Abstract
Ulcerative colitis (UC) is a chronic recurrent intestinal inflammatory disease characterized by high incidence and young onset age. Recently, there have been some interesting findings in the pathogenesis of UC. The mucus barrier, which is composed of a mucin complex rich in O-glycosylation, not only provides nutrients and habitat for intestinal microbes but also orchestrates the taming of germs. In turn, the gut microbiota modulates the production and secretion of mucins and stratification of the mucus layers. Active bidirectional communication between the microbiota and its ‘slimy’ partner, the mucus barrier, seems to be a continually performed concerto, maintaining homeostasis of the gut ecological microenvironment. Any abnormalities may induce a disorder in the gut community, thereby causing inflammatory damage. Our review mainly focuses on the complicated communication between the mucus barrier and gut microbiome to explore a promising new avenue for UC therapy.
Similar content being viewed by others
Introduction
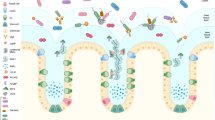
In recent years, the incidence of ulcerative colitis (UC), an inflammatory bowel disease (IBD) of unknown etiology, has been increasing globally, especially in some newly industrialized countries, including India and China1. Microbial infections such as those by Clostridium difficile have been described as a mono-associated cause of UC flare-ups2; however, there is growing evidence that UC is an overly robust mucosal immune response to dysbiosis of particular gut flora that is characterized by abnormal microbiota composition and bacterial products3,4. A balanced microbiome community is vital for maintaining mucus barrier homeostasis, which involves a dynamic balance of production, secretion, expansion, and proteolysis of mucus components. Commensal bacteria and their fermentation products (short-chain fatty acids, SCFAs) are implicated in the regulation of the production and secretion of mucin 2 (Muc2), the major component of mucus, in sentinel goblet cells (sGCs) at crypt opening5 (Fig. 1a). The gut microbiome also influences the mucosal structure. Carbon dioxide (CO2) generated from β-oxidation of SCFAs in colonocytes is converted by carbonic anhydrase to bicarbonate (HCO3−), which in turn dictates the stratification of the mucus layers, such as the unfolding of mucin and resultant inner-towards-outer conversion of the mucus layer6 (Fig. 1b).
a During homeostasis, the gut microbiome at the outer mucus layer modulates mucin production and secretion and mucus stratification mediated by HCO3− to maintain mucus barrier integrity. Dysbiosis induces impairment of the mucus barrier, accompanied by increased epithelium damage, bacterial translocation, goblet cell depletion, and host inflammation. b Gut microbiome-generated short-chain fatty acids enter colonocytes and are oxidized to generate CO2 that can be converted by carbonic anhydrase into HCO3−, which is the ideal physiological solution for precipitating calcium and raising the pH at the epithelial surface. This in turn promotes the stratification of the mucus layer. c Intestinal bacteria have evolved several strategies to adhere to the mucus barrier, including the use of adhesins, flagella, and fimbriae; achieve cross-feeding by mucin degradation; and maintain colonization resistance by means of a commensal type VI secretion system.
While the secreted, attached, hydrated, and stratified mucus barrier is mostly considered a simple lubricant layer overlying the epithelium, it also provides an environment for bacterial colonization and nourishes the commensal microbiota, thereby stabilizing the microbial community and promoting symbiotic interactions, resulting in microbial commensalism7. Mucus barrier abnormalities, including depleted upper crypt GCs, bacterial penetration of the inner mucus layer, and decreased core mucus components, such as FCGBP (human IgGFc binding protein), CLCA1 (calcium-activated chloride channel regulator 1), and ZG16 (zymogen granule protein 16), in active UC support the notion that an impaired mucus barrier may occur prior to the onset of inflammation in the pathogenesis of UC8. Environmental factors such as diet and lifestyle factors may shape the human gut microbiome composition, thereby influencing mucus homeostasis and the development of intestinal inflammatory lesions9. Dietary fiber-deprived intestinal microbiota consume components of the mucus layer, leading to intestinal barrier dysfunction and increased susceptibility to pathogens and colitis occurrence10. It is obvious that the interplay between the microbiota and its ‘slimy’ partner, the mucus barrier, in the gut is constitutive. Therefore, any attempt to simply explore the underlying mechanism of UC from any single part of the biosystem (the mucus barrier and gut microbiota) is unwise. Currently, the development of microbiome-targeted therapeutic strategies for mild to moderate UC is growing11, and mucus barrier-associated colonization resistance involves commensal bacteria out-competing foreign microbes for space, trophic resources and bactericidal factors in the mucus barrier and decreasing the efficacy of fecal microbiome transplantation (FMT) therapy. This review provides insight into mucus barrier-gut microbiome interactions.
The gut microbiome: orchestrator of the mucus barrier
The gut microbiome adheres to mucus
Compared with the small intestine, the colonic epithelium is covered by mucus layers composed of a firm inner layer and loose outer layer that function to separate microbes from epithelial cells and provide a diffusion barrier to maintain a balanced community. The outer mucus layer is colonized with an abundance of commensal microbes, while the inner layer is relatively sterile (Fig. 1a). The combination of the mucus barrier and gut microbiome, composed of approximately 100 trillion symbiotic microbial cells and more than 9000 carbohydrate-degrading enzymes, is described as “the last human body organ”12. Commensal bacteria and pathogens have evolved several strategies to occupy a narrowly defined niche within the mucus barrier.
The first strategy is to adhere to the mucus by surface display of adhesins and extracellular appendages (fimbria) that bind to specific mucin glycans (Fig. 1c). Mucus-binding proteins (MUBs) are one class of effectors involved in the adherence of lactobacilli, abundant commensal bacteria in the human gut and the best studied example of mucus adhesins that confine commensal/probiotic bacteria to the outer mucus layer13. Phylogenetically, adhesins are proteins characterized by the MUB domain, which shares homology with the Pfam-MucBP (mucin-binding protein) domains14. MUB and MucBP domain-containing proteins contain a C-terminal recognition motif (LPxTG) that is recognized by a family of enzymes called sortases for covalent attachment to peptidoglycan of the bacterial cell wall and an N-terminal region for protein secretion, in addition to a signal peptide (Table 1). A number of proteins containing MUB homologs and MucBP domains have been found; for instance, the mucin/mucin-binding protein of Lactobacillus fermentum BCS87 (32-Mmubp), S-layer protein in L. acidophilus (SlpA), MucBP-containing mannose-specific adhesin (Msa), and elongation factor Tu (EF-Tu) are highly prevalent in lactobacilli naturally existing in intestinal niches. Competitive adhesion studies have shown that MUB interacts with specific muco-oligosaccharides and that MUB binding has little to no host specificity regarding mucus components15. The second strategy of mucus adhesion is mediated by fimbrial adhesion of commensal bacteria. For example, Escherichia coli, a commensal bacterium residing in the human gut, has the potential to act as an opportunistic pathogen. E. coli strains use extracellular fimbriae, which have a two-domain organization: lectin at the most external N-terminal domain and pilin at the C-terminus connecting to the rest of the fimbria. The affinity and specificity of the adhesion by fimbrial proteins are governed by recognition of mucus glycan epitopes, which are age-, organ-, and species-specific16. However, for many bacterial pathogens, binding to mucus is a crucial step in their colonization. Flagella, composed of flagellin arranged in helical chains, are an important evolved strategy for mucus adhesion during infection by some pathogens, and they play a critical role in biofilm formation17. Enterotoxigenic E. coli (ETEC) strains are major causes of morbidity and mortality due to diarrheal illness in developing countries. ETEC-secreted pathovar-specific proteins (such as EtpA, a two-partner adhesin conserved within the ETEC pathovar) can interact with both the tips of ETEC flagella and mucus glycans to form molecular bridges promoting bacterial adhesion and intestinal colonization of pathogens18. Flagella are used as virulence factors by many enteropathogenic bacteria (e.g., Listeria monocytogenes, Vibrio cholerae, E. coli, and Salmonella typhimurium) to traverse the mucus barrier, resulting in infection. Flagella-driven motility propels pathogens towards the epithelium and accelerates disease progression19. Many human pathogens, including C. difficile, pathogenic E. coli, Neisseria meningitidis, and Streptococcus pneumoniae, also employ phase-variable flagella and fimbriae to evade the host immune system and promote host colonization, persistence, motility, and virulence20.
