Abstract
One of the pathological hallmarks of amyotrophic lateral sclerosis (ALS) is mislocalized, cytosolic aggregation of TAR DNA-Binding Protein-43 (TDP-43). Not only TDP-43 per se is a causative gene of ALS but also mislocalization and aggregation of TDP-43 seems to be a common pathological change in both sporadic and familial ALS. The mechanism how nuclear TDP-43 transforms into cytosolic aggregates remains elusive, but recent studies using optogenetics have proposed that aberrant liquid–liquid phase separation (LLPS) of TDP-43 links to the aggregation process, leading to cytosolic distribution. Although LLPS plays an important role in the aggregate formation, there are still several technical problems in the optogenetic technique to be solved to progress further in vivo study. Here we report a chemically oligomerizable TDP-43 system. Oligomerization of TDP-43 was achieved by a small compound AP20187, and oligomerized TDP-43 underwent aggregate formation, followed by cytosolic mislocalization and induction of cell toxicity. The mislocalized TDP-43 co-aggregated with wt-TDP-43, Fused-in-sarcoma (FUS), TIA1 and sequestosome 1 (SQSTM1)/p62, mimicking ALS pathology. The chemically oligomerizable TDP-43 also revealed the roles of the N-terminal domain, RNA-recognition motif, nuclear export signal and low complexity domain in the aggregate formation and mislocalization of TDP-43. The aggregate-prone properties of TDP-43 were enhanced by a familial ALS-causative mutation. In conclusion, the chemically oligomerizable TDP-43 system could be useful to study the mechanisms underlying the droplet-aggregation phase transition and cytosolic mislocalization of TDP-43 in ALS and further study in vivo.
Similar content being viewed by others
Introduction
Amyotrophic lateral sclerosis (ALS) is a progressive motor neuron disease affecting more than 200,000 people worldwide [1]. Over 30 causative genes have been identified as causes of familial ALS including TAR DNA-binding protein-43 (TDP-43) which is a cause of familial ALS10 occupying 3-4% of familial ALS [2, 3]. The involvement of TDP-43 in the pathogenesis of ALS is also apparent due to cytosolic TDP-43-positive aggregates seen in most cases of both familial and sporadic ALS, and it is one of hallmarks of ALS pathology [4, 5]. TDP-43 is a highly conserved DNA/RNA-binding protein widely expressed [3]. TDP-43 is involved in a variety of biological processes including RNA transcription [6], RNA splicing [7,8,9], RNA editing [10], formation of nuclear bodies [11], regulation of stress granules [12], and DNA repair [13, 14], and its functional disturbance is supposed to trigger neuronal cell death [9, 15]. It has been shown that TDP-43 is prone to self-associate via its N-terminal domain as well as C-terminal low complexity domain (LCD) [16,17,18,19]. However, the mechanisms how nuclear TDP-43 transforms into cytosolic aggregates have remained elusive due to the technological difficulty in the regulation of oligomerization in live cells. The breakthrough to control oligomerization was achieved by optogenetic technology using the Arabidopsis photoreceptor cryptochrome 2 (CRY2) [20]. By modification and optimization of the oligomer-promoting nature of wt-CRY2, CRY2olig becomes a powerful tool to investigate the biochemical behavior of protein of interest upon clustering by light. Fusing this optogenetic module to TDP-43 enables control of oligomerization status of TDP-43 in live cells and it has revealed that abnormal phase separation of TDP-43 linked to aggregate formation followed by cytosolic mislocalization [21, 22]. However, the Cry2-olig system is still not a perfect system for inducing and monitoring of TDP-43 phase separation in several aspects. First, it is impossible to selectively induce dimer and oligomer which undergoes clustering, thus it is difficult to assess the role of intrinsic domains which harbor self-associating property, participating in the phase-separation processes. Second, it is challenging to induce Cry2olig-mediated clustering in whole body of mammalian ALS models due to absorbance of blue-green light by hemoglobin [23]. Third, light-mediated photoactivation is applicable for limited number of cells and simultaneous activation of large quantity of cells requires special equipment [24]. Fourth, longer irradiation of blue light might cause photo-toxicity [25]. To conquer these scientific problems, we developed a chemically inducible oligomerization system using the dimer domain of FK506 binding protein (FKBP)-F36V mutant and its homodimerizing ligand, AP20187 [26,27,28] to study the role of LLPS in TDP-43 aggregate formation and subsequent localization changes. Addition of AP20187 induced aggregate formation of TDP-43 followed by mislocalization from nucleus, indicating its usefulness for studying molecular mechanisms underlying TDP-43 mislocalization seen in ALS.
Material and methods
Cell culture
HeLa cells, a motor neuronal cell line NSC34 or HEK293 cells were cultured in Dulbecco’s-modified Eagle Medium (DMEM) supplemented with 10% fetal bovine serum and penicillin/streptomycin. The cells were cultured in a standard incubator at 37 °C with 5% CO2. Transfection of plasmids was done with LipofectAmine2000 (ThermoFisher) following the manufacturer’s protocol.
