Abstract
Long non-coding RNAs (lncRNAs) have been found to play regulatory roles in cancers; for example, UCC was reported to promote colorectal cancer progression. However, the function of UCC in non-small-cell lung cancer (NSCLC) remains unclear. Therefore, mRNA and protein levels were assessed using qPCR and western blots. Cell viability was assessed by colony-formation assays. The interaction between lncRNAs and miRNAs was detected by dual-luciferase reporter and RIP assays. The tumorigenesis of NSCLC cells in vivo was determined by xenograft assays. LncRNA UCC was highly expressed in both NSCLC tissues and cells. Knockdown of UCC expression suppressed the proliferation of NSCLC cells. In addition, a dual-luciferase reporter system and RIP assays showed that UCC specifically bound to miR-143-3p and acted as a sponge of miR-143-3p in NSCLC cells. The miR-143-3p inhibitor rescued the inhibitory effect of sh-UCC on the proliferation of NSCLC cells. Moreover, miR-143-3p and UCC showed opposite effects on the expression of SOX5, which promoted EMT in NSCLC cells. In addition, in a mouse model, knockdown of UCC expression alleviated EMT and NSCLC progression in vivo, which was consistent with the in vitro results. In the current study, we found that UCC induced the proliferation and migration of NSCLC cells both in vitro and in vivo by inducing the expression of SOX5 via miR-143-3p and subsequently promoted EMT in NSCLC.
Similar content being viewed by others
Introduction
As the most prevalent cancer worldwide, lung cancer has the highest mortality rate among all cancers, and ~1.8 million new cases are reported per year [1]. According to the pathological morphology and clinical characteristics, lung cancer is divided into small cell lung cancer and non-small-cell lung cancer (NSCLC), with the latter accounting for 80–85% of total cases [2]. Although the increase in new therapies has provided more choices for NSCLC patients, surgery and chemotherapy are still the mainstay of NSCLC treatment, and the 5-year survival rate of NSCLC patients is still unsatisfactory [3, 4]. Consequently, exploring the underlying mechanisms and finding new targets are urgently needed for the early diagnosis, monitoring and treatment of NSCLC.
Long non-coding RNAs (lncRNAs) were found to play a crucial role in the development of several diseases [5]. The regulatory function of lncRNAs in cancers has also become a hot issue in this field. Several lncRNAs have been reported to participate in the growth, apoptosis and migration of tumour cells: lncRNA LUNAR1 was reported to promote the growth of cancer cells in T-cell acute lymphoblastic leukaemia by inducing the expression of insulin-like growth factor 1 [6]; lncRNA growth arrest-specific 5 (Gas5) could act as a competitor of glucocorticoid response elements and inhibit the apoptosis of tumour cells [7, 8]; and in liver cancer, the expression of lncRNA ATB was activated by TGF-β and then promoted the invasion, migration and EMT of hepatoma cells [9]. Colon-cancer-associated transcript 2 was also identified as a novel oncogenic lncRNA in hepatocellular carcinoma [10]. However, the regulatory mechanisms of lncRNAs have not been fully elucidated. Recently, a new mechanism of lncRNA regulation, miRNA sponges or competing endogenous RNAs (ceRNAs) was reported. MiRNAs are known as critical regulators in numerous processes, especially tumour progression [11,12,13,14]. LncRNAs can function as ceRNAs, competitively bind with miRNAs in a sequence-specific manner and subsequently regulate downstream target genes of miRNAs. To date, several lncRNAs have been reported to act as miRNA sponges in osteosarcoma [15,16,17,18], gastric cancer [19,20,21,22], lung cancer [23, 24], hepatocellular cancer [25, 26] and breast cancer [27,28,29]. LncRNA UCC (ENST00000602992) was reported to promote colorectal cancer progression by acting as a sponge for miR-143-3p, which is an important regulator in diverse physiological and pathological processes [30,31,32,33]. However, the function of UCC/miR-143-3p in NSCLC progression remains unclear.
The SOX5 gene, which is universally expressed, belongs to the SRY-related HMG-box family of transcription factors [34]. SOX5 participates in the maintenance of normal physiological functions [35]. Moreover, SOX5 was reported to play important roles in the progression of various cancers, such as colorectal cancer, melanoma and hepatocellular carcinoma [36,37,38]. However, the regulatory mechanism of SOX5 in NSCLC still needs to be elucidated.
In the current study, we investigated the regulatory function of lncRNA UCC in the progression of NSCLC, determined the interaction between UCC and miR-143-3p and further explored the regulatory effect of UCC on SOX5 and EMT in NSCLC.
Methods and materials
NSCLC tissues
NSCLC tissues and the corresponding adjacent normal tissues were obtained from 20 patients at Xiangya Hospital. All patients were well-informed and provided written informed consent. The current study was approved by the medical ethics committee of Xiangya Hospital Central South University. All tissue samples were kept in liquid nitrogen.
Cell culture
A549 cells were cultured in Ham’s F-12K (Kaighn’s) (Gibco, Thermo Fisher Scientific, Inc., Waltham, MA, USA). HCC827, H1299, H460 and 95D cells were cultured in RPMI-1640 (Gibco), and 16HBE cells were cultured in DMEM (high glucose, Gibco). All media were supplemented with 10% foetal bovine serum (HyClone), 100-U/mL penicillin (Gibco) and 100-μg/mL streptomycin (Gibco). All cell lines were obtained from the Cell Bank of Chinese Academy of Sciences (Beijing, China) and cultured under 5% CO2 at 37 °C.
