Abstract
Background/Objectives
Obesity is an important risk factor for the development of diseases such as diabetes mellitus, hypertension, and dyslipidemia; however, a small number of individuals with long-standing obesity do not present with these cardiometabolic diseases. Such individuals are referred to as metabolically healthy obese (MHO) and potentially represent a subgroup of the general population with a protective genetic predisposition to obesity-related diseases. We hypothesized that individuals who were metabolically healthy, but significantly obese (BMI ≥ 35 kg/m2) would represent a highly homogenous subgroup, with which to investigate potential genetic associations to obesity. We further hypothesized that such a cohort may lend itself well to investigate potential genotypes that are protective with respect to the development of cardiometabolic disease.
Subjects/Methods
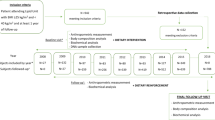
In the present study, we implemented this novel selection strategy by screening 892 individuals diagnosed as Class 2 or Class 3 obese and identified 38 who presented no manifestations of cardiometabolic disease. We then assessed these subjects for single-nucleotide polymorphisms (SNPs) that associated with this phenotype.
Results
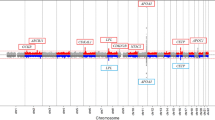
Our analysis identified 89 SNPs that reach statistical significance (p < 1 × 10−5), some of which are associated with genes of biological pathways that influences dietary behavior; others are associated with genes previously linked to obesity and cardiometabolic disease as well as neuroimmune disease. This study, to the best of our knowledge, represents the first genetic screening of a cardiometabolically healthy, but significantly obese population.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Ogden CL, Carroll, MD, Fryar, CD, Flegal, KM Prevalence of Obesity Among Adults and Youth: United States, 2011–2014. Centers for Disease Control and Prevention, November 2015;288:1-2
Hruby A, Manson JE, Qi L, Malik VS, Rimm EB, Sun Q, et al. Determinants and consequences of obesity. Am J Public Health. 2016;106:1656–62.
Stunkard AJ, Sorensen TI. Obesity and socioeconomic status--a complex relation. N Engl J Med. 1993;329:1036–7.
Reddon H, Gueant JL, Meyre D. The importance of gene-environment interactions in human obesity. Clin Sci. 2016;130:1571–97.
Goodarzi MO. Genetics of obesity: what genetic association studies have taught us about the biology of obesity and its complications. Lancet Diabetes Endocrinol. 2018;6:223–36
Sims EA. Are there persons who are obese, but metabolically healthy? Metabolism. 2001;50:1499–504.
Karelis AD, St-Pierre DH, Conus F, Rabasa-Lhoret R, Poehlman ET. Metabolic and body composition factors in subgroups of obesity: what do we know? J Clin Endocrinol Metab. 2004;89:2569–75.
Garcia-Moll X. Obesity and prognosis: time to forget about metabolically healthy obesity. Eur Heart J. 2018;39:407–9
Munoz-Garach A, Cornejo-Pareja I, Tinahones FJ. Does metabolically healthy obesity exist? Nutrients. 2016;8:1–10
Espinosa De Ycaza AE, Donegan D, Jensen MD. Long-term metabolic risk for the metabolically healthy overweight/obese phenotype. Int J Obesity. 2018;42:302–9
Iglesias Molli AE, Penas Steinhardt A, Lopez AP, Gonzalez CD, Vilarino J, Frechtel GD, et al. Metabolically healthy obese individuals present similar chronic inflammation level but less insulin-resistance than obese individuals with metabolic syndrome. PLoS ONE. 2017;12:e0190528.
Bluher M. The distinction of metabolically ‘healthy’ from ‘unhealthy’ obese individuals. Curr Opin Lipidol. 2010;21:38–43.
National Cholesterol Education Program Expert Panel on Detection, Evaluation, and Treatment of High Blood Cholesterol in Adults. Third Report of the National Cholesterol Education Program (NCEP) expert panel on detection, evaluation, and treatment of high blood cholesterol in adults (Adult Treatment Panel III) final report. Circulation. 2002;106:3143–421.
Smith AK, Fang H, Whistler T, Unger ER, Rajeevan MS. Convergent genomic studies identify association of GRIK2 and NPAS2 with chronic fatigue syndrome. Neuropsychobiology. 2011;64:183–94.