Plasmid constructions
Yellow fluorescent protein (YFP)-conjugated TDP-43 was obtained from Addgene (#84911). YFP was PCR amplified and subcloned into pcDNA3.1 vector to create the YFP control. For constructing D1-YFP, D2-YFP, D1-TDP-43-YFP, and D2-TDP-43-YFP, one or two tandem FKBPF36V sequences were inserted into the N-terminus of YFP or TDP-43-YFP. mCherry-Sequestosome1 (SQSTM1)-N-18 was obtained from Addgene (#55132). TDP-43-mCherry was constructed by subcloning of TDP-43 into mCherry2-N1 vector (#54517, Addgene). TDP-43 deletion mutants were constructed by the PCR-based mutagenesis with following primer sets: a sense primer (AAAAGAAAAATGGATGAGACAGATGCTTCATCAGC) and an anti-sense primer (GCGGCCGCTTTCCAGTTTTAGAAG) for TDP-43-ΔN; a sense primer (AAGCACAATAGCAATAGACAG) and an anti-sense primer (GCTTCTCAAAGGCTCATC) for TDP-43-ΔRRM2; a sense primer (ATTGCGCAGTCTCTTTGTG) and an anti-sense primer (GCTTCTCAAAGGCTCATC) for TDP-43-ΔRRM2 w/NES; a sense primer (AAGCTTATCGATAGCAAGGGCGAG) and an anti-sense primer (TCTTTCTAACTGTCTATTGCTATTGTGC) for TDP-43-ΔLCD, respectively. The Q331K mutation was created by a site-directed mutagenesis with a sense primer (AAGAGCAGTTGGGGTATGATGGGCATGTTAG) and an anti-sense primer (TAGTGCTGCCTGGGCGGCAGCCATCATG). FUS cDNA was a kind gift from Dr. Masaaki Matsuoka (Tokyo Medical University). TIA1 cDNA was PCR amplified from HeLa cell cDNA with a sense primer (ATGGAGGACGAGATGCCC) and an anti-sense primer (TCACTGGGTTTCATACCCTGCC).
Induction of oligomerization
Oligomerization of YFP or TDP-43 was achieved by the addition of AP20187 (Clontech, 635059) into the culture medium of transfected cells at 0.5 μM otherwise mentioned, and the cells were incubated in the presence of AP20187 for the indicated time period.
Confocal imaging
HeLa cells, NSC34 cells, or HEK293 cells were plated onto a chambered slide glass (Matsunami glass). After transfection followed by induction of oligomerization, the cells were fixed with 4%-formalin-PBS for 15 min at room temperature. The nuclei were counterstained with DAPI, and then the cells were mounted with VECTASHIELD antifade mounting medium (Vectorlabs). Confocal imaging for fixed cells was performed with FV-10i (Olympus). The image analysis was performed with ImageJ software.
FRAP analysis
HeLa cells were plated onto a chambered coverglass (Matsunami glass). After transfection followed by induction of oligomerization, the cells were observed by LSM710 confocal microscopy (Zeiss). FRAP analysis was performed with ZEN software (Zeiss). For the washout experiments, HeLa cells expressing TDP-43 variants were treated with AP20187 for 24 h, followed by incubation for another 24 h of washout period.
Gel-filtration analysis
To evaluate oligomerization status of TDP-43, Chroma spin column-based gel-filtration analysis was performed as previously described [29, 30]. Briefly, HEK293 cells overexpressing D1-TDP-43-YFP or D2-TDP-43-YFP were treated with 0.5 μM AP20187 for 6 h. After the treatment, the cells were harvested and lysed in the lysis buffer (150 mM NaCl, 20 mM Tris-HCl pH7.4, 0.5% Nonidet-P40 and protease inhibitor cocktail). The lysates were applied to a Chroma spin column followed by sequential centrifugation, and 14 fractions were collected and analyzed by Western blotting analysis.
Fractionation into nuclear and cytosolic fractions
Fractionation of nuclear and cytosolic fractions from HEK293 cells expressing D1-TDP-43-YFP or D2-TDP-YFP treated with 0.5 μM AP20187 for 2 h or 6 h was performed with ProteoExtract subcellular proteome extraction kit (Millipore), following the manufacture’s protocol. After fractionation, the lysates were crosslinked by dimethyl pimelimidate (DMP) (ThermoFisher) before Western blot analysis.
Fractionation into soluble and insoluble fractions
HEK293 cells expressing D1-YFP, D2-YFP, D1-TDP-43-YFP or D2-TDP-43-YFP, cultured in the presence or absence of 0.5 μM AP20187 for 24 h, were harvested for lysis with the RIPA buffer (10 mM Tris-HCl, 1% Nonidet-P40, 0.1% sodium deoxycholate, 0.1% SDS, 0.15 M NaCl, 1 mM EDTA and protease inhibitor cocktails). The soluble fraction was defined as the supernatant fraction after centrifugation at 15000 rpm for 15 min. After removal of supernatant, the pellets were resuspended in RIPA Buffer and re-centrifuged. The supernatants were discarded and the residual pellets were resuspended in the Urea lysis buffer (7 M Urea, 30 mM Tris-HCl, 0.5% Nonidet-P40 and protease inhibitor cocktails) followed by sonication. The solutions were centrifugated at 15000 rpm for 15 min and the pellets were defined as the insoluble fractions. The soluble and insoluble fractions were then analyzed by Western blot analysis.
Western blotting
The samples were mixed with equal volume of 2x Laemmli SDS-PAGE sample buffer (Biorad) and boiled at 95 °C for 5 min. The samples were next subjected to SDS-PAGE and blotted onto Immobilon-P polyvinylidene fluoride (PVDF) membrane (Millipore). Immunoblotting was performed by a standard protocol, and immunoreactive signals were detected by ChemiDoc Touch (Biorad) using ECL-select detection reagent (GE Amersham). The following antibodies were used; anti-TDP-43 antibody (Proteintech: 10782-2-AP); anti-GFP antibody (Cell Signaling: #2956); anti-GAPDH antibody (Cell Signaling: #2118).
