulated only in dic(1;7) (Supplementary Tables 2-5). Among them, we found one group of lipid metabolism genes related to oxidative/inflammatory response: the apolipoprotein APOM and the cholesterol esterifying enzyme LCAT that have been reported to decrease upon oxidative and/or inflammatory response [30, 31]; the cytidine deaminase APOBEC2, found to be regulated by the inflammatory transforming growth factor-beta signaling [32] and the nuclear receptor NR1H2 that was involved in lipid homeostasis and inflammation [33]. A second group of genes is encoding for lipids related to cancer: LPL, lipoprotein lipase hypo-expression that was shown to predict evolution in chronic lymphocytic leukemia patients [34]; PLA2G7, a phospholipase which expression in the peripheral blood was predictive of survival in melanoma patients [35]; apolipoprotein APOC1 that was associated with poor prognosis in pancreas cancer patients; moreover, when inhibited, it induced apoptosis in pancreatic cancer cell lines [36].
Only two specifically upregulated pathways emerged in dic(1;7), the most important of which was the TP53 signaling pathway, which included 9 of the DEGs (Supplementary Tables 2-5). In particular, the four TP53 target genes upregulated in dicentrics, compared to both t-MNs and controls, were: ZMAT3, STAT1, SUMO1 and CCNG1 (Supplementary Table 5). Moreover, SHISA5 and MYC were upregulated only against controls and SERPINE1, FAS and ATR only against t-MNs. By contrast, t-MNs versus controls showed TP53 signaling downregulation (Supplementary Table 4). The TP53 gene by itself was not significantly deregulated in any subgroups, although there was a trend towards RNA hyper-expression in dic(1, 7) and hypo-expression in t-MNs (data not shown). We validated the upregulation of the TP53 target gene ZMAT3 and the downregulation of NR1H2 gene, involved in lipid homeostasis and inflammation, by qRT-PCR (Fig. 2e).
Both gene dosage effect and DNA methylation profile contributed to the dic(1;7) signature
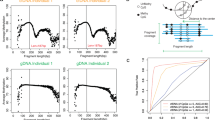
We investigated gene dosage effect of 1q trisomy and 7q monosomy from RNA-seq. Compared with controls and t-MNs, 97.5% (117/120) and 95.9% (118/123), respectively, of 7q DEGs were downregulated in dic(1;7), in keeping with 7q loss (Fig. 3a). Interestingly, downregulation also involved EZH2 and MLL3, previously identified as haploinsufficient putative suppressor genes on chromosome 7. Surprisingly, among 198 1q DEGs included in the signature of dicentrics against controls, 90 were upregulated and 108 downregulated (Fig. 3b). Also, when dic(1;7) was compared to t-MNs, the 157 1q DEGs resulted to be 52 up- and 105 down-regulated (Fig. 3b). These findings showed that, despite of trisomy, more than 50% of 1q genes were always hypo-expressed in the dic(1;7). Consistent with this, 85/119 1q DMRs were hypermethylated in dicentrics compared with controls (67/109 compared with t-MNs) (Fig. 3c) and they were annotated to 83 genes, 31 of which were downregulated. Interestingly, 61/85 hypermethylated 1q DMRs (71.8%) were annotated to enhancers, thus linking 1q hypermethylation to functional DNA region (Supplementary Table 6).