Colony-formation assay
Cells in logarithmic phase were trypsinized to generate single cells and suspended in complete medium. After gradient dilution and counting, cells were seeded in 10 cm dishes (100 cell/dish) and cultured at 37 °C with 5% CO2 for 2 weeks. When the colonies were visible, we removed the medium and washed the cells with methanol for 15 min. The colonies were then stained with Giemsa stain for 15 min, washed gently, air dried at room temperature and counted.
Methyl thiazolyl tetrazolium (MTT) assay
Cells were seeded at a concentration of 1800 cells/well in 96-well plates 24 h before the experiment. Media were removed, and the cells were incubated with 20 μL of MTT reagent (5 mg/mL; Sigma-Aldrich) at 37 °C. After 4 h, the supernatant was removed, and 150 μL of dimethyl sulfoxide (Sigma-Aldrich) was added to each well to dissolve the formazan crystals. The samples were then detected for absorbance at 490 nm using a microplate spectrophotometer.
Scratch assay
Cells were grown in six-well plates to a monolayer and then cultured with serum-free medium for 24 h. A scratch was induced in a monolayer for each well with a sterile 200-μL pipette tip. Serum-free medium was changed to remove cell fragments. The cells were cultured at 37 °C with 5% CO2 for 24 h. The scratches were imaged using an inverted phase contrast microscope (CKX31, Olympus, 100×) at 10 min and 24 h after scratching. The width of each scratch wound was determined by using ImageJ software.
Quantitative polymerase chain reaction (qPCR) assay
Total RNA was isolated using TRIzol reagent (Invitrogen, Thermo Fisher Scientific, Inc., Waltham, MA, USA) following the manual. Complementary DNA (cDNA) was reverse-transcribed using an EasyScript® kit (Transgene, Beijing, China) with random primers or miRNA reverse-transcription primers. qPCR cycling was performed at 98 °C for 5 min and 40 cycles at 95 °C for 15 s and 60 °C for 30 s. Primers were designed and ordered from Sangon Biotech (Shanghai, China). GAPDH and U6 were used as internal control. The primer sequences are listed below:
UCC-F: 5′-GAAAGCATTTTGAAAGCCACTG-3′
UCC-R: 5′-GAAACTCACCAACCCAAACCTC-3′
SOX5-F: 5′-CGATCATAGGTGGCTGCTGT-3′
SOX5-R: 5′-CAGCTGACCTTGAACCTGGA-3′
GAPDH-F: 5′-GCTCTCTGCTCCTCCTGTTC-3′
GAPDH-R: 5′-ACGACCAAATCCGTTGACTC-3′
miR-143-3p-F: 5′-CGGCTGAGATGAAGCACTG-3′
miR-143-3p-R: 5′-GTCGTATCCAGTGCAGGGTCCGAGGTATTCGCACTGGATACGAC ACCAGA-3′
U6-F: 5′-CTCGCTTCGGCAGCACA-3′
U6-R: 5′-AACGCTTCACGAATTTGCGT-3′
Cell transfection and lentiviral infection
The target sh-UCC was selected using BLOCK-iTTM RNAi Designer (http://rnaidesigner.thermofisher.com/rnaiexpress/). Lentivirus vectors with sh-UCC (LV-sh-UCC) were ordered from GenePharma Co., Ltd. (Shanghai) based on pGLVU6/GFP, with the optimal target tested. A scramble sequence (GenePharma Co., Ltd., Shanghai) was used to construct the negative control lentivirus vector (LV-sh-NC). Recombinant lentiviral vectors were transfected into HEK293T cells using Lipofectamine™ 2000 (Invitrogen, Thermo Fisher; Shanghai, China) following the manual. The culture supernatant was collected and filtered (0.45-mm filter) after 48 h to collect lentivirus. In addition, 95D cells were infected with LV-sh-UCC and LV-sh-NC with 8-μg/mL polybrene. Cells stably expressing shRNAs were used in the following experiments.
MiR-143-3p mimics (chemosynthetic simulant of endogenous miR-143-3p) and miR-143-3p inhibitor (a modified chemosynthetic nucleotide sequence, which inhibits the activity of the corresponding miRNAs due to the complementary sequence) were ordered from Shanghai GenePharma Co., Ltd. (Shanghai, China). DNA fragments containing UCC sequences were obtained by PCR with the cDNA of the transcriptome as the PCR template. The DNA fragments were then inserted into pcDNA3.1 (Invitrogen), and pcDNA3.1-UCC was constructed. Then, 50-nM miR-143-3p mimics/inhibitor and 200-ng pcDNA3.1-UCC were transfected with Lipofectamine 3000 (Invitrogen, Thermo Fisher; Shanghai, China). The original medium was replaced with serum-free medium before transfection. Fresh complete medium was added 6 h after transfection. Cells were harvested 24 h after transfection.
Dual-luciferase reporter assay
The predicted target region of miR-143-3p in UCC and SOX5 was synthesized and inserted into the 3′-UTR (untranslated region) of the psiCHECK-2 luciferase reporter vector (Promega) via restriction enzyme sites to obtain dual-luciferase reporter vectors of UCC and SOX5. For UCC, sequences containing both predicted binding sequences were synthesized (positions 450–650); for SOX5, only the predicted binding sequence (positions 4405–4411) was inserted. Mutated target sequences were used as negative controls. Then, 95D and HCC827 cells were seeded in 24-well plates, cultured for 1 day, and transfected with miRNA mimics (50 nM) and corresponding luciferase reporters (200 ng/well) using Lipofectamine 3000 (Thermo) following the manual in serum-free medium. Complete medium was added after 4 h, and the cells were lysed after 24 h. The luciferase activity was assessed with a dual-luciferase kit (Promega) using a microplate spectrophotometer. The relative expression of luciferase is presented as the ratio of Renilla luciferase levels to firefly luciferase levels.