Immunostaining
HEK293 cells expressing D2-YFP, D1-TDP-43-YFP, or D2-TDP-43-YFP were treated with 0.5 μM AP20187 for 24 h and then fixed by formalin-PBS. The cells were permeabilized by 0.5% Triton-X100 and blocking was performed with Blocking-one reagent (Nacalai). Immunostaining was performed with anti-cleaved poly(ADP-ribose) polymerase-1 (PARP) antibody (Cell Signaling: #5625).
Statistics
All data are represented as mean ± standard deviation. Statistical analysis of the data was performed with SPSS software 26 (IBM).
Results
To precisely control of oligomerization status of TDP-43 in live cells, we adopted and modified the chemically inducible dimerization (CID) system consisting of the dimer domain of FK506 binding protein (FKBP)-F36V mutant and its homodimerizing ligand, AP20187 (Fig. 1A, B) [26,27,28]. AP20187 is a cell-permeable, dumbbell-shaped chemical compound which can connect two FKBP dimer domains, inducing homodimerization. AP20187 is also known to penetrate blood–brain barrier without significant toxicity, substantiating its usefulness in mammalian neurodegenerative disease models [31]. We designed constructs of YFP and TDP-43-YFP harboring one or two dimer domains (D1- or D2-YFP and D1- or D2-TDP-43-YFP, respectively) (Fig. 1C). Theoretically, D1 constructs can form monomer or dimer, and D2 constructs can form monomer, dimer, or oligomer under the existence of AP20187 (Fig. 1D).
A Scheme of chemically inducible dimerization (CID) system using the dimer domain of FKBP-F36V and AP20187. The dimer domain fused to protein of interest triggers dimerization upon binding to AP20187. B The dumbbell-shaped structure of AP20187. C Schematic structures of constructs used in this study. D1: one dimer domain, D2: two dimer domains. D Theoretically possible dimerization and oligomerization of each construct.
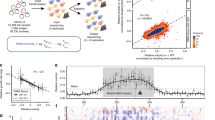
Next, we tested if this system works as designed in live cells. We introduced the D2 constructs in HEK293 cells, followed by induction of oligomerization by adding AP20187. The D2-YFP localized to cytosol under the existence of AP20187 in most cells, but less than 10% of cells contained cytosolic droplet-like aggregates of D2-YFP (Fig. 2A). On the other hand, most of D2-TDP-43-YFP formed cytosolic aggregates upon AP20187 treatment (Fig. 2B). To evaluate the oligomerization status of TDP-43 with or without AP20187, we performed a gel-filtration analysis. The gel-filtration analysis indicated that treatment with AP20187 increase the proportion of TDP-43-YFP in the high molecular weight fractions (Fig. 2C, D). Importantly, both D1-TDP-43-YFP and D2-TDP-43-YFP showed a similar distribution in the fractions, suggesting that D1-TDP-43-YFP also formed oligomer by AP20187. Next, we tested the effect of oligomerization on the cellular distribution of D1-TDP-43-YFP and D2-TDP-43-YFP. Under normal condition, D1-YFP and D2-YFP diffusely localized to the nucleus and cytosol. On the other hand, D1-TDP-43-YFP and D2-TDP-43-YFP localized to nucleus when overexpressed in HEK293 cells (Fig. 3A, B). Because endogenous TDP-43 is known to localize to the nucleus, we could confirm that tagging TDP-43 with D1 or D2 did not affect its subcellular localization. Theoretically, D1-YFP would only form dimer by AP20187. The addition of AP20187 did not affect the localization D1-YFP even 24 h after treatment (Fig. 3A, upper panels). Upon treatment with AP20187 for 4 h, most of D2-YFP diffusely localized to cytosol, but some cells began to form round-shaped condensates in nucleus and cytosol (Fig. 3A, lower panels). Twenty-four hour after treatment, some cells formed large round droplets of D2-YFP in cytosol (Fig. 3A, lower panels). On the other hand, D1-TDP-43-YFP which was expected to exclusively form dimer, condensed in nucleus 2 h after treatment, began leaking from nucleus to cytosol at 4 h, and eventually aggregated in cytosol 24 h after the treatment (Fig. 3B, upper panels). D2-TDP-43-YFP also condensed in nucleus at 2 h, started leaking from the nucleus at 4 h and aggregated 24 h after the treatment (Fig. 3B, lower panels). Titration with different dose of AP20187 revealed that D2-TDP-43-YFP was more prone to aggregate and mislocalize than D1-TDP-43-YFP (Fig. 3C). Furthermore, a familial ALS-associated mutant, Q331K, enhanced the aggregate-prone properties (Fig. 3C) [32]. Line plot profiles also showed that both D1-TDP-43-YFP and D2-TDP-43-YFP were depleted from the nucleus, and the degree of depletion was higher in D2-TDP-43-YFP after 24 h of incubation with AP20187 (Fig. 3D). The aggregate formation and mislocalization of TDP-43 by induction of oligomerization were also confirmed in a motor neuronal cell line, NSC34 cells (Fig. 3E). In NSC34 cells, we could not find D2-YFP aggregates. These observations were further substantiated by the time-course fractionation followed by Western blot analyses. In the nucleus, AP20187 increased oligomeric TDP-43 and decreased monomeric TDP-43, and in cytosol, AP20187 increased oligomeric TDP43 in a time-dependent manner (Fig. 3F). Importantly, monomeric TDP-43 was not detected in the cytosol even after the treatment, indicating that oligomeric TDP-43 was specifically mislocalized.
A, B Confocal imaging of HEK293 cells expressing D2-YFP (A) or D2-TDP-43-YFP (B) treated with 0.5 μM AP20187 for 6 h. C, D Gel-filtration analysis followed by immunoblot of D1-TDP 43- YFP (C) or D2-TDP-43-YFP (D) in the presence or absence of AP20187. Endo. TDP43: endogenous TDP-43. The scale bar shows 10 μm.
