Abstract
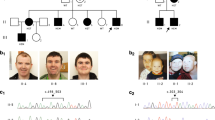
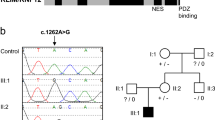
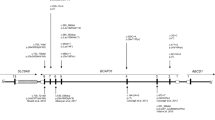
HCFC1, a global transcriptional regulator, has been shown to associate with MMACHC expression. Pathogenic variants in HCFC1 cause X-linked combined methylmalonic acidemia and hyperhomocysteinemia, CblX type (MIM# 309541). Recent studies showed that certain variants in HCFC1 are associated with X-linked intellectual disability with mild or absent metabolic abnormalities. Here, we report five subjects (three males, two females) from the same family with a novel predicted loss of function HCFC1 variant. All five patients exhibit developmental delay or intellectual disability/learning difficulty and some dysmorphic features; findings were milder in the female as compared to male subjects. Biochemical studies in all patients did not show methylmalonic acidemia or hyperhomocysteinemia but revealed elevated vitamin B12 levels. Trio exome sequencing of the proband and his parents revealed a maternally inherited novel variant in HCFC1 designated as c.1781_1803 + 3del26insCA (NM_005334). Targeted testing confirmed the presence of the same variant in two half-siblings and maternal great uncle. In silico analysis showed that the variant is expected to reduce the quality of the splice donor site in intron 10 and causes abnormal splicing. Sequencing of proband’s cDNA revealed exon 10 skipping. Further molecular studies in the two manifesting females revealed moderate and high skewing of X inactivation. Our results support previous observation that HCFC1 variants located outside the Kelch domain exhibit dissociation of the clinical and biochemical phenotype and cause milder or no metabolic changes. We also show that this novel variant can be associated with a phenotype in females, although with milder severity, but further studies are needed to understand the role of skewed X inactivation among females in this rare disorder. Our work expands the genotypes and phenotypes associated with HCFC1-related disorder.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
References
Zablotsky B, Black LI, Blumberg SJ. Estimated prevalence of children with diagnosed developmental disabilities in the United States, 2014–2016. NCHS Data Brief. 2017;291:1–8
Karam SM, Riegel M, Segal SL, et al. Genetic causes of intellectual disability in a birth cohort: a population-based study. Am J Med Genet A. 2015;167:1204–1214
Harripaul R, Noor A, Ayub M, Vincent JB. The use of next-generation sequencing for research and diagnostics for intellectual disability. Cold Spring Harb Perspect Med. 2017;7:a026864 https://doi.org/10.1101/cshperspect.a026864. Published 2017 Mar 1
Carvill GL, Mefford HC. Next-generation sequencing in intellectual disability. J Pediatr Genet. 2015;4:128–135. https://doi.org/10.1055/s-0035-1564439
Yu HC, Sloan JL, Scharer G, et al. An X-linked cobalamin disorder caused by mutations in transcriptional coregulator HCFC1. Am J Hum Genet. 2013;93:506–514. https://doi.org/10.1016/j.ajhg.2013.07.022
Gérard M, Morin G, Bourillon A, et al. Multiple congenital anomalies in two boys with mutation in HCFC1 and cobalamin disorder. Eur J Med Genet. 2015;58:148–153. https://doi.org/10.1016/j.ejmg.2014.12.015
Scalais E, Osterheld E, Weitzel C, et al. X-linked cobalamin disorder (HCFC1) mimicking nonketotic hyperglycinemia with increased both cerebrospinal fluid glycine and methylmalonic acid. Pediatr Neurol. 2017;71:65–69. https://doi.org/10.1016/j.pediatrneurol.2016.12.003
Vercauteren K, Gleyzer N, Scarpulla RC. PGC-1-related coactivator complexes with HCF-1 and NRF-2beta in mediating NRF-2(GABP)-dependent respiratory gene expression. J Biol Chem. 2008;283:12102–12111. https://doi.org/10.1074/jbc.M710150200
