Abstract
Lipids foster energy production and their altered levels have been coupled with metabolic ailments. Indians feature high prevalence of metabolic diseases, yet uncharacterized for genes regulating lipid homeostasis. We performed first GWAS for quantitative lipids (total cholesterol, LDL, HDL, and triglycerides) exclusively in 5271 Indians. Further to corroborate our genetic findings, we investigated DNA methylation marks in peripheral blood in Indians at the identified loci (N = 233) and retrieved gene regulatory features from public domains. Recurrent GWAS loci—CELSR2, CETP, LPL, ZNF259, and BUD13 cropped up as lead signals in Indians, reflecting their universal applicability. Besides established variants, we found certain unreported variants at sub-genome-wide level—QKI, REEP3, TMCC2, FAM129C, FAM241B, and LOC100506207. These variants though failed to attain GWAS significance in Indians, but largely turned out to be active CpG sites in human subcutaneous adipose tissue and showed robust association to two or more lipid traits. Of which, QKI variants showed significant association to all four lipid traits and their designated region was observed to be a key gene regulatory segment denoting active transcription particularly in human subcutaneous adipose tissue. Both established and novel loci were observed to be significantly associated with altered DNA methylation in Indians for specific CpGs that resided in key regulatory elements. Further, gene-based association analysis pinpointed novel GWAS loci—LINC01340 and IQCJ-SCHIP1 for TC; IFT27, IFT88, and LINC02141 for HDL; and TEX26 for TG. Present study ascertains universality of selected known genes and also identifies certain novel loci for lipids in Indians by integrating data from various levels of gene regulation.
Similar content being viewed by others
Introduction
Lipids are major biomolecules that fuel energy production of the human body. During energy-rich conditions, lipids get stored in various human tissues and are utilized in energy insufficiency. Abnormal lipid metabolism is associated with obesity, type 2 diabetes (T2D), cardiovascular diseases, Alzheimer’s disease, sleep apnea, cancers, etc. [1,2,3,4,5,6]. Total cholesterol (TC), low-density lipoproteins (LDL), high-density lipoproteins (HDL), and triglycerides (TG) are vital constituents of lipid fraction in blood. Lipid traits exhibit strong genetic control [7,8,9,10,11,12] and are largely modifiable by dietary intake and physical activity [13,14,15].
Individuals from North India, speaking the Indo-European language are genetically a diverse population. Their diet constitutes high intake of carbohydrates [16] and thus are specifically vulnerable to atherogenic dyslipidemia that is characterized by lower TC, lower HDL and higher TG levels [17]. Atherogenic dyslipidemia is strongly associated to several metabolic diseases [18, 19]. Indians have been previously studied for replication of known lipid-associated variants [20,21,22], however, there was a lack of comprehensive genome-wide association study (GWAS) conduced exclusively on Indians.
We performed a two-staged genome-wide association study (GWAS) on 5271 healthy individuals and replicated earlier GWAS findings—CELSR2, CETP, LPL, ZNF259, and BUD13 and correspondingly determined independent signals within these established lipid loci. Besides, we identified certain novel variants that exhibited strong association to lipid parameters in Indians but fail to attain GWAS significance. Majority of these variants were found to be dynamic CpG sites in human subcutaneous adipose tissue and pancreas. Of which, QKI variants showed noteworthy association to all four lipid traits and their designated region was observed to be a key gene regulatory segment denoting dynamic transcription particularly in human subcutaneous adipose tissue. To substantiate our genetic findings, we investigated DNA methylation marks in peripheral blood in Indians at the identified loci and spotted robust associations to important genic CpG sites. Identified signals featured key control regions of the genome regulating lipid homeostasis.
Participants and methods
The study was conducted in accordance with principles of Helsinki Declaration and approved by Ethics Committee of All India Institute of Medical Sciences, New Delhi, India and CSIR-Institute of Genomics and Integrative Biology, New Delhi, India. Written informed consent was taken from all study participants.
Study population
The individuals included in the study are Indo-European speakers randomly sampled from the Northern part of India. These individuals are also a part of the INdian DIabetes Consortium (INDICO) [23] and were included in the control group (normoglycemic) in T2D GWAS conducted previously in the lab [24]. Apparently healthy individuals were recruited through health awareness camps piloted in/around Delhi. Their anthropometric and clinical characteristics are provided in Supplementary Table 1.
Serum TC, LDL, HDL, and TG levels were measured enzymatically using COBAS Integra 400 plus (Roche Diagnostics, Germany). Genomic DNA was extracted from peripheral blood using salt precipitation method.
Genome-wide association study
Discovery phase
Illumina Human610-Quad BeadChips (Illumina Inc., San Diego, CA) were used for the genome-wide scan as part of GWAS studies earlier conducted for T2D and related metabolic traits in our laboratory [24,25,26,27]. GenCall algorithm was employed to ascertain genotype calls (GenomeStudio, Illumina Inc.). Quality control and association analysis is charted in Supplementary Fig. 1. In brief, individuals under lipid-lowering medication, samples with genotype call rate <95%, extremely low or high heterozygosity (3 SD from mean value) and discordant sex were removed. SNP calls with missing rate >5%, MAF <0.01 or with MAF 0.01–0.05 but Hardy–Weinberg equilibrium (HWE) p < 10−6 were removed. Identity-by-descent was checked using the following parameters (pi_hat >0.1875 {relatedness} and pi_hat >0.98 {duplication}). Principal component analysis (PCA) was used to detect population outliers. We ran PCA on a LD-pruned, reduced set of markers. LD pruning of SNPs was performed with autosomal SNPs exercising the “–indep-pairwise” option of PLINK v1.07 (http://pngu.mgh.harvard.edu/~purcell/plink) [28] with r² of 0.2 and window size of 50 SNPs. First ten principal components were used to identify 31 subjects as population outliers (6 SD away from mean value) and were removed. Serum lipid values were inverse normalized using R (http://www.r-project.org/). After QC, association testing was performed for 551825 SNPs with inverse-normalized lipid values in 1036 individuals by linear regression analysis under additive model adjusting for age, sex, BMI, and first two principal components using PLINK. Median χ²-statistics were used to compute genomic inflation factor λ. QQ and Manhattan plots were constructed using qqman package in R (http://www.r-project.org/).