A Time-course confocal imaging of D1-YFP and D2-YFP in the presence of 0.5 μM AP20187. The YFP signals were merged to nuclear staining by DAPI. B Time-course confocal imaging of D1-TDP-43-YFP and D2-TDP-43-YFP in the presence of AP20187. C Titration of concentration of AP20187 on subcellular localization of D1-wt-TDP-43-YFP, D1-Q331K-TDP-43-YFP, D2-wt-TDP-43-YFP, and D2-Q331K-TDP-43-YFP. The cells were treated with indicated concentrations of AP20187 for 6 h. N = 150 cells/condition. D The line profile along the white lines were shown in (B). The orange lines indicate the signal of YFP along the white line and the blue lines indicate the signal of DAPI staining. E Confocal imaging of NSC34 cells expressing D2-YFP, D1-TDP-43-YFP, or D2-TDP-43-YFP treated with 0.5 μM AP20187 for 6 h. F Time-course fractionation into cytosolic and nuclear fractions followed by Western blot analyses of D1-TDP-43-YFP or D2-TDP-43-YFP treated with 0.5 μM AP20187 for indicated periods. The samples were crosslinked with DMP before application to SDS-PAGE. GAPDH was used as a loading control. The scale bar shows 10 μm.
Although D1-YFP did not change its diffuse distribution all over the nucleus and cytoplasm in the presence of AP20187, D1-TDP-43-YFP formed aggregates and mislocalized from nucleus, similar to D2-TDP-43-YFP (Fig. 3A, B). The discrepancy of the oligomeric status between D1-YFP and D1-TDP-43-YFP prompted us to speculate that the oligomerization-prone domain of TDP-43 triggers oligomerization of D1-TDP-43-YFP. Several functional domains in TDP-43 are reportedly play roles in the oligomerization of TDP-43; N-terminal domain (NTD) of TDP-43 plays a central role in homodimerization of TDP-43; [17,18,19] binding to RNA via RRM2 domain has inhibitory effects on aggregate formation; [21] C-terminal LCD is responsible for LLPS [16]. To clarify the role of these domains in the aggregate formation of TDP-43, we made a series of deletion mutants and examined their aggregate formation and subcellular localization. In this series of experiments, we used D1-TDP-43-YFP constructs. Deletion of NTD diminished aggregate formation seen in D1-TDP-43-YFP (Fig. 4A, B). On the other hand, TDP-43 lacking RRM2 resulted in the aggregate formation, but the aggregates stayed in the nucleus (Fig. 4C). The RRM2 domain of TDP-43 contains nuclear export signal (NES)[33], and adding the NES to the mutant shifted the localization of TDP-43 aggregates from nucleus to cytosol, indicating that the NES is essential for exporting D1-TDP-43 aggregates from the nucleus (Fig. 4D). Removal of LCD from TDP-43 diminished the aggregate formation of D1-TDP-43-YFP (Fig. 4E). Deletion of either NTD or LCD resulted in the loss of aggregate formation, suggesting their roles in aggregate formation are distinct and not redundant.
A The structures of TDP-43-YFP and its mutants. NLS: Nuclear localizing signal; RRM1: RNA-recognition motif-1; RRM2: RNA recognition motif-2; LCD: Low complexity domain. B–E Effects of deletion of N-terminal domain (ΔN) (B); deletion of RRM2 (ΔRRM2) (C); deletion of RRM2 rescued with NES (ΔRRM2-w/NES) (D); or deletion of LCD (ΔLCD) (E) on aggregate formation and nuclear depletion of D1-TDP-43-YFP. F–I Incorporation of TDP-43-mCherry into the chemically induced aggregates. The HeLa cells expressing indicated constructs were treated with 0.5 μM of AP20187 for 6 h for (B–E) and 24 h for (F–I). The scale bar shows 10 μm.
In the residual motor neurons in the spinal cord of ALS patients with mutations in TDP-43, complete depletion of nuclear TDP-43 is often observed even though the patients harbor one allele of mutant TDP-43 [4]. This suggests that when a part of TDP-43 obtains aggregate-prone feature, it may become seeds for further aggregation of soluble TDP-43 [34, 35]. To examine if wt-TDP-43 is incorporated into the chemically induced TDP-43 aggregates, we overexpressed mCherry-tagged wt-TDP-43 with D1 or D2 constructs. Wt-TDP-43 was incorporated into both D1-TDP-43-YFP or D2-TDP-43-YFP aggregates, but it was not detected in the D2-YFP condensates, indicating that chemically oligomerizable TDP-43 is capable of trapping coexpressed wt-TDP-43 into the aggregates (Fig. 4F–I). Recently, a number of studies reported that LLPS of TDP-43 play a pivotal role in the aggregate formation and mislocalization [16, 17, 19, 21, 36]. In those studies, cytosolic TDP-43 condensates still contain some mobile fractions which evidences these condensates harbor liquid-like features. To clarify if chemically induced oligomer of TDP-43-YFP have liquid-like properties, we performed fluorescence recovery after photo-bleaching (FRAP) analysis. Without AP20187, both D1-TDP-43-YFP and D2-TDP-43-YFP diffusely localized to the nucleus, and their prompt FRAP recovery indicated their mobility in the nucleus was high (Fig. 5A–D). Upon AP20187 treatment, their localization shifted from nucleus to cytosol, and their FRAP recovery rate was strongly suppressed (Fig. 5A–D). However, slow but significant recovery reflected their fluidity in the condensates. Importantly, when we washed out the AP20187, D2-YFP aggregates were diminished (data not shown) whereas D1-TDP-43-YFP and D2-TDP-43-YFP remained unaffected, and the fluidity inside the aggregates were also not affected (Fig. 5C, D), indicating that once TDP-43 was induced to form the aggregates, the intrinsic domains of TDP-43 were responsible for maintaining oligomeric status. Furthermore, D2-YFP formed round-shaped condensates in the presence of AP20187, its fluidity was much higher than that of D1-TDP-43-YFP or D2-TDP-43-YFP, showing that D2-YFP was still freely mobile in the D2-YFP droplets after addition of AP20187 and TDP-43 conferred stickiness to the oligomer (Fig. 5E, F).