The gut microbiome feeds on mucin glycans
After adhesion to mucins, colonization by colonic bacteria is initiated, while the degradation of diverse and structurally complex mucin glycans depends on the cooperative action of sialidases, sulfatases, proteases, and glycoside hydrolases (GHs) encoded by the genomes of mucin-degrading bacteria (Fig. 1c). Mucin-degrading carbohydrate-active enzyme (CAZyme) families include sialidases (GH33), fucosidases (GH29, GH95), blood-group endo-β-1,4-galactosidases (GH98), mucin core GHs (GH101, GH129, GH84, GH85, and GH89), and sulfatases (GH20, GH2, GH42, unclassified)21 (Table 2). Carbohydrate-binding modules (CBMs) in CAZymes mediate their adherence to carbohydrate substrates in mucin polymers22.
The adult gut microbiome consists of hundreds to thousands of different species of bacteria, with two predominant bacterial phyla: gram-positive Firmicutes and gram-negative Bacteroidetes23. Bacteroides spp. are prominent members of this microbial ecosystem and widely studied commensal bacteria24. They degrade a vast range of dietary and endogenous glycans by utilizing a complex transenvelope machinery known as starch utilization system (Sus)-like systems, which are encoded by coregulated clusters of genes known as polysaccharide utilization loci (PULs)25. Bacteroides spp., in particular B. thetaiotaomicron containing PULs, encode highly specific CAZymes and degrade a wide range of glycan substrates, thereby stratifying the niche space with different orders of substrate preferences, which is why they are sometimes referred to as “generalists”26. Akkermansia muciniphila can hydrolyze up to 85% of mucin structures using different enzyme combinations27, strengthen intestinal epithelial integrity, and fortify damaged gut barriers28. Interestingly, the abundances of A. muciniphila in both fecal samples and mucosal biopsies of UC patients are markedly reduced29. Butyrate, an SCFA produced by commensal bacteria, is the main energy source of colonocytes and exerts various beneficial effects, such as enhancement of intestinal barrier function. The production of butyrate using complex mucin glycans as a substrate is generally restricted to some Clostridium clusters (IV and XIVa) from the Firmicutes phylum. In addition, the butyrogenic effect of A. muciniphila30 is related to its cross-feeding with mucus-degrading Clostridium clusters (IV and XIVa).
Notably, continual glycan degradation mediated by bacterial glycosidases may cause the disappearance of host-specific glycan epitopes and degradation of the protein backbone (Table 1). Dietary fiber-deprived intestinal microbiota have been shown to actively forage on the mucus layer, leading to dysfunction of the intestinal barrier and increased host susceptibility to pathogens and inflammation10 (Fig. 1c). It was reported that pathogenic Proteobacteria and Firmicutes species, including Salmonella enterica serovar Typhimurium, E. coli, and C. difficile, can benefit from cross-feeding through consumption of sialic acids from mucin molecules released by B. thetaiotaomicron. The expansion of pathogens during colitis is directly dependent on sialic acid released from host glycans catalyzed by sialidases31. Oral administration of a sialidase inhibitor and low levels of intestinal α2,3-linked sialic acid decreased E. coli outgrowth and colitis severity in mice32. The cleavage site of the zinc metalloprotease zmpB from C. perfringens was established to be next to the mucus glycoprotein backbone (Ser and/or Thr residues), with optimal splicing of GlcNAcβ1–3 (Neu5Acα2–6), GalNAcα1, or GalNAcα1 (α2,6-sialylated core 1 or core-3 O-glycan)33.
The gut microbiome modulates mucus layer dynamics
Mucin production was reported to be induced by the gut microbiome. SCFAs such as acetate, propionate, and butyrate, the fermentation products of commensal bacteria, enhance the synthesis of mucin and stimulated mucin secretion in mice34. Moreover, the stimulating effect of butyrate on Muc2 expression is mediated via AP-1 at the Muc2 promoter35. Lactic acid-based probiotics, containing Lactobacilli and Bifidobacteria, increase mucin production in human intestinal epithelial cells and block enteropathogenic E. coli invasion and adherence in vitro36. Bifidobacterium species colonizing the intestinal mucus barrier modulate mucus production and expulsion by increasing the expression of GC markers such as Krüppel-like factor 4 (KLF4), trefoil factor 3 (TFF3), resistin-like molecule-beta (Relm-β), and Muc 237. A randomized, placebo-controlled trial tested the efficacy and safety of a highly concentrated mixture of probiotic bacterial strains (VSL#3) in active UC and its role in the maintenance of UC remission38 and demonstrated that the protective effect of VSL#3 was related to enhanced colonic mucin expression and secretion in vivo and in vitro39. Several bacterial Toll-like receptor (TLR) ligands or effectors (e.g., lipopolysaccharide (LPS), flagellin, probiotic agents, commensal bacteria, and bacterial fermentation products) have been shown to trigger Muc2 expression in colonic sGCs5,40. In addition, Muc2 production can also be enhanced by several stimuli, including T-helper type 1 (Th1)- and Th2 cell-mediated cytokines, acute phase responses (colonic ischemia), and viral infection41,42.
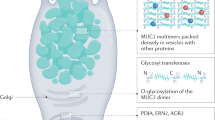
The gut microbiome is also involved in the modulation of mucus secretion by GCs. Non-O-glycosylated mucins with molecular weights of approximately 500 kDa are synthesized in the endoplasmic reticulum of GCs and dimerized via disulfide bonds between the cystine knot (CK) domains. Mucin dimers transported to the Golgi apparatus are subjected to O-glycosylation and then multimerization by disulfide bonds at N-terminal von Willebrand factor type D3 (vWF D3) domains. The resulting polymers reach molecular weights of up to 2.5 million Da43. Mucin multimers of 10–50 MDa [extended rods 1–10 μm in length] are then packaged in an ordered state within secretory vesicles (<1 μm) in the presence of low pH and high calcium44 (Fig. 1a). Upon secretion, the densely packed mucins can expand >1,000-fold, resulting in the formation of enormous net-like polymeric sheets45. Secretion of mucin can occur in at least two ways: regulated vesicle secretion and compound exocytosis. During regulated vesicle secretion (also called regulated exocytosis), the membrane of a secretory vesicle fuses with the plasma membrane by mediating the actions of typical vesicle exocytosis components such as syntaxins, mammalian uncoordinated-18 (Munc-18), vesicle-associated membrane proteins (VAMP), and synaptosome-associated proteins (SNAP), and this is a tightly controlled process most often triggered by calcium46. In compound exocytosis, storage vesicles rapidly fuse with the GC membrane after fusion with each other and empty all thecal contents47. The inner mucus layer is continuously renewed by mucin secretion of the surface GCs, and renewal of the inner mucus layer is estimated to occur every 1–2 h in live murine distal colonic tissue48. In general, spontaneous mucus production occurs at a rate of 240 μm/h in humans and 100 μm/h in the mouse colon; thus, the colonic mucus is continuously renewed at an average of 5–10 L per day49. Recently, sGCs have been shown to endocytose bacteria-derived TLR agonists such as LPS, lipid A, and flagellin but not lipotechoic acid, bacterial DNA, muramyl dipeptide, or γ-D-glutamylmeso-diaminopimelic acid and activate TLR- and MyD88-dependent NOD-like receptor family pyrin domain containing 6 (NLRP6) signaling to facilitate the exocytosis of mucin and flush bacteria away from crypt openings ex vivo5.