Gauderman WJMJ. QUANTO 1.2.4: a computer program for power and sample size calculations for genetic–epidemiology studies. Los Angeles, CA: University of Southern California; 2009.
den Hoed M, Luan J, Langenberg C, Cooper C, Sayer AA, Jameson K, et al. Evaluation of common genetic variants identified by GWAS for early onset and morbid obesity in population-based samples. Int J Obes. 2013;37:191–6.
Schlauch KA, Khaiboullina SF, De Meirleir KL, Rawat S, Petereit J, Rizvanov AA, et al. Genome-wide association analysis identifies genetic variations in subjects with myalgic encephalomyelitis/chronic fatigue syndrome. Transl Psychiatry. 2016;6:e730.
Carvalho B, Bengtsson H, Speed TP, Irizarry RA. Exploration, normalization, and genotype calls of high-density oligonucleotide SNP array data. Biostatistics. 2007;8:485–99.
Gonzalez JR, Armengol L, Sole X, Guino E, Mercader JM, Estivill X, et al. SNPassoc: an R package to perform whole genome association studies. Bioinformatics. 2007;23:644–5.
Jorgenson E, Witte JS. Genome-wide association studies of cancer. Future Oncol. 2007;3:419–27.
Benjamini Y, Hochberg Y. Controlling the false discovery rate: a practical and powerful approach to multiple testing. J R Stat Soc. 1995;Series B:289–300.
Devlin B, Roeder K. Genomic control for association studies. Biometrics. 1999;55:997–1004.
Lehne B, Lewis CM, Schlitt T. From SNPs to genes: disease association at the gene level. PLoS ONE. 2011;6:e20133.
Radkowski P, Wator G, Skupien J, Bogdali A, Wolkow P. Analysis of gene expression to predict dynamics of future hypertension incidence in type 2 diabetic patients. BMC Proc. 2016;10(Suppl 7):113–7.
Al-Shammari MS, Al-Ali R, Al-Balawi N, Al-Enazi MS, Al-Muraikhi AA, Busaleh FN, et al. Type 2 diabetes associated variants of KCNQ1 strongly confer the risk of cardiovascular disease among the Saudi Arabian population. Genet Mol Biol. 2017;40:586–90.
van Vliet-Ostaptchouk JV, van Haeften TW, Landman GW, Reiling E, Kleefstra N, Bilo HJ, et al. Common variants in the type 2 diabetes KCNQ1 gene are associated with impairments in insulin secretion during hyperglycaemic glucose clamp. PLoS ONE. 2012;7:e32148.
Anveden A, Sjoholm K, Jacobson P, Palsdottir V, Walley AJ, Froguel P, et al. ITIH-5 expression in human adipose tissue is increased in obesity. Obesity. 2012;20:708–14.
Yeo GS, Connie Hung CC, Rochford J, Keogh J, Gray J, Sivaramakrishnan S, et al. A de novo mutation affecting human TrkB associated with severe obesity and developmental delay. Nat Neurosci. 2004;7:1187–9.
Dong C, Wong ML, Licinio J. Sequence variations of ABCB1, SLC6A2, SLC6A3, SLC6A4, CREB1, CRHR1 and NTRK2: association with major depression and antidepressant response in Mexican-Americans. Mol Psychiatry. 2009;14:1105–18.
International Multiple Sclerosis Genetics C, Beecham AH, Patsopoulos NA, Xifara DK, Davis MF, Kemppinen A, et al. Analysis of immune-related loci identifies 48 new susceptibility variants for multiple sclerosis. Nat Genet. 2013;45:1353–60.
Sturm R, Hattori A. Morbid obesity rates continue to rise rapidly in the United States. Int J Obes. 2013;37:889–91.
Flegal KM, Carroll MD, Kit BK, Ogden CL. Prevalence of obesity and trends in the distribution of body mass index among US adults, 1999-2010. JAMA. 2012;307:491–7.
Behrens M, Bartelt J, Reichling C, Winnig M, Kuhn C, Meyerhof W. Members of RTP and REEP gene families influence functional bitter taste receptor expression. J Biol Chem. 2006;281:20650–9.
Tepper BJ, Banni S, Melis M, Crnjar R, Tomassini Barbarossa I. Genetic sensitivity to the bitter taste of 6-n-propylthiouracil (PROP) and its association with physiological mechanisms controlling body mass index (BMI). Nutrients. 2014;6:3363–81.
Dotson CD, Zhang L, Xu H, Shin YK, Vigues S, Ott SH, et al. Bitter taste receptors influence glucose homeostasis. PLoS ONE. 2008;3:e3974.