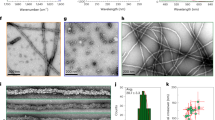
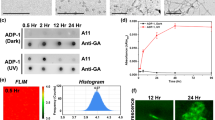
A, B FRAP analysis of aggregates of D1-TDP-43-YFP and D2-TDP-43-YFP; before bleaching (pre), bleached (Bleach) and 120 s after FRAP (Recover) were shown. The aggregate formation was induced by 24 h treatment of AP20187. C–E FRAP analysis of D1-TDP-YFP (C), D2-TDP-43-YFP (D), or D2-YFP (E) in the presence (red) or absence (blue) or after washout (green) of AP20187. Note that we could not find D2-YFP aggregate after washout of AP20187. F Comparison of the FRAP recovery rate between D2- YFP and D2-TDP-43-YFP in the presence of AP20187. For FRAP analyses, the cells were treated with AP20187 for 24 h before the analyses. * p < 0.05; **** p < 0.001. N = 10 cells/condition. The error bars indicate S.D.
Next, we characterized biochemical features of cytosolic TDP-43 aggregates to test whether they present typical pathological properties of TDP43-aggreagtes seen in ALS. In the ALS patient tissues, TDP-43 accumulates in the cytosol, making SQSTM1/p62-positive aggregates [37, 38]. SQSTM1/p62 is not only a component of cytosolic aggregates seen in ALS patients but also its mutations cause familial ALS [39]. SQSTM1/p62 interacts with misfolded, ubiquitin-positive proteins [40] to degrade them through selective autophagy [41], and optogenetic Cry2olig-TDP-43 forms cytosolic SQSTM1/p62-positive aggregates [21]. Upon induction of oligomerization, D2-YFP, D1-TDP-43-YFP and D2-TDP-43-YFP formed cytosolic aggregates, and all of these were positive for SQSTM1/p62 (Fig. 6A–C). These indicated that the SQSTM/p62 accumulation to the aggregate was not specific for TDP-43 and D2-YFP aggregates were also recognized as misfolded by SQSTM1/p62. Next, we examined whether fused-in-sarcoma (FUS) protein was incorporated into the TDP-43 aggregates [42]. FUS is an RNA-binding protein whose mutations cause familial ALS (ALS6) [43]. When we induced aggregates formation using D1- or D2- constructs, D1-TDP-43-YFP and D2-TDP-43-YFP aggregates, but not D2-YFP aggregates, contained mCherry-FUS, indicating that these aggregates mimic the pathological findings seen in ALS (Fig. 6D–F). Many studies have shown that TDP-43 aggregation has characteristics of stress granules/RNA granules and affects their formation [12, 44]. Therefore, we next tested if TDP-43 aggregates contain TIA-1, which is also an ALS-causative gene and an essential component of stress granule [45]. When we induced aggregate formation, D1-TDP-43-YFP and D2-TDP-43-YFP aggregates, but not D2-YFP aggregates, contained mCherry-TIA1, indicating that these TDP-43 aggregates can interfere dynamics of stress granules (Fig. 6G–I). We also tested whether chemically oligomerized TDP-43 was contained in the detergent-insoluble fractions. In ALS patient tissues, TDP-43 becomes detergent-insoluble, indicating the misfolded and aggregated status of the protein [34, 46]. Especially, cleaved forms of TDP-43 are reported to gain aggregative properties, mainly contained in the insoluble fractions [47]. When fractionated the lysates of HEK293 cells overexpressing D2-YFP, D1-TDP-43-YFP, or D2-TDP-43-YFP in the presence or absence of AP20187 into detergent soluble and insoluble fractions, we confirmed that the AP20187 treatment increased the amount of full-length TDP-43-YFP contained in the insoluble fraction, especially in the cells expressing D2-TDP-43-YFP (Fig. 6J). Finally, we tested if induction of oligomerization of TDP-43 induces cell toxicity. After induction of oligomerization, we observed increase of apoptotic cleaved-PARP1 positive cells in D2-TDP-43-YFP expressing cells but not in the cells expressing D2-YFP or D1-TDP-43-YFP (Fig. 6K). These results indicate that our chemically oligomerized TDP-43 mimics at least some molecular signatures of TDP-43 aggregates seen in the ALS.
A–C Confocal imaging of aggregates of D1-construct and D2- constructs, coexpressed with mCherry-SQSTM1/p62 (p62) after 24 h treatment of AP20187. The arrows show colocalized signals. D–F Confocal imaging of aggregates of D1- construct and D2- constructs, coexpressed with mCherry-FUS. The arrows show colocalized signals. G–I Confocal imaging of aggregates of D1- construct and D2- constructs, coexpressed with TIA1-mCherry. J Immunoblot analysis of HEK293 lysates fractionated into soluble and insoluble fractions. The white arrow shows full-length of D2-TDP-43-YFP whose signal in the insoluble fraction was enhanced by 24 h treatment of AP20187 (upper panel). Ponceau S staining of the blotted membrane as a loading control (lower panel). K The apoptotic cell population in HEK293 cells expressing D2-YFP, D1-TDP-43-YFP, or D2-TDP-43-YFP treated with or without 0.5 μM AP20187 for 24 h. The cells were immunostained with anti-cleaved PARP1 antibody, an established apoptosis marker. Approximately 100–200 GFP + cells/frame were counted, and the bar graph shows the average % of c-PARP1 positive cells obtained from 6 frames/each condition. **p < 0.01. n.s. not significant. The scale bar shows 10 μm.