Gene dosage effect and DNA methylation pattern on 7q and 1q regions of dic(1;7). a Chromosome view of regions significantly enriched in genes differentially expressed using PGE (Positional Gene Enrichment) on 7q (left). Percentage of differentially expressed gene (DEG) distribution within down- and up-regulated genes in dic(1;7) compared to controls (up-right) and to therapy-related myeloid neoplasms (t-MNs; bottom right). Compared to controls, 117/120 of 7q DEGs were downregulated in dic(1;7), while compared to t-MNs, 118/123 7q DEGs were downregulated. b Chromosome view of regions significantly enriched in genes differentially expressed using PGE on 1q (left). Percentage of DEG distribution within down- and up-regulated genes in dic(1;7) compared to controls (up-right) and to t-MNs (bottom right). The signature of dic(1;7) contained 90 up- and 108 (54.5%) down-regulated genes on 1q compared to controls; 52 up- and 105 (66.8%) down-regulated genes on 1q compared to t-MNs. c Differentially methylated region (DMR) distribution along chromosome 1q involved in the translocation. Dicentric cases showed hypermethylation of DMRs on 1q compared to both controls (left) and t-MN (middle). By contrast, t-MNs showed a prevalent hypomethylated pattern on 1q (right)
Global DNA methylation profiling captured 3.2 M CpGs across the genome. Principal component analysis of high-variance CpGs identified dic(1;7) and t-MN cases from controls (Fig. 4a, b). Unsupervised analysis using hierarchical clustering segregated dicentrics with additional anomalies in a separate cluster with the t-MN cases with complex karyotype (Fig. 4c; Table 1). Supervised analysis showed both global hyper- and hypo-methylation in dic(1;7), compared with the other subgroups, while t-MNs showed global hypomethylation (Fig. 4b, c). We identified a total of 3261 DMRs between dic(1;7) and controls, and 4568 DMRs between dicentric cases and t-MNs (Fig. 5a). Genome-wide distribution of DMRs and annotation to genomic regions showed that, compared to controls, dicentrics were significantly depleted at promoter regions (15% vs. background 30%, binomial test p < 2.2e−16) and significantly enriched in non-promoter regions (Fig. 5a, b). These results were also maintained in the comparison against the t-MN group. Moreover, in comparing dicentrics to controls, 33% of DMRs had CTCF binding sites (Supplementary Figure 3). By contrast, the analysis of hematopoietic transcription factors revealed that less than 1% of DMRs had binding sites for the specific transcription factor (TF) motifs analyzed (Supplementary Figure 3).
Summary of multiplex enhanced reduced representation bisulfite sequencing (mERRBS) analysis. a Unsupervised multivariate analysis using principal component analysis (PCA) was able to separate controls from all tumor samples. The scree plot line indicates the amount of the total variance preserved by a principal component (middle). Unsupervised analysis of DNA methylation using hierarchical clustering (left). Red indicates control (CTL), blue indicatestherapy-related myeloid neoplasms (t-MNs) and green dic(1;7) (DIC). b Circos plot showing the hyper- and hypomethylated differentially methylated regions (DMRs) found for each group comparison: DIC/CTL (left), DIC/t-MN (middle) and t-MN/CTL (right). The outer circle indicates the G-banded human chromosomes, red and blue colors refers to hyper- and hypomethylated DMR distribution, respectively. c Volcano plots of all DMR distributions between each group. The plot shows the DNA methylation difference on percentage (x-axis) with respect to the −log10 of the adjusted p value for each DMR. Thresholds are shown as red lines: dot vertical lines indicate DNA methylation difference cut-off (−25% and +25%), dot horizontal line indicates statistically significant cut-off (q-value ≤0.1). Significantly different hypo- and hyper-DMRs are highlighted as blue and red dots, respectively. DIC vs. CTL (left), DIC vs. t-MN (middle) and t-MN vs. CTL (right)
Summary of annotation analysis. a Table summarizing the distribution of annotated differentially methylated regions (DMRs) on genomic regions for each group. We used Methylkit package on R to annotate previously identified DMRs on Methylsig to hg19. Statistical results for binomial enrichment/depletion test were conduct on R. For all tests, we used 95% confidence interval and percentage of background annotation for each category (promoters, exons, introns, intergenic) as true probability of success. b Pie charts illustrate DMRs annotated to promoter (red), exonic (green), intronic (blue) and intergenic regions (purple) in dic(1;7) (DIC) vs. control (CTL; left) and vs. therapy-related myeloid neoplasm (t-MN; right). As background, we used total identified DMRs without filters (top left). c Distribution of hypo- and hypermethylated DMRs on promoter regions for each comparison (top) and relative number of hypo- and hyper-promoter DMRs for each group (bottom). d Venn diagram showed the overlap between genes annotated to promoter DMRs and all differentially expressed genes (DEGs) identified on RNA-sequencing (RNA-seq) analysis in DIC vs. CTL (i) and vs. t-MN (ii)
Focusing on promoter regions, t-MNs showed gene-specific promoter hypermethylation (Fig. 5), confirming previous studies [37]. dic(1;7) cases compared with controls showed hypermethylation of 356/502 (68%) promoters (Fig. 5c). However, promoter hypermethylation correlated with significant changes in expression level only for 31 genes, suggesting that its contribution to dic(1;7) expression profile was limited (Fig. 5d).