Stratification of the mucus layer has been shown to be indirectly influenced by the gut microbiome (Fig. 1a). An increase in pH and removal of N-terminally bound single calcium ions are necessary for the conversion of the inner firm mucus layer to the outer loose mucus layer, the so-called mucus layer stratification50. In general, colonocytes are mainly dependent on adenosine triphosphate produced by the β-oxidation of butyrate, a metabolite of the gut microbiome, which is accompanied by the generation of CO2 that can be converted by carbonic anhydrase into HCO3−51; this is the ideal physiological solution for precipitating calcium and raising the pH at the epithelial surface52. The absence of HCO3− at the intestinal epithelial surface or inhibition of HCO3− transepithelial transport decreases the amounts and rates of stimulated mucus release in vitro and in vivo53. For instance, facultative anaerobic bacteria such as pathogenic E. coli and Salmonella expand and invade the surface epithelium, thereby subverting colonocyte metabolism from β-oxidation of SCFAs to anaerobic glycolysis to promote their own luminal growth in competition against the gut microbiota by increasing the luminal bioavailability of oxygen (O2), lactate, and additional electron acceptors, including tetrathionate (S4O62−) and nitrate (NO3−)51,54. The resultant decrease in HCO3− in the lumen creates a high-H+ environment, enhancing the Ca2+-binding of mucin polymers and making them more adhesive to each other in condensed mucin granules55. As a result, the structure of mucus layers is impaired, and host susceptibility to pathogens and even UC incidence increases; therefore, UC was postulated to be an energy-deficient disease resulting from a failure to utilize butyrate56.
The mucus barrier regulates bacterial colonization
The mucus layer creates a habitat for commensal bacterial colonization
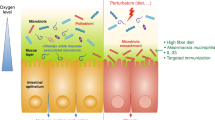
Hosts have evolved multiple strategies to maintain homeostasis of the intestinal microbiota (Fig. 2a). The best strategy is a highly adaptable protective mucus barrier exhibiting a heterogeneous spatial structure that establishes a habitat for commensal bacteria (Fig. 2b). The mucus barrier is a natural defense at the interface between host tissue and the luminal microbial community. Muc2 is the basic component of mucus that is continuously secreted and replenished by GCs in the large intestine. In the endoplasmic reticulum, the amino-terminal vWF and carboxy-terminal cystine knot (CK) domains of Muc2 mediate disulfide crosslinking of mucins to build a much larger mucin fishnet comprising thousands of monomers57,58. Muc2 consists of multiple domains, including the PTS [proline (Pro), threonine (Thr), and serine (Ser)] domain, a hallmark of the mucin family that is composed of a variable number of tandem repeats (VNTRs) that allow for heavy O-glycosylation with great heterogeneity in the Golgi apparatus and a stretched, brush-like arrangement of mucin. Neutral or negatively charged sugars, including N-acetylgalactosamine (GalNAc), sulfated acetyl-D-glucosamine (GlcNAc), D-galactose (Gal), sulfated Gal, sialic acid (Neu5Ac), and fucose, are attached to the PTS domains under catalysis by glycosyltransferases in the Golgi apparatus. Ultimately, these glycans account for up to 80% of the total mucin mass21. Importantly, the vast repertoire of O-glycosylated epitopes derived from the peripheral terminus of mucins (such as sialic acid and fucose) creates a habitat for unique bacterial ecosystems that thrive in proximity to host tissue59,60. Species of Bacteroides, the most abundant genus of the human gut microbiome, have a unique class of polysaccharide-utilizing loci that are referred to as commensal colonization factors (CCFs). Bacteroides fragilis can penetrate the colonic mucus and reside deep within crypt channels, whereas strains with CCF mutations are defective in crypt invasion61. It is known that reestablishment and resilience are fundamental characteristics of the gut microbial community61,62. The recolonization of gut B. fragilis following microbiome disruption caused by Citrobacter rodentium infection or antibiotic treatment is also dependent on CCFs61,63. Sulfatase (BF3086) and glycosyl hydrolase (BF3134) were annotated as mucosal colonization factors in B. fragilis. BF3086 is also important for B. fragilis to metabolize host mucus O-glycans64. During colonic mucus colonization, B. fragilis upregulates the expression of a set of candidate colonization factors, including BF3086 and BF3134, while in-frame deletions of these factors reduce its colonization abilities, which are fully or partially recovered by transcomplementation of BF3134 or BF308664.
a The mucus barrier forms a fundamental niche for gut microbiome colonization, where the major O-glycan epitopes are sialic acid, fucose, N-acetylneuraminic acid (Neu5Ac), type A antigen [GalNAcα1,3(Fucα1,2)Galβ], and type 1 H antigens [Fucα1,2Galβ1,3(GlcNAc)]. b The mucus barrier dictates the spatial organization of microbes, forming a steric and orderly microorganism network to inhibit pathogen colonization. c The mucus barrier is also a scaffold containing antimicrobial agents [including RELM-β (purple solid circle), ZG16 (blue solid circle), Ang4 (red solid circle), Lypd8 (green solid circle), sIgA (orange solid circle), and bacteriophages] protecting epithelial cells against microbes.
The inhibition of symbiotic bacterial colonization by pathogens is mediated by degradation of mucosal glycosylation and includes decreasing fucosylation and increasing the release of sialic acid, which promotes the outgrowth and colonization of pathogenic E. coli32. LPS induces an increase in the expression of microbial virulence genes, such as RtxA (K10953) and hemolysin III (K11068), which enhance intestinal colonization of pathogenic microbes in fucosyltransferase 2 (Fut2)-deficient mice65. Enterohemorrhagic E. coli (EHEC) encodes a two-component sensing system (FusKR) consisting of a histidine sensor kinase (FusK) and response regulator (FusR). During colonization, EHEC cleaves fucose from mucin, thereby activating the FusKR signaling cascade and increasing the expression of virulence genes66. It was observed that S. typhimurium had significantly increased expression of genes (nan, fuc, and pdu) that utilize host mucin monosaccharides such as sialic acid, fucose, and propanediol, the catabolite of fucose, in gnotobiotic mice colonized with sialidase-expressing B. thetaiotaomicron67. Furthermore, antibiotic-treated conventional mice exhibited a transient surge in free sialic acid liberated by the resident microbiota from host mucus, promoting the expansion of Salmonella and C. difficile expressing sialic acid catabolic signaling67. As a result, it was concluded that antibiotic-associated pathogens such as S. typhimurium and C. difficile catabolize fucose and sialic acid liberated by the resident microbiota from mucin glycans in a resident microbiota-dependent manner67. Pathogens have also evolved a range of mucin-hydrolyzing enzymes called mucinases (glycosidases, proteases, and sulfatases) to degrade mucin complexes due to the mucus net-like nature. Notably, some commensal bacteria also produce mucinases, but their expression levels are much lower (Fig. 1c). Compared to pathogenic E. coli, commensal E. coli strains generate a lower amount of YghJ68, a lipoprotein with a zinc metalloprotease domain that is involved in mucin degradation as well as proinflammatory responses.
The colonization of commensals at the mucus layer also renders host resistance to pathogen colonization. CCFs mediate the production of a polysaccharide capsule around B. fragilis, thereby initiating an IL-36γ response in mucosal macrophages of the gut to prevent colonization and infection by Klebsiella pneumoniae, which is a multidrug-resistant pathogen with high lethality69. Pathogens can be directly killed or inhibited by commensals that produce several antibacterial compounds. For example, bacteriocins produced by commensal E. coli inhibit EHEC70, microbicides secreted by Enterobacteriaceae mediate interspecies competition in the inflamed gut71, the bacteriocin thuricin produced by Bacillus thuringiensis inhibits the proliferation of C. difficile and L. monocytogenes72, and lantibiotics produced by lactic acid bacteria are used to target pathogens73. In addition, mucin was found to affect microbial behavior. For instance, gram-negative pathogens V. cholerae74 and S. Typhimurium75 as well as commensals from the Bacteroides genus76 were reported to exert bactericidal effects mediated by the Type VI secretory system (T6SS) (Fig. 1c). It was recently revealed that mucin-associated glycans activate RetS, the sensor kinase of Pseudomonas aeruginosa, thereby inhibiting T6SS-dependent bacterial killing action77,78.