Discussion
Mislocalized, aggregated TDP-43 is found in most cases of ALS tissues and thus it is considered as an important pathological property of this terrible motor neuron disease. TDP-43 is a DNA/RNA-binding protein involved in many biological processes, and its loss of function due to aggregate formation followed by subcellular mislocalization is supposed to lead to motor neuronal cell death. Recent advances in optogenetics have unveiled the enigmatic mechanism in which LLPS triggers aggregation and mislocalization of TDP-43. Here, we used a novel approach to induce oligomerization of TDP-43 by a small compound AP20187. The advantages of this method overcoming optogenetics are: (1) application to mammalian animal model is established; (2) it is possible to selectively induce dimer and oligomer; (3) simultaneous induction of oligomerization of TDP-43 in a large quantity of cells; (4) AP20187 has no substantial toxicity. On the other hand, there are also several disadvantages: (1) the optogenetic response is much faster than that by AP20187; (2) repetitive treatment is easy by the optogenetic approach; (3) the optogenetic approach enables region-specific activation at subcellular level. Therefore, it is important to choose an appropriate method for dissecting biological phenomena involving TDP-43 phase separation.
In this study, we showed that the induction of dimer formation of TDP-43 by AP20187 triggered unexpected aggregate formation, indicating TDP-43 has intrinsic dimerizing/oligomerizing domain(s), playing a role under some conditions. Both deletion of NTD and deletion of LCD impaired aggregate formation of D1-TDP-43-YFP, indicating that NTD and LCD are involved in the aggregate formation in a non-redundant way. The NTD of TDP-43 is shown to be involve in the dimer formation of TDP-43 [17, 18], and the LCD reportedly play a central role in phase separation. We are still looking for the difference of the roles between the NTD and LCD, but possible explanation will be that dimer formation via NTD makes two TDP-43 molecules in very close distance, where the LCD domain can intervene each other, entangling these molecules to form aggregates. There are still biological problems to be solved: (1) TDP-43 itself is an aggregate-prone protein, but how it transforms into aggregates under physiological condition has remained uncertain; (2) the mechanism how accumulated TDP-43 translocates to cytosol remains unclear. For the first question, several pathological stressors are reported to induce phase separation of TDP-43, but which is responsible for the development of ALS remains unclear, and some experimental conditions are supraphysiological which will not occur in vivo. For the second question, we demonstrated that the NES domain was essential for the cytosolic distribution. However, which molecule is responsible for the translocation is unknown and further study is warranted.
In conclusion, we established a novel chemically oligomerizable TDP-43 system which complement the missing part of optogenetic approaches. Complemental usage of both techniques will help to clarify the pathophysiology underlying ALS and to find therapeutic modalities for the disease.
Data availability
All data are available in the main text. Further information and requests for resources and reagents should be addressed by Kohsuke Kanekura (kanekura@tokyo-med.ac.jp).
References
Logroscino G, Piccininni M, Marin B, Nichols E, Abd-Allah F, Abdelalim A, et al. Global, regional, and national burden of motor neuron diseases 1990–2016: a systematic analysis for the Global Burden of Disease Study 2016. Lancet Neurol. 2018;17:1083–97.
Maurel C, Dangoumau A, Marouillat S, Brulard C, Chami A, Hergesheimer R, et al. Causative Genes in Amyotrophic Lateral Sclerosis and Protein Degradation Pathways: a Link to Neurodegeneration. Mol. Neurobiol. 2018;55:6480–99.
Sreedharan J, Blair IP, Tripathi VB, Hu X, Vance C, Rogelj B, et al. TDP-43 mutations in familial and sporadic amyotrophic lateral sclerosis. Science. 2008;319:1668–72.
Neumann M, Sampathu DM, Kwong LK, Truax AC, Micsenyi MC, Chou TT, et al. Ubiquitinated TDP-43 in frontotemporal lobar degeneration and amyotrophic lateral sclerosis. Science. 2006;314:130–3.
Arai T, Hasegawa M, Akiyama H, Ikeda K, Nonaka T, Mori H, et al. TDP-43 is a component of ubiquitin-positive tau-negative inclusions in frontotemporal lobar degeneration and amyotrophic lateral sclerosis. Biochem. Biophys. Res. Commun. 2006;351:602–11.
Morera AA, Ahmed NS, Schwartz JC. TDP-43 regulates transcription at protein-coding genes and Alu retrotransposons. Biochim Biophys Acta Gene Regul Mech. 2019;1862:194434.
Afroz T, Hock EM, Ernst P, Foglieni C, Jambeau M, Gilhespy LAB, et al. Functional and dynamic polymerization of the ALS-linked protein TDP-43 antagonizes its pathologic aggregation. Nat Commun. 2017;8:45.
Arnold ES, Ling SC, Huelga SC, Lagier-Tourenne C, Polymenidou M, Ditsworth D, et al. ALS-linked TDP-43 mutations produce aberrant RNA splicing and adult-onset motor neuron disease without aggregation or loss of nuclear TDP-43. Proc Natl Acad Sci U S A. 2013;110:E736–745.
Highley JR, Kirby J, Jansweijer JA, Webb PS, Hewamadduma CA, Heath PR, et al. Loss of nuclear TDP-43 in amyotrophic lateral sclerosis (ALS) causes altered expression of splicing machinery and widespread dysregulation of RNA splicing in motor neurones. Neuropathol. Appl. Neurobiol. 2014;40:670–85.