Clark AA, Dotson CD, Elson AE, Voigt A, Boehm U, Meyerhof W, et al. TAS2R bitter taste receptors regulate thyroid function. FASEB J. 2015;29:164–72.
Vong QP, Leung WH, Houston J, Li Y, Rooney B, Holladay M, et al. TOX2 regulates human natural killer cell development by controlling T-BET expression. Blood. 2014;124:3905–13.
Stolarczyk E, Vong CT, Perucha E, Jackson I, Cawthorne MA, Wargent ET, et al. Improved insulin sensitivity despite increased visceral adiposity in mice deficient for the immune cell transcription factor T-bet. Cell Metab. 2013;17:520–33.
Kasuga M. KCNQ1, a susceptibility gene for type 2 diabetes. J Diabetes Investig. 2011;2:413–4.
Tavira B, Coto E, Diaz-Corte C, Ortega F, Arias M, Torres A, et al. KCNQ1 gene variants and risk of new-onset diabetes in tacrolimus-treated renal-transplanted patients. Clin Transplant. 2011;25:E284–91.
Al-Haggar M, Madej-Pilarczyk A, Kozlowski L, Bujnicki JM, Yahia S, Abdel-Hadi D, et al. A novel homozygous p.Arg527Leu LMNA mutation in two unrelated Egyptian families causes overlapping mandibuloacral dysplasia and progeria syndrome. Eur J Hum Genet. 2012;20:1134–40.
Cordovado SK, Hendrix M, Greene CN, Mochal S, Earley MC, Farrell PM, et al. CFTR mutation analysis and haplotype associations in CF patients. Mol Genet Metab. 2012;105:249–54.
Farh KK, Marson A, Zhu J, Kleinewietfeld M, Housley WJ, Beik S, et al. Genetic and epigenetic fine mapping of causal autoimmune disease variants. Nature. 2015;518:337–43.
San Francisco IF, Rojas PA, Torres-Estay V, Smalley S, Cerda-Infante J, Montecinos VP, et al. Association of RNASEL and 8q24 variants with the presence and aggressiveness of hereditary and sporadic prostate cancer in a Hispanic population. J Cell Mol Med. 2014;18:125–33.
Judge AD, Zhang X, Fujii H, Surh CD, Sprent J. Interleukin 15 controls both proliferation and survival of a subset of memory-phenotype CD8(+) T cells. J Exp Med. 2002;196:935–46.
Farooqi S, O’Rahilly S. Genetics of obesity in humans. Endocr Rev. 2006;27:710–18.
Frayling TM, Timpson NJ, Weedon MN, Zeggini E, Freathy RM, Lindgren CM, et al. A common variant in the FTO gene is associated with body mass index and predisposes to childhood and adult obesity. Science. 2007;316:889–94.
Yako YY, Guewo-Fokeng M, Balti EV, Bouatia-Naji N, Matsha TE, Sobngwi E, et al. Genetic risk of type 2 diabetes in populations of the African continent: a systematic review and meta-analyses. Diabetes Res Clin Pract. 2016;114:136–50.
Manriquez V, Aviles J, Salazar L, Saavedra N, Seron P, Lanas F et al. Polymorphisms in genes involved in the leptin-melanocortin pathway are associated with obesity-related cardiometabolic alterations in a Southern Chilean Population. Mol Diagnosis Therapy. 2017.
Suzuki R, Matsumoto M, Fujikawa A, Kato A, Kuboyama K, Yonehara K, et al. SPIG1 negatively regulates BDNF maturation. J Neurosci. 2014;34:3429–42.
Abbott GW, Tai KK, Neverisky DL, Hansler A, Hu Z, Roepke TK, et al. KCNQ1, KCNE2, and Na+-coupled solute transporters form reciprocally regulating complexes that affect neuronal excitability. Sci Signal. 2014;7:ra22.
Acknowledgements
We are grateful to the Coriell Genotyping and Microarray Center at Coriell Institute for Medical Research in Camden, NJ, and to its former Director, Dr. Norman P. Gerry, for their assistance in processing our specimens. This work was funded in part by the National Institute of General Medical Sciences (GM103440).
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Electronic supplementary material
Rights and permissions
About this article
Cite this article
Schlauch, K.A., Kulick, D., Subramanian, K. et al. Single-nucleotide polymorphisms in a cohort of significantly obese women without cardiometabolic diseases. Int J Obes 43, 253–262 (2019). https://doi.org/10.1038/s41366-018-0181-3
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41366-018-0181-3