Epithelial surface pH modulates the gut microbiota composition
There are two key transport systems for HCO3− extrusion into the colonic lumen: Cl−/HCO3− and SCFA/HCO3− exchangers79 (Fig. 1b). Several lines of evidence indicate that SCFA/HCO3− exchangers mediate ionized SCFA entry into colonocytes concomitant with an increase in luminal pH and a decrease in oxygen tension in both human and rodent colons80, which are vital for the stratification of the secreted mucin complex and colonization of obligate anaerobes, respectively. Treatment with live Bifidobacterium and its culture supernatants stimulated the expression of Slc26a3, a Cl−/HCO3− exchanger81. Inflammation in the mid-distal82 or distal colon83 in Slc26a3-deficient mice was related to the loss of mucus secretion resulting from a remarkably low surface pH microclimate83, a more aggressive microbiota82 and/or reduced microbiome diversity83. A luminal microenvironment with higher oxygen and lower pH could change the gut microbiota composition and drive an uncontrolled luminal expansion of E. coli and Salmonella84.
Mucus viscosity determines the spatial organization of the gut microbiota
The intestinal microflora is not evenly mixed but is spatially organized (Fig. 2c). Some mechanisms for the spatial organization of gut bacteria have been elucidated. Mucus is mainly composed of water (95% w/w), mucins (0.2–5.0% w/v), globular proteins (0.5% w/v), salts (0.5–1.0% w/w), lipids (1–2% w/w), DNA, cells, and cellular debris that form a dense, viscoelastic layer over epithelial cells85. There is a longitudinal (proximal to distal colon) viscosity gradient that increases progressively towards the distal colon in murine models, which restricts bacterial motility and confers spatial organization of bacterial populations. As a result, bacteria are selectively separated from the mucosa in the proximal colon and completely separated in the mid-distal colon86. Of note, uncovered cecum epithelium tips are a hotspot for S. typhimurium infection in mice due to the lack of a continuous mucus layer19. In the proximal murine colon, select bacterial populations intimately contact the mucosa and enter the crypts, thereby concentrating and forming a 20–240-μm thick film flanking the mucosa. The existence of vertical (surface to lumen) viscosity gradients within the colonic mucus layer was further demonstrated by low mucus viscosity at the crypt base and high viscosity at sites adjacent to the columnar epithelium or close to the intestinal lumen. A viscosity-dependent spatial distribution of bacteria in the murine colon revealed that short rods and cocci moved best in low viscosity, while long curly bacteria preferred a moderately viscous environment, and all bacteria were immobilized by high viscosity87. The lower viscosity of mucus at the crypt base makes intestinal cells more vulnerable to invasion by potential pathogens. In general, mucins contain several crosslinking domains to form dimers and larger-order structures via disulfide bonds that may be broken by sulfate-reducing bacteria (SRB), particularly Desulfovibrio desulfuricans88. Many studies have described a high abundance of SRB detected in the mucosa of UC patients89,90. The resultant mucus barrier becomes less viscous and more permeable, allowing the gut microbiota in the gut lumen to interact with epithelial cells, thereby causing an aberrant immune response91. Recent studies have revealed the importance of site-specific gene expression for robust host-microbial symbiosis. B. fragilis near the epithelium upregulates the expression of genes involved in protein synthesis; moreover, compared to bacteria in the lumen, B. fragilis in mucus and tissue has high levels of sulfatase (BF3086) and glycosyl hydrolase (BF3134)64. Intestinal mechanics are a host spatial control measure capable of regulating the abundance and persistence of gut bacteria. A V. cholerae symbiont native to zebrafish that governs its spatial organization using swimming motility and chemotaxis displayed strong localization to the foregut region, an anatomical region comparable to the mammalian small intestine with close contact with the intestinal epithelium to counter intestinal flow. In contrast, motility-deficient mutants that are susceptible to host spatial control largely aggregated within the intestinal mucus and were confined to the lumen, whereas chemotaxis-deficient mutants were restricted to the lumen of the midgut, and two mutants were susceptible to intestinal expulsion. Wild-type V. cholerae actively escapes mucus through regular changes in swimming direction mediated by chemotactic signaling92.
There are some factors influencing the viscosity of the mucus layer, TFF3 and HCO3−. TFF3, as a component of mucus, is essential for protection of the gastrointestinal mucosa93. It is a small cysteine-rich acidic secreted protein that is covalently bound to the C-terminal domain of Muc294. Mucus viscosity has been shown to increase after the introduction of TFF3 dimers (0.3% w/v) compared with no treatment95. Tff3-knockout mice are more susceptible to dextran sulfate sodium (DSS)-induced colitis96,97, while oral treatment with TFF3 protected against DSS-induced colitis in mice93.
There are two separate signaling pathways vital for normal mucus formation: Ca2+-mediated exocytosis of mucin granules of GCs and independent cAMP-mediated, cystic fibrosis transmembrane conductance regulator (CFTR)-dependent HCO3− secretion, which helps discharge sulfated and sialylated glycosylated domains85 and stratifies exocytosed mucus98. Additionally, HCO3− also participates in mucin expansion and hydration mechanisms by reducing Ca2+ cross-linking in mucins, thereby decreasing the viscosity55. CFTR is the secretory chloride/HCO3− channel; its dysfunction causes acidification of the mucus layer (pH < 6.5) due to defective HCO3− release, resulting in increased mucus viscoelasticity and the formation of a stationary mucus layer in cystic fibrosis99.
The mucus barrier generates a protective shield
Colonic mucus is a key component of the colonic barrier, as it is located at the interface between luminal microflora and the colonic mucosa. The mucus barrier effectively partitions the enteric epithelium from the microbiota as the first line of defense and supports the growth of intestinal commensals as an energy source. The development of colitis in animals lacking a functional mucus layer closely reflects clinical and cellular features in patients with active UC. Penetration of the inner mucus layer in the distal colon by pathogens and/or commensals often found in mice with colitis is related to impaired mucus barrier structure and function caused by genetic deficiency in Muc2100, inactivation of glycosyltransferase-mediated O-glycosylation of Muc2101,102, deficiency of the NLRP6 inflammasome, or exposure to colitis-inducing chemicals103. Some pathogens such as enterohemorrhagic or enteropathogenic E. coli (EHEC or EPEC), C. rodentium, and S. typhimurium disrupt the protective mucus barrier, causing dysbiosis characterized by decreased abundances of Firmicutes and Verrucomicrobia and increased abundances of Bacteroidetes and facultative anaerobes104, which adhere to or invade host epithelial cells beneath the mucus layer. The vicious cycle of dysbiosis and colonic inflammation is characterized by destruction of the mucus barrier and persistent overstimulation of the immune system by the microflora19. Chronic or intermittent dietary fiber deficiency pushes the resident microbiota to rely more heavily on endogenous nutrients (host-secreted mucin glycoproteins), leading to erosion of the colonic mucus barrier and exacerbation of colitis triggered by the mucosal pathogen C. rodentium10.