Quinones-Valdez G, Tran SS, Jun HI, Bahn JH, Yang EW, Zhan L, et al. Regulation of RNA editing by RNA-binding proteins in human cells. Commun Biol. 2019;2:19.
Wang C, Duan Y, Duan G, Wang Q, Zhang K, Deng X, et al. Stress induces dynamic, cytotoxicity-antagonizing TDP-43 nuclear bodies via paraspeckle LncRNA NEAT1-mediated liquid-liquid phase separation. Mol. Cell. 2020;79:443–58.e447.
Khalfallah Y, Kuta R, Grasmuck C, Prat A, Durham HD, Vande Velde C. TDP-43 regulation of stress granule dynamics in neurodegenerative disease-relevant cell types. Sci. Rep. 2018;8:7551.
Konopka A, Whelan DR, Jamali MS, Perri E, Shahheydari H, Toth RP, et al. Impaired NHEJ repair in amyotrophic lateral sclerosis is associated with TDP-43 mutations. Mol. Neurodegener. 2020;15:51.
Mitra J, Guerrero EN, Hegde PM, Liachko NF, Wang H, Vasquez V, et al. Motor neuron disease-associated loss of nuclear TDP-43 is linked to DNA double-strand break repair defects. Proc Natl Acad Sci U S A. 2019;116:4696–705.
Iguchi Y, Katsuno M, Niwa J, Takagi S, Ishigaki S, Ikenaka K, et al. Loss of TDP-43 causes age-dependent progressive motor neuron degeneration. Brain. 2013;136:1371–82.
Babinchak WM, Haider R, Dumm BK, Sarkar P, Surewicz K, Choi JK, et al. The role of liquid-liquid phase separation in aggregation of the TDP-43 low-complexity domain. J. Biol. Chem. 2019;294:6306–17.
Wang, A, Conicella, AE, Schmidt, HB, Martin, EW, Rhoads, SN, Reeb, AN et al. A single N-terminal phosphomimic disrupts TDP-43 polymerization, phase separation, and RNA splicing. EMBO J. 2018;37:e97452.
Jiang LL, Xue W, Hong JY, Zhang JT, Li MJ, Yu SN, et al. The N-terminal dimerization is required for TDP-43 splicing activity. Sci. Rep. 2017;7:6196.
Wang L, Kang J, Lim L, Wei Y, Song J. TDP-43 NTD can be induced while CTD is significantly enhanced by ssDNA to undergo liquid-liquid phase separation. Biochem. Biophys. Res. Commun. 2018;499:189–95.
Taslimi A, Vrana JD, Chen D, Borinskaya S, Mayer BJ, Kennedy MJ, et al. An optimized optogenetic clustering tool for probing protein interaction and function. Nat Commun. 2014;5:4925.
Mann JR, Gleixner AM, Mauna JC, Gomes E, DeChellis-Marks MR, Needham PG, et al. RNA binding antagonizes neurotoxic phase transitions of TDP-43. Neuron. 2019;102:321–38.e328.
Asakawa K, Handa H, Kawakami K. Optogenetic modulation of TDP-43 oligomerization accelerates ALS-related pathologies in the spinal motor neurons. Nat Commun. 2020;11:1004.
Polesskaya O, Baranova A, Bui S, Kondratev N, Kananykhina E, Nazarenko O, et al. Optogenetic regulation of transcription. BMC Neurosci. 2018;19:12.
Pouzet, S, Banderas, A, Le Bec, M, Lautier, T, Truan, G & Hersen, P. The promise of optogenetics for bioproduction: dynamic control strategies and scale-up instruments. Bioengineering (Basel). 2020;7:151.
Stockley JH, Evans K, Matthey M, Volbracht K, Agathou S, Mukanowa J, et al. Surpassing light-induced cell damage in vitro with novel cell culture media. Sci. Rep. 2017;7:849.
Clackson T, Yang W, Rozamus LW, Hatada M, Amara JF, Rollins CT, et al. Redesigning an FKBP-ligand interface to generate chemical dimerizers with novel specificity. Proc Natl Acad Sci U S A. 1998;95:10437–42.
Fegan A, White B, Carlson JC, Wagner CR. Chemically controlled protein assembly: techniques and applications. Chem Rev. 2010;110:3315–36.
Yang W, Rozamus LW, Narula S, Rollins CT, Yuan R, Andrade LJ, et al. Investigating protein-ligand interactions with a mutant FKBP possessing a designed specificity pocket. J. Med. Chem. 2000;43:1135–42.
Minaki H, Sasaki K, Honda H, Iwaki T. Prion protein oligomers in Creutzfeldt-Jakob disease detected by gel-filtration centrifuge columns. Neuropathology. 2009;29:536–42.
Kanekura K, Ma X, Murphy JT, Zhu LJ, Diwan A, Urano F. IRE1 prevents endoplasmic reticulum membrane permeabilization and cell death under pathological conditions. Sci Signal. 2015;8:ra62.
Lin W, Lin Y, Li J, Fenstermaker AG, Way SW, Clayton B, et al. Oligodendrocyte-specific activation of PERK signaling protects mice against experimental autoimmune encephalomyelitis. J. Neurosci. 2013;33:5980–91.
Johnson BS, Snead D, Lee JJ, McCaffery JM, Shorter J, Gitler AD. TDP-43 is intrinsically aggregation-prone, and amyotrophic lateral sclerosis-linked mutations accelerate aggregation and increase toxicity. J. Biol. Chem. 2009;284:20329–39.