Enhancer hypermethylation enriched on TF binding sites emerged in dic(1;7)
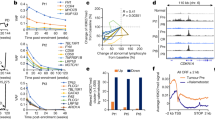
Analysis of the non-promoter regions showed that 43% of the dic(1;7) DMRs were enriched for enhancers (background 34%, binomial test p < 2.2e−16) when compared with controls, while t-MNs had 38% enrichment (background 34%, binomial test p = 0.046) (Fig. 6a). In silico analysis using ChIP-seq data sets for histone repressive marks (H3K27me3 and H3K9me3) and histone active marks (H3K36me3, H3K4me1 and H3K4me3) from CD34+ primary cells [17] confirmed that aberrant DNA methylation in dic(1;7) is mainly associated with regions marked by active histone marks specifically linked to enhancers (72% of DMRs overlap with H3K4me1, background 68.6%; binomial test p = 3.255e−07). Using the Hnisz D data set of 234 super-enhancers identified in CD34+ adult cells [38], in our series we found 5 super-enhancers out of the 744 hypermethylated enhancers (Supplementary Table 6).
Summary of enhancers analysis. a Pie chart illustrates differentially methylated region (DMR) enrichment on enhancer regions. Red indicates feat1, enhancers; and light blue indicates others, non-enhancers. Compared with controls (CTL), dic(1;7) (DIC) DMRs showed an enrichment on enhancers with p value < 2.2e−16 (left), compared with therapy-related myeloid neoplasms (t-MNs) that showed an enrichment on enhancers with p value of 6.617e−06 (middle) and t-MNs compared with controls that showed an enrichment on enhancers with p = 0.046 (right). For statistical analysis, we used binomial test with 95% confidence interval and percentage of background annotation as true probability of success. b Venn diagrams illustrate the overlap between hypermethylated enhancer target genes and downregulated signature on DIC/CTL (left) and DIC/t-MN (right) groups
In both dicentric and t-MNs, enhancer DMRs were predominantly found in intronic regions. A direct enriched test confirmed that the enrichment at enhancers was stronger in dicentric cases than in t-MNs, with a prevalent hypermethylation (Fig. 6a). Since DNA methylation negatively correlates with enhancer activity and transcription of target genes [39], we used the Predicting Enhancer Target by modules tool to identify putative enhancer-target genes [40]. Indeed, compared to controls, dic(1;7) RNA-seq revealed that 1747 out of 4860 DEGs were target genes of hypermethylated enhancers (Fig. 6b). Moreover, 1377/1747 DEGs had decreased gene expression, suggesting that enhancer hypermethylation directly accounts for 34.7% of the downregulated signature in dicentric cases (Fig. 6b). These results were maintained by dic(1;7) against t-MNs, with enhancer hypermethylation accounting for 40% of the downregulated profile (Fig. 6b). By contrast, the t-MN group showed 128 out of 192 enhancer DMRs (66%) to be hypomethylated (Fig. 6a). Since DNA methylation has been shown to prevent binding of TFs to their target sites [41], we analyzed DMRs with HOMER. This analysis revealed that DMRs were enriched for TF binding sites (Supplementary Figures 4-7), with those regions differentially methylated in dic(1;7) compared to t-MNs showing enrichment for the key hematopoietic TF factors, GATA1 (p-value = 1e−4), GATA2 (p-value = 1e−3), GATA3 (p value = 1e−2), ETV1 (p value = 1e−2), STAT3 (p value = 1e−2) and RUNX1 (p value = 1e−2) (Supplementary Figure 5). Specifically, GATA and RUNX1 transcription factors binding sites were hypermethylated while ETV1 and STAT3 binding sites were hypomethylated. With respect to hypermethylated enhancers, dic(1;7) showed enrichment of 28 and 27 TFs, compared to controls and t-MNs, respectively (Supplementary Figures 6 and 7). Interestingly, dic(1;7) hypermethylated enhancers were strongly enriched in motifs for Krüppel-like factor (KLFs) protein family transcription factors. Specifically, KLF5, KLF6 and KLF4 were included among the top enriched transcription factors in the analysis against controls (Supplementary Table 6), while KLF3, KLF5 and KLF4 emerged in the analysis against t-MNs (Supplementary Table 7). By contrast, none of these KLF transcription factors emerged as enriched in t-MNs enhancers when compared to controls (Supplementary Figures 8).
Discussion
This is the first in-depth genetic and epigenetic characterization of dic(1;7) which provided us with new insights into the biological features of a recurrent leukemic chromosome rearrangement involving repetitive DNA at centromeric regions.
In our patients, breakpoints of dic(1;7) fell within α-satellite DNA at chromosome 1 and chromosome 7, in agreement with previously reported data [42]. As a result, the affected cells had genomic imbalances involving both euchromatic regions, due to entire 1q trisomy and 7q monosomy, and heterochromatic regions, with an excess of α-satellite and pericentromeric sequences of chromosome 1. Interestingly, centromeric juxtaposition in dic(1;7) did not alter centromere action on chromosome segregation and the der(1;7) behaved as a stable clonal marker along mitotic divisions of many cell generations, even during long-lasting cytogenetic monitoring, for up to 5 years (data not shown). Moreover, the dic(1;7) chromosome behaved as a cytogenetically primary abnormality since additional karyotypic changes, when present, were individually different, in both de novo and therapy-induced MDS. Accordingly, data from 258 cases of dic(1;7) collected from Mitelman database showed 82.5% of cases with the dic(1;7) as sole cytogenetic aberration or plus only one/two additional abnormalities [2]. In addition, here we excluded the presence of cytogenetically cryptic changes by using SNP-arrays. Altogether, these data highlight dic(1;7) as the cytogenetic hallmark of a biological entity including both de novo and t-MDS strikingly different from other t-MNs typically associated with complex karyotypes [43].
Common somatic mutations did not emerge in our cases with dic(1;7) in which we found only private variations affecting GGPS1; CCDC8; and PSMF1 genes, still not reported in MDS/AML, and ANKS1B/EB1 gene, recently annotated in two AML cases [44].
Interestingly, when compared to both controls and t-MNs, the dic(1;7) subgroup was characterized by a signature with prevalence of gene downregulation that was supported by a specific epigenetic program. Notably, promoter hypermethylation contributed to downregulation to a lower extent than hypermethylation at the level of non-promoter regions, particularly at intronic enhancers which target genes represented at around 35% of the entire downregulated expression profile. Both enhancers and super-enhancers aberrant DNA methylation have been identified in malignancies [45, 46]. Interestingly, in multiple myeloma, Agirre et al. [47] found hypermethylated intronic enhancers to be associated with binding sites for transcription factors related to B-cell differentiation. Similarly, the hypermethylated enhancers in dic(1;7) included binding sites for important transcription factors in normal hematopoiesis, such as GATA1, GATA2, GATA3 and RUNX1. In addition, in dic(1;7) the KLF protein family specifically emerged with four top enriched members involved in the regulation of: stem/progenitor proliferation and localization in the bone marrow (KLF5) [48], erythroid differentiation (KLF5, KLF6, KLF3) [49], stem cell pluripotency (KLF4) [50] and tumor suppression (KLF4) [51]. Intriguingly, a checkpoint role following DNA damage was attributed to KLF4 loss of function in a murine model, in which its inactivation was associated with high level of apoptosis and genetic aberrations, including dicentric chromosomes [52]. Altogether, these data first emphasize a pathogenetic role of the KLF family in a distinct cytogenetic subgroup of MDS/AML. Future studies are required to investigate potentialities of their modulation for new therapeutic strategies.
Dosage effect due to chromosome number variations is a well-known mechanism altering gene expression in leukemic cells with a strong relationship between gene hypo-expression and chromosome loss [53]. Accordingly, in the dicentric signature we found the expected gene dosage effect due to 7q monosomy, as more than 95% of differentially expressed genes belonging to the long arm of chromosome 7 were downregulated. Among them, EZH2 and MLL3 downregulation might have an impact on global gene expression.
In contrast, a gene dosage effect by 1q trisomy acted only partially, given that more than 50% of 1q differentially expressed genes were downregulated in the cases with dicentric chromosome. Indeed, DNA methylation profile clarified that discrepancies between 1q trisomy and gene hypo-expression were related to high level of 1q hypermethylation in dic(1;7), compared to both controls and t-MNs. Whether gain of the 1q heterochromatin is related to this epigenetic effect is still unknown.
Pathway analysis of the signature showed that ABC transporters were significantly downregulated in dic(1;7) when compared to both controls and t-MNs. ABC transporters, a large family of proteins with localization at mitochondria, endoplasmic reticulum, Golgi, endosomes and cytoplasmic membranes, are involved in multiple cellular functions, including protection against genetic damage caused by xenobiotic and chemotherapeutical compounds, and transmembrane transport of amino acids, ions, polysaccharides and lipids [54]. In particular, ABCB1, alias MDR1/p-glycoprotein, and ABCG2, alias BCRP/MXR, have been extensively studied in drug resistance in AML [55]. The ABC transporters specifically involved in the dic(1, 7) signature are known to play a role in the transport of phospholipids, cholesterol and phosphatidylcholine. Accordingly, only in the dic(1;7) subgroup, our GO analysis showed a significant enrichment for genes regulating the lipid metabolism involved in stress and inflammatory responses [37, 58]. Among them the downregulated gene NR1H2 is.an ubiquitous isoform of the liver X receptor (LXR), a targetable candidate in leukemic cells with deregulation of cholesterol homeostasis [56, 57]. Whether the ABC involvement impacts the chemo-resistance/sensitivity of dic(1;7)-affected cells remains to be elucidated.
The p53 signaling pathway emerged as a specifically upregulated pathway in dic(1;7) versus both controls and t-MNs. In the last subgroup, loss-of-function p53 mutations accompanied complex karyotypes, as expected [58]. Interestingly, upregulation of p53 target genes has been previously found in AML with wild-type p53 gene, suggesting that, in tumor cells, downstream targets may mediate p53 dysfunction regardless of gene mutations [59]. In line with this hypothesis, both our series of dicentrics and the published AML series [59], sharing wild-type p53, showed upregulation of ZMAT3, a p53 target gene encoding for an RNA-binding zinc-finger protein regulating cell cycle arrest [60]. Altogether, these data suggest that dic(1;7) MDS belong to a subgroup of myeloid malignancies in which, in the absence of mutations at the gene, p53 is involved through deregulation of its targets.
In conclusion, in this study we showed that MDS with dic(1;7) are a distinct cytogenetic–epigenetic entity characterized by a downregulated expression profile closely connected to gene dosage effect and site-specific hypermethylation, mostly at intronic enhancers enriched for hematopoietic transcription factors. Downregulation of pathways and genes involved in lipid homeostasis and upregulation of TP53 signaling emerged as specific biological features, and separated dic(1;7)-positive MDS from t-MN.
References
Huret JL, Dessen P, Bernheim A. Atlas of genetics and cytogenetics in oncology and haematology, year 2003. Nucleic Acids Res. 2003;31:272–4.
Mitelman F, Johansson B, Mertens F. Mitelman database of chromosome aberrations and gene fusions in cancer. 2014. http://cgap.nci.nih.gov/Chromosomes/Mitelman
Miniou P, Jeanpierre M, Bourc’his D, Coutinho Barbosa AC, Blanquet V, Viegas-Pequignot E. alpha-satellite DNA methylation in normal individuals and in ICF patients: heterogeneous methylation of constitutive heterochromatin in adult and fetal tissues. Hum Genet. 1997;99:738–45.
Sawyer JR, Tricot G, Mattox S, Jagannath S, Barlogie B. Jumping translocations of chromosome 1q in multiple myeloma: evidence for a mechanism involving decondensation of pericentromeric heterochromatin. Blood. 1998;91:1732–41.
Li H, Durbin R. Fast and accurate long-read alignment with Burrows-Wheeler transform. Bioinformatics. 2010;26:589–95.
Gambacorti-Passerini CB, Donadoni C, Parmiani A, Pirola A, Redaelli S, Signore G, et al. Recurrent ETNK1 mutations in atypical chronic myeloid leukemia. Blood. 2015;125:499–503.
Wang K, Li M, Hakonarson H. ANNOVAR: functional annotation of genetic variants from high-throughput sequencing data. Nucleic Acids Res. 2010;38:e164.
Trapnell C, Pachter L, Salzberg SL. TopHat: discovering splice junctions with RNA-Seq. Bioinformatics. 2009;25:1105–11.
Anders S, Pyl PT, Huber W. HTSeq--a Python framework to work with high-throughput sequencing data. Bioinformatics. 2015;31:166–9.
Love MI, Huber W, Anders S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 2014;15:550.
Garrett-Bakelman FE, Sheridan CK, Kacmarczyk TJ, Ishii J, Betel D, Alonso A, et al. Enhanced reduced representation bisulfite sequencing for assessment of DNA methylation at base pair resolution. J Vis Exp. 2015;96:e52246.
Krueger F, Andrews SR. Bismark: a flexible aligner and methylation caller for Bisulfite-Seq applications. Bioinformatics. 2011;27:1571–2.
Akalin A, Kormaksson M, Li S, Garrett-Bakelman FE, Figueroa ME, Melnick A, et al. methylKit: a comprehensive R package for the analysis of genome-wide DNA methylation profiles. Genome Biol. 2012;13:R87.
Park Y, Figueroa ME, Rozek LS, Sartor MA. MethylSig: a whole genome DNA methylation analysis pipeline. Bioinformatics. 2014;30:2414–22.
Meldi K, Qin T, Buchi F, Droin N, Sotzen J, Micol JB, et al. Specific molecular signatures predict decitabine response in chronic myelomonocytic leukemia. J Clin Invest. 2015;125:1857–72.
Welch RP, Lee C, Imbriano PM, Patil S, Weymouth TE, Smith RA, et al. ChIP-Enrich: gene set enrichment testing for ChIP-seq data. Nucleic Acids Res. 2014;42:e105.
Kundaje A, Meuleman W, Ernst J, Bilenky M, Yen A, Heravi-Moussavi A, et al. Integrative analysis of 111 reference human epigenomes. Nature. 2015;518:317–30.
Huang dW, Sherman BT, Lempicki RA. Systematic and integrative analysis of large gene lists using DAVID bioinformatics resources. Nat Protoc. 2009;4:44–57.
Lee C, Patil S, Sartor MA. RNA-Enrich: a cut-off free functional enrichment testing method for RNA-seq with improved detection power. Bioinformatics. 2016;32:1100–2.
Heinz S, Benner C, Spann N, Bertolino E, Lin YC, Laslo P, et al. Simple combinations of lineage-determining transcription factors prime cis-regulatory elements required for macrophage and B cell identities. Mol Cell. 2010;38:576–89.
Steiner LA, Schulz V, Makismova Y, Lezon-Geyda K, Gallagher PG. CTCF and cohesin SA-1 mark active promoters and boundaries of repressive chromatin domains in primary human erythroid cells. PLoS ONE. 2016;11:e0155378.
Beck D, Thoms JA, Perera D, Schutte J, Unnikrishnan A, Knezevic K, et al. Genome-wide analysis of transcriptional regulators in human HSPCs reveals a densely interconnected network of coding and noncoding genes. Blood. 2013;122:e12–22.
Huret JL, Ahmad M, Arsaban M, Bernheim A, Cigna J, Desangles F, et al. Atlas of genetics and cytogenetics in oncology and haematology in 2013. Nucleic Acids Res. 2013;41(Database issue):D920–4.
Fu X, McGrath S, Pasillas M, Nakazawa S, Kamps MP. EB-1, a tyrosine kinase signal transduction gene, is transcriptionally activated in the t(1;19) subset of pre-B ALL, which express oncoprotein E2a-Pbx1. Oncogene. 1999;18:4920–9.
Li J, Bi L, Shi Z, Sun Y, Lin Y, Shao H, et al. RNA-Seq analysis of non-small cell lung cancer in female never-smokers reveals candidate cancer-associated long non-coding RNAs. Pathol Res Pract. 2016;212:549–54.
Kataoka K, Nagata Y, Kitanaka A, Shiraishi Y, Shimamura T, Yasunaga J, et al. Integrated molecular analysis of adult T cell leukemia/lymphoma. Nat Genet. 2015;47:1304–15.
Hanson D, Murray PG, O’Sullivan J, Urquhart J, Daly S, Bhaskar SS, et al. Exome sequencing identifies CCDC8 mutations in 3-M syndrome, suggesting that CCDC8 contributes in a pathway with CUL7 and OBSL1 to control human growth. Am J Hum Genet. 2011;89:148–53.
Chen C, Bartenhagen C, Gombert M, Okpanyi V, Binder V, Rottgers S, et al. Next-generation-sequencing of recurrent childhood high hyperdiploid acute lymphoblastic leukemia reveals mutations typically associated with high risk patients. Leuk Res. 2015;39:990–1001.
Aye IL, Singh AT, Keelan JA. Transport of lipids by ABC proteins: interactions and implications for cellular toxicity, viability and function. Chem Biol Interact. 2009;180:327–39.
Xu N, Hurtig M, Zhang XY, Ye Q, Nilsson-Ehle P. Transforming growth factor-beta down-regulates apolipoprotein M in HepG2 cells. Biochim Biophys Acta. 2004;1683:33–7.
Petropoulou PI, Berbee JF, Theodoropoulos V, Hatziri A, Stamou P, Karavia EA, et al. Lack of LCAT reduces the LPS-neutralizing capacity of HDL and enhances LPS-induced inflammation in mice. Biochim Biophys Acta. 2015;1852(10 Pt A):2106–15.
Vonica A, Rosa A, Arduini BL, Brivanlou AH. APOBEC2, a selective inhibitor of TGFbeta signaling, regulates left-right axis specification during early embryogenesis. Dev Biol. 2011;350:13–23.
Fessler MB. Liver X receptor: crosstalk node for the signaling of lipid metabolism, carbohydrate metabolism, and innate immunity. Curr Signal Transduct Ther. 2008;3:75–81.
Kristensen L, Kristensen T, Abildgaard N, Royo C, Frederiksen M, Mourits-Andersen T, et al. LPL gene expression is associated with poor prognosis in CLL and closely related to NOTCH1 mutations. Eur J Haematol. 2016;97:175–82.
Saenger Y, Magidson J, Liaw B, de Moll E, Harcharik S, Fu Y, et al. Blood mRNA expression profiling predicts survival in patients treated with tremelimumab. Clin Cancer Res. 2014;20:3310–8.
Takano S, Yoshitomi H, Togawa A, Sogawa K, Shida T, Kimura F, et al. Apolipoprotein C-1 maintains cell survival by preventing from apoptosis in pancreatic cancer cells. Oncogene. 2008;27:2810–22.
Uehara E, Takeuchi S, Tasaka T, Matsuhashi Y, Yang Y, Fujita M, et al. Aberrant methylation in promoter-associated CpG islands of multiple genes in therapy-related leukemia. Int J Oncol. 2003;23:693–6.
Hnisz D, Abraham BJ, Lee TI, Lau A, Saint-Andre V, Sigova AA, et al. Super-enhancers in the control of cell identity and disease. Cell. 2013;155:934–47.
Aran D, Sabato S, Hellman A. DNA methylation of distal regulatory sites characterizes dysregulation of cancer genes. Genome Biol. 2013;14:R21.
Zhao C, Li X, Hu H. PETModule: a motif module based approach for enhancer target gene prediction. Sci Rep. 2016;6:30043.
Tate PH, Bird AP. Effects of DNA methylation on DNA-binding proteins and gene expression. Curr Opin Genet Dev. 1993;3:226–31.
Wang L, Ogawa S, Hangaishi A, Qiao Y, Hosoya N, Nanya Y, et al. Molecular characterization of the recurrent unbalanced translocation der(1;7)(q10; p10). Blood. 2003;102:2597–604.
Olney HJ, Le Beau MM. The cytogenetics of myelodysplastic syndromes. Best Pract Res Clin Haematol. 2001;14:479–95.
Cancer Genome Atlas Research N, Weinstein JN, Collisson EA, Mills GB, Shaw KR, Ozenberger BA, et al. The Cancer Genome Atlas Pan-Cancer analysis project. Nat Genet. 2013;45:1113–20.
Heyn H, Vidal E, Ferreira HJ, Vizoso M, Sayols S, Gomez A, et al. Epigenomic analysis detects aberrant super-enhancer DNA methylation in human cancer. Genome Biol. 2016;17:11.
Vidal E, Sayols S, Moran S, Guillaumet-Adkins A, Schroeder MP, Royo R, et al. A DNA methylation map of human cancer at single base-pair resolution. Oncogene. 2017;36:5648–57.
Agirre X, Castellano G, Pascual M, Heath S, Kulis M, Segura V, et al. Whole-epigenome analysis in multiple myeloma reveals DNA hypermethylation of B cell-specific enhancers. Genome Res. 2015;25:478–87.
Shahrin NH, Diakiw S, Dent LA, Brown AL, D’Andrea RJ. Conditional knockout mice demonstrate function of Klf5 as a myeloid transcription factor. Blood. 2016;128:55–9.
Humbert M, Halter V, Shan D, Laedrach J, Leibundgut EO, Baerlocher GM, et al. Deregulated expression of Kruppel-like factors in acute myeloid leukemia. Leuk Res. 2011;35:909–13.
Takahashi K, Yamanaka S. Induction of pluripotent stem cells from mouse embryonic and adult fibroblast cultures by defined factors. Cell. 2006;126:663–76.
Shen Y, Chen TJ, Lacorazza HD. Novel tumor-suppressor function of KLF4 in pediatric T-cell acute lymphoblastic leukemia. Exp Hematol. 2017;53:16–25.
Hagos EG, Ghaleb AM, Dalton WB, Bialkowska AB, Yang VW. Mouse embryonic fibroblasts null for the Kruppel-like factor 4 gene are genetically unstable. Oncogene. 2009;28:1197–205.
Schoch C, Kohlmann A, Dugas M, Kern W, Hiddemann W, Schnittger S, et al. Genomic gains and losses influence expression levels of genes located within the affected regions: a study on acute myeloid leukemias with trisomy 8, 11, or 13, monosomy 7, or deletion 5q. Leukemia. 2005;19:1224–8.
Raaijmakers MH. ATP-binding-cassette transporters in hematopoietic stem cells and their utility as therapeutical targets in acute and chronic myeloid leukemia. Leukemia. 2007;21:2094–102.
Fukuda Y, Lian S, Schuetz JD. Leukemia and ABC transporters. Adv Cancer Res. 2015;125:171–96.
Ceroi A, Masson D, Roggy A, Roumier C, Chague C, Gauthier T, et al. LXR agonist treatment of blastic plasmacytoid dendritic cell neoplasm restores cholesterol efflux and triggers apoptosis. Blood. 2016;128:2694–707.
Sanchez PV, Glantz ST, Scotland S, Kasner MT, Carroll M. Induced differentiation of acute myeloid leukemia cells by activation of retinoid X and liver X receptors. Leukemia. 2014;28:749–60.
Wong TN, Ramsingh G, Young AL, Miller CA, Touma W, Welch JS, et al. Role of TP53 mutations in the origin and evolution of therapy-related acute myeloid leukaemia. Nature. 2015;518:552–5.
Parikh N, Hilsenbeck S, Creighton CJ, Dayaram T, Shuck R, Shinbrot E, et al. Effects of TP53 mutational status on gene expression patterns across 10 human cancer types. J Pathol. 2014;232:522–33.
Bersani C, Xu LD, Vilborg A, Lui WO, Wiman KG. Wig-1 regulates cell cycle arrest and cell death through the p53 targets FAS and 14-3-3sigma. Oncogene. 2014;33:4407–17.
Acknowledgements
This work was supported by AIRC 5 × 1000 call “Metastatic disease: the key unmet need in oncology” to MYNERVA project, #21267 (MYeloid NEoplasms Research Venture Airc). A detailed description of the MYNERVA project is available at http://www.progettoagimm.it; Sergio Luciani Association, Fabriano, Italy; Fondazione Cassa di Risparmio Perugia (2016.0113.021 Ricerca Scientifica e Tecnologica).
Author contributions
AGLF performed methylation and expression experiments, the analysis and wrote the article; BC and VP performed cytogenetic studies and FISH experiments; GB performed SNPa experiments and analysis; GR and VDB provided clinical–hematological data; DDG and VDB performed mutational analysis; RP and FP performed and analyzed whole exome sequencing; ERA contributed to methylation and expression analysis; MEF supervised all epigenetic experiments and contributed to data interpretation and writing; CM conceived and designed the study, contributed to data interpretation and wrote the article. All authors approved the final article.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Fernandez, A.G.L., Crescenzi, B., Pierini, V. et al. A distinct epigenetic program underlies the 1;7 translocation in myelodysplastic syndromes. Leukemia 33, 2481–2494 (2019). https://doi.org/10.1038/s41375-019-0433-9
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41375-019-0433-9