Western blot assay
Cells were washed with phosphate-buffered saline (PBS), scraped off and then lysed with lysis buffer (Sigma-Aldrich) on ice. Cell lysate was centrifuged, and cell debris was removed. The protein concentration was assessed using a Thermo Scientific Pierce BCA Protein Assay Kit (Thermo). Fifty micrograms of each sample were loaded and analysed using a quantitative electrophoresis assay with Tris-glycine polyacrylamide gels and nitrocellulose membranes. The membranes were blocked in 2.5% BSA for 1 h and then incubated with primary antibodies against SOX5, Vimentin, Twist (Cell Signaling; 1:1000 dilution), E-cadherin, N-cadherin (Abcam; 1:1000 dilution) and β-actin (Cell Signaling; 1:5000 dilution) at 4 °C overnight. The membranes were then washed with PBS and incubated with secondary antibodies (Bioworld Technology, 1:5000 dilution) for 1 h. Protein bands were detected with a chemiluminescence detection system.
Xenograft assay
Animal experiments in the current study were approved by the ethics committee of Xiangya Hospital Central South University. BALB/C-nu/nu nude mice (male, 8 weeks) were obtained from the SJA Laboratory Animal Company (Hunan, China) and grown in a sterile environment. In brief, 1 × 107 HCC827 cells expressing sh-UCC or sh-NC were subcutaneously injected into nude mice (six for each group) in the flanks. The tumour volumes were analysed weekly. Thirty-two days after the injection, the mice were sacrificed, and the tumours were taken for further analysis. The xenograft assay was conducted with six biological duplications.
Immunohistochemistry assay
Paraffin sections were washed with xylene three times, 100% ethanol two times, 95% ethanol two times and dH2O two times. The slides were then boiled for 10 min in 10-mM sodium citrate buffer. After three washes in dH2O, the sections were incubated with 3% hydrogen peroxide for 10 min and washed twice. The sections were blocked in 5% goat serum for 1 h at room temperature and incubated with each primary antibody overnight at 4 °C. Antibody solution was then removed, and the slides were washed and incubated with biotinylated secondary antibodies (Cell Signaling, Danvers, MA, USA) for 30 min at room temperature. After washing, the sections were successively incubated with ABC reagent (Vectastain ABC Kit) and DAB reagent and then counterstained in haematoxylin. After washing and dehydration in 95% ethanol, 100% ethanol and xylene, the sections were imaged using a microscope (CKX31, Olympus, 100×).
RNA immunoprecipitation (RIP) assay
The RIP assay was conducted using the Magna RIP™ RNA-Binding Protein Immunoprecipitation Kit (Millipore, Merck, Darmstadt, Germany). In brief, cell lysate was incubated with beads and anti-Ago2 or IgG, shaken gently for 2 h, eluted and treated with DNase and Proteinase K. After purification, precipitated RNAs were isolated and reverse-transcribed using random primers. Follow-up quantification was conducted using qPCR amplification.
Statistical analysis
Data are presented as the mean ± SD of at least three repeated experiments. For 2-group comparisons, Student’s t test was conducted; for multiple comparisons, one-way ANOVA followed by the Tukey post hoc test was conducted. P < 0.05 was considered statistically significant. All statistical analyses were conducted using GraphPad Prism 8.0 (GraphPad Software, Inc., La Jolla, CA, USA).
Results
Effect of lncRNA UCC in NSCLC tissues and cells
To investigate the function of lncRNA UCC in NSCLC, we determined the UCC levels in NSCLC tissues and cell lines. qPCR assays showed that the expression of UCC in NSCLC tissues was significantly higher than that in adjacent normal tissues (Fig. 1A), and UCC was highly expressed in all five NSCLC cell lines, especially 95D and HCC827 (Fig. 1B). Therefore, the following experiments in the current study were mainly based on 95D and HCC827 cells. The cell lines expressing shRNAs targeting UCC, 95D-sh-UCC and HCC827-sh-UCC were then developed to further explore the effect of UCC on the proliferation and migration of NSCLC cells. The expression of lncRNA UCC in the 95D-sh-UCC and HCC827-sh-UCC cells is shown in Fig. 1C. UCC expression was effectively suppressed in the 95D-sh-UCC and HCC827-sh-UCC cells. In the colony-formation assay, the numbers of clones formed by the 95D-sh-UCC and HCC827-sh-UCC cells were significantly lower than those formed by the 95D-sh-NC and HCC827-sh-NC cells. The MTT assay also showed the same trend: the proliferation rates of the 95D-sh-UCC and HCC827-sh-UCC cells were lower than those in the control groups (Fig. 1D), indicating that a low level of UCC decreased the proliferation of NSCLC cells. In addition, a scratch assay showed that the migration of NSCLC cells was significantly inhibited by sh-UCC (Fig. 1E). Thus, with the demonstrated high baseline expression level in NSCLC tissues and cell lines, knockdown of lncRNA UCC expression suppressed the proliferation and migration of NSCLC cells, suggesting that UCC may play a critical role in NSCLC cells.
A Plots of the relative expression of UCC in NSCLC tissues and adjacent tissues. B The expression of UCC in NSCLC cell lines. C The expression of UCC in the 95D and HCC827 cells transfected with sh-UCC or sh-NC. D MTT assays of 95D and HCC827 cells. E Colony formation of 95D and HCC827 cells. F Scratch assays of the 95D and HCC827 cells transfected sh-UCC or sh-NC. *P < 0.05, **P < 0.01, ***P < 0.001.