Replication phase
The present study was conducted as a part of a large study to identify genetic determinants of different quantitative traits (glycemic, lipids, anthropometric, nitrogen metabolites, etc.) in Indians [25,26,27]. To select a decent and creditable number of variants for replication as per our available resources, we selected all markers that were strongly associated (p < 10−4) to any of the studied quantitative trait, entailing that certain markers with discovery p > 10−4 for lipids were also genotyped for replication in an independent sample-set (N = 4235) using Illumina Golden Gate assay. Overall 204 samples (6%) were genotyped as replicates to assess experimental accuracy (replication error <0.01 was detected between technical replicates). Samples with call rate <90% were excluded. SNPs with genotype call rate <90%, genotype confidence score <0.25, GenTrans score <0.60, cluster separation score <0.40, MAF <0.01 and HWE p < 1 × 10−6 were discarded. Lipid values were inverse normalized. Association analysis was performed using linear regression model adjusted for age, sex, and BMI.
An inverse variance method was used to meta-analyze discovery and replication phase results under a fixed effect model by METAL (www.sph.umich.edu/csg/abecasis/Metal) [29]. Conditional analysis for signals in CETP, LPL, and ZNF259 loci was carried in combined data (discovery and replication phases) using additive linear model and was adjusted for age, sex, BMI, and respective SNP genotypes as covariates using PLINK.
Statistical power of the study
Statistical power of study was calculated for meta-analysis for allele frequencies ranging from 0.01 to 0.50 at different effect sizes assuming log-additive model. Power has been plotted in y-axis for corresponding MAF in x-axis. Two-tailed test at significance level of 0.05 and effect sizes ranging from 0.001 to 8.63 (TC), 0.001 to 7.01 (LDL), 0.0001 to 3.18 (HDL), and 0.009 to 18.69 (TG) obtained from literature was utilized for power calculation. Average TC levels of 183.42 mg/dl, LDL levels of 115.74 mg/dl, HDL levels of 46.13 mg/dl and TG levels of 121.82 mg/dl and a mean standard deviation of 42.32 mg/dl (TC), 33.32 mg/dl (LDL), 12.25 mg/dl (HDL) and 71.26 mg/dl (TG) were used.
In silico replication
We performed in silico replication of novel signals by combining summary statistics in Indians and Global Lipids Genetics Consortium (GLGC) dataset that constituted GWAS data of quantitative lipids from 188,577 Europeans and 7898 non-Europeans (East-Asians, South-Asians, and Africans) [11]. The effect sizes were converted to uniform unit by absolute conversion factor before meta-analysis by METAL.
Imputation analysis
Imputation was carried out as described previously [24]. In brief, 1000Genomes Phase 3 was used as the reference panel. Pre-phasing was done using SHAPEIT [30] for the respective chromosomes. A region of 2 Mb on either side of the variant was imputed using IMPUTE2 [31] which also covered its respective LD block. Imputed SNPs were subjected to a stringent QC: Certainty ≥0.90, Info ≥0.5, and MAF ≥0.01. Finally, the QC passed SNPs were tested for association in Indians using PLINK, adjusting for age, sex, BMI, PC1, and PC2.
Correlation among lipid traits and SNPs
Correlations among lipid traits were computed using R corrplot package (http://www.r-project.org/). R corrplot function was used to plot the graph of the correlation matrix. Correlation among association p-values for SNPs were calculated for at least 2 lipid traits in combined genotype dataset of discovery and replication phases exercising available commands in Microsoft Excel.
Pathway analysis
We used the shared genes among at least 2 lipid traits as an input for STRING analysis (version 10.5 {https://string-db.org}) [32] to explore known or predicted protein–protein interactions between query genes and their additional functional interactors. All interactions were scored at the highest confidence (0.90) and only 10 more interactors of strongest interaction evidence with the query genes were allowed in the network. Identified genes were also analyzed using Reactome [33] and GeneMANIA [34].
Gene-based association analysis
We also performed Univariate gene-based association analysis using effective chi-squared test (ECS) implemented in knowledge-based mining system for genome-wide genetic studies (KGG v4) available at http://statgenpro.psychiatry.hku.hk/limx/kgg/download.php. SNPs within each gene for the respectively associated lipid trait were used as an input for the KGG v4.
For QKI, we performed multivariate gene-based association test by extended Simes procedure (MGAS) in our GWAS dataset [35]. Association p-values of markers within 2 Mb region of QKI loci for all 4 lipid traits and trait correlation information were incorporated in MGAS based model using KGG v4.
1000Genomes Phase III data that constitutes Europeans, Africans, Americans, East-Asians, and South-Asians was used for computing LD between all tested markers within the studied region.
