Abstract
Background
Breast cancer (BC) is among the most common cause of cancer 10.4% and one of the leading causes of death among 20–50 years old women in the world. Tamoxifen drug is the first line therapy for BC however tamoxifen resistance (TR) has shown in 30–50% of cases that may face BC recurrence. Hence, TR early detection reduces BC recurrence and fatalities. The epigenetic alteration that happens by hypermethylation of tumor suppressor genes and hypomethylation of oncogenes has been suggested to be useful in early cancer or drug resistance diagnosis.
Methods
This is the first study to investigate DOK7 CpG hypermethylation in blood leukocytes of 31 TR (ER+) BC compared to 29 tamoxifen sensitive BC to evaluate DOK7 as a potential TR biomarker. DNA was extracted from blood samples of all participants and MSRE-PCR and real-time PCR were used for quantification of CpG methylation alterations.
Results
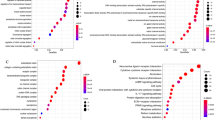
The means of DOK7 CpG hypermethylation were obtained as 85.03%, 29.1% and 57.34% in TR, TS and normal control respectively. Significant hypermethylation were found among TR vs. TS (p < 0.001), TS vs. normal (p < 0.001) and TR vs. normal controls (p < 0.03). Online databases expression and survival analysis of DOK7 showed increasing expression in TS groups vs. TR groups which have consistency with our methylation alteration results. The sensitivity and specificity of the TR epigenetic test were determined using ROC analysis showed 89.66% and 96.77% respectively and showed that 37.5% above hypermethylation is at risk for TR and breast cancer recurrence.
Conclusion
There is a significant difference in the methylation ratio of DOK7 between tamoxifen resistant and tamoxifen sensitive groups that may be useful in the early diagnosis of tamoxifen resistance in BC cases and cancer recurrence prevention.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Siegel RL, Miller KD, Jemal A. Cancer statistics, 2020. CA Cancer J Clin. 2020;70:7–30.
Lumachi F, Brunello A, Maruzzo M, Basso U, Basso SM. Treatment of estrogen receptor-positive breast cancer. Curr Med Chem. 2013;20:596–604.
Tsang JYS, Tse GM. Molecular classification of breast cancer. Adv Anat Pathol. 2020;27:27–35.
Hon JD, Singh B, Sahin A, Du G, Wang J, Wang VY, et al. Breast cancer molecular subtypes: from TNBC to QNBC. Am J Cancer Res. 2016;6:1864–72.
Eroles P, Bosch A, Pérez-Fidalgo JA, Lluch A. Molecular biology in breast cancer: intrinsic subtypes and signaling pathways. Cancer Treat Rev. 2012;38:698–707.
Matthews J, Gustafsson JA. Estrogen signaling: a subtle balance between ER alpha and ER beta. Mol Inter. 2003;3:281–92.
Hayashi S, Yamaguchi Y. Estrogen signaling pathway and hormonal therapy. Breast Cancer. 2008;15:256–61.
Saha Roy S, Vadlamudi RK. Role of estrogen receptor signaling in breast cancer metastasis. Int J Breast Cancer. 2012;2012:654698.
Levin ER, Pietras RJ. Estrogen receptors outside the nucleus in breast cancer. Breast Cancer Res Treat. 2008;108:351–61.
Yaşar P, Ayaz G, User SD, Güpür G, Muyan M. Molecular mechanism of estrogen-estrogen receptor signaling. Reprod Med Biol. 2017;16:4–20.
Komm BS, Mirkin S. An overview of current and emerging SERMs. J Steroid Biochem Mol Biol. 2014;143:207–22.
Hultsch S, Kankainen M, Paavolainen L, Kovanen R-M, Ikonen E, Kangaspeska S, et al. Association of tamoxifen resistance and lipid reprogramming in breast cancer. BMC Cancer. 2018;18:850.
Hu R, Hilakivi-Clarke L, Clarke R. Molecular mechanisms of tamoxifen-associated endometrial cancer (Review). Oncol Lett. 2015;9:1495–501.
Cook KL, Shajahan AN, Clarke R. Autophagy and endocrine resistance in breast cancer. Expert Rev Anticancer Ther. 2011;11:1283–94.
Mills JN, Rutkovsky AC, Giordano A. Mechanisms of resistance in estrogen receptor positive breast cancer: overcoming resistance to tamoxifen/aromatase inhibitors. Curr Opin Pharm. 2018;41:59–65.
Fan W, Chang J, Fu P. Endocrine therapy resistance in breast cancer: current status, possible mechanisms and overcoming strategies. Future Med Chem. 2015;7:1511–9.
Gutierrez MC, Detre S, Johnston S, Mohsin SK, Shou J, Allred DC, et al. Molecular changes in tamoxifen-resistant breast cancer: relathe tionship between estrogen receptor, HER-2, and p38 mitogen-activated protein kinase. J Clin Oncol. 2005;23:2469–76.
Li L, Choi JY, Lee KM, Sung H, Park SK, Oze I, et al. DNA methylation in peripheral blood: a potential biomarker for cancer molecular epidemiology. J Epidemiol. 2012;22:384–94.
Rauch TA, Wang Z, Wu X, Kernstine KH, Riggs AD, Pfeifer GP. DNA methylation biomarkers for lung cancer. Tumor Biol. 2012;33:287–96.
Yi JM. DNA methylation change profiling of colorectal disease: screening towards clinical use. Life. 2021;11:5.