Goto H, Motomura S, Wilson AC, et al. A single-point mutation in HCF causes temperature-sensitive cell-cycle arrest and disrupts VP16 function. Genes Dev. 1997;11:726–737. https://doi.org/10.1101/gad.11.6.726
Zargar Z, Tyagi S. Role of host cell factor-1 in cell cycle regulation. Transcription 2012;3:187–192. https://doi.org/10.4161/trns.20711
Wilson AC, Boutros M, Johnson KM, Herr W. HCF-1 amino- and carboxy-terminal subunit association through two separate sets of interaction modules: involvement of fibronectin type 3 repeats. Mol Cell Biol. 2000;20:6721–6730. https://doi.org/10.1128/mcb.20.18.6721-6730.2000
Jolly LA, Nguyen LS, Domingo D, et al. HCFC1 loss-of-function mutations disrupt neuronal and neural progenitor cells of the developing brain. Hum Mol Genet. 2015;24:3335–3347. https://doi.org/10.1093/hmg/ddv083
Quintana AM, Geiger EA, Achilly N, et al. Hcfc1b, a zebrafish ortholog of HCFC1, regulates craniofacial development by modulating mmachc expression. Dev Biol. 2014;396:94–106. https://doi.org/10.1016/j.ydbio.2014.09.02
Huang L, Jolly LA, Willis-Owen S, et al. A noncoding, regulatory mutation implicates HCFC1 in nonsyndromic intellectual disability. Am J Hum Genet. 2012;91:694–702. https://doi.org/10.1016/j.ajhg.2012.08.011
Koufaris C, Alexandrou A, Tanteles GA, Anastasiadou V, Sismani C. A novel HCFC1 variant in male siblings with intellectual disability and microcephaly in the absence of cobalamin disorder. Biomed Rep. 2016;4:215–218.
Allen RC, Zoghbi HY, Moseley AB, Rosenblatt HM, Belmont JW. Methylation of HpaII and HhaI sites near the polymorphic CAG repeat in the human androgen-receptor gene correlates with X chromosome inactivation. Am J Hum Genet. 1992;51:1229–39.
Richards S, Aziz N, Bale S, et al. Standards and guidelines for the interpretation of sequence variants: a joint consensus recommendation of the American College of Medical Genetics and Genomics and the Association for Molecular Pathology. Genet Med. 2015;17:405–24. https://doi.org/10.1038/gim.2015.30. Epub 2015 Mar 5
Wysocka J, Myers MP, Laherty CD, Eisenman RN, Herr W. Human Sin3 deacetylase and trithorax-related Set1/Ash2 histone H3-K4 methyltransferase are tethered together selectively by the cell-proliferation factor HCF-1. Genes Dev. 2003;17:896–911. https://doi.org/10.1101/gad.252103
Karczewski KJ, Francioli LC, Tiao G, et al. The mutational constraint spectrum quantified from variation in 141,456 humans. Nature. 2020;581:434–43. https://doi.org/10.1038/s41586-020-2308-7. Epub 2020 May 27
Migeon BR. X-linked diseases: susceptible females. Genet Med. 2020;22:1156–74. https://doi.org/10.1038/s41436-020-0779-4. Epub 2020 Apr 14
Plenge RM, Stevenson RA, Lubs HA, Schwartz CE, Willard HF. Skewed X-chromosome inactivation is a common feature of X-linked mental retardation disorders. Am J Hum Genet. 2002;71:168–73. https://doi.org/10.1086/341123. Epub 2002 May 30
Daou S, Mashtalir N, Hammond-Martel I, et al. Crosstalk between O-GlcNAcylation and proteolytic cleavage regulates the host cell factor-1 maturation pathway. Proc Natl Acad Sci USA. 2011;108:2747–2752. https://doi.org/10.1073/pnas.1013822108
Lazarus MB, Jiang J, Kapuria V, et al. HCF-1 is cleaved in the active site of O-GlcNAc transferase. Science. 2013;342:1235–1239. https://doi.org/10.1126/science.1243990
Capotosti F, Guernier S, Lammers F, et al. O-GlcNAc transferase catalyzes site-specific proteolysis of HCF-1. Cell. 2011;144:376–388. https://doi.org/10.1016/j.cell.2010.12.030
Bhuiyan T, Waridel P, Kapuria V, Zoete V, Herr W. Distinct OGT-binding sites promote HCF-1 cleavage. PLoS One. 2015;10:e0136636 https://doi.org/10.1371/journal.pone.0136636. Published 2015 Aug 25