DNA methylation analysis
We searched for associated meth-QTLs in peripheral blood using Infinium HumanMethylation450 BeadChips of 233 normoglycemic Indians studied in the discovery phase of GWAS. Data generation has been described previously [36]. QC method is charted in Supplementary Fig. 1. Briefly, sample QC involved sex disparity checks and samples with failed bisulphite conversion (samples having intensity 3 SD away from mean intensity for C1, C2, C3, and C4 probes). CpGs with bead count less than 3 in 5% of samples and detection p-value >0.01 for less than 1% of samples were excluded. CpGs in sex chromosomes (X and Y), established cross-hybridization probes and polymorphic CpGs were also removed. CpGs with 100% call rate in all the samples have only been considered for analysis. Methylation outlier value for CpGs was fixed by fixMeth-Outliers command in minifi. Methylation data were regressed for confounders such as cell composition, age, sex, BMI, bisulphite conversion efficiency, and plate number. Methylation data were extracted for identified GWAS variants and tested for SNP-CpG association using linear regression model in PLINK.
Gene regulatory information
Global expression-QTL (eQTL) data was retrieved from GTEx-portal-v7 (The Broad-institute of MIT and Harvard) [37]. Whole Genome Bisulphite Sequencing (WGBS) data was retrieved from ENCODE dataset [38] for human subcutaneous adipose tissue and pancreas of a female aged 30 years and blood monocytes of a male aged 37 years. Human ATAC-seq and histone marks H3K36me3 or H3K27me3 data is obtained from female individuals aged 49 years (subcutaneous adipose tissue) and 30 years (pancreas) [38]. ChIP-seq data of CTCF binding was acquired from a female aged 51 years for both the tissues [38]. ENCODE data for K562 cell-line (DNase I hypersensitivity, Histone modifications, chromatin state segmentation, Transcription Factor (TF) binding sites, etc.) was also examined. Predicted sites for TF binding were retrieved from JASPAR database [39]. All gene regulatory data was plotted in UCSC genome-browser [40].
Results
The present study was observed to be sufficiently powered to detect truly associated variants for quantitative lipid traits in Indians (Supplementary Fig. 2). Under the null distribution QQ plots of all four lipid traits reflected a good agreement (Supplementary Fig. 3). Genomic inflation factor (λ) was observed to be 1 thereby indicating homogeneity of the studied population.
Genome-wide association analysis of total cholesterol (TC)
In discovery phase, the strongest association was observed for variant rs134221 that resided in a strong enhancer element within a long non-coding RNA gene LINC01399 (p = 2.05 × 10−6) (Fig. 1). However, in replication phase, the association with LINC01399 appeared nominal (rs134221 p = 0.04 and rs80731 p = 0.02). Meta-analysis of discovery and replication phases revealed genome-wide significance at a previously known locus—CELSR2 harboring SNP rs646776 (p = 1.03 × 10−9) (Table 1). Loci QKI (rs9458854, rs9458855) and REEP3 (rs7083226) showed sub-genome-wide significance and surfaced for the first time in association with TC (respective p-values = 3.71 × 10−5, 9.78 × 10−5 and 9.89 × 10−5, respectively) (Table 2).
Manhattan plots of associated P-values for lipid traits. The −log10 p-values for the association of directly genotyped SNPs are plotted as a function of genomic position (National Center for Biotechnology Information Build 37). P-values were determined using linear regression adjusted for age, sex, BMI, PC1, and PC2 in discovery phase analysis
Amongst the variants that were tested only in replication phase, earlier reported SNPs rs10401969 (SUGP1 gene) and rs599839 (PSRC1 locus) featured genome-wide association to TC levels in Indians (Supplementary Table 2). Additionally, a novel variant rs16996148 falling near CILP2 gene turned out to be nearly genome-wide significant (p = 6.47 × 10−8) (Supplementary Table 2).
Genome-wide association analysis of LDL
In discovery phase of LDL GWAS, variant rs4979205, a highly significant cis-eQTL for ZNF833 in human brain, appeared as lead signal (p = 4.52 × 10−6) (Fig. 1). The association with ZNF833 was reinforced by other variants rs10981574, rs1886525 and an exonic variant rs10981592 (p-values = 3.05 × 10−5, 3.26 × 10−5 and 3.27 × 10−5). However, this association was lost after meta-analysis of discovery and replication phases (rs4979205, p = 0.12). Another locus LYZL1, represented by four SNPs rs1858571, rs12412488, rs2256713, and rs10491049, showed up strongly in discovery phase (p-values = 5.64 × 10−6, 5.88 × 10−6, 1.54 × 10−5 and 2.6 × 10−5, respectively) but not when meta-analyzed (rs12412488, p = 0.31). Besides, variants rs1862859 and rs1846697 residing near microRNA gene MIR4426 (respective p = 7.65 × 10−6, 1.87 × 10−5), SNPs rs10519042 and rs7180578 in SQOL locus (p-values = 7.9 × 10−6, 3.37 × 10−5), and cis-eQTLs rs11168524 and rs1471997 for H1FNT (p = 8.96 × 10−6 and 2 × 10−5, respectively) featured strong association to LDL levels in discovery phase (Fig. 1) but not after meta-analysis.
Established GWAS signal CELSR2 (rs646776) was the strongest signal after meta-analysis (p = 1.08 × 10−13) (Table 1). Two novel loci—REEP3 (rs7083226) and TMCC2 (rs2290265) presented robust association to LDL levels but failed to attain GWAS significance (Table 2). Previously known variants—rs599839 (PSRC1) and rs10401969 (SUGP1) staged genome-wide significance when tested in the replication phase (Supplementary Table 2).
