Abstract
Background
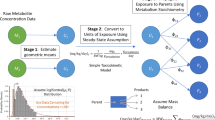
Toxicokinetic (TK) data needed for chemical risk assessment are not available for most chemicals. To support a greater number of chemicals, the U.S. Environmental Protection Agency (EPA) created the open-source R package “httk” (High Throughput ToxicoKinetics). The “httk” package provides functions and data tables for simulation and statistical analysis of chemical TK, including a population variability simulator that uses biometrics data from the National Health and Nutrition Examination Survey (NHANES).
Objective
Here we modernize the “HTTK-Pop” population variability simulator based on the currently available data and literature. We provide explanations of the algorithms used by “httk” for variability simulation and uncertainty propagation.
Methods
We updated and revised the population variability simulator in the “httk” package with the most recent NHANES biometrics (up to the 2017–18 NHANES cohort). Model equations describing glomerular filtration rate (GFR) were revised to more accurately represent physiology and population variability. The model output from the updated “httk” package was compared with the current version.
Results
The revised population variability simulator in the “httk” package now provides refined, more relevant, and better justified estimations.
Significance
Fulfilling the U.S. EPA’s mission to provide open-source data and models for evaluations and applications by the broader scientific community, and continuously improving the accuracy of the “httk” package based on the currently available data and literature.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 6 print issues and online access
$259.00 per year
only $43.17 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
The data from this study are provided within the article and its Supplementary Files.
References
Egeghy PP, Judson R, Gangwal S, Mosher S, Smith D, Vail J, et al. The exposure data landscape for manufactured chemicals. Sci total Environ. 2012;414:159–66.
Breyer S. Breaking the vicious circle: toward effective risk regulation. Cambridge (MA): Harvard University Press; 2009.
Judson R, Richard A, Dix DJ, Houck K, Martin M, Kavlock R, et al. The toxicity data landscape for environmental chemicals. Environ Health Perspect. 2009;117:685.
Kavlock RJ, Bahadori T, Barton-Maclaren TS, Gwinn MR, Rasenberg M, Thomas RS. Accelerating the pace of chemical risk assessment. Chem Res Toxicol. 2018;31:287–90.
Dix DJ, Houck KA, Martin MT, Richard AM, Setzer RW, Kavlock RJ. The toxcast program for prioritizing toxicity testing of environmental chemicals. Toxicological Sci. 2006;95:5–12.
Collins FS, Gray GM, Bucher JR. Transforming environmental health protection. Science. 2008;319:906.
Breen M, Ring CL, Kreutz A, Goldsmith MR, Wambaugh JF. High-throughput PBTK models for in vitro to in vivo extrapolation. Expert Opin Drug Metab Toxicol. 2021;17:903–21.
Coecke S, Pelkonen O, Leite SB, Bernauer U, Bessems JG, Bois FY, et al. Toxicokinetics as a key to the integrated toxicity risk assessment based primarily on non-animal approaches. Toxicol Vitr. 2013;27:1570–7.
Bessems JG, Loizou G, Krishnan K, Clewell HJ III, Bernasconi C, Bois F, et al. PBTK modelling platforms and parameter estimation tools to enable animal-free risk assessment: recommendations from a joint EPAA–EURL ECVAM ADME workshop. Regulatory Toxicol Pharmacol. 2014;68:119–39.
Wambaugh JF, Bare JC, Carignan CC, Dionisio KL, Dodson RE, Jolliet O, et al. New approach methodologies for exposure science. Curr Opin Toxicol. 2019;15:76–92.
Pearce RG, Setzer RW, Strope CL, Sipes NS, Wambaugh JF. Httk: R package for high-throughput toxicokinetics. J Stat Softw. 2017;79:1–26.
Health Canada. Science approach document—bioactivity exposure ratio: application in priority setting and risk assessment. 2021.
Paul Friedman K, Gagne M, Loo LH, Karamertzanis P, Netzeva T, Sobanski T, et al. Utility of in vitro bioactivity as a lower bound estimate of in vivo adverse effect levels and in risk-based prioritization. Toxicol Sci. 2020;173:202–25.
U.S. Congress, Frank R. Lautenberg chemical safety for the 21st century act. Public Law 114–182 (114th Congress). Washington, DC: U.S. Congress; 2016.
Ring CL, Pearce RG, Setzer RW, Wetmore BA, Wambaugh JF. Identifying populations sensitive to environmental chemicals by simulating toxicokinetic variability. Environ Int. 2017;106:105–18.
Levey AS, Stevens LA, Schmid CH, Zhang YL, Castro AF 3rd, Feldman HI, et al. A new equation to estimate glomerular filtration rate. Ann Intern Med. 2009;150:604–12.
Eneanya ND, Yang W, Reese PP. Reconsidering the consequences of using race to estimate kidney function. JAMA. 2019;322:113–4.
UW Medicine to exclude race from calculation of eGFR (measure of kidney function) [press release]. Seattle (WA): University of Washington Department of Medicine; 2020.
Delgado C, Baweja M, Burrows NR, Crews DC, Eneanya ND, Gadegbeku CA, et al. Reassessing the inclusion of race in diagnosing kidney diseases: an interim report from the NKF-ASN task force. Am J Kidney Dis. 2021;78:103–15.