Mangone M, Myers MP, Herr W. Role of the HCF-1 basic region in sustaining cell proliferation. PLoS One. 2010;5:e9020 https://doi.org/10.1371/journal.pone.0009020. Published 2010 Feb 2
Vaidyanathan K, Niranjan T, Selvan N, et al. Identification and characterization of a missense mutation in the O-linked β-N-acetylglucosamine (O-GlcNAc) transferase gene that segregates with X-linked intellectual disability. J Biol Chem. 2017;292:8948–8963. https://doi.org/10.1074/jbc.M116.771030
Witteveen JS, Willemsen MH, Dombroski TC, et al. Haploinsufficiency of MeCP2-interacting transcriptional co-repressor SIN3A causes mild intellectual disability by affecting the development of cortical integrity. Nat Genet. 2016;48:877–887. https://doi.org/10.1038/ng.3619
Michaud J, Praz V, James Faresse N, et al. HCFC1 is a common component of active human CpG-island promoters and coincides with ZNF143, THAP11, YY1, and GABP transcription factor occupancy. Genome Res. 2013;23:907–916. https://doi.org/10.1101/gr.150078.112
Lu R, Yang P, Padmakumar S, Misra V. The herpesvirus transactivator VP16 mimics a human basic domain leucine zipper protein, luman, in its interaction with HCF. J Virol. 1998;72:6291–6297.
Freiman RN, Herr W. Viral mimicry: common mode of association with HCF by VP16 and the cellular protein LZIP. Genes Dev. 1997;11:3122–3127. https://doi.org/10.1101/gad.11.23.3122
Quintana AM, Yu HC, Brebner A, et al. Mutations in THAP11 cause an inborn error of cobalamin metabolism and developmental abnormalities. Hum Mol Genet. 2017;26:2838–2849. https://doi.org/10.1093/hmg/ddx157
Fujita J, Freire P, Coarfa C, et al. Ronin governs early heart development by controlling core gene expression programs. Cell Rep. 2017;21:1562–1573. https://doi.org/10.1016/j.celrep.2017.10.036
Gregor A, Oti M, Kouwenhoven EN, et al. De novo mutations in the genome organizer CTCF cause intellectual disability. Am J Hum Genet. 2013;93:124–131. https://doi.org/10.1016/j.ajhg.2013.05.007
Konrad EDH, Nardini N, Caliebe A, et al. CTCF variants in 39 individuals with a variable neurodevelopmental disorder broaden the mutational and clinical spectrum. Genet Med. 2019;21:2723–2733. https://doi.org/10.1038/s41436-019-0585-z
Acknowledgements
We thank the patients and their families for their cooperation with this case report. We also thank Mrs. Marisa Andrews, CGC, who coordinated patients’ care and genetic testing.
Author information
Authors and Affiliations
Contributions
PW and MS designed and conceptualized the study and performed clinical analysis of the patients. PW drafted the manuscript. DW performed molecular studies and interpreted the molecular data. All authors were involved with revising the manuscript. MS supervised the study and obtained consents.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Ethical approval
The guardian of patients IV-1 and IV-2 signed a consent form of Washington University Institutional Review Board (IRB) approved protocol entitled: “Molecular and Cytogenetic Testing for Genetic Diseases; IRB ID #: 201102465). All subjects or their guardians signed a consent form for publication approved by Washington University IRB (Media Authorization for the Use and Disclosure of Protected Health Information).
A patient consent statement
Consent was obtained from the patient’s family for publication of this report.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Wongkittichote, P., Wegner, D.J. & Shinawi, M.S. Novel exon-skipping variant disrupting the basic domain of HCFC1 causes intellectual disability without metabolic abnormalities in both male and female patients. J Hum Genet 66, 717–724 (2021). https://doi.org/10.1038/s10038-020-00892-9
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s10038-020-00892-9