Genome-wide association analysis of HDL
Known HDL signal—CETP was observed as the strongest signal in Indians throughout the study (Fig. 1, Table 1). In discovery phase, the locus was represented by five variants rs3764261 (p = 6.29 × 10−12), rs1800775 (p = 3.97 × 10−10), rs1532624 (p = 5.28 × 10−9), rs708272 (p = 3.41 × 10−7), and rs7499892 (p = 3.12 × 10−6). In meta-analysis, variants—rs3764261, rs1532624, rs1800775, rs9989419, and rs4783961 defined CETP locus (p-values ≤ 10−8). Novel variants rs2009667 and rs1048015, highly significant cis-eQTLs for IFT27 gene in several human tissues, were observed amongst lead signals in the discovery phase (respective p = 1.63 × 10−6 and 1.41 × 10−5) but persisted after meta-analysis (rs2009667 p = 1.22 × 10−4). Further, the variants falling in NCS1 gene (rs7873936 and rs7852859) sustained significance after meta-analysis (p-values = 1.23 × 10−4 and 6.32 × 10−4). However, association of other discovery phase leads diminished after meta-analysis—rs4804386 (MUC16); rs3094471 and rs1859137 (KCTD5); rs1151452 and rs1151457 (LINC02337); and rs6068695 (SUMO1P1). Besides CETP, LPL surfaced as another strong GWAS signal after meta-analysis (Table 1) and also amongst variants tested only in the replication phase (Supplementary Table 2). Though not genome-wide significant, we observed novel variant (rs4544358) residing within FAM129C to show strong association to HDL levels after meta-analysis (p = 9.96 × 10−5) (Table 2).
Genome-wide association analysis of triglycerides (TG)
The strongest association with triglyceride levels was seen for GUCY1A2 variant rs12574588 in the discovery phase (p = 1.48 × 10−7) (Fig. 1). This association however receded in meta-analysis (p = 0.003). Reported TG variants- rs7350481 (BUD13), rs964184 and rs6589567 (ZNF259) and rs10096633 (LPL) persisted as lead signals throughout the study (respective meta-analysis p-values = 1.12 × 10−20, 7.64 × 10−30, 1.01 × 10−16, 1.71 × 10−9) (Table 1). Conversely, association of another discovery phase lead rs6857945 (LDB2) regressed in meta-analysis (p = 4.2 × 10−4). Strong discovery phase SNPs in VAPB and GNPTAB perished in meta-analysis. We identified an unreported SNP (rs4128744) near LPL to feature robust genome-wide association to serum TG levels in Indians (p = 2.32 × 10−9) (Table 1). Variant rs4128744 is in strong linkage-disequilibrium (r2 > 0.9) with earlier reported TG GWAS variants rs9644568 and rs115849089. All through study, novel variants rs12771265 and rs4746882 (FAM241B), and rs9393071 (LOC100506207) exhibited strong association to TG but could not reach genome-wide significance (respective meta-analysis p values = 1.16 × 10−6, 3.1 × 10−5 and 4.07 × 10−5) (Table 2).
Conditional analysis to identify independent GWAS signals
To identify effective independent variants within CETP, LPL, and ZNF259 loci, we performed conditional analysis of the combined data (discovery and validation phase) using additive linear model. We found rs3764261 as the leading signal in the HDL-associated CETP locus (Table 3). Association testing after adjusting for variant rs3764261 resulted in loss of genome-wide significance of other four variants in CETP locus (rs1532624, rs1800775, rs4783961, and rs9989419), though nominal significance retained (rs1532624, p = 8.13 × 10−4; rs1800775 p = 1.03 × 10−4; rs4783961, p = 2.99 × 10−4 and rs9989419, p = 0.03) (Table 3). Conditional analysis of LPL signals pinpointed SNP rs10096633 as the key variant to dismiss the genetic significance of variants rs12678919 and rs4128744 with HDL levels (Table 3). Furthermore, this LPL variant turned out to be a central variant for TG levels also. Correspondingly, for ZNF259 locus, we noticed that variant rs6589567 fail to attain genetic significance upon conditioning of association analysis for genotypes of variant rs964184 (p = 0.25) (Table 3).
In silico replication of novel variants in GLGC population
Meta-analysis of summary statistics of the identified novel variants in publicly available high-throughput dataset of quantitative lipid traits in diverse human populations (Global Lipids Genetics Consortium Data-GLGC) did not improve their association status (Supplementary Table 3). All identified novel variants (Table 2) were observed to lack association in other populations but few (QKI and TMCC2 variants) were nevertheless seen to follow a similar directionality for the effect.