Jones N, Dumont DJ. Recruitment of Dok-R to the EGF receptor through its PTB domain is required for attenuation of Erk MAP kinase activation. Curr Biol. 1999;9:1057–60.
Bedirian A, Baldwin C, Abe J, Takano T, Lemay S. Pleckstrin homology and phosphotyrosine-binding domain-dependent membrane association and tyrosine phosphorylation of Dok-4, an inhibitory adapter molecule expressed in epithelial cells. J Biol Chem. 2004;279:19335–49.
Okada K, Inoue A, Okada M, Murata Y, Kakuta S, Jigami T, et al. The muscle protein Dok-7 is essential for neuromuscular synaptogenesis. Science. 2006;312:1802–5.
Bergamin E, Hallock PT, Burden SJ, Hubbard SR. The cytoplasmic adaptor protein Dok7 activates the receptor tyrosine kinase MuSK via dimerization. Mol Cell. 2010;39:100–9.
Alsallum MS, Alshareef A, Abuzinadah AR, Bamaga AK, Dallol A. A novel DOK7 mutation causing congenital myasthenic syndrome with limb-girdle weakness: case series of three family members. Heliyon. 2021;7:e06869.
Zhang L, Li R, Hu K, Dai Y, Pang Y, Jiao Y, et al. Prognostic role of DOK family adapters in acute myeloid leukemia. Cancer Gene Ther. 2019;26:305–12.
Chen G, Yu H, Satherley L, Zabkiewicz C, Resaul J, Zhao H, et al. The downstream of tyrosine kinase 7 is reduced in lung cancer and is associated with poor survival of patients with lung cancer. Oncol Rep. 2017;37:2695–701.
Hua C-D, Bian E-B, Chen E-F, Yang Z-H, Tang F, Wang H-L, et al. Repression of Dok7 expression mediated by DNMT1 promotes glioma cells proliferation. Biomed Pharmacother. 2018;106:678–85.
Yang S-M, Li S-Y, Yu H-B, Li J-R, Sun L-L. Repression of DOK7 mediated by DNMT3A promotes the proliferation and invasion of KYSE410 and TE-12 ESCC cells. Biomed Pharmacother. 2017;90:93–9.
Yue C, Bai Y, Piao Y, Liu H. DOK7 inhibits cell proliferation, migration, and invasion of breast cancer via the PI3K/PTEN/AKT pathway. J Oncol. 2021;2021:4035257.
Heyn H, Carmona FJ, Gomez A, Ferreira HJ, Bell JT, Sayols S, et al. DNA methylation profiling in breast cancer discordant identical twins identifies DOK7 as novel epigenetic biomarker. Carcinogenesis. 2013;34:102–8.
Oakes CC, La Salle S, Robaire B, Trasler JM. Evaluation of a quantitative DNA methylation analysis technique using methylation-sensitive/dependent restriction enzymes and real-time PCR. Epigenetics. 2006;1:146–52.
Győrffy B. Survival analysis across the entire transcriptome identifies biomarkers with the highest prognostic power in breast cancer. Comput Struct Biotechnol J. 2021;19:4101–9.
Fekete JT, Győrffy B. ROCplot.org: validating predictive biomarkers of chemotherapy/hormonal therapy/anti-HER2 therapy using transcriptomic data of 3,104 breast cancer patients. Int J Cancer. 2019;145:3140–51.
Riggins RB, Schrecengost RS, Guerrero MS, Bouton AH. Pathways to tamoxifen resistance. Cancer Lett. 2007;256:1–24.
Flanagan JM, Munoz-Alegre M, Henderson S, Tang T, Sun P, Johnson N, et al. Gene-body hypermethylation of ATM in peripheral blood DNA of bilateral breast cancer patients. Hum Mol Genet. 2009;18:1332–42.
Iwamoto T, Yamamoto N, Taguchi T, Tamaki Y, Noguchi S. BRCA1 promoter methylation in peripheral blood cells is associated with increased risk of breast cancer with BRCA1 promoter methylation. Breast Cancer Res Treat. 2011;129:69–77.
Yin Y, Lei S, Li L, Yang X, Yin Q, Xu T, et al. RPTOR methylation in the peripheral blood and breast cancer in the Chinese population. Genes Genomics. 2022;44:435–43.
Yang R, Pfütze K, Zucknick M, Sutter C, Wappenschmidt B, Marme F, et al. DNA methylation array analyses identified breast cancer-associated HYAL2 methylation in peripheral blood. Int J Cancer. 2015;136:1845–55.
Acknowledgements
This work was carried out at the National Institute of Genetic Engineering and Biotechnology (NIGEB), and the authors would like to express their appreciation to NIGEB. We are grateful to Arash Moradi, a student at the National Institute of Genetic Engineering and Biotechnology (NIGEB) for his participation in the study.
Funding
Our study was supported by grants number 766 from National Institute of Genetic Engineering and Biotechnology (NIGEB).
Author information
Authors and Affiliations
Contributions
SAA designed experiments, support and helped write the manuscript; EG carried out some experiments and write the manuscript; NR carried out some experiments; NN and MT contributed to sample collection and patients data extraction.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Gowdini, E., Aleyasin, S.A., Ramezani, N. et al. DOK7 CpG hypermethylation in blood leukocytes as an epigenetic biomarker for acquired tamoxifen resistant in breast cancer. J Hum Genet 68, 33–38 (2023). https://doi.org/10.1038/s10038-022-01092-3
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s10038-022-01092-3
This article is cited by
-
Tamoxifen induces eryptosis through calcium accumulation and oxidative stress
Medical Oncology (2023)