Delgado C, Baweja M, Crews DC, Eneanya ND, Gadegbeku CA, Inker LA, et al. A unifying approach for GFR estimation: recommendations of the NKF-ASN task force on reassessing the inclusion of race in diagnosing kidney disease. Am J Kidney Dis. 2022;79:268–88.e1.
Kaufman JD, Hajat A. Confronting environmental racism. Environ Health Perspect. 2021;129:51001.
U.S. Environmental Protection Agency. Environmental Justice. 2022. https://www.epa.gov/environmentaljustice.
Wambaugh JF, Hughes MF, Ring CL, MacMillan DK, Ford J, Fennell TR, et al. Evaluating in vitro-in vivo extrapolation of toxicokinetics. Toxicological Sci. 2018;163:152–69.
Wambaugh JF, Wetmore BA, Ring CL, Nicolas CI, Pearce RG, Honda GS, et al. Assessing toxicokinetic uncertainty and variability in risk prioritization. Toxicol Sci. 2019;172:235–51.
Tonnelier A, Coecke S, Zaldívar J-M. Screening of chemicals for human bioaccumulative potential with a physiologically based toxicokinetic model. Arch Toxicol. 2012;86:393–403.
Wetmore BA, Wambaugh JF, Allen B, Ferguson SS, Sochaski MA, Setzer RW, et al. Incorporating high-throughput exposure predictions with dosimetry-adjusted in vitro bioactivity to inform chemical toxicity testing. Toxicol Sci. 2015;148:121–36.
Johnson TN, Rostami-Hodjegan A, Tucker GT. Prediction of the clearance of eleven drugs and associated variability in neonates, infants and children. Clin Pharmacokinet. 2006;45:931–56.
Bernstein AS, Kapraun DF, Schlosser PM. A model template approach for rapid evaluation and application of physiologically based pharmacokinetic models for use in human health risk assessments: a case study on per- and polyfluoroalkyl substances. Toxicol Sci. 2021;182:215–28.
Kim SJ, Choi EJ, Choi GW, Lee YB, Cho HY. Exploring sex differences in human health risk assessment for PFNA and PFDA using a PBPK model. Arch Toxicol. 2019;93:311–30.
Kriz W, Bankir L. A standard nomenclature for structures of the kidney. The Renal Commission of the International Union of Physiological Sciences (IUPS). Kidney Int. 1988;33:1–7.
Strope CL, Mansouri K, Clewell HJ, Rabinowitz JR, Stevens C, Wambaugh JF. High-throughput in-silico prediction of ionization equilibria for pharmacokinetic modeling. Sci Total Environ. 2018;615:150–60.
Mansouri K, Cariello NF, Korotcov A, Tkachenko V, Grulke CM, Sprankle CS, et al. Open-source QSAR models for pKa prediction using multiple machine learning approaches. J Cheminformatics. 2019;11:1–20.
Mansouri K, Grulke CM, Judson RS, Williams AJ. OPERA models for predicting physicochemical properties and environmental fate endpoints. J Cheminformatics. 2018;10:10.
Williams WW, Hogan JW, Ingelfinger JR. Time to eliminate health care disparities in the estimation of kidney function. N Engl J Med. 2021;385:1804–6.
Duggal V, Thomas IC, Montez-Rath ME, Chertow GM, Kurella, Tamura M. National estimates of CKD prevalence and potential impact of estimating glomerular filtration rate without race. J Am Soc Nephrol. 2021;32:1454–63.
Young BA. Removal of race from estimation of kidney function. Nat Rev Nephrol. 2022;18:201–2.
Hsu CY, Yang W, Parikh RV, Anderson AH, Chen TK, Cohen DL, et al. Race, genetic ancestry, and estimating kidney function in CKD. N. Engl J Med. 2021;385:1750–60.
Levey AS, Titan SM, Powe NR, Coresh J, Inker LA. Kidney disease, race, and GFR estimation. Clin J Am Soc Nephrol. 2020;15:1203–12.
Acknowledgements
The authors thank Drs. Xiaoqing Chang and Kristin Eccles for their helpful U.S. EPA internal reviews of the manuscript. We greatly appreciate Dr. Sarah Davidson-Fritz for support with software engineering for the “httk” R package. We thank Drs. Peter Egeghy and Risa Sayre for useful conversations. Although the manuscript was reviewed by the US EPA and approved for publication, it may not necessarily reflect official Agency policy. Mention of trade names or commercial products does not constitute endorsement or recommendation for use.
Funding
The United States Environmental Protection Agency (EPA) through its Office of Research and Development (ORD) funded the research described here. This project was supported by appointments to the Internship/Research Participation Program at ORD and administered by the Oak Ridge Institute for Science and Education through an interagency agreement between the U.S. Department of Energy and U.S. EPA.
Author information
Authors and Affiliations
Contributions
Conceptualization, methodology, investigation, writing: MB, JFW, AB, MS, and CLR; software, data curation, validation, and formal analysis, project administration: MB, JFW, and CLR. All authors have read and agreed to the published version of the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
About this article
Cite this article
Breen, M., Wambaugh, J.F., Bernstein, A. et al. Simulating toxicokinetic variability to identify susceptible and highly exposed populations. J Expo Sci Environ Epidemiol 32, 855–863 (2022). https://doi.org/10.1038/s41370-022-00491-0
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41370-022-00491-0
Keywords
This article is cited by
-
Exposure forecasting – ExpoCast – for data-poor chemicals in commerce and the environment
Journal of Exposure Science & Environmental Epidemiology (2022)