Imputation of novel loci
For QKI, we identified certain variants that displayed higher significance for association to total cholesterol (discovery p = 10−3) than index variants rs9458854 and rs9458855 (Supplementary Table 4a, Supplementary Fig. 4). These imputed variants resided in key regulatory elements of QKI gene (other introns and an upstream enhancer). An interesting variant rs76779527 within an upstream bivalent enhancer of QKI gene in human adipose tissue featured a strong binding site for GATA2, a TF that inhibits adipogenesis. Apart from this, some imputed intronic variants also constituted strong binding sites for important TFs involved in lipid metabolism (STAT1, STAT2, FOXD3, IRF1, and TGIF1). Further for REEP3 locus, in association to total cholesterol, p-value significance sustained 10−3 after imputation (Supplementary Table 4b) but marginally improved from 10−2 to 10−3 for LDL (Supplementary Table 4c). None of the later variants seemed to constitute regions vital for a gene’s activity. At TMCC2 locus, two intergenic variants were found to be associated with LDL at a marginally higher significance than the index SNP rs2290265 (Supplementary Table 4d). These imputed variants along with the two other genotyped variants in the region were observed as highly significant cis-eQTLs for TMCC2 gene in human subcutaneous adipose and skeletal muscle tissues. For loci FAM129C, FAM241B, and LOC100506207, no variant other than the index SNPs turned out to be strongly associated to the respective trait after imputation.
Shared variants for lipid traits
We next looked for shared associated variants among studied lipid traits and observed significant correlations among all four lipid traits (Supplementary Fig. 5) that remained unaffected by the transformations. TC levels were seen to strongly correlate with LDL levels (correlation coefficient: 0.88) and modestly with HDL and TG levels (correlation coefficients: 0.21{HDL} and 0.33{TG}). LDL levels showed suggestive correlation with TG levels (correlation coefficient: 0.22) and a relatively weaker one with HDL levels (correlation coefficient: 0.09). Furthermore, as reported in earlier studies, HDL and TG levels featured a reasonable inverse correlation (correlation coefficient: −0.27).
Taking into consideration that lipid traits are well-correlated and can dictate the likely overlap of certain variants in Indians, we next calculated the pair-wise correlation between the association p-values for all the SNPs for at least 2 traits in the combined genotype data. We observed very little overall correlation (data not shown). Interestingly, three novel variants within the QKI gene—rs9458854, rs9458855, and rs9458846 showed robust association to all the four studied lipid traits (Fig. 2a). Also, we found that there are minimum of 99 SNPs that are simultaneously associated to at least 2 lipid traits (p-value <0.05) (Supplementary Table 5), reflecting that these genomic regions administer the levels of multiple quantitative lipids at a time.
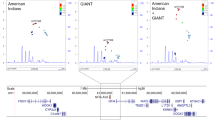
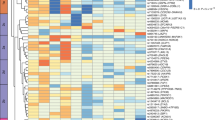
QKI as a novel locus for lipid traits. a Association analysis of QKI variants with all four lipid traits in Indians. TC: total cholesterol, TG: triglycerides. b Reported genetic variants for TC in QKI locus. The highlighted area shows the associated region in Indians (present study). Information has been obtained from T2D knowledge portal. c Gene expression of QKI in human subcutaneous adipose tissue and pancreas (mRNA levels). [GTEx portal]. d Comparison of gene regulatory signatures in human subcutaneous adipose and pancreas tissue. H3K36me3: active gene body mark signifies active transcription; H3K27me3: mark for repressed transcription; ATAC-seq peaks: open chromatin region; CTCF: TF that binds to repressed genic regions. [ENCODE data]. e QKI region harboring associated variants rs9458854 and rs9458855. Predicted motifs for respective TFs. The gray scale reflects enrichment where black color denotes strongest binding and light gray as weak binding [JASPAR data]
QKI as a novel locus for lipid traits
Subsequent to observing QKI locus to be associated with all lipid traits, we looked for all reported genetic variants in the region. The strongest reported association for TC was for SNP rs190262954 (p = 1.14 × 10−4) (Fig. 2b). There was no other stronger variant in the region documented for any of four blood lipids (Supplementary Fig. 6). QKI gene was observed to be highly expressed in human subcutaneous adipose tissue (Fig. 2c). To speculate the functional role for our identified QKI variants, we studied the open chromatin signatures, active and repressive histone marks (H3K36me3 and H3K27me3, respectively), CTCF binding and TF binding in human subcutaneous adipose tissue and pancreas. Both tissues featured an open chromatin at QKI gene (Fig. 2d). However, in comparison to pancreas where the expression of QKI is spare (Fig. 2c), adipose tissue exhibited remarkably higher enrichment for H3K36me3 marks and an absence of H3K27me3 marks, which is essentially a signature for active gene transcription (Fig. 2d). Additionally, the region located immediate downstream of the associated variants featured an abolition of CTCF binding in adipose tissue although present in the pancreas (Fig. 2d). The associated variants were seen to display high TF bit-scores at their respective positions (Supplementary Fig. 7), thereby denoting highly conserved elements of the strong predicted motifs for key transcription factors—GCM1, GCM2, SPIC, FOXK1, Foxo1, FOXP2 (Fig. 2e), and ESR2 (data not shown).
Exploring the WGBS data, we observed that the variant rs9458854 is a dynamic hypermethylated CpG site in both the human adipose tissue and the pancreas (Fig. 2e). Indeed, 100% of the sequenced were found to be methylated. Besides QKI, majority of the other identified novel variants were also found to be active CpG sites in human adipose or pancreas tissue (Supplementary Fig. 8).
Pathway analysis of shared genes
In view of a substantial number of shared SNPs among lipid traits, we then performed a gene-set enrichment analysis to identify associated biological pathways. We used a total of 80 genes (listed in Supplementary Table 5) as an input for a STRING database search. The analysis revealed significant enrichment of crucial biological processes such as citric acid cycle (TCA cycle), lipid homeostasis or remodeling, acyl-CoA metabolism, macrophage colony-stimulating factor signaling pathway and RIG-I signaling (Supplementary Fig. 9, Supplementary Table 6a). Besides STRING, we analyzed the identified genes using two other high-performance tools – pathway analysis by Reactome and biological function prediction by GeneMANIA. Here too we observed significant enrichment (FDR ≤ 0.05) of lipid-related processes pertaining to lipid homeostasis, lipoprotein remodeling, foam cell differentiation and assembly of active LPL and LIPC lipase complexes (Supplementary Tables 6b, 6c).
