Abstract
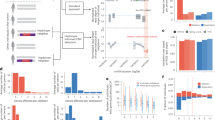
Bicuspid aortic valve (BAV) is the most common congenital heart defect with a high index of heritability. Patients with BAV have different clinical courses and disease progression. Herein, we report three siblings with BAV and clinical differences. Their clinical presentations include moderate to severe aortic regurgitation, aortic stenosis, and ascending aortic aneurysm. Genetic investigation was carried out using Whole-Exome Sequencing for the three patients. We identified two non-synonymous variants in ROBO1 and GATA5 genes. The ROBO1: p.(Ser327Pro) variant is shared by the three BAV-affected siblings. The GATA5: p.(Gln3Arg) variant is shared only by the two brothers who presented BAV and ascending aortic aneurysm. Their sister, affected by BAV without aneurysm, does not harbor the GATA5: p.(Gln3Arg) variant. Both variants were absent in the patients’ fourth brother who is clinically healthy with tricuspid aortic valve. To our knowledge, this is the first association of ROBO1 and GATA5 variants in familial BAV with a potential genotype-phenotype correlation. Our findings are suggestive of the implication of ROBO1 gene in BAV and the GATA5: p.(Gln3Arg) variant in ascending aortic aneurysm. Our family-based study further confirms the intrafamilial incomplete penetrance of BAV and the complex pattern of inheritance of the disease.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
References
Michelena HI, Prakash SK, Della Corte A, Bissell MM, Anavekar N, Mathieu P, et al. Bicuspid aortic valve: identifying knowledge gaps and rising to the challenge from the International Bicuspid Aortic Valve Consortium (BAVCon). Circulation. 2014;129:2691–704.
Martín M, Lorca R, Rozado J, Alvarez-Cabo R, Calvo J, Pascual I, et al. Bicuspid aortic valve syndrome: a multidisciplinary approach for a complex entity. J Thorac Dis. 2017;9:S454–64.
Tessler I, Albuisson J, Goudot G, Carmi S, Shpitzen S, Messas E, et al. Bicuspid aortic valve: genetic and clinical insights. AORTA J. 2021;9:139–46.
Martin LJ, Pilipenko V, Kaufman KM, Cripe L, Kottyan LC, Keddache M, et al. Whole exome sequencing for familial bicuspid aortic valve identifies putative variants. Circ Cardiovasc Genet. 2014;7:677–83.
Tessler I, Goudot G, Albuisson J, Reshef N, Zwas DR, Carmi S, et al. Is bicuspid aortic valve morphology genetically determined? A family-based study. Am J Cardiol. 2022;163:85–90.
Galian-Gay L, Carro Hevia A, Teixido-Turà G, Rodríguez Palomares J, Gutiérrez-Moreno L, Maldonado G, et al. Familial clustering of bicuspid aortic valve and its relationship with aortic dilation in first-degree relatives. Heart. 2018;heartjnl-2018-313802.
Hui DS, Bonow RO, Stolker JM, Braddock SR, Lee R. Discordant aortic valve morphology in monozygotic twins: a clinical case series. JAMA Cardiol. 2016;1:1043–7.
Saravanan P, Kadir I. Apolipoprotein E alleles and bicuspid aortic valve stenosis in monozygotic twins. Interact Cardiovasc Thorac Surg. 2009;8:687–8.
Foffa I, Ait Alì L, Panesi P, Mariani M, Festa P, Botto N, et al. Sequencing of NOTCH1, GATA5, TGFBR1 and TGFBR2 genes in familial cases of bicuspid aortic valve. BMC Med Genet. 2013;14:44.
Musfee FI, Guo D, Pinard AC, Hostetler EM, Blue EE, Nickerson DA, et al. Rare deleterious variants of NOTCH1, GATA4, SMAD6, and ROBO4 are enriched in BAV with early onset complications but not in BAV with heritable thoracic aortic disease. Mol Genet Genom Med. 2020;8:e1406.
Bonachea EM, Chang S-W, Zender G, LaHaye S, Fitzgerald-Butt S, McBride KL, et al. GATA5 sequence variants identified in individuals with bicuspid aortic valve. Pediatr Res. 2014;76:211–6.
Laforest B, Nemer M. GATA5 interacts with GATA4 and GATA6 in outflow tract development. Dev Biol. 2011;358:368–78.
Zhao J, Mommersteeg MTM Slit–Robo signalling in heart development. Cardiovasc Res. 2018;114:794–804.
Medioni C, Bertrand N, Mesbah K, Hudry B, Dupays L, Wolstein O, et al. Expression of slit and robo genes in the developing mouse heart. Dev Dyn Publ Am Assoc Anat. 2010;239:3303–11.