LncRNA UCC inhibited the level of miR-143-3p
The complementarity between miR-143-3p and the target sequences in UCC is shown in Fig. 2A. To investigate the effect of UCC on the level of miR-143-3p, we assessed the level of miR-143-3p in the 95D-sh-UCC and HCC827-sh-UCC cells by qPCR. The levels of UCC and miR-143-3p were inversely correlated in NSCLC cells. The inhibition of UCC expression induced the miR-143-3p level, and UCC overexpression suppressed the miR-143-3p level (Fig. 2B). Moreover, in NSCLC tissues, the level of miR-143-3p was significantly lower than that in the controls (Fig. 2C). UCC was negatively correlated with miR-143-3p in NSCLC tissues (Fig. 2D). The same trend showed that the levels of miR-143-3p were significantly lower in NSCLC cells than in 16HBE cells (Fig. 2E). Given that UCC was shown to be involved in the proliferation and migration of NSCLC cells, exogenous miR-143-3p mimics or inhibitors were then used to explore the effect of miR-143-3p on these processes. The transfection efficacy was tested, and the results showed that miR-143-3p mimics and inhibitor effectively induced or suppressed the level of miR-143-3p in 95D and HCC827 cells (Supplementary Fig. 1). To explore miR-143-3p recognition of the target, we inserted target sequences and mutated control sequences into the 3′-UTR of the luciferase reporter to assess the binding of miR-143-3p to the target sequence. In both 95D and HCC827 cells, the relative expression of firefly luciferase was significantly inhibited by the cotransfection of miR-143-3p mimics and UCC-WT reporter but not the UCC-MUT reporter (Fig. 2F). Furthermore, the RIP assay demonstrated that UCC and miR-143-3p levels were enriched by AGO2 (Fig. 2G), suggesting the coexistence of UCC and miR-143-3p in NSCLC cells. These results indicated that UCC specifically binds with miR-143-3p and acts as a sponge of miR-143-3p in NSCLC cells.
A Schematic diagram of the complementarity between miR-143-3p and two potential target sequences in UCC. B MiR-143-3p expression in the 95D and HCC827 cells expressing sh-UCC or pcDNA3.1-UCC. C The relative expression of miR-143-3p in NSCLC tissues and adjacent tissues. D The correlation analysis of UCC and miR-143-3p. E The relative expression of miR-143-3p in NSCLC cell lines. F The relative luciferase activity of vectors carrying wild-type or mutated miR-143-3p target sequences from UCC in the 95D and HCC827 cells transfected with miR-143-3p mimics. G Plots of the relative level of UCC and miR-143-3p binding with Ago2 assessed using RIP assays. UCC expression was assessed by qRT-PCR. *P < 0.05, **P < 0.01, ***P < 0.001.
LncRNA UCC induced the proliferation and migration of NSCLC cells by inhibiting miR-143-3p
The miR-143-3p inhibitor was transfected into the 95D and HCC827 cells expressing sh-UCC to investigate its involvement in the regulatory effect of UCC. The level of miR-143-3p significantly decreased in the 95D-sh-UCC and HCC827-sh-UCC cells transfected with the miR-143-3p inhibitor compared with the inhibitor NC (Fig. 3A). In addition, the miR-143-3p inhibitor reversed the suppression of 95D-sh-UCC and HCC827-sh-UCC cell proliferation (Fig. 3B, C). Consistently, in the scratch assay, the transfection of the miR-143-3p inhibitor also rescued the suppression of migration in the 95D-sh-UCC and HCC827-sh-UCC cells caused by sh-UCC expression (Fig. 3D). Therefore, these results suggested that miR-143-3p plays an important role in the regulatory effect of UCC on the proliferation and migration of NSCLC cells.
A The expression of miR-143-3p in the 95D and HCC827 cells cotransfected with sh-UCC and miR-143-3p inhibitor. UCC expression was assessed by qRT-PCR. B MTT assays of the 95D and HCC827 cells expressing sh-UCC or sh-NC, with or without transfection of miR-143-3p inhibitor. C Representative images and plots of colony-formation assays of the 95D and HCC827 cells expressing sh-UCC or sh-NC, with or without transfection of miR-143-3p inhibitor. D Scratch assays of the 95D and HCC827 cells expressing sh-UCC or sh-NC, with or without transfection of miR-143-3p inhibitor. *P < 0.05, **P < 0.01, ***P < 0.001.
MiR-143-3p suppressed the expression of SOX5
To further investigate the function of miR-143-3p and its downstream factor in the regulatory process of UCC, we searched the potential target genes of miR-143-3p by TargetScan (http://www.targetscan.org/mamm_31/) and found specific target sequences in the 3′-UTR of the SOX5 gene (Fig. 4A). SOX5 mRNA and protein levels were inhibited by miR-143-3p mimics and promoted by miR-143-3p inhibitor in 95D and HCC827 cells (Fig. 4B, C), showing that SOX5 was as a downstream gene of miR-143-3p. In addition, the high level of SOX5 in NSCLC tissues and cell lines was determined by qPCR and western blotting (Fig. 4D, E). Moreover, in tumour tissues, the levels of SOX5 and miR-143-3p showed an inverse correlation (Fig. 4F). The mRNA and protein levels of SOX5 were also measured in NSCLC cells. The results showed that SOX5 was upregulated in NSCLC cells (Fig. 4G, H). In the luciferase assay, the relative expression of firefly luciferase was significantly inhibited by cotransfection of miR-143-3p mimics and SOX5-WT reporter but not by cotransfection of the SOX5-MUT reporter (Fig. 4I), indicating the binding between miR-143-3p and the 3′-UTR of SOX5. Thus, miR-143-3p inhibits the expression of SOX5 by recognizing the target sequence in the 3′-UTR of SOX5.