Antimicrobial agents fortify the mucus barrier
Importantly, the dense gel-forming structure of the mucus layer acts as a trap to stabilize numerous molecules, such as RELM-β and zymogen granule protein 16 (ZG16), angiogenin 4 (Ang4), Ly6/PLAUR domain containing 8 (Lypd8), and secretory immunoglobulin A (sIgA) (Fig. 2b). RELM-β exerts a microbicidal effect predominantly on gram-positive pathogens penetrating the mucus layer104. ZG16 prevents the adherence of bacteria to the epithelium by binding to the peptidoglycan of the bacterial cell wall105. Ang 4, another antimicrobial agent derived from GCs, is associated with Trichuris muris expulsion from the colonic epithelium of mice during inflammation106. B. thetaiotaomicron promotes Ang 4 expression, which inhibits the growth of some bacterial species, such as L. monocytogenes and Enterococcus faecalis107. Lypd 8, a highly glycosylated glycosylphosphatidylinositol-anchored protein selectively expressed in enterocytes, can bind to flagellated bacteria to inhibit bacterial invasion into the colonic epithelia when secreted into the lumen. Lypd8 strongly causes early-phase defense against C. rodentium, which can induce colitis by triggering attachment and effacement (A/E) lesions on colonic epithelia. Mechanistically, Lypd8 inhibits C. rodentium attachment to intestinal epithelial cells by binding to intimin, thereby protecting against enteric bacterial pathogens108. sIgA secreted as a dimer by colonocytes and integrated into the mucus layer exerts a critical function in trapping luminal bacteria to prevent unrestricted access of the microbiota to the epithelial surface109. The decreasing gradient of antimicrobial agents from the epithelial surface to the lumen is positively correlated with mucin concentration in the bilayered mucus matrix, which is why the intestinal mucus layers harbor significant antibacterial activity, whereas only low activity is detected in the luminal content. Because of the anti-inflammatory and antimicrobial nature of mucosal contents, the mucus layer generates a protective shield to prevent bacterial translocation and inappropriate immune stimulation of the epithelium110. However, when a functional mucus layer is absent, the gradient of antimicrobial agents is diminished, and the related defense system is eliminated from the intestine with fecal flow23.
Bacteriophage attachment to mucus strengthens mucus defense
Under homeostatic conditions, mucus provides protection against dysbiosis by bacteriophage deployment (Fig. 2b). Bacteriophage, a resident member of the gut microbiome, interacts with mucin glycoproteins in the mucus barrier though immunoglobulin-like domains that are exposed on the capsid, triggering nonhost-derived immunity, which is considered part of the innate immune system111. Adherent invasive E. coli (AIEC) strain LF82 has type 1 pili mediating its binding to the host adhesion receptor carcinoembryonic antigen-related cell adhesion molecule 6 (CEACAM6), which is more strongly expressed in the ileal tissues of patients with Crohn’s disease (CD)112. A single day of oral treatment with a cocktail of bacteriophages was found to induce significantly decreased intestinal colonization by AIEC strain LF82 in CEABAC10 transgenic mice113. Moreover, this single dose of bacteriophage inhibited DSS-induced colitis symptoms over a two-week period in conventional mice colonized with LF82114. Bacteriophage intervention is planned to be evaluated in patients with IBD in the United States113. Data from UC mouse models have revealed that some bacteriophages that infect bacteria with pathogenic potential (pathobionts) are elevated during colitis115. Specifically, an increased abundance of bacteriophages predicted to infect Streptococcus sp. and Alistipes and Clostridiales phages predicted to infect C. difficile were observed during colitis116. This elevated abundance of specific phages could be postulated as a proxy for strain-level resolution of disease-causing bacteria during IBD116. It has been reported that intestinal microbiota-associated phages attach to mucins and protect underlying epithelial cells from invading bacteria117. Spatial organization of the mucus generates a gradient of phage replication with lysogeny at the top mucosal layer and lytic predation in the bacteria-sparse intermediary layers117. However, animals with bacteriophage expansion, such as Caudovirales phages, exhibit a significant exacerbation of intestinal colitis118. This inconsistency indicates a complex role of phages in IBD.
Conclusion
Massive advances in the etiology of UC over the past few decades have improved our understanding of the importance of active communication between the gut microbiota and the mucus barrier. It is evident that disturbance of this interplay is a vital pathological factor for UC development. From the perspective of intricate interactions between the mucus barrier and the gut microbiome in the gut microenvironment, it is important to explore interventional approaches to control inflammation or promote FMT. Hence, exploring promising therapeutic agents from the viewpoint of ‘slimy’ partners is necessary to effectively treat UC.
References
Alatab, S. et al. The global, regional, and national burden of inflammatory bowel disease in 195 countries and territories, 1990–2017: a systematic analysis for the Global Burden of Disease Study 2017. Lancet Gastroenterol. Hepatol. 5, 17–30 (2020).
Clayton, E. M. et al. The vexed relationship between Clostridium difficile and inflammatory bowel disease: an assessment of carriage in an outpatient setting among patients in remission. Am. J. Gastroenterol. 104, 1162–1169 (2009).
Round, J. L. & Mazmanian, S. K. The gut microbiota shapes intestinal immune responses during health and disease. Nat. Rev. Immunol. 9, 313–323 (2009).
Tremaroli, V. & Bäckhed, F. Functional interactions between the gut microbiota and host metabolism. Nature 489, 242–249 (2012).
Birchenough, G. M., Nyström, E. E., Johansson, M. E. & Hansson, G. C. A sentinel goblet cell guards the colonic crypt by triggering Nlrp6-dependent Muc2 secretion. Science 352, 1535–1542 (2016).
Gad, S. C. Toxicology of the gastrointestinal tract 2th edn. (CRC Press, Boca Raton, 2018).
Paone, P. & Cani, P. D. Mucus barrier, mucins and gut microbiota: the expected slimy partners? Gut 69, 2232–2243 (2020).
Van der Post, S. et al. Structural weakening of the colonic mucus barrier is an early event in ulcerative colitis pathogenesis. Gut 68, 2142–2151 (2019).
Rothschild, D. et al. Environment dominates over host genetics in shaping human gut microbiota. Nature 555, 210–215 (2018).
Desai, M. S. et al. A dietary fiber-deprived gut microbiota degrades the colonic mucus barrier and enhances pathogen susceptibility. Cell 167, 1339–1353. e1321 (2016).
Costello, S. P. et al. Effect of fecal microbiota transplantation on 8-week remission in patients with ulcerative colitis: a randomized clinical trial. JAMA 321, 156–164 (2019).
Proctor, L. Priorities for the next 10 years of human microbiome research. Nature 569, 623–625 (2019).
Etzold, S. & Juge, N. Structural insights into bacterial recognition of intestinal mucins. Curr. Opin. Struct. Biol. 28, 23–31 (2014).
Juge, N. Microbial adhesins to gastrointestinal mucus. Trends Microbiol. 20, 30–39 (2012).
Roos, S. & Jonsson, H. A high-molecular-mass cell-surface protein from Lactobacillus reuteri 1063 adheres to mucus componentsThe GenBank accession number for the sequence reported in this paper is AF120104. Microbiology 148, 433–442 (2002).
Colomb, F. et al. Epithelial mucins and bacterial adhesion. in Carbohydr. Chem., Vol. 40 (ed. Amelia Pilar Rauter, A. P. R., Thisbe Lindhorst, Yves Queneau) 596–623 (Royal Society of Chemistry, 2014).
Lam, W. W.-L., Sun, K., Zhang, H. & Au, S. W.-N. Crystal structure of flagellar export chaperone FliS in complex with flagellin and HP1076 of Helicobacter pylori. Front Microbiol. 11, 787 (2020).
Kumar, P. et al. Dynamic interactions of a conserved enterotoxigenic Escherichia coli adhesin with intestinal mucins govern epithelium engagement and toxin delivery. Infect. Immun. 84, 3608–3617 (2016).
Furter, M., Sellin, M. E., Hansson, G. C. & Hardt, W.-D. Mucus architecture and near-surface swimming affect distinct Salmonella Typhimurium infection patterns along the murine intestinal tract. Cell Rep. 27, 2665–2678. e2663 (2019).
Garrett, E. M. et al. Phase variation of a signal transduction system controls Clostridioides difficile colony morphology, motility, and virulence. PLoS Biol. 17, e3000379 (2019).
Tailford, L. E., Crost, E. H., Kavanaugh, D. & Juge, N. Mucin glycan foraging in the human gut microbiome. Front Genet. 6, 81 (2015).
Sidar, A. et al. Carbohydrate Binding Modules: Diversity of Domain Architecture in Amylases and Cellulases From Filamentous Microorganisms. Front Bioeng. Biotechnol. 8, 871–886 (2020).
Donaldson, G. P., Lee, S. M. & Mazmanian, S. K. Gut biogeography of the bacterial microbiota. Nat. Rev. Microbiol. 14, 20–32 (2016).