Gene-based association analysis
In order to avoid missing out any important genetic loci for the studied lipid traits, we also performed gene-based analysis based on associated marker accumulation on whole genes. The analysis revealed certain novel GWAS loci which were not captured previously by SNP-based association testing. TC lead signal CELSR2 as captured by SNP-based GWAS, retained significance in gene-based testing (p = 0.02). Additionally, genes—LINC01340 and IQCJ-SCHIP1 turned out be genome-wide significant for TC in gene-based testing (Supplementary Table 7). For LDL levels, both CELSR2 and TMCC2 remained significant (p = 0.01). Gene PCBP3 showed near genome-wide significance to LDL levels in Indians (p = 5.52 × 10−7) (Supplementary Table 7). In agreement with SNP-based testing for HDL, gene-based analysis also featured CETP as the lead signal (p = 1.03 × 10−16), and LPL sustained nominal significance (p = 0.04). Besides, previously unreported genes—FT27, IFT88, and LINC02141 exhibited genome-wide significance for HDL levels in Indians (Supplementary Table 7). For TG, ZNF259 maintained strong association in gene-based analysis (p = 1.90 × 10−6) (Supplementary Table 7). Furthermore, genes BUD13 and LPL also sustained significance for TG in Indians (respective p = 1.15 × 10−3 and 0.02). The analysis also revealed TEX26 as a strong GWAS loci for TG for the first time (p = 9.0 × 10−8) (Supplementary Table 7).
In conjunction with our findings indicating QKI as an important locus for all four studied quantitative lipids in Indians, we performed a multivariate gene-based association test for QKI by extended Simes procedure (MGAS) implementing gene-based testing of multiple correlated phenotypes in unrelated individuals. This gene-based testing also set out QKI as the lead gene in the associated loci (2 Mb genomic region) retaining statistical significance (p = 0.05) (Supplementary Table 8).
DNA methylation study in Indians
To pinpoint putative functional variants in identified GWAS and sub-GWAS signals, we investigated and integrated DNA methylation data from peripheral blood in Indians. Both, known and novel genetic variants were profoundly enriched as meth-QTLs that tend to regulate methylation pattern at related CpG sites within associated genes (Supplementary Table 9, Table 4). Amongst novel signals, FAM129C variant rs4544358 was observed to be significantly associated with differential methylation of five different CpG sites in Indians (Table 4). TMCC2 SNP rs2290265 and REEP3 variant rs7083226 portrayed robust association to DNA methylation at specific genic CpG sites.
We observed that all three unreported genes (FAM129C, TMCC2, and REEP3) are expressed moderately in human blood (Supplementary Fig. 10). Therefore, we layered our genetic and epigenetic data from peripheral human blood with gene regulatory information from leukemia cell-line K562. We found that the associated CpG sites for meth-QTL rs4544358 resided in FAM129C promoter, exons, and 3′UTR; and constituted strong binding-sites for REST, ZBTB7A, RAD21, and NRSF (Supplementary Table 10). Likewise, meth-QTLs rs2290265 (TMCC2) and rs7083226 (REEP3) were significantly associated with CpGs that comprise key regulatory elements.
Though QKI is fairly expressed in human blood (8.12 transcripts per million), we did not find any association between alternative alleles of variant rs9458854 and level of DNA methylation at the overlapping CpG site in blood in Indians. To validate our observation, we looked at the publicly available WGBS data of classical monocytes (CD14+) in human blood. Consistent with our observation in peripheral blood in Indians, classical monocytes in blood also showed lack of DNA methylation at variant rs9458854 wherein none of the sequenced reads were found to be methylated (Supplementary Fig. 11) in contrast to be a definite hypermethylated site in adipose tissue.
Discussion
This is first GWAS conducted for serum lipid traits (total cholesterol, LDL, HDL, and triglycerides) exclusively in Indians. Recurrent GWAS signals found in studies in European, Hispanic, African, South Asian, and East Asian populations, namely—CELSR2, CETP, LPL, ZNF259, and BUD13 [9, 11, 12, 41,42,43], also constituted the most significantly associated SNPs in our study as well. It signifies universality of these variants, thereby confirming the existence of an independent genetic control that governs lipid traits regardless of vast genetic diversity and differential food habits in varied human populations.
CELSR2 (Cadherin EGF LAG seven-pass G-type receptor 2) is a receptor protein in which cadherin domains act as homophilic-binding regions and EGF-like domains exhibit cell adhesion and receptor-ligand interactions [44]. CETP is a Cholesteryl Ester Transfer Protein, involved in transfer of neutral lipids among lipoprotein particles [44]. LPL encodes for lipoprotein lipase that breaks down stored triglycerides for energy production whereas BUD13 and ZNF259 are involved in modification and cellular transport of pre-mRNAs [44]. Previously, a few candidate studies have individually replicated these variants in Indians [20,21,22] but not at genome-wide before.