Gould RA, Aziz H, Woods CE, Seman-Senderos MA, Sparks E, Preuss C, et al. ROBO4 variants predispose individuals to bicuspid aortic valve and thoracic aortic aneurysm. Nat Genet. 2019;51:42–50.
Kruszka P, Tanpaiboon P, Neas K, Crosby K, Berger SI, Martinez AF, et al. Loss of function in ROBO1 is associated with tetralogy of Fallot and septal defects. J Med Genet. 2017;54:825–9.
Desvignes J-P, Bartoli M, Delague V, Krahn M, Miltgen M, Béroud C, et al. VarAFT: a variant annotation and filtration system for human next generation sequencing data. Nucleic Acids Res. 2018;46:W545–53.
Desmet F-O, Hamroun D, Lalande M, Collod-Béroud G, Claustres M, Béroud C. Human Splicing Finder: an online bioinformatics tool to predict splicing signals. Nucleic Acids Res. 2009;37:e67.
Ackerman C, Locke AE, Feingold E, Reshey B, Espana K, Thusberg J, et al. An excess of deleterious variants in VEGF-A pathway genes in Down-syndrome-associated atrioventricular septal defects. Am J Hum Genet. 2012;91:646–59.
Padang R, Bagnall RD, Richmond DR, Bannon PG, Semsarian C. Rare non-synonymous variations in the transcriptional activation domains of GATA5 in bicuspid aortic valve disease. J Mol Cell Cardiol. 2012;53:277–81.
Tong M, Jun T, Nie Y, Hao J, Fan D. The role of the slit/robo signaling pathway. J Cancer. 2019;10:2694–705.
Le Bras A. ROBO4 variants linked to congenital heart defects. Nat Rev Cardiol. 2019;16:70.
Mommersteeg MTM, Yeh ML, Parnavelas JG, Andrews WD. Disrupted Slit-Robo signalling results in membranous ventricular septum defects and bicuspid aortic valves. Cardiovasc Res. 2015;106:55–66.
Phillips HM, Mahendran P, Singh E, Anderson RH, Chaudhry B, Henderson DJ. Neural crest cells are required for correct positioning of the developing outflow cushions and pattern the arterial valve leaflets. Cardiovasc Res. 2013;99:452–60.
Odelin G, Faure E, Coulpier F, Di Bonito M, Bajolle F, Studer M, et al. Krox20 defines a subpopulation of cardiac neural crest cells contributing to arterial valves and bicuspid aortic valve. Dev Camb Engl. 2018;145:dev151944.
Wei D, Bao H, Zhou N, Zheng G-F, Liu X-Y, Yang Y-Q. GATA5 loss-of-function mutation responsible for the congenital ventriculoseptal defect. Pediatr Cardiol. 2013;34:504–11.
Jiang J-Q, Li R-G, Wang J, Liu X-Y, Xu Y-J, Fang W-Y, et al. Prevalence and spectrum of GATA5 mutations associated with congenital heart disease. Int J Cardiol. 2013;165:570–3.
Morrisey EE, Ip HS, Tang Z, Parmacek MS. GATA-4 activates transcription via two novel domains that are conserved within the GATA-4/5/6 subfamily. J Biol Chem. 1997;272:8515–24.
Acknowledgements
We would like to thank the family for their collaboration. This work was supported by the “Association Française contre les Myopathies-AFM Telethon” [MoThARD-Project], and the “Institut National de la Santé et de la Recherche Médicale” to SZ. HJ received Postdoctoral fellowships from the “Association Française contre les Myopathies-AFM Telethon”.
Author information
Authors and Affiliations
Contributions
Study concept and design: JFA, SZ; Clinical Investigation of the patients and family members; HG, AT, FC, JFA; Analysis and interpretation of data: GCB, HJ. Molecular investigation: HJ.; Writing—Original draft preparation: HJ; HG; Critical—review & editing: AT, JFA, SZ. Supervision: SZ; Validation: SZ, JFA.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Ethical approval
This study was performed according to the principles of the Declaration of Helsinki and to the ethical standards of the first author’s institutional review board. The patients provided their written informed consent to participate in this study.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
About this article
Cite this article
Jaouadi, H., Gérard, H., Théron, A. et al. Identification of non-synonymous variations in ROBO1 and GATA5 genes in a family with bicuspid aortic valve disease. J Hum Genet 67, 515–518 (2022). https://doi.org/10.1038/s10038-022-01036-x
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s10038-022-01036-x
This article is cited by
-
Insights into the Inherited Basis of Valvular Heart Disease
Current Cardiology Reports (2024)
-
Expanding the phenome and variome of the ROBO-SLIT pathway in congenital heart defects: toward improving the genetic testing yield of CHD
Journal of Translational Medicine (2023)