Bloom, S. M. et al. Commensal bacteroides species induce colitis in host-genotype-specific fashion in a mouse model of inflammatory bowel disease. Cell Host Microbe 9, 390–403 (2011).
Martens, E. C., Chiang, H. C. & Gordon, J. I. Mucosal glycan foraging enhances fitness and transmission of a saccharolytic human gut bacterial symbiont. Cell Host Microbe 4, 447–457 (2008).
Walter, J. & Ley, R. The human gut microbiome: ecology and recent evolutionary changes. Annu. Rev. Microbiol. 65, 411–429 (2011).
Trastoy, B., Naegeli, A., Anso, I., Sjögren, J. & Guerin, M. E. Structural basis of mammalian mucin processing by the human gut O-glycopeptidase OgpA from Akkermansia muciniphila. Nat. Commun. 11, 4844 (2020).
Reunanen, J. et al. Akkermansia muciniphila adheres to enterocytes and strengthens the integrity of the epithelial cell layer. Appl. Environ. Microbiol. 81, 3655–3662 (2015).
Png, C. W. et al. Mucolytic bacteria with increased prevalence in IBD mucosa augment in vitro utilization of mucin by other bacteria. Am. J. Gastroenterol. 105, 2420–2428 (2010).
Naito, Y., Uchiyama, K. & Takagi, T. A next-generation beneficial microbe: Akkermansia muciniphila. J. Clin. Biochem. Nutr. 63, 33–35 (2018).
Sorbara, M. T. & Pamer, E. G. Interbacterial mechanisms of colonization resistance and the strategies pathogens use to overcome them. Mucosal Immunol. 12, 1–9 (2019).
Huang, Y.-L., Chassard, C., Hausmann, M., Von Itzstein, M. & Hennet, T. Sialic acid catabolism drives intestinal inflammation and microbial dysbiosis in mice. Nat. Commun. 6, 1–11 (2015).
Noach, I. et al. Recognition of protein-linked glycans as a determinant of peptidase activity. Proc. Natl Acad. Sci. USA 114, E679–E688 (2017).
Gaudier, E. et al. Butyrate specifically modulates MUC gene expression in intestinal epithelial goblet cells deprived of glucose. Am. J. Physiol. Gastrointest. Liver Physiol. 287, G1168–G1174 (2004).
Burger-van Paassen, N. et al. The regulation of intestinal mucin MUC2 expression by short-chain fatty acids: implications for epithelial protection. Biochem. J. 420, 211–219 (2009).
Mack, D. R., Michail, S., Wei, S., McDougall, L. & Hollingsworth, M. A. Probiotics inhibit enteropathogenic E. coli adherence in vitro by inducing intestinal mucin gene expression. Am. J. Physiol. Gastrointest. Liver Physiol. 276, G941–G950 (1999).
Engevik, M. A. et al. Bifidobacterium dentium fortifies the intestinal mucus layer via autophagy and calcium signaling pathways. mBio 10, e01087–01019 (2019).
Miele, E. et al. Effect of a probiotic preparation (VSL# 3) on induction and maintenance of remission in children with ulcerative colitis. Am. J. Gastroenterol. 104, 437–443 (2009).
Caballero-Franco, C., Keller, K., De Simone, C. & Chadee, K. The VSL# 3 probiotic formula induces mucin gene expression and secretion in colonic epithelial cells. Am. J. Physiol. Gastrointest. Liver Physiol. 292, G315–G322 (2007).
Sicard, J.-F., Le Bihan, G., Vogeleer, P., Jacques, M. & Harel, J. Interactions of intestinal bacteria with components of the intestinal mucus. Front Cell Infect. Microbiol. 7, 387 (2017).
Nyström, E. Colonic mucus structure and processing. Sweden: University of Gothenburg (2018).
Grootjans, J. et al. Ischaemia-induced mucus barrier loss and bacterial penetration are rapidly counteracted by increased goblet cell secretory activity in human and rat colon. Gut 62, 250–258 (2013).
Thornton, D. J., Rousseau, K. & McGuckin, M. A. Structure and function of the polymeric mucins in airways mucus. Annu. Rev. Physiol. 70, 459–486 (2008).
Bansil, R. & Turner, B. S. The biology of mucus: composition, synthesis and organization. Adv. Drug Deliv. Rev. 124, 3–15 (2018).
Ambort, D. et al. Calcium and pH-dependent packing and release of the gel-forming MUC2 mucin. Proc. Natl Acad. Sci. USA 109, 5645–5650 (2012).
Birchenough, G. M., Johansson, M. E., Gustafsson, J. K., Bergström, J. H. & Hansson, G. C. New developments in goblet cell mucus secretion and function. Mucosal Immunol. 8, 712–719 (2015).
Specian, R. D. & Neutra, M. R. Mechanism of rapid mucus secretion in goblet cells stimulated by acetylcholine. J. Cell Biol. 85, 626–640 (1980).
Johansson, M. E. Fast renewal of the distal colonic mucus layers by the surface goblet cells as measured by in vivo labeling of mucin glycoproteins. PLoS ONE 7, e41009 (2012).
Gustafsson, J. K. et al. An ex vivo method for studying mucus formation, properties, and thickness in human colonic biopsies and mouse small and large intestinal explants. Am. J. Physiol. Gastrointest. Liver Physiol. 302, G430–G438 (2012).
Ridley, C. et al. Assembly of the respiratory Mucin MUC5B a new model for a gel-forming Mucin. J. Biol. Chem. 289, 16409–16420 (2014).
Von Engelhardt, W. et al. Functional role of bicarbonate in propionate transport across guinea‐pig isolated caecum and proximal colon. J. Physiol. 477, 365–371 (1994).
Gustafsson, J. K. et al. Bicarbonate and functional CFTR channel are required for proper mucin secretion and link cystic fibrosis with its mucus phenotype. J. Exp. Med. 209, 1263–1272 (2012).
Garcia, M. A. S., Yang, N. & Quinton, P. M. Normal mouse intestinal mucus release requires cystic fibrosis transmembrane regulator–dependent bicarbonate secretion. J. Clin. Invest. 119, 3497–3497 (2009).
Cani, P. D. Gut cell metabolism shapes the microbiome. Science 357, 548–549 (2017).
Chen, E. Y., Yang, N., Quinton, P. M. & Chin, W.-C. A new role for bicarbonate in mucus formation. Am. J. Physiol. Lung Cell Mol. Physiol. 299, L542–L549 (2010).
Roediger, W. The colonic epithelium in ulcerative colitis: an energy-deficiency disease? Lancet 316, 712–715 (1980).
Martens, E. C., Neumann, M. & Desai, M. S. Interactions of commensal and pathogenic microorganisms with the intestinal mucosal barrier. Nat. Rev. Microbiol. 16, 457–470 (2018).
Werlang, C., Cárcarmo-Oyarce, G. & Ribbeck, K. Engineering mucus to study and influence the microbiome. Nat. Rev. Mater. 4, 134–145 (2019).
Cai, R. et al. Interactions of commensal and pathogenic microorganisms with the mucus layer in the colon. Gut Microbes, 11, 680–690 (2020).
Wagner, C., Wheeler, K. & Ribbeck, K. Mucins and their role in shaping the functions of mucus barriers. Annu. Rev. Cell. Dev. Biol. 34, 189–215 (2018).
Lee, S. M. et al. Bacterial colonization factors control specificity and stability of the gut microbiota. Nature 501, 426–429 (2013).
Cullen, T. et al. Antimicrobial peptide resistance mediates resilience of prominent gut commensals during inflammation. Science 347, 170–175 (2015).
Donaldson, G. et al. Gut microbiota utilize immunoglobulin A for mucosal colonization. Science 360, 795–800 (2018).
Donaldson, G. P. et al. Spatially distinct physiology of Bacteroides fragilis within the proximal colon of gnotobiotic mice. Nat. Microbiol. 5, 746–756 (2020).
Pickard, J. M. et al. Rapid fucosylation of intestinal epithelium sustains host–commensal symbiosis in sickness. Nature 514, 638–641 (2014).