Within these ubiquitously associated loci, we identified independent variants. SNP rs3764261 residing within repressed element of CETP promoter, is highly significant cis-eQTL for CETP in human lungs, liver, and stomach [37]. Prime variant rs10096633 is significant cis-eQTL for LPL in blood and SNP rs964184 located in 3′UTR of ZNF259 is cis-eQTL for pseudogene RP11-109L13.1 situated downstream of BUD13 [37]. Evidently, eQTL findings justify observed genetic associations on biological grounds.
Besides established variants, we found certain novel variants at sub-genome-wide significance in Indians—QKI, REEP3, TMCC2, FAM129C, FAM241B, and LOC100506207. It is well known that genetic variance is mainly contributed by multiple SNPs with small effects that may often be missed out due to stringent GWAS p-value thresholds and multiple testing corrections [45].
We recognized convincing genetic variants that simultaneously presented association to at least two of the four studied lipid traits. Majority of such variants were often seen associated with HDL, TG levels and LDL, TC levels simultaneously. This trend resembles our previous reported observation in African-Americans where we employed a completely different disease gene mapping method (admixture mapping) to study quantitative lipids [12]. This concurrent finding in two entirely diverse populations in terms of both the genetic and environmental influences, further conforms the universality of lipid variants. The discovered novel variants though not associated to respective lipid traits in other populations yet revealed similarity in the direction of the effect and robust association to multiple other related traits such as BMI, adipose tissue volume, adiponectin, hip circumference, fasting insulin, fasting glucose, HbA1c, and creatinine [46].
QKI gene surfaced as a lead hit featuring significant associations with all the four studied lipid traits in Indians. QKI or Protein Quaking is an RNA binding protein that regulates pre-mRNA splicing, mRNA nuclear export, stability, and protein translation [44]. Changes in QKI expression are reported to significantly alter the expression of known genes involved in lipid metabolism (IDOL, ABCG1, PPARG, NR1H3/LXRA, CD36, LDLR, etc.) in primary monocytes in humans and mouse [47]. Expression of QKI is remarkably higher in macrophages [47] and its depletion impairs differentiation of monocytes into macrophages by triggering alternative splicing events [47]. Macrophages, the key cells of innate immunity, differ extremely from monocytes in terms of lipid metabolism [48] and their chronic accumulation is at the site of tissue injury is a hallmark of numerous complex diseases [47]. Liver and adipose tissue are the primary sites for resident macrophages. Adipose tissue macrophages (ATM) administer obesity-related metabolic dysfunction by secreting LPL (lipoprotein lipase) to enhance their ability to sequester excess lipids [49]. We observed drastically higher enrichment of gene activation histone mark (H3K36me3) for QKI transcription in human subcutaneous adipose tissue encompassing the identified variants. In addition, QKI variants were observed to be strong seats for known transcription factors that are well-established in lipid-related-diseases, especially FOXO1, FOXK1, and GCM2. FOXO1 regulates lipid metabolism by promoting lipolysis and governing adipocyte differentiation and its activity is pivotal in obesity, non-alcoholic fatty liver disease and type 2 diabetes [50, 51]. FOXK1 is a critical mediator of mTORC1-driven gene expression, a process that plays an important role in metabolic diseases [51, 52]. Activity of GCM2 is associated with hyperactivity of parathyroid gland wherein an excessive amount of parathyroid hormone has been proved to adversely affect lipid metabolism [53, 54]. Alleles of QKI variants can effectively influence the binding of these TFs to govern their activity to regulate a gene’s expression which eventually will result in a drastic variability in the levels of expressed QKI protein. A higher expression of QKI in adipose tissue will lead to the production of more macrophages and thus attribute to an enhanced quarantine for free lipids.
Previously, genetic variants within QKI locus have been documented to portray genome-wide significance for height, BMI and T2D [46] but not with quantitative lipids. Certain QKI variants show strong association to classical lipid-related diseases—type 2 diabetes, coronary artery disease, bipolar disorder and measures of obesity (BMI and WHR) [46]. Our identified QKI variants rs9458854 and rs9458855 earlier presented robust association with bipolar disorder (p ≤ 10−4) in a mixed ancestry GWAS comprising 16,731 individuals [55]. Many studies associate state of dyslipidemia with bipolar disorder. A recent study in Asians suffering from bipolar disorder [56], revealed drastic state-dependent alterations in blood lipid profiles (especially cholesterol and triglyceride levels). Considering independently, a strong association with bipolar disease and lipid metabolism, we anticipate that our identified QKI variants do play an important role in certain dyslipidemia-related diseases and hence propose QKI as an essential gene involved in lipid metabolism.
Aside from QKI, our gene-based association analysis revealed certain novel GWAS loci for quantitative lipids in Indians—LINC01340 and IQCJ-SCHIP1 for TC; IFT27, IFT88, and LINC02141 for HDL; and TEX26 for TG. These genes have never been earlier reported as a GWAS hit for the respective lipid trait [46] and shall now be explored further for replication in different human populations.
Further to infer the downstream effects of the identified genetic variants, we followed an integrative approach. We layered our GWAS findings with DNA methylation data from peripheral blood in Indians and examined the gene regulatory signatures. We found that the presence of alternate alleles of these variants significantly influenced methylation levels of several genic CpG sites in Indians. Alterations in DNA methylation modifies binding of specific transcriptional factors and recruitment of methyl CpG binding proteins that affect gene transcription. Gene-specific DNA methylation has been documented to explain inter-individual variability in blood lipid profiles. In context, methylation levels of CpG sites within specific lipid metabolism genes correlate with plasma lipid levels. For instance, DNA methylation in the promoter region of LPL gene is positively correlated with HDL levels in blood leukocytes, however negatively correlated with relative LPL mRNA levels in visceral adipose tissue [57]. CETP DNA methylation is negatively associated with LDL levels [57]. Further, we observed an overlap of meth-QTLs and eQTLs indicating that certain variants within these globally associated loci relate to both DNA methylation and gene expression variation. Associated CpG sites of FAM129C, TMCC2, and REEP3 variants reside in active chromatin regions enriched with regulatory histone modifications and embody confirmed binding sites for essential TFs. Thence, intensive fine mapping in different ethnicities accompanied by functional studies will be valuable in untangling the relevance of such loci in lipid biology.