Pinarbasi ES, Cagatay T, Fung HYJ, Li YC, Chook YM, Thomas PJ. Active nuclear import and passive nuclear export are the primary determinants of TDP-43 localization. Sci. Rep. 2018;8:7083.
Nonaka T, Masuda-Suzukake M, Arai T, Hasegawa Y, Akatsu H, Obi T, et al. Prion-like properties of pathological TDP-43 aggregates from diseased brains. Cell Rep. 2013;4:124–34.
Shimonaka S, Nonaka T, Suzuki G, Hisanaga S, Hasegawa M. Templated Aggregation of TAR DNA-binding Protein of 43 kDa (TDP-43) by Seeding with TDP-43 Peptide Fibrils. J. Biol. Chem. 2016;291:8896–907.
Li HR, Chiang WC, Chou PC, Wang WJ, Huang JR. TAR DNA-binding protein 43 (TDP-43) liquid-liquid phase separation is mediated by just a few aromatic residues. J. Biol. Chem. 2018;293:6090–8.
Maekawa S, Leigh PN, King A, Jones E, Steele JC, Bodi I, et al. TDP-43 is consistently co-localized with ubiquitinated inclusions in sporadic and Guam amyotrophic lateral sclerosis but not in familial amyotrophic lateral sclerosis with and without SOD1 mutations. Neuropathology. 2009;29:672–83.
Tanji K, Zhang HX, Mori F, Kakita A, Takahashi H, Wakabayashi K. p62/sequestosome 1 binds to TDP-43 in brains with frontotemporal lobar degeneration with TDP-43 inclusions. J. Neurosci. Res. 2012;90:2034–42.
Fecto F, Yan J, Vemula SP, Liu E, Yang Y, Chen W, et al. SQSTM1 mutations in familial and sporadic amyotrophic lateral sclerosis. Arch. Neurol. 2011;68:1440–6.
Seibenhener ML, Babu JR, Geetha T, Wong HC, Krishna NR, Wooten MW. Sequestosome 1/p62 is a polyubiquitin chain binding protein involved in ubiquitin proteasome degradation. Mol. Cell. Biol. 2004;24:8055–68.
Bjorkoy G, Lamark T, Johansen T. p62/SQSTM1: a missing link between protein aggregates and the autophagy machinery. Autophagy. 2006;2:138–9.
Watanabe S, Inami H, Oiwa K, Murata Y, Sakai S, Komine O, et al. Aggresome formation and liquid-liquid phase separation independently induce cytoplasmic aggregation of TAR DNA-binding protein 43. Cell Death Dis. 2020;11:909.
Vance C, Rogelj B, Hortobagyi T, De Vos KJ, Nishimura AL, Sreedharan J, et al. Mutations in FUS, an RNA processing protein, cause familial amyotrophic lateral sclerosis type 6. Science. 2009;323:1208–11.
McDonald KK, Aulas A, Destroismaisons L, Pickles S, Beleac E, Camu W, et al. TAR DNA-binding protein 43 (TDP-43) regulates stress granule dynamics via differential regulation of G3BP and TIA-1. Hum. Mol. Genet. 2011;20:1400–10.
Mackenzie IR, Nicholson AM, Sarkar M, Messing J, Purice MD, Pottier C, et al. TIA1 mutations in amyotrophic lateral sclerosis and frontotemporal dementia promote phase separation and alter stress granule dynamics. Neuron. 2017;95:808–16. e809.
Bosque PJ, Boyer PJ, Mishra P. A 43-kDa TDP-43 species is present in aggregates associated with frontotemporal lobar degeneration. PLoS ONE. 2013;8:e62301.
Yamashita T, Hideyama T, Hachiga K, Teramoto S, Takano J, Iwata N, et al. A role for calpain-dependent cleavage of TDP-43 in amyotrophic lateral sclerosis pathology. Nat Commun. 2012;3:1307.
Acknowledgements
The authors thank Addgene and Dr. Aaron Gitler and Dr. Michael Davidson for providing us TDP-43-YFP (Addgene plasmid #84911) and mCherry-Sequestosome1 (SQSTM1)-N-18 (Addgene plasmid #55132). We also thank Dr. Masaaki Matsuoka for providing us wt-FUS cDNA.
Funding
This work was supported by grants from the JSPS KAKENHI Grant numbers (16H06247, 17H03923 and 20H03593 to K.K., 17K15671 to Y.H., and 17H04067 and 21H02706 to M.K.). This work was also supported in part by the Japan Agency for Medical Research and Development (AMED) (16ek0109180h0001 and 17ae0101016s0904), Strategic Research Foundation Grant-aided Project for Private Universities from the Ministry of Education, Culture, Sports, Science and Technology of Japan (M.K.), Takeda Science Foundation (K.K.), Japan Intractable Diseases (Nanbyo) Research Foundation (K.K.), the Tokyo Biochemistry Research Foundation (K.K.), and the Ichiro Kanehara Foundation (K.K.).
Author information
Authors and Affiliations
Contributions
Y.Y., M.K. and K.K. designed the research and wrote the paper. Y.Y., T.M., Y.H. and K.K. designed and performed all the imaging studies, vector constructions, immunoblot analyses, and FRAP analyses.
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Yamanaka, Y., Miyagi, T., Harada, Y. et al. Establishment of chemically oligomerizable TAR DNA-binding protein-43 which mimics amyotrophic lateral sclerosis pathology in mammalian cells. Lab Invest 101, 1331–1340 (2021). https://doi.org/10.1038/s41374-021-00623-4
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41374-021-00623-4
This article is cited by
-
How can we interpret the relationship between liquid-liquid phase separation and amyotrophic lateral sclerosis?
Laboratory Investigation (2022)