Pacheco, A. R. et al. Fucose sensing regulates bacterial intestinal colonization. Nature 492, 113–117 (2012).
Ng, K. M. et al. Microbiota-liberated host sugars facilitate post-antibiotic expansion of enteric pathogens. Nature 502, 96–99 (2013).
Luo, Q. et al. Enterotoxigenic Escherichia coli secretes a highly conserved mucin-degrading metalloprotease to effectively engage intestinal epithelial cells. Infect. Immun. 82, 509–521 (2014).
Sequeira, R. P., McDonald, J. A., Marchesi, J. R. & Clarke, T. B. Commensal bacteroidetes protect against Klebsiella pneumoniae colonization and transmission through IL-36 signalling. Nat. Microbiol. 5, 304–313 (2020).
Etcheverría, A. I., Arroyo, G. H., Perdigon, G. & Parma, A. E. Escherichia coli with anti‐O157: H7 activity isolated from bovine colon. J. Appl. Microbiol. 100, 384–389 (2006).
Sassone-Corsi, M. et al. Microcins mediate competition among Enterobacteriaceae in the inflamed gut. Nature 540, 280–283 (2016).
Rea, M. C. et al. Thuricin CD, a posttranslationally modified bacteriocin with a narrow spectrum of activity against Clostridium difficile. Proc. Natl Acad. Sci. USA 107, 9352–9357 (2010).
Rea, M. C. et al. Antimicrobial activity of lacticin 3147 against clinical Clostridium difficile strains. J. Med. Microbiol. 56, 940–946 (2007).
MacIntyre, D. L., Miyata, S. T., Kitaoka, M. & Pukatzki, S. The Vibrio cholerae type VI secretion system displays antimicrobial properties. Proc. Natl Acad. Sci. USA 107, 19520–19524 (2010).
Sana, T. G. et al. Salmonella typhimurium utilizes a T6SS-mediated antibacterial weapon to establish in the host gut. Proc. Natl Acad. Sci. USA 113, E5044–E5051 (2016).
Chatzidaki-Livanis, M., Geva-Zatorsky, N. & Comstock, L. E. Bacteroides fragilis type VI secretion systems use novel effector and immunity proteins to antagonize human gut Bacteroidales species. Proc. Natl Acad. Sci. USA 113, 3627–3632 (2016).
Wheeler, K. M. et al. Mucin glycans attenuate the virulence of Pseudomonas aeruginosa in infection. Nat. Microbiol. 4, 2146–2154 (2019).
Wang, B. X. et al. Mucin glycans signal through the sensor kinase RetS to inhibit virulence-associated traits in Pseudomonas aeruginosa. Curr. Biol. 31, 90–102 (2020).
Kunzelmann, K. & Mall, M. Electrolyte transport in the mammalian colon: mechanisms and implications for disease. Physiol. Rev. 82, 245–289 (2002).
Binder, H. J. Role of colonic short-chain fatty acid transport in diarrhea. Annu. Rev. Physiol. 72, 297–313 (2010).
Kumar, A. et al. Probiotic Bifidobacterium species stimulate human SLC26A3 gene function and expression in intestinal epithelial cells. Am. J. Physiol. Cell Physiol. 307, C1084–C1092 (2014).
Singh, A. K. et al. Tu1846 SLC26A3 (DRA)-deficient mice display dramatically low surface pH, normal mucus secretion but loss of firmly adherent mucus layer, altered colonic microbiome and low grade intestinal inflammation. Gastroenterology 148, 918 (2015).
Kini, A. et al. Slc26a3 deletion alters pH‐microclimate, mucin biosynthesis, microbiome composition and increases the TNFα expression in murine colon. Acta Physiol., 230, e13498 (2020).
Byndloss, M. X. et al. Microbiota-activated PPAR-γ signaling inhibits dysbiotic Enterobacteriaceae expansion. Science 357, 570–575 (2017).
Leal, J., Smyth, H. D. & Ghosh, D. Physicochemical properties of mucus and their impact on transmucosal drug delivery. Int. J. Pharm. 532, 555–572 (2017).
Swidsinski, A., Loening-Baucke, V., Lochs, H. & Hale, L. P. Spatial organization of bacterial flora in normal and inflamed intestine: a fluorescence in situ hybridization study in mice. World J. Gastroenterol. 11, 1131 (2005).
Swidsinski, A. et al. Viscosity gradient within the mucus layer determines the mucosal barrier function and the spatial organization of the intestinal microbiota. Inflamm. Bowel. Dis. 13, 963–970 (2007).
Ijssennagger, N., van der Meer, R. & van Mil, S. W. Sulfide as a mucus barrier-breaker in inflammatory bowel disease? Trends Mol. Med. 22, 190–199 (2016).
Rowan, F. et al. Desulfovibrio bacterial species are increased in ulcerative colitis. Dis. Colon Rectum 53, 1530–1536 (2010).
Zinkevich, V. & Beech, I. B. Screening of sulfate-reducing bacteria in colonoscopy samples from healthy and colitic human gut mucosa. FEMS Microbiol Ecol. 34, 147–155 (2000).
Swidsinski, A. et al. Comparative study of the intestinal mucus barrier in normal and inflamed colon. Gut 56, 343–350 (2007).
Wiles, T. J. et al. Swimming motility of a gut bacterial symbiont promotes resistance to intestinal expulsion and enhances inflammation. PLoS Biol. 18, e3000661 (2020).
Podolsky, D. K., Gerken, G., Eyking, A. & Cario, E. Colitis-associated variant of TLR2 causes impaired mucosal repair because of TFF3 deficiency. Gastroenterology 137, 209–220 (2009).
Yu, H. et al. The rat IgGFcγBP and Muc2 C-terminal domains and TFF3 in two intestinal mucus layers bind together by covalent interaction. PLoS ONE 6, e20334 (2011).
Thim, L., Madsen, F. & Poulsen, S. S. Effect of trefoil factors on the viscoelastic properties of mucus gels. Eur. J. Clin. Invest. 32, 519–527 (2002).
Mashimo, H., Wu, D.-C., Podolsky, D. K. & Fishman, M. C. Impaired defense of intestinal mucosa in mice lacking intestinal trefoil factor. Science 274, 262–265 (1996).
Belle, N. M. et al. TFF3 interacts with LINGO2 to regulate EGFR activation for protection against colitis and gastrointestinal helminths. Nat. Commun. 10, 1–13 (2019).
Yang, N., Garcia, M. A. S. & Quinton, P. M. Normal mucus formation requires cAMP‐dependent HCO3− secretion and Ca2+‐mediated mucin exocytosis. J. Physiol. 591, 4581–4593 (2013).
Yoon, S. S. et al. Anaerobic killing of mucoid Pseudomonas aeruginosa by acidified nitrite derivatives under cystic fibrosis airway conditions. J. Clin. Invest. 116, 436–446 (2006).
Van der Sluis, M. et al. Muc2-deficient mice spontaneously develop colitis, indicating that MUC2 is critical for colonic protection. Gastroenterology 131, 117–129 (2006).
Sommer, F. & Bäckhed, F. The gut microbiota—masters of host development and physiology. Nat. Rev. Microbiol. 11, 227–238 (2013).
Bergstrom, K. et al. Core 1–and 3–derived O-glycans collectively maintain the colonic mucus barrier and protect against spontaneous colitis in mice. Mucosal Immunol. 10, 91–103 (2017).
Okayasu, I. et al. A novel method in the induction of reliable experimental acute and chronic ulcerative colitis in mice. Gastroenterology 98, 694–702 (1990).
Morampudi, V. et al. The goblet cell-derived mediator RELM-β drives spontaneous colitis in Muc2-deficient mice by promoting commensal microbial dysbiosis. Mucosal Immunol. 9, 1218–1233 (2016).
Bergström, J. H. et al. Gram-positive bacteria are held at a distance in the colon mucus by the lectin-like protein ZG16. Proc. Natl Acad. Sci. USA 113, 13833–13838 (2016).