Our study elucidates universal applicability of certain known GWAS variants for quantitative lipid traits in human population irrespective of diverse genetic or environmental influences and proposes QKI as an important gene regulating lipid biology.
References
Mooradian AD. Dyslipidemia in type 2 diabetes mellitus. Nat Clin Pract Endocrinol Metab. 2009;5:150–9.
Benjamin EJ, Virani SS, Callaway CW, Chamberlain AM, Chang AR, Cheng S, et al. Heart disease and stroke statistics - 2018 update: a report from the American Heart Association. Circulation. 2018;137:E67–E492.
Burnett JR. Lipids, lipoproteins, atherosclerosis and cardiovascular disease. Clin Biochem Rev. 2004;25:2.
Reitz C. Dyslipidemia and the risk of Alzheimer’s disease. Curr Atheroscler Rep. 2013;15:e307.
Adedayo AM, Olafiranye O, Smith D, Hill A, Zizi F, Brown C, et al. Obstructive sleep apnea and dyslipidemia: evidence and underlying mechanism. Sleep Breath. 2014;18:13–18.
Bielecka-Dąbrowa A, Hannam S, Rysz J, Banach M. Malignancy-associated dyslipidemia. Open Cardiovasc Med J. 2011;5:35–40.
Gao C, Tabb KL, Dimitrov LM, Taylor KD, Wang N, Guo X, et al. Exome sequencing identifies genetic variants associated with circulating lipid levels in Mexican Americans: The Insulin Resistance Atherosclerosis Family Study (IRASFS). Sci Rep. 2018;8:5603.
Knoblauch H, Busjahn A, Münter S, Nagy Z, Faulhaber HD, Schuster H, et al. Heritability analysis of lipids and three gene loci in twins link the macrophage scavenger receptor to HDL cholesterol concentrations. Arterioscler Thromb Vasc Biol. 1997;17:2054–60.
Teslovich TM, Musunuru K, Smith AV, Edmondson AC, Stylianou IM, Koseki M, et al. Biological, clinical and population relevance of 95 loci for blood lipids. Nature. 2010;466:707–13.
Dumitrescu L, Carty CL, Taylor K, Schumacher FR, Hindorff LA, Ambite JL, et al. Genetic Determinants of Lipid Traits in Diverse Populations from the Population Architecture using Genomics and Epidemiology (PAGE) Study. PLoS Genet. 2011;7:e1002138.
Willer CJ, Schmidt EM, Sengupta S, Peloso GM, Gustafsson S, Kanoni S, et al. Discovery and refinement of loci associated with lipid levels. Nat Genet. 2013;45:1274–83.
Basu A, Tang H, Lewis CE, North K, Curb JD, Quertermous T, et al. Admixture mapping of quantitative trait loci for blood lipids in African-Americans. Hum Mol Genet. 2009;18:2091–8.
Stamler J. Diet and coronary heart disease. Biometrics. 1982;38(Suppl):95–118.
MacAuley D. Exercise, cardiovascular disease and lipids. Br J Clin Pract. 1993;47:323–7.
Lalonde L, Gray-Donald K, Lowensteyn I, Marchand S, Dorais M, Michaels G, et al. Comparing the benefits of diet and exercise in the treatment of dyslipidemia. Prev Med (Baltim). 2002;35:16–24.
Misra A, Khurana L, Isharwal S, Bhardwaj S. South Asian diets and insulin resistance. Br J Nutr. 2009;101:465–73.
Misra A, Wasir JS, Vikram NK. Carbohydrate diets, postprandial hyperlipidaemia, abdominal obesity & Asian Indians: A recipe for atherogenic disaster. Indian J Med Res. 2005;121:5–8.
Valensi P, Avignon A, Sultan A, Chanu B, Nguyen MT, Cosson E. Atherogenic dyslipidemia and risk of silent coronary artery disease in asymptomatic patients with type 2 diabetes: A cross-sectional study. Cardiovasc Diabetol. 2016;15:104.
Tenenbaum A, Fisman EZ, Motro M, Adler Y. Atherogenic dyslipidemia in metabolic syndrome and type 2 diabetes: Therapeutic options beyond statins. Cardiovasc Diabetol. 2006;5:20.
Braun TR, Been LF, Singhal A, Worsham J, Ralhan S, Wander GS, et al. A replication study of GWAS-derived lipid genes in asian indians: The chromosomal region 11q23.3 harbors loci contributing to triglycerides. PLoS One. 2012;7:e37056.
Arvind P, Nair J, Jambunathan S, Kakkar VV, Shanker J. CELSR2-PSRC1-SORT1 gene expression and association with coronary artery disease and plasma lipid levels in an Asian Indian cohort. J Cardiol. 2014;64:339–46.
Walia GK, Gupta V, Aggarwal A, Asghar M, Dudbridge F, Timpson N, et al. