Abstract
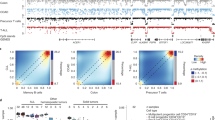
Acute myeloid leukemia (AML) often harbors mutations in epigenetic regulators, and also has frequent DNA hypermethylation, including the presence of CpG island methylator phenotypes (CIMPs). Although global hypomethylation is well known in cancer, the question of whether distinct demethylator phenotypes (DMPs) exist remains unanswered. Using Illumina 450k arrays for 194 patients from The Cancer Genome Atlas, we identified two distinct DMPs by hierarchical clustering: DMP.1 and DMP.2. DMP.1 cases harbored mutations in NPM1 (94%), FLT3 (71%) and DNMT3A (61%). Surprisingly, only 40% of patients with DNMT3A mutations were DMP.1, which has implications for mechanisms of transformation by this mutation. In contrast, DMP.2 AML was comprised of patients with t(8;21), inv(16) or t(15;17), suggesting common methylation defects connect these disparate rearrangements. RNA-seq revealed upregulated genes functioning in immune response (DMP.1) and development (DMP.2). We confirmed these findings by integrating independent 450k data sets (236 additional cases), and found prognostic effects by DMP status, independent of age and cytogenetics. The existence of DMPs has implications for AML pathogenesis and may augment existing tools in risk stratification.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Ehrlich M, Gama-Sosa MA, Huang LH, Midgett RM, Kuo KC, McCune RA, et al. Amount and distribution of 5-methylcytosine in human DNA from different types of tissues of cells. Nucleic Acids Res. 1982;10:2709–21.
Razin A, Riggs AD. DNA methylation and gene function. Science. 1980;210:604–10.
Toyota M, Ahuja N, Ohe-Toyota M, Herman JG, Baylin SB, Issa JP. CpG island methylator phenotype in colorectal cancer. Proc Natl Acad Sci USA. 1999;96:8681–6.
Toyota M, Ahuja N, Suzuki H, Itoh F, Ohe-Toyota M, Imai K, et al. Aberrant methylation in gastric cancer associated with the CpG island methylator phenotype. Cancer Res. 1999;59:5438–42.
Hill VK, Shinawi T, Ricketts CJ, Krex D, Schackert G, Bauer J, et al. Stability of the CpG island methylator phenotype during glioma progression and identification of methylated loci in secondary glioblastomas. BMC Cancer. 2014;14:506.
Kelly AD, Kroeger H, Yamazaki J, Taby R, Neumann F, Yu S, et al. A CpG island methylator phenotype in acute myeloid leukemia independent of IDH mutations and associated with a favorable outcome. Leukemia. 2017;31:2011–9.
van den Bent MJ, Gravendeel LA, Gorlia T, Kros JM, Lapre L, Wesseling P, et al. A hypermethylated phenotype is a better predictor of survival than MGMT methylation in anaplastic oligodendroglial brain tumors: a report from EORTC study 26951. Clin Cancer Res. 2011;17:7148–55.
Hegi ME, Diserens AC, Godard S, Dietrich PY, Regli L, Ostermann S, et al. Clinical trial substantiates the predictive value of O-6-methylguanine-DNA methyltransferase promoter methylation in glioblastoma patients treated with temozolomide. Clin Cancer Res. 2004;10:1871–4.
Feinberg AP, Vogelstein B. Hypomethylation distinguishes genes of some human cancers from their normal counterparts. Nature. 1983;301:89–92.
Cancer Genome Atlas Research Network. Genomic and epigenomic landscapes of adult de novo acute myeloid leukemia. N Engl J Med. 2013;368:2059–74.
Ferreira HJ, Heyn H, Vizoso M, Moutinho C, Vidal E, Gomez A, et al. DNMT3A mutations mediate the epigenetic reactivation of the leukemogenic factor MEIS1 in acute myeloid leukemia. Oncogene. 2015;36:4233.
Yang L, Rodriguez B, Mayle A, Park HJ, Lin X, Luo M, et al. DNMT3A loss drives enhancer hypomethylation in FLT3-ITD-associated leukemias. Cancer Cell. 2016;29:922–34.
Figueroa ME, Abdel-Wahab O, Lu C, Ward PS, Patel J, Shih A, et al. Leukemic IDH1 and IDH2 mutations result in a hypermethylation phenotype, disrupt TET2 function, and impair hematopoietic differentiation. Cancer Cell. 2010;18:553–67.
Yamazaki J, Taby R, Jelinek J, Raynal NJ, Cesaroni M, Pierce SA, et al. Hypomethylation of TET2 Target Genes Identifies a Curable Subset of Acute Myeloid Leukemia. J Natl Cancer Inst. 2016;108:2.
Barrett T, Wilhite SE, Ledoux P, Evangelista C, Kim IF, Tomashevsky M, et al. NCBI GEO: archive for functional genomics data sets--update. Nucleic Acids Res. 2013;41(database issue):D991–5.
Fortin JP, Labbe A, Lemire M, Zanke BW, Hudson TJ, Fertig EJ, et al. Functional normalization of 450k methylation array data improves replication in large cancer studies. Genome Biol. 2014;15:503.
Qu Y, Lennartsson A, Gaidzik VI, Deneberg S, Karimi M, Bengtzén S, et al. Differential methylation in CN-AML preferentially targets non-CGI regions and is dictated by DNMT3A mutational status and associated with predominant hypomethylation of HOX genes. Epigenetics. 2014;9:1108–19.
Cauchy P, James SR, Zacarias-Cabeza J, Ptasinska A, Imperato MR, Assi SA, et al. Chronic FLT3-ITD signaling in acute myeloid leukemia is connected to a specific chromatin signature. Cell Rep. 2015;12:821–36.
Farrar JE, Schuback HL, Ries RE, Wai D, Hampton OA, Trevino LR, et al. Genomic profiling of pediatric acute myeloid leukemia reveals a changing mutational landscape from disease diagnosis to relapse. Cancer Res. 2016;76:2197–205.
Reinius LE, Acevedo N, Joerink M, Pershagen G, Dahlén SE, Greco D, et al. Differential DNA methylation in purified human blood cells: implications for cell lineage and studies on disease susceptibility. PLoS ONE. 2012;7:e41361.
Karolchik D, Hinrichs AS, Furey TS, Roskin KM, Sugnet CW, Haussler D, et al. The UCSC Table Browser data retrieval tool. Nucleic Acids Res. 2004;32(database issue):D493–6.
Rosenbloom KR, Sloan CA, Malladi VS, Dreszer TR, Learned K, Kirkup VM, et al. ENCODE data in the UCSC Genome Browser: year 5 update. Nucleic Acids Res. 2013;41(database issue):D56–63.
Smit A, Hubley R, Green P. RepeatMasker at http://repeatmasker.org. (cited 2017).
Robinson MD, McCarthy DJ, Smyth GK. edgeR: a Bioconductor package for differential expression analysis of digital gene expression data. Bioinformatics. 2010;26:139–40.
McCarthy DJ, Chen Y, Smyth GK. Differential expression analysis of multifactor RNA-Seq experiments with respect to biological variation. Nucleic Acids Res. 2012;40:4288–97.
Nogales-Cadenas R, Carmona-Saez P, Vazquez M, Vicente C, Yang X, Tirado F, et al. GeneCodis: interpreting gene lists through enrichment analysis and integration of diverse biological information. Nucleic Acids Res. 2009;37(Web Server issue):W317–322.
Carmona-Saez P, Chagoyen M, Tirado F, Carazo JM, Pascual-Montano A. GENECODIS: a web-based tool for finding significant concurrent annotations in gene lists. Genome Biol. 2007;8:R3.
Tabas-Madrid D, Nogales-Cadenas R, Pascual-Montano A. GeneCodis3: a non-redundant and modular enrichment analysis tool for functional genomics. Nucleic Acids Res. 2012;40(Web Server issue):W478–83.
R Core Development Team. R: a language and environment for statistical computing. Vienna, Austria: R Foundation for Statistical Computing; 2014.
Therneau T. A package for survival analysis in S. R package version 2.37-7 ed; 2014.
Itonaga H, Imanishi D, Wong YF, Sato S, Ando K, Sawayama Y, et al. Expression of myeloperoxidase in acute myeloid leukemia blasts mirrors the distinct DNA methylation pattern involving the downregulation of DNA methyltransferase DNMT3B. Leukemia. 2014;28:1459–66.
Hock H, Hamblen MJ, Rooke HM, Traver D, Bronson RT, Cameron S, et al. Intrinsic requirement for zinc finger transcription factor Gfi-1 in neutrophil differentiation. Immunity. 2003;18:109–20.
Skokowa J, Cario G, Uenalan M, Schambach A, Germeshausen M, Battmer K, et al. LEF-1 is crucial for neutrophil granulocytopoiesis and its expression is severely reduced in congenital neutropenia. Nat Med. 2006;12:1191–7.
Yamazaki J, Jelinek J, Lu Y, Cesaroni M, Madzo J, Neumann F, et al. TET2 mutations affect non-CpG island DNA methylation at enhancers and transcription factor-binding sites in chronic myelomonocytic leukemia. Cancer Res. 2015;75:2833–43.
Glass JL, Hassane D, Wouters BJ, Kunimoto H, Avellino R, Garrett-Bakelman FE, et al. Epigenetic identity in AML depends on disruption of nonpromoter regulatory elements and is affected by antagonistic effects of mutations in epigenetic modifiers. Cancer Discov. 2017;7:868–83.
Meazza R, Basso S, Gaggero A, Detotero D, Trentin L, Pereno R, et al. Interleukin (IL)-15 induces survival and proliferation of the growth factor-dependent acute myeloid leukemia M-07e through the IL-2 receptor beta/gamma. Int J Cancer. 1998;78:189–95.
Kittang AO, Hatfield K, Sand K, Reikvam H, Bruserud Ø. The chemokine network in acute myelogenous leukemia: molecular mechanisms involved in leukemogenesis and therapeutic implications. Curr Top Microbiol Immunol. 2010;341:149–72.
Acknowledgements
We would like to thank Drs. Humberto Ferreira and Manel Esteller for providing additional clinical data to supplement their 450k GEO submission.
Funding
This work was supported by National Institutes of Health grants R01CA158112 and P50CA100632. J-PJI is an American Cancer Society Clinical Research Professor supported by a generous gift from the FM Kirby Foundation.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Electronic supplementary material
Rights and permissions
About this article
Cite this article
Kelly, A.D., Madzo, J., Madireddi, P. et al. Demethylator phenotypes in acute myeloid leukemia. Leukemia 32, 2178–2188 (2018). https://doi.org/10.1038/s41375-018-0084-2
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41375-018-0084-2
This article is cited by
-
DNA hypomethylation patterns and their impact on the tumor microenvironment in colorectal cancer
Cellular Oncology (2024)