Abstract
Objective
A causal obesity risk variant in the FTO locus was recently shown to inhibit adipocyte thermogenesis via increased adipose expression of the homeobox transcription factors IRX3 and IRX5. However, causal effects of IRX5 on fat storage remain to be shown in vivo, and discovery of downstream mediators may open new therapeutic avenues.
Methods
17 WT and 13 Irx5 knockout (KO) mice were fed low-fat control (Ctr) or high-fat (HF) diet for 10 weeks. Body weight, energy intake and fat mass were measured. Irx5-dependent gene expression was explored by transcriptome analysis of epididymal white adipose tissue (eWAT), confirmatory obesity-dependent expression in human adipocytes in vivo, and in vitro knock-down, overexpression and transcriptional activation assays.
Results
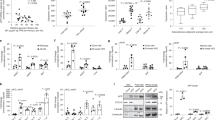
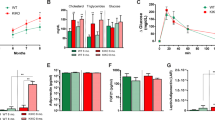
Irx5 knock-out mice weighed less, had diminished fat mass, and were protected from diet-induced fat accumulation. Key adipose mitochondrial genes Pparγ coactivator 1-alpha (Pgc-1α) and uncoupling protein 1 (Ucp1) were upregulated, and a gene network centered on amyloid precursor protein (App) was downregulated in adipose tissue of knock-out mice and in isolated mouse adipocytes with stable Irx5 knock-down. An APP-centered network was also enriched in isolated adipocytes from obese compared to lean humans. IRX5 overexpression increased APP promoter activity and both IRX5 and APP inhibited transactivation of PGC-1α and UCP1. Knock-down of Irx5 or App increased mitochondrial respiration in adipocytes.
Conclusion
Irx5-KO mice were protected from obesity and this can partially be attributed to reduced adipose App and improved mitochondrial respiration. This novel Irx5-App pathway in adipose tissue is a possible therapeutic entry point against obesity.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Allison DB, Kaprio J, Korkeila M, Koskenvuo M, Neale MC, Hayakawa K. The heritability of body mass index among an international sample of monozygotic twins reared apart. Int J Obes Relat Metab Disord. 1996;20:501–6.
Speliotes EK, Willer CJ, Berndt SI, Monda KL, Thorleifsson G, Jackson AU, et al. Association analyses of 249,796 individuals reveal 18 new loci associated with body mass index. Nat Genet. 2010;42:937–48.
Hertel JK, Johansson S, Sonestedt E, Jonsson A, Lie RT, Platou CGP, et al. FTO, type 2 diabetes, and weight gain throughout adult life: a meta-analysis of 41,504 subjects from the scandinavian HUNT, MDC, and MPP studies. Diabetes. 2011;60:1637–44.
Claussnitzer M, Dankel SN, Kim K-H, Quon G, Meuleman W, Haugen C, et al. FTO obesity variant circuitry and adipocyte browning in humans. N Engl J Med. 2015;373:895–907.
Dankel SN, Fadnes DJ, Stavrum AK, Stansberg C, Holdhus R, Hoang T, et al. Switch from stress response to homeobox transcription factors in adipose tissue after profound fat loss. PLoS ONE 2010;5.
Bürglin TR, Affolter M. Homeodomain proteins: an update. Chromosoma. 2016;125:497–521.
Hueber SD, Lohmann I. Shaping segments: Hox gene function in the genomic age. Bioessays. 2008;30:965–79.
Bonnard C, Strobl AC, Shboul M, Lee H, Merriman B, Nelson SF, et al. Mutations in IRX5 impair craniofacial development and germ cell migration via SDF1. Nat Genet. 2012;44:709–13.
Cheng CW, Chow RL, Lebel M, Sakuma R, Cheung HOL, Thanabalasingham V, et al. The Iroquois homeobox gene, Irx5, is required for retinal cone bipolar cell development. Dev Biol. 2005;287:48–60.
Costantini DL, Arruda EP, Agarwal P, Kim KH, Zhu Y, Zhu W, et al. The homeodomain transcription factor Irx5 establishes the mouse cardiac ventricular repolarization gradient. Cell. 2005;123:347–58.
Gaborit N, Sakuma R, Wylie JN, Kim K-H, Zhang S-S, Hui C-C, et al. Cooperative and antagonistic roles for Irx3 and Irx5 in cardiac morphogenesis and postnatal physiology. Development. 2012;139:4007–19.
Smemo S, Tena JJ, Kim K-H, Gamazon ER, Sakabe NJ, Gómez-Marín C, et al. Obesity-associated variants within FTO form long-range functional connections with IRX3. Nature. 2014;507:371–5.
Thomas EL, Saeed N, Hajnal JV, Brynes A, Goldstone AP, Frost G. et al. Magnetic resonance imaging of total body fat. J Appl Physiol. 1998;85:1778–85.
Lonowski LA, Narimatsu Y, Riaz A, Delay CE, Yang Z, Niola F, et al. Genome editing using FACS enrichment of nuclease-expressing cells and indel detection by amplicon analysis. Nat Protoc. 2017;12:581.
Lukas J, Bartkova J, Rohde M, Strauss M, Bartek J. Cyclin D1 is dispensable for G1 control in retinoblastoma gene-deficient cells independently of cdk4 activity. Mol Cell Biol. 1995;15:2600–11.
Hansen JB, Jørgensen C, Petersen RK, Hallenborg P, De Matteis R, Bøye HA, et al. Retinoblastoma protein functions as a molecular switch determining white versus brown adipocyte differentiation. Proc Natl Acad Sci USA. 2004;101:4112–7.
Hakim-Weber R, Krogsdam A-M, Jørgensen C, Fischer M, Prokesch A, Bogner-Strauss JG, et al. Transcriptional regulatory program in wild-type and retinoblastoma gene-deficient mouse embryonic fibroblasts during adipocyte differentiation. BMC Res Notes. 2011;4:157.
Rim JS, Kozak LP. Regulatory motifs for CREB-binding protein and Nfe2l2 transcription factors in the upstream enhancer of the mitochondrial uncoupling protein 1 gene. J Biol Chem. 2002;277:34589–600.
Handschin CC, Rhee JJ, Lin JJ, PTPT Tarr, BMBM. Spiegelman. An autoregulatory loop controls peroxisome proliferator-activated receptor gamma coactivator 1alpha expression in muscle. Pnas. 2003;100:7111–6.
Ye J, Coulouris G, Zaretskaya I, Cutcutache I, Rozen S, Madden TL. Primer-BLAST: a tool to design target-specific primers for polymerase chain reaction. BMC Bioinform. 2012;13:134.
Bø TH, Dysvik B, Jonassen I. LSimpute: accurate estimation of missing values in microarray data with least squares methods. Nucleic Acids Res. 2004;32:e34–e34.
Bolstad BM, Irizarry Ra, Astrand M, Speed TP. A comparison of normalization methods for high density oligonucleotide array data based on variance and bias. Bioinformatics. 2003;19:185–93.
Puig KL, Brose SA, Zhou X, Sens MA, Combs GF, Jensen MD, et al. Amyloid precursor protein modulates macrophage phenotype and diet-dependent weight gain. Sci Rep. 2017;7:43725.
Balakrishnan K, Verdile G, Mehta PD, Beilby J, Nolan D, Galvão DA, et al. Plasma Aβ42 correlates positively with increased body fat in healthy individuals. J Alzheimers Dis. 2005;8:269–82.
Lee Y-H, Tharp WG, Maple RL, Nair S, Permana PA, Pratley RE. Amyloid precursor protein expression is upregulated in adipocytes in obesity. Obesity. 2008;16:1493–500.
Ghanim H, Monte SV, Sia CL, Abuaysheh S, Green K, Caruana Ja, et al. Reduction in inflammation and the expression of amyloid precursor protein and other proteins related to Alzheimer’s disease following gastric bypass surgery. J Clin Endocrinol Metab. 2012;97:E1197–201.
Lee Y-H, Martin JM, Maple RL, Tharp WG, Pratley RE. Plasma amyloid-beta peptide levels correlate with adipocyte amyloid precursor protein gene expression in obese individuals. Neuroendocrinology. 2009;90:383–90.
Puig KL, Floden AM, Adhikari R, Golovko MY, Combs CK. Amyloid precursor protein and proinflammatory changes are regulated in brain and adipose tissue in a murine model of high fat diet-induced obesity. PLoS ONE. 2012;7.
Freeman LR, Zhang L, Dasuri K, Fernandez-Kim SO, Bruce-Keller AJ, Keller JN. Mutant amyloid precursor protein differentially alters adipose biology under obesogenic and non-obesogenic conditions. PLoS ONE. 2012;7.
Wang J, Gu BJ, Masters CL, Wang YJ. A systemic view of Alzheimer disease—insights from amyloid-β metabolism beyond the brain. Nat Rev Neurol. 2017;13:612–23.
Sun Q, Li J, Gao F. New insights into insulin: the anti-inflammatory effect and its clinical relevance. World J Diabetes. 2014;5:89–96.
Dandona P, Mohamed I, Ghanim H, Sia CL, Dhindsa S, Dandona S, et al. Insulin suppresses the expression of amyloid precursor protein, presenilins, and glycogen synthase kinase-3beta in peripheral blood mononuclear cells. J Clin Endocrinol Metab. 2011;96:1783–8.
Mattson MP. Pathways towards and away from Alzheimer’s disease. Nature. 2004;430:631–9.
Anandatheerthavarada HK, Devi L. Amyloid precursor protein and mitochondrial dysfunction in Alzheimer’s disease. Neurosci. 2007;13:626–38.
Chen JX, Yan SDu. Amyloid-beta-induced mitochondrial dysfunction. J Alzheimers Dis. 2007;12:177–84.
Shi C, Zhu XM, Wang J, Long D. Intromitochondrial IκB/NF-κB signaling pathway is involved in amyloid β peptide-induced mitochondrial dysfunction. J Bioenerg Biomembr. 2014;46:371–6.
Schöttl T, Kappler L, Fromme T, Klingenspor M. Limited OXPHOS capacity in white adipocytes is a hallmark of obesity in laboratory mice irrespective of the glucose tolerance status. Mol Metab. 2015;4:631–42.
Acknowledgements
We thank Carol Cook, Margit Solsvik, Rita Holdhus, Alba Kaci, Regine Åsen Jersin, André Madsen, Mona Nilsen and Tone Flølo for excellent technical assistance. We thank the patients and hospital staff at Voss and Haraldsplass Hospitals for the adipose tissue biopsies. The Irx5-KO mice were a gift from Drs. Kyoung-Han Kim and Chi-chung Hui. The pGL4-phAPP-luc reporter construct was provided by the RIKEN BRC through the National Bio-Resource Project of the MEXT, Japan. The mPgc-1α-luc construct was a gift from Dr. Bruce Spiegelman and the pGL3-UCP1-luc construct was a gift from Dr. Leslie P. Kozak. The ME3 cells were a gift from Dr. Karsten Kristiansen.
Author contributions
SD and GM conceived the study and designed the experiments. JIB, CH, and OG designed and conducted the experiments. HJN and VV obtained the human adipose tissue biopsies and performed the bariatric surgery. OPN made the CRISPR-cells. SD and JIB performed the data analyses. JIB, CH and SD drafted the manuscript. JIB, CH, OG, OPN, HJN, VV, PRN, JVS, SD and GM contributed to data interpretation, discussion and reviewed and edited the manuscript.
Funding
Funding for this project was provided by Persontilpasset medisin for barn og voksne med diabetes mellitus (PERSON-MED-DIA), the Western Norway Regional Health Authority and Meltzerfondet. PRN was supported by the European Research Council (AdG #293574).
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Electronic Information
Rights and permissions
About this article
Cite this article
Bjune, JI., Haugen, C., Gudbrandsen, O. et al. IRX5 regulates adipocyte amyloid precursor protein and mitochondrial respiration in obesity. Int J Obes 43, 2151–2162 (2019). https://doi.org/10.1038/s41366-018-0275-y
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41366-018-0275-y
This article is cited by
-
MRI adipose tissue segmentation and quantification in R (RAdipoSeg)
Diabetology & Metabolic Syndrome (2022)
-
IRX5 promotes adipogenesis of hMSCs by repressing glycolysis
Cell Death Discovery (2022)
-
Deficiency of Irx5 protects mice from obesity and associated metabolic abnormalities
International Journal of Obesity (2022)
-
Irx3 and Irx5 in Ins2-Cre+ cells regulate hypothalamic postnatal neurogenesis and leptin response
Nature Metabolism (2021)
-
Dysregulation of amyloid precursor protein impairs adipose tissue mitochondrial function and promotes obesity
Nature Metabolism (2019)