Forman, R. A. et al. The goblet cell is the cellular source of the anti-microbial angiogenin 4 in the large intestine post Trichuris muris infection. PLoS ONE 7, e42248 (2012).
Hooper, L. V., Stappenbeck, T. S., Hong, C. V. & Gordon, J. I. Angiogenins: a new class of microbicidal proteins involved in innate immunity. Nat. Immunol. 4, 269–273 (2003).
Okumura, R. et al. Lypd8 inhibits attachment of pathogenic bacteria to colonic epithelia. Mucosal Immunol. 13, 75–85 (2020).
Kaetzel, C. S. Cooperativity among secretory IgA, the polymeric immunoglobulin receptor, and the gut microbiota promotes host–microbial mutualism. Immunol. Lett. 162, 10–21 (2014).
Dupont, A., Heinbockel, L., Brandenburg, K. & Hornef, M. W. Antimicrobial peptides and the enteric mucus layer act in concert to protect the intestinal mucosa. Gut microbes 5, 761–765 (2014).
Barr, J. J. et al. Bacteriophage adhering to mucus provide a non–host-derived immunity. Proc. Natl Acad. Sci. USA 110, 10771–10776 (2013).
Barnich, N. et al. CEACAM6 acts as a receptor for adherent-invasive E. coli, supporting ileal mucosa colonization in Crohn disease. J. Clin. Invest. 117, 1566–1574 (2007).
Galtier, M. et al. Bacteriophages targeting adherent invasive Escherichia coli strains as a promising new treatment for Crohn’s disease. J. Crohns Colitis 11, 840–847 (2017).
Chervy, M., Barnich, N. & Denizot, J. Adherent-invasive E. coli: update on the lifestyle of a troublemaker in Crohn’s disease. Int. J. Mol. Sci. 21, 3734 (2020).
Thoma, C. Bacteriophage virome in IBD. Nat. Rev. Gastroenterol. Hepatol. 15, 520–520 (2018).
Duerkop, B. A. et al. Murine colitis reveals a disease-associated bacteriophage community. Nat. Microbiol. 3, 1023–1031 (2018).
Silveira, C. B. & Rohwer, F. L. Piggyback-the-Winner in host-associated microbial communities. NPJ Biofilms Microbiomes 2, 1–5 (2016).
Gogokhia, L. et al. Expansion of bacteriophages is linked to aggravated intestinal inflammation and colitis. Cell Host Microbe 25, 285–299. e288 (2019).
Nishiyama, K., et al. Bifidobacterium bifidum extracellular sialidase enhances adhesion to the mucosal surface and supports carbohydrate assimilation. mBio. 8, e00928-17 (2017).
Garrido, D., Kim, J. H., German, J. B., Raybould, H. E. & Mills, D. A. Oligosaccharide binding proteins from Bifidobacterium longum subsp. infantis reveal a preference for host glycans. PLoS ONE 6, e17315 (2011).
Troge, A. et al. More than a marine propeller–the flagellum of the probiotic Escherichia coli strain Nissle 1917 is the major adhesin mediating binding to human mucus. Int. J. Med. Microbiol. 302, 304–314 (2012).
Owen, C. D. et al. Unravelling the specificity and mechanism of sialic acid recognition by the gut symbiont Ruminococcus gnavus. Nat. Commun. 8, 1–15 (2017).
Ho, J. G., Greco, A., Rupnik, M. & Ng, K. K.-S. Crystal structure of receptor-binding C-terminal repeats from Clostridium difficile toxin A. Proc. Natl Acad. Sci. USA 102, 18373–18378 (2005).
Ruiz-Palacios, G. M., Cervantes, L. E., Ramos, P., Chavez-Munguia, B. & Newburg, D. S. Campylobacter jejuni binds intestinal H (O) antigen (Fucα1, 2Galβ1, 4GlcNAc), and fucosyloligosaccharides of human milk inhibit its binding and infection. J. Biol. Chem. 278, 14112–14120 (2003).
Conover, M. S. et al. Inflammation-induced adhesin-receptor interaction provides a fitness advantage to uropathogenic E. coli during chronic infection. Cell Host Microbe 20, 482–492 (2016).
Erdem, A. L., Avelino, F., Xicohtencatl-Cortes, J. & Girón, J. A. Host protein binding and adhesive properties of H6 and H7 flagella of attaching and effacing Escherichia coli. J. Bacteriol. 189, 7426–7435 (2007).
Dodson, K. W. et al. Structural basis of the interaction of the pyelonephritic E. coli adhesin to its human kidney receptor. Cell 105, 733–743 (2001).
Buts, L. et al. The fimbrial adhesin F17-G of enterotoxigenic Escherichia coli has an immunoglobulin-like lectin domain that binds N-acetylglucosamine. Mol. Microbiol. 49, 705–715 (2003).
Moonens, K. et al. Structural insight in histo-blood group binding by the F18 fimbrial adhesin FedF. Mol. Microbiol. 86, 82–95 (2012).
Hung, C. S. et al. Structural basis of tropism of Escherichia coli to the bladder during urinary tract infection. Mol. Microbiol. 44, 903–915 (2002).
Chessa, D., Winter, M. G., Jakomin, M. & Bäumler, A. J. Salmonella enterica serotype Typhimurium Std fimbriae bind terminal α (1, 2) fucose residues in the cecal mucosa. Mol. Microbiol. 71, 864–875 (2009).
Mariscotti, J. F., Quereda, J. J., García-del Portillo, F. & Pucciarelli, M. G. The Listeria monocytogenes LPXTG surface protein Lmo1413 is an invasin with capacity to bind mucin. Int. J. Med. Microbiol. 304, 393–404 (2014).
Diderrich, R. et al. Structural hot spots determine functional diversity of the Candida glabrata epithelial adhesin family. J. Biol. Chem. 290, 19597–19613 (2015).
Henrissat, B. A classification of glycosyl hydrolases based on amino acid sequence similarities. Biochem. J. 280, 309–316 (1991).
Ndeh, D. & Gilbert, H. J. Biochemistry of complex glycan depolymerisation by the human gut microbiota. FEMS Microbiol Rev. 42, 146–164 (2018).
Acknowledgements
We would like to thank Editage (www.editage.cn) for English language editing. This study was supported by the National Natural Science Foundation of China (NSFC; Grant No. 81872620, 81673163), the Ningbo Natural Science Foundation (2018A610370), the Natural Science Foundation of Zhejiang Province (LY21H260001), the Zhejiang Key Laboratory of Pathophysiology (201802), the Zhejiang Public Welfare Technology Applied Research Project (LGF18H060006) and the Zhejiang Provincial Medical and Health Science and Technology plan (2018KY819). This study was sponsored by the K. C. Wong Magna Fund at Ningbo University, Shaoxing Municipal Science and Technology Project of Medical and Health (2020A13064) and the Scientific Research Projects of Shaoxing University (2019SK003).
Author information
Authors and Affiliations
Contributions
J.F., H.W., Y.Z. and H.Z. contributed equally to this work.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Fang, J., Wang, H., Zhou, Y. et al. Slimy partners: the mucus barrier and gut microbiome in ulcerative colitis. Exp Mol Med 53, 772–787 (2021). https://doi.org/10.1038/s12276-021-00617-8
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s12276-021-00617-8
This article is cited by
-
Muc2 mucin o-glycosylation interacts with enteropathogenic Escherichia coli to influence the development of ulcerative colitis based on the NF-kB signaling pathway
Journal of Translational Medicine (2023)
-
Mucin glycans drive oral microbial community composition and function
npj Biofilms and Microbiomes (2023)
-
Impact of structurally diverse polysaccharides on colonic mucin O-glycosylation and gut microbiota
npj Biofilms and Microbiomes (2023)
-
ERdj5 protects goblet cells from endoplasmic reticulum stress-mediated apoptosis under inflammatory conditions
Experimental & Molecular Medicine (2023)
-
Non-alcoholic fatty liver disease and gut microbial dysbiosis- underlying mechanisms and gut microbiota mediated treatment strategies
Reviews in Endocrine and Metabolic Disorders (2023)