Abstract
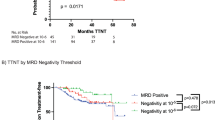
The iliac crest is the sampling site for minimal residual disease (MRD) monitoring in multiple myeloma (MM). However, the disease distribution is often heterogeneous, and imaging can be used to complement MRD detection at a single site. We have investigated patients in complete remission (CR) during first-line or salvage therapy for whom MRD flow cytometry and the two imaging modalities positron emission tomography (PET) and diffusion-weighted magnetic resonance imaging (DW-MRI) were performed at the onset of CR. Residual focal lesions (FLs), detectable in 24% of first-line patients, were associated with short progression-free survival (PFS), with DW-MRI detecting disease in more patients. In some patients, FLs were only PET positive, indicating that the two approaches are complementary. Combining MRD and imaging improved prediction of outcome, with double-negative and double-positive features defining groups with excellent and dismal PFS, respectively. FLs were a rare event (12%) in first-line MRD-negative CR patients. In contrast, patients achieving an MRD-negative CR during salvage therapy frequently had FLs (50%). Multi-region sequencing and imaging in an MRD-negative patient showed persistence of spatially separated clones. In conclusion, we show that DW-MRI is a promising tool for monitoring residual disease that complements PET and should be combined with MRD.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Avet-Loiseau H, Lauwers-Cances V, Corre J, Moreau P, Attal M, Munshi N. Minimal residual disease in multiple myeloma: final analysis of the IFM2009 Trial, Abstract 435. ASH 59th Annual Meeting & Exposition, December 2017.
Anderson KC. Should minimal residual disease negativity be the end point of myeloma therapy? Blood Adv. 2017;1:517–21.
Munshi NC, Avet-Loiseau H, Rawstron AC, Owen RG, Child JA, Thakurta A, et al. Association of minimal residual disease with superior survival outcomes in patients with multiple myeloma: a meta-analysis. JAMA Oncol. 2017;3:28–35.
Schinke C, Hoering A, Wang H, Carlton V, Thanandrarajan S, Deshpande S, et al. The prognostic value of the depth of response in multiple myeloma depends on the time of assessment, risk status and molecular subtype. Haematologica. 2017;102:e313–6.
Lahuerta JJ, Paiva B, Vidriales MB, Cordon L, Cedena MT, Puig N, et al. Depth of response in multiple myeloma: a pooled analysis of three PETHEMA/GEM clinical trials. J Clin Oncol. 2017;35:2900–10.
Landgren O, Devlin S, Boulad M, Mailankody S. Role of MRD status in relation to clinical outcomes in newly diagnosed multiple myeloma patients: a meta-analysis. Bone Marrow Transplant. 2016;51:1565–8.
Kis O, Kaedbey R, Chow S, Danesh A, Dowar M, Li T, et al. Circulating tumour DNA sequence analysis as an alternative to multiple myeloma bone marrow aspirates. Nat Commun. 2017;8:15086.
Lohr JG, Kim S, Gould J, Knoechel B, Drier Y, Cotton MJ, et al. Genetic interrogation of circulating multiple myeloma cells at single-cell resolution. Sci Transl Med. 2016;8:363ra147.
Rasche L, Chavan SS, Stephens OW, Patel PH, Tytarenko R, Ashby C, et al. Spatial genomic heterogeneity in multiple myeloma revealed by multi-region sequencing. Nat Commun. 2017;8:268.
Raab MS, Lehners N, Xu J, Ho AD, Schirmacher P, Goldschmidt H, et al. Spatially divergent clonal evolution in multiple myeloma: overcoming resistance to BRAF inhibition. Blood. 2016;127:2155–7.
Lopez-Anglada L, Gutierrez NC, Garcia JL, Mateos MV, Flores T, San Miguel JF. P53 deletion may drive the clinical evolution and treatment response in multiple myeloma. Eur J Haematol. 2010;84:359–61.
Mithraprabhu S, Khong T, Ramachandran M, Chow A, Klarica D, Mai L, et al. Circulating tumour DNA analysis demonstrates spatial mutational heterogeneity that coincides with disease relapse in myeloma. Leukemia. 2017;31:1695–705.
Moreau P, Attal M, Caillot D, Macro M, Karlin L, Garderet L, et al. Prospective evaluation of magnetic resonance imaging and [(18)F]fluorodeoxyglucose positron emission tomography-computed tomography at diagnosis and before maintenance therapy in symptomatic patients with multiple myeloma included in the IFM/DFCI 2009 trial: results of the IMAJEM study. J Clin Oncol. 2017;35:2911–8.
Pawlyn C, Fowkes L, Otero S, Jones JR, Boyd KD, Davies FE, et al. Whole-body diffusion-weighted MRI: a new gold standard for assessing disease burden in patients with multiple myeloma? Leukemia. 2016;30:1446–8.
Rasche L, Angtuaco E, McDonald JE, Buros A, Stein C, Pawlyn C, et al. Low expression of hexokinase-2 is associated with false-negative FDG-positron emission tomography in multiple myeloma. Blood. 2017;130:30–34.
Weinhold N, Heuck CJ, Rosenthal A, Thanendrarajan S, Stein CK, Van Rhee F, et al. Clinical value of molecular subtyping multiple myeloma using gene expression profiling. Leukemia. 2016;30:423–30.
Dimopoulos MA, Oriol A, Nahi H, San-Miguel J, Bahlis NJ, Usmani SZ, et al. Daratumumab, lenalidomide, and dexamethasone for multiple myeloma. N Engl J Med. 2016;375:1319–31.
Chari A, Suvannasankha A, Fay JW, Arnulf B, Kaufman JL, Ifthikharuddin JJ, et al. Daratumumab plus pomalidomide and dexamethasone in relapsed and/or refractory multiple myeloma. Blood. 2017;130:974–81.
Kumar S, Paiva B, Anderson KC, Durie B, Landgren O, Moreau P, et al. International Myeloma Working Group consensus criteria for response and minimal residual disease assessment in multiple myeloma. Lancet Oncol. 2016;17:e328–46.
Lecouvet FE. Whole-body MR imaging: musculoskeletal applications. Radiology. 2016;279:345–65.
Bartel TB, Haessler J, Brown TL, Shaughnessy JD Jr, van Rhee F, Anaissie E, et al. F18-fluorodeoxyglucose positron emission tomography in the context of other imaging techniques and prognostic factors in multiple myeloma. Blood. 2009;114:2068–76.
Waheed S, Mitchell A, Usmani S, Epstein J, Yaccoby S, Nair B, et al. Standard and novel imaging methods for multiple myeloma: correlates with prognostic laboratory variables including gene expression profiling data. Haematologica. 2013;98:71–78.
Paiva B, Cedena MT, Puig N, Arana P, Vidriales MB, Cordon L, et al. Minimal residual disease monitoring and immune profiling in multiple myeloma in elderly patients. Blood. 2016;127:3165–74.
Zhan F, Huang Y, Colla S, Stewart JP, Hanamura I, Gupta S, et al. The molecular classification of multiple myeloma. Blood. 2006;108:2020–8.
Shaughnessy J, Tian E, Sawyer J, Bumm K, Landes R, Badros A, et al. High incidence of chromosome 13 deletion in multiple myeloma detected by multiprobe interphase FISH. Blood. 2000;96:1505–11.
Greipp PR, San Miguel J, Durie BG, Crowley JJ, Barlogie B, Blade J, et al. International staging system for multiple myeloma. J Clin Oncol. 2005;23:3412–20.
Shaughnessy JD Jr, Zhan F, Burington BE, Huang Y, Colla S, Hanamura I, et al. A validated gene expression model of high-risk multiple myeloma is defined by deregulated expression of genes mapping to chromosome 1. Blood. 2007;109:2276–84.
Weinhold N, Ashby C, Rasche L, Chavan SS, Stein C, Stephens OW, et al. Clonal selection and double-hit events involving tumor suppressor genes underlie relapse in myeloma. Blood. 2016;128:1735–44.
Cibulskis K, Lawrence MS, Carter SL, Sivachenko A, Jaffe D, Sougnez C, et al. Sensitive detection of somatic point mutations in impure and heterogeneous cancer samples. Nat Biotechnol. 2013;31:213–9.
Saunders CT, Wong WS, Swamy S, Becq J, Murray LJ, Cheetham RK. Strelka: accurate somatic small-variant calling from sequenced tumor-normal sample pairs. Bioinformatics. 2012;28:1811–7.
Chen X, Schulz-Trieglaff O, Shaw R, Barnes B, Schlesinger F, Kallberg M, et al. Manta: rapid detection of structural variants and indels for germline and cancer sequencing applications. Bioinformatics. 2016;32:1220–2.
Robinson JT, Thorvaldsdottir H, Winckler W, Guttman M, Lander ES, Getz G, et al. Integrative genomics viewer. Nat Biotechnol. 2011;29:24–26.
Wang K, Li M, Hakonarson H. ANNOVAR: functional annotation of genetic variants from high-throughput sequencing data. Nucleic Acids Res. 2010;38:e164.
Cingolani P, Platts A, Wang le L, Coon M, Nguyen T, Wang L, et al. A program for annotating and predicting the effects of single nucleotide polymorphisms, SnpEff: SNPs in the genome of Drosophila melanogaster strain w1118; iso-2; iso-3. Fly (Austin). 2012;6:80–92.
Favero F, Joshi T, Marquard AM, Birkbak NJ, Krzystanek M, Li Q, et al. Sequenza: allele-specific copy number and mutation profiles from tumor sequencing data. Ann Oncol. 2015;26:64–70.
Zamagni E, Nanni C, Mancuso K, Tacchetti P, Pezzi A, Pantani L, et al. PET/CT improves the definition of complete response and allows to detect otherwise unidentifiable skeletal progression in multiple myeloma. Clin Cancer Res. 2015;21:4384–90.
Davies FE, Rosenthal A, Rasche L, Petty NM, McDonald JE, Ntambi JA, et al. Treatment to suppression of focal lesions on positron emission tomography-computed tomography is a therapeutic goal in newly diagnosed multiple myeloma. Haematologica. 2018;103:1047–53.
Koh DM, Collins DJ. Diffusion-weighted MRI in the body: applications and challenges in oncology. AJR Am J Roentgenol. 2007;188:1622–35.
Provenzale JM, Engelter ST, Petrella JR, Smith JS, MacFall JR. Use of MR exponential diffusion-weighted images to eradicate T2 “shine-through” effect. AJR Am J Roentgenol. 1999;172:537–9.
Rasche L, Buros A, Weinhold N, Stein CK, McDonald JE, Chavan SS, et al. The clinical impact of macrofocal disease in multiple myeloma differs between presentation and relapse. Blood. 2016;128:4431.
Landgren O, Lu SX, Hultcrantz M. MRD testing in multiple myeloma: the main future driver for modern tailored treatment. Semin Hematol. 2018;55:44–50.
Anderson KC, Auclair D, Kelloff GJ, Sigman CC, Avet-Loiseau H, Farrell AT, et al. The role of minimal residual disease testing in myeloma treatment selection and drug development: current value and future applications. Clin Cancer Res. 2017;23:3980–93.
Hillengass J, Merz M, Delorme S. Minimal residual disease in multiple myeloma: use of magnetic resonance imaging. Semin Hematol. 2018;55:19–21.
Pandit-Taskar N. Functional imaging methods for assessment of minimal residual disease in multiple myeloma: current status and novel immunoPET based methods. Semin Hematol. 2018;55:22–32.
Messiou C, Giles S, Collins DJ, West S, Davies FE, Morgan GJ, et al. Assessing response of myeloma bone disease with diffusion-weighted MRI. Br J Radiol. 2012;85:e1198–1203.
Rasche L, Angtuaco EJ, Alpe TL, Gershner GH, McDonald JE, Samant RS, et al. The presence of large focal lesions is a strong independent prognostic factor in multiple myeloma. Blood. 2018;132:59–66.
Padhani AR, van Ree K, Collins DJ, D’Sa S, Makris A. Assessing the relation between bone marrow signal intensity and apparent diffusion coefficient in diffusion-weighted MRI. AJR Am J Roentgenol. 2013;200:163–70.
Messiou C, Collins DJ, Morgan VA, Desouza NM. Optimising diffusion weighted MRI for imaging metastatic and myeloma bone disease and assessing reproducibility. Eur Radiol. 2011;21:1713–8.
Roshal M. Minimal residual disease detection by flow cytometry in multiple myeloma: why and how? Semin Hematol. 2018;55:4–12.
Flores-Montero J, Sanoja-Flores L, Paiva B, Puig N, Garcia-Sanchez O, Bottcher S, et al. Next generation flow for highly sensitive and standardized detection of minimal residual disease in multiple myeloma. Leukemia. 2017;31:2094–103.
Lapa C, Garcia-Velloso MJ, Luckerath K, Samnick S, Schreder M, Otero PR, et al. 11C-methionine-PET in multiple myeloma: a combined study from two different institutions. Theranostics. 2017;7:2956–64.
Caserta E, Chea J, Minnix M, Viola D, Vonderfecht S, Yazaki P, et al. Copper 64-labeled daratumumab as a PET/CT imaging tracer for multiple myeloma. Blood. 2018;131:741–5.
https://www.adaptivebiotech.com/clonoseq/clonoseq-assay. ClonoSEQ Revealing Previously Unseen Disease.
Acknowledgements
We thank the patients and staff of the Myeloma Institute, UAMS. We also thank the Department of Radiology, UAMS. This work was supported by P01 CA 55819 from the National Cancer Institute. LR was supported by the Deutsche Forschungsgemeinschaft (DFG). NW was supported by the National Institute of General Medical Sciences of the National Institutes of Health under Award Number P20GM125503.
Author contributions
Conception and design: NW, LR, GJM. Provision of study material or patients: DA, GJM, BB, FvR, MZ, ST, CS, FED, JE, AFW. Reporting imaging: MK, JM, RS, RVH. Data analysis: NW, LR, GG, DA, CA, MB, CPW, BAW. Wrote the paper: LR, NW, GJM. Reviewed and approved the paper: All authors.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
BB is a co-inventor on patents and patent applications related to use of GEP in cancer medicine that have been licensed to Quest diagnostics. The other authors declare that they have no conflict of interest.
Supplementary information
Rights and permissions
About this article
Cite this article
Rasche, L., Alapat, D., Kumar, M. et al. Combination of flow cytometry and functional imaging for monitoring of residual disease in myeloma. Leukemia 33, 1713–1722 (2019). https://doi.org/10.1038/s41375-018-0329-0
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41375-018-0329-0
This article is cited by
-
Implications and prognostic impact of mass spectrometry in patients with newly-diagnosed multiple myeloma
Blood Cancer Journal (2023)
-
Dual assessment with multiparameter flow cytometry and 18F-FDG PET/CT scan provides enhanced prediction of measurable residual disease after autologous haemopoietic stem cell transplant in myeloma—a prospective study
Bone Marrow Transplantation (2023)
-
The spatio-temporal evolution of multiple myeloma from baseline to relapse-refractory states
Nature Communications (2022)
-
Characteristics of measurable residual disease assessment in myeloma: a review of clinical trials from 2015–2020
Blood Cancer Journal (2022)
-
Progress of modern imaging modalities in multiple myeloma
International Journal of Hematology (2022)