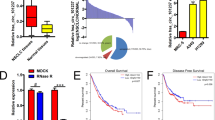
A Schematic diagram of the complementarity between miR-143-3p and the potential target sequence on SOX5. B The expression of SOX5 in the 95D and HCC827 cells transfected with mimics-NC, miR-143-3p mimics, inhibitor NC or miR-143-3p inhibitor. C The protein level of SOX5 assessed by western blots of the 95D and HCC827 cells transfected with mimics-NC, miR-143-3p mimics, inhibitor NC or miR-143-3p inhibitor. D The relative expression of SOX5 in NSCLC tissues and adjacent tissues. E Representative images and plots of the protein levels of SOX5 assessed by western blotting in NSCLC tissues and adjacent tissues. F Correlation analysis of SOX5 and miR-143-3p levels. G SOX5 expression detected by qPCR in NSCLC cell lines. H The protein level of SOX5 assessed by western blotting in NSCLC cell lines. I The relative luciferase activity of vector carrying wild-type or mutated miR-143-3p target sequence from SOX5 in the 95D or HCC827 cells transfected with miR-143-3p mimics. *P < 0.05, **P < 0.01, ***P < 0.001.
LncRNA UCC promoted EMT in NSCLC cells by inducing the expression of SOX5
We further determined the level of SOX5 in NSCLC cells with low baseline UCC expression (Fig. 5A, B). SOX5 was downregulated in the 95D-sh-UCC and HCC827-sh-UCC cells. Given that miR-143-3p was determined to be involved in the UCC-mediated regulation of the proliferation and migration of NSCLC cells, the expression of key factors of EMT was further assessed. The levels of E-cadherin, N-cadherin, Vimentin and Twist were assessed in the 95D-sh-UCC and HCC827-sh-UCC cells. N-cadherin, Vimentin and Twist were suppressed by sh-UCC, whereas the level of E-cadherin was induced (Fig. 5B). To further investigate the function of miR-143-3p in this process, we transfected a miR-143-3p inhibitor into the 95D-sh-UCC and HCC827-sh-UCC cells. qPCR and western blot assays showed that the miR-143-3p inhibitor rescued the downregulation of SOX5 expression triggered by sh-UCC (Fig. 5C, D), and the differential expression of E-cadherin, N-cadherin, Vimentin and Twist triggered by sh-UCC was also reversed (Fig. 5D). These results showed that lncRNA UCC promoted EMT in NSCLC cells and that miR-143-3p and SOX5 played significant roles in this process.
A Expression of SOX5 in the UCC knockdown 95D and HCC827 cells. B Protein levels of SOX5, E-cadherin, N-cadherin, Twist and Vimentin in 95D and HCC827 cells expressing sh-UCC. C Expression of SOX5 in the 95D and HCC827 cells transfected with UCC and transfected with miR-143-3p inhibitor. SOX5 expression was assessed by qRT-PCR. D The protein levels of SOX5, E-cadherin, N-cadherin, Twist and Vimentin in the 95D and HCC827 cells expressing sh-UCC transfected with miR-143-3p inhibitor. *P < 0.05, **P < 0.01, ***P < 0.001.
Knockdown of lncRNA UCC expression inhibited the proliferation and EMT of NSCLC in vivo
A xenograft assay was performed to investigate the effect of UCC on the tumorigenesis and EMT of NSCLC in vivo. First, the 95D-sh-UCC cells were subcutaneously injected into the flanks of nude mice, and the mice were observed for 32 days. The average weight and volume of the tumours were found to be significantly lower in the sh-UCC group than in the sh-NC group (Fig. 6A–C), indicating that the tumorigenic capacity was suppressed by the decrease in UCC expression. The expression of UCC, SOX5 and miR-143-3p in the tumour tissues was found to be consistent with the results in cell lines (Fig. 6D, E). The downregulation of N-cadherin, Twist and Vimentin and the upregulation of E-cadherin were found in the 95D-sh-UCC-derived tumours (Fig. 6D, E), showing an inhibited level of EMT, which is consistent with the results in cell lines. Moreover, a lower level of Ki67 was found by immunohistochemical assays of the 95D-sh-UCC cells, which further indicated that sh-UCC suppressed the progression of NSCLC. In summary, knockdown of lncRNA UCC alleviated EMT and NSCLC progression in vivo, which is consistent with the in vitro results.
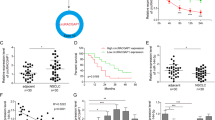
A xenograft assay was performed, and the sh-UCC- or sh-NC-expressing HCC827 cells were subcutaneously injected into the flanks of nude mice. A Picture of the tumours formed in the nude mice injected with HCC827 cells. B Tumour weight. C Plots of the tumour volumes. The tumour volumes were assessed every 4 days. D The relative expression of UCC, miR-143-3p and SOX5 in the tumours. E The protein levels of SOX5, E-cadherin, N-cadherin, Twist and Vimentin in the tumours. F Ki67 expression in the tumours was determined by immunohistochemical assays. *P < 0.05, **P < 0.01, ***P < 0.001.
Discussion
According to the data reported by the National Cancer Institute of the US in 2015, the 5-year survival rates of lung cancer patients were 54.8% for those with a limited stage, 27.4% for those with regional metastasis, 4.2% for those with distant metastasis and 7.5% for those with an unknown stage [39]. Hence, novel effective therapies or early diagnostic methods are needed. In this study, we determined that lncRNA UCC competitively bound to miR-143-3p, weakened the inhibition of SOX5 triggered by miR-143-3p, induced the level of SOX5 and ultimately promoted EMT in NSCLC cells.
The function of lncRNAs in suppressing tumour progression has been studied in several studies for more than 10 years, and many have been shown to function as miRNA sponges. MALAT1 was reported to promote osteosarcoma cell growth through the inhibition of miR-376a, leading to increased expression of TGFA, which was suppressed by miR-376a [40]. In addition, Li et al. reported that GAS5 suppresses breast cancer proliferation by absorbing miR-21 and rescues the suppression of phosphatase and tensin homologues (PTEN) [41]. In NSCLC, lncRNA XIST functions as a sponge of miR-449a, which is a negative regulator of Bcl-2 and is associated with the proliferation rate and metastatic potential of NSCLC cell lines [42]. As a novel lncRNA, UCC was reported to be highly expressed in colorectal cancer and contributed to tumour progression by specifically suppressing miR-143 [31]. In the current study, we first demonstrated that lncRNA UCC is highly expressed in NSCLC tissue and cell lines, suggesting that UCC could be a potential novel carcinogenic lncRNA or a novel biomarker for the early diagnosis of NSCLC. In addition, we determined that UCC inhibits the level of miR-143-3p in NSCLC both in vitro and in vivo, indicating that miR-143-3p also deserves to be noted in target exploration for new cancer therapies.
SOX5 is a transcription factor of the SRY-related HMG-box family, which plays critical roles in the maintenance of functions [35, 43,44,45,46]. Several studies have shown that SOX5 participates in the progression of colorectal cancer [38], melanoma [37], lymphoma [47] and liver cancer [36]. Moreover, Zou et al. found that SOX5 drives the malignant potential in NSCLC by interacting with YAP1 [34], and Hu et al. showed the regulatory function of SOX5 in the TGF-b-induced EMT of prostate cancer by controlling Twist1 expression [48]. Moreover, SOX5 was found to be a direct target of miRNAs, such as miR‐497‐5p [49], miR-132-3p [50] and miR-539 [51], which enabled miRNAs to play significant roles by regulating SOX5. In the present study, we found that miR-143-3p inhibited SOX5 specifically by recognizing the target sequence in the 3′-UTR of SOX5 mRNA in NSCLC cell lines. Suppression was then confirmed in an animal model. In addition, loss-of-function experiments showed that SOX5 was essential for the regulatory function of lncRNA UCC in EMT. Therefore, lncRNA UCC promoted the EMT of NSCLC by inhibiting miR-143-3p and then enhanced SOX5 expression.
In the current study, we found that lncRNA UCC is highly expressed in NSCLC tissues and cell lines. In addition, lncRNA UCC induced the proliferation and migration of NSCLC cells both in vitro and in vivo. Moreover, we found that lncRNA UCC acts as a ceRNA of miR-143-3p and could upregulate SOX5 by absorbing miR-143-3p. Thus, lncRNA UCC decreased the level of miR-143-3p, induced the expression of SOX5 and subsequently promoted EMT in NSCLC. Our findings in this study also provide new insights into therapeutic strategies for NSCLC.
Data availability
All data generated during this study are available within the articel.
References
Siegel RL, Miller KD, Jemal A. Cancer statistics, 2015. CA Cancer J Clin. 2015;65:5–29.
Seve P, Reiman T, Dumontet C. The role of betaIII tubulin in predicting chemoresistance in non-small cell lung cancer. Lung Cancer. 2010;67:136–43.
Raungrut P, Wongkotsila A, Lirdprapamongkol K, Svasti J, Geater SL, Phukaoloun M, et al. Prognostic significance of 14-3-3γ overexpression in advanced non-small cell lung cancer. Asian Pac J Cancer Prev. 2014;15:3513–8.
Miller KD, Siegel RL, Lin CC, Mariotto AB, Kramer JL, Rowland JH, et al. Cancer treatment and survivorship statistics, 2016. CA Cancer J Clin. 2016;66:271–89.
Beermann J, Piccoli MT, Viereck J, Thum T. Non-coding RNAs in development and disease: background, mechanisms, and therapeutic approaches. Physiol Rev. 2016;96:1297–325.
Trimarchi T, Bilal E, Ntziachristos P, Fabbri G, Dalla-Favera R, Tsirigos A, et al. Genome-wide mapping and characterization of Notch-regulated long noncoding RNAs in acute leukemia. Cell. 2014;158:593–606.
Kino T, Hurt DE, Ichijo T, Nader N, Chrousos GP. Noncoding RNA gas5 is a growth arrest- and starvation-associated repressor of the glucocorticoid receptor. Sci Signal. 2010;3:ra8.
Hudson WH, Pickard MR, de Vera IM, Kuiper EG, Mourtada-Maarabouni M, Conn GL, et al. Conserved sequence-specific lincRNA-steroid receptor interactions drive transcriptional repression and direct cell fate. Nat Commun. 2014;5:5395.
Yuan JH, Yang F, Wang F, Ma JZ, Guo YJ, Tao QF, et al. A long noncoding RNA activated by TGF-β promotes the invasion-metastasis cascade in hepatocellular carcinoma. Cancer Cell. 2014;25:666–81.
Zhou N, Si Z, Li T, Chen G, Zhang Z, Qi H. Long non-coding RNA CCAT2 functions as an oncogene in hepatocellular carcinoma, regulating cellular proliferation, migration and apoptosis. Oncol Lett. 2016;12:132–8.
Liu L, Zhao X, Zhu X, Zhong Z, Xu R, Wang Z, et al. Decreased expression of miR-430 promotes the development of bladder cancer via the upregulation of CXCR7. Mol Med Rep. 2013;8:140–6.
Chen GS, Zhou N, Li JQ, Li T, Zhang ZQ, Si ZZ. Restoration of miR-20a expression suppresses cell proliferation, migration, and invasion in HepG2 cells. Onco Targets Ther. 2016;9:3067–76.
Shen ED, Liu B, Yu XS, Xiang ZF, Huang HY. The effects of miR-1207-5p expression in peripheral blood on cisplatin-based chemosensitivity of primary gallbladder carcinoma. Onco Targets Ther. 2016;9:3633–42.
Xiao P, Liu W, Zhou H. miR-200b inhibits migration and invasion in non-small cell lung cancer cells via targeting FSCN1. Mol Med Rep. 2016;14:1835–40.
Wang Y, Kong D. Knockdown of lncRNA MEG3 inhibits viability, migration, and invasion and promotes apoptosis by sponging miR-127 in osteosarcoma cell. J Cell Biochem. 2018;119:669–79.
Xu R, Feng F, Yu X, Liu Z, Lao L. LncRNA SNHG4 promotes tumour growth by sponging miR-224-3p and predicts poor survival and recurrence in human osteosarcoma. Cell Prolif. 2018;51:e12515.
Liu K, Hou Y, Liu Y, Zheng J. LncRNA SNHG15 contributes to proliferation, invasion and autophagy in osteosarcoma cells by sponging miR-141. J Biomed Sci. 2017;24:46.
Kong D, Wang Y. Knockdown of lncRNA HULC inhibits proliferation, migration, invasion, and promotes apoptosis by sponging miR-122 in osteosarcoma. J Cell Biochem. 2018;119:1050–61.
Zhang H, Lu W. LncRNA SNHG12 regulates gastric cancer progression by acting as a molecular sponge of miR‑320. Mol Med Rep. 2018;17:2743–9.
Zhang L, Kang W, Lu X, Ma S, Dong L, Zou B. LncRNA CASC11 promoted gastric cancer cell proliferation, migration and invasion in vitro by regulating cell cycle pathway. Cell Cycle. 2018;17:1886–900.
Liu J, Liu L, Wan JX, Song Y. Long noncoding RNA SNHG20 promotes gastric cancer progression by inhibiting p21 expression and regulating the GSK-3β/ β-catenin signaling pathway. Oncotarget. 2017;8:80700–8.
Sun M, Nie F, Wang Y, Zhang Z, Hou J, He D, et al. LncRNA HOXA11-AS promotes proliferation and invasion of gastric cancer by scaffolding the chromatin modification factors PRC2, LSD1, and DNMT1. Cancer Res. 2016;76:6299–310.
Wang C, Han C, Zhang Y, Liu F. LncRNA PVT1 regulate expression of HIF1α via functioning as ceRNA for miR‑199a‑5p in non‑small cell lung cancer under hypoxia. Mol Med Rep. 2018;17:1105–10.
Wang X, Zhang G, Cheng Z, Dai L, Jia L, Jing X, et al. Knockdown of LncRNA-XIST suppresses proliferation and TGF-β1-induced EMT in NSCLC through the Notch-1 Pathway by regulation of miR-137. Genet Test Mol Biomark. 2018;22:333–42.
Chen Z, Xu D, Zhang T. Inhibition of proliferation and invasion of hepatocellular carcinoma cells by lncRNA-ASLNC02525 silencing and the mechanism. Int J Oncol. 2017;51:851–8.
Lv J, Kong Y, Gao Z, Liu Y, Zhu P, Yu Z. LncRNA TUG1 interacting with miR-144 contributes to proliferation, migration and tumorigenesis through activating the JAK2/STAT3 pathway in hepatocellular carcinoma. Int J Biochem Cell Biol. 2018;101:19–28.
Gu J, Wang Y, Wang X, Zhou D, Shao C, Zhou M, et al. Downregulation of lncRNA GAS5 confers tamoxifen resistance by activating miR-222 in breast cancer. Cancer Lett. 2018;434:1–10.
Hu HB, Chen Q, Ding SQ. LncRNA LINC01116 competes with miR-145 for the regulation of ESR1 expression in breast cancer. Eur Rev Med Pharmacol Sci. 2018;22:1987–93.
Ma Y, Bu D, Long J, Chai W, Dong J. LncRNA DSCAM-AS1 acts as a sponge of miR-137 to enhance Tamoxifen resistance in breast cancer. J Cell Physiol. 2019;234:2880–94.
Cordes KR, Sheehy NT, White MP, Berry EC, Morton SU, Muth AN, et al. miR-145 and miR-143 regulate smooth muscle cell fate and plasticity. Nature. 2009;460:705–10.
Huang FT, Chen WY, Gu ZQ, Zhuang YY, Li CQ, Wang LY, et al. The novel long intergenic noncoding RNA UCC promotes colorectal cancer progression by sponging miR-143. Cell Death Dis. 2017;8:e2778.
Pagliuca A, Valvo C, Fabrizi E, di Martino S, Biffoni M, Runci D, et al. Analysis of the combined action of miR-143 and miR-145 on oncogenic pathways in colorectal cancer cells reveals a coordinate program of gene repression. Oncogene. 2013;32:4806–13.
Zhang JX, Song W, Chen ZH, Wei JH, Liao YJ, Lei J, et al. Prognostic and predictive value of a microRNA signature in stage II colon cancer: a microRNA expression analysis. Lancet Oncol. 2013;14:1295–306.
Zou H, Wang S, Wang S, Wu H, Yu J, Chen Q, et al. SOX5 interacts with YAP1 to drive malignant potential of non-small cell lung cancer cells. Am J Cancer Res. 2018;8:866–78.
Wunderle VM, Critcher R, Ashworth A, Goodfellow PN. Cloning and characterization of SOX5, a new member of the human SOX gene family. Genomics. 1996;36:354–8.
Wang D, Han S, Wang X, Peng R, Li X. SOX5 promotes epithelial–mesenchymal transition and cell invasion via regulation of Twist1 in hepatocellular carcinoma. Med Oncol. 2015;32:461.
Kordaß T, Weber CE, Oswald M, Ast V, Bernhardt M, Novak D, et al. SOX5 is involved in balanced MITF regulation in human melanoma cells. BMC Med Genom. 2016;9:10.
Wu K, Zhao Z, Liu K, Zhang J, Li G, Wang L. Long noncoding RNA lnc-sox5 modulates CRC tumorigenesis by unbalancing tumor microenvironment. Cell Cycle. 2017;16:1295–301.
Howlader N,NA, Krapcho M, Garshell J, Miller D, Altekruse SF, Kosary CL, et al. SEER cancer statistics review, 1975–2012. National Cancer Institute; 2015.
Luo W, He H, Xiao W, Liu Q, Deng Z, Lu Y, et al. MALAT1 promotes osteosarcoma development by targeting TGFA via MIR376A. Oncotarget. 2016;7:54733–43.
Li W, Zhai L, Wang H, Liu C, Zhang J, Chen W, et al. Downregulation of LncRNA GAS5 causes trastuzumab resistance in breast cancer. Oncotarget. 2016;7:27778–86.
Zhang YL, Li XB, Hou YX, Fang NZ, You JC, Zhou QH. The lncRNA XIST exhibits oncogenic properties via regulation of miR-449a and Bcl-2 in human non-small cell lung cancer. Acta Pharmacol Sin. 2017;38:371–81.
Lai T, Jabaudon D, Molyneaux BJ, Azim E, Arlotta P, Menezes JR, et al. SOX5 controls the sequential generation of distinct corticofugal neuron subtypes. Neuron. 2008;57:232–47.
Ikeda T, Zhang J, Chano T, Mabuchi A, Fukuda A, Kawaguchi H, et al. Identification and characterization of the human long form of Sox5 (L-SOX5) gene. Gene. 2002;298:59–68.
Stolt CC, Schlierf A, Lommes P, Hillgärtner S, Werner T, Kosian T, et al. SoxD proteins influence multiple stages of oligodendrocyte development and modulate SoxE protein function. Dev Cell. 2006;11:697–709.
Hersh CP, Silverman EK, Gascon J, Bhattacharya S, Klanderman BJ, Litonjua AA, et al. SOX5 is a candidate gene for chronic obstructive pulmonary disease susceptibility and is necessary for lung development. Am J Respir Crit Care Med. 2011;183:1482–9.
Shiseki M, Masuda A, Yoshinaga K, Mori N, Okada M, Motoji T, et al. Identification of the SOX5 gene as a novel IGH-involved translocation partner in BCL2-negative follicular lymphoma with t(12;14)(p12.2;q32). Int J Hematol. 2015;102:633–8.
Hu J, Tian J, Zhu S, Sun L, Yu J, Tian H, et al. Sox5 contributes to prostate cancer metastasis and is a master regulator of TGF-beta-induced epithelial mesenchymal transition through controlling Twist1 expression. Br J Cancer. 2018;118:88–97.
Li G, Wang K, Wang J, Qin S, Sun X, Ren H. miR-497-5p inhibits tumor cell growth and invasion by targeting SOX5 in non-small-cell lung cancer. J Cell Biochem. 2019;120:10587–95.
Liu P, Li X, Guo X, Chen J, Li C, Chen M, et al. Circular RNA DOCK1 promotes bladder carcinoma progression via modulating circDOCK1/hsa-miR-132-3p/Sox5 signalling pathway. Cell Prolif. 2019;52:e12614.
Ding S, Zhang Y. MicroRNA‑539 inhibits the proliferation and migration of gastric cancer cells by targeting SRY‑box 5 gene. Mol Med Rep. 2019;20:2533–40.
Acknowledgements
We would like to give our sincere gratitude to the reviewers for their constructive comments.
Funding
This work was supported by funding Natural Science Foundation of Hunan Province (2019JJ80090), National Multidisciplinary Cooperative Diagnosis and Treatment Capacity Building Project for Major Diseases (Lung Cancer).
Author information
Authors and Affiliations
Contributions
RC: study design, definition of intellectual content, data acquisition, data analysis, statistical analysis, manuscript preparation, manuscript editing, manuscript review. CZ: guarantor of integrity of the entire study, study concepts, study design. YC: literature research, clinical studies, experimental studies, data acquisition, statistical analysis, manuscript preparation, manuscript editing. SW: clinical studies, experimental studies, data acquisition, statistical analysis. HL: clinical studies, experimental studies, data acquisition. HZ: guarantor of integrity of the entire study, study concepts, study design, literature research, data analysis, manuscript editing, manuscript review.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no competing interests.
Ethics approval and consent to participate
All patients were well-informed and provided written informed consent. Current study was approved by the medical ethics committee of Xiangya Hospital Central South University.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
About this article
Cite this article
Chen, R., Zhang, C., Cheng, Y. et al. LncRNA UCC promotes epithelial–mesenchymal transition via the miR-143-3p/SOX5 axis in non-small-cell lung cancer. Lab Invest 101, 1153–1165 (2021). https://doi.org/10.1038/s41374-021-00586-6
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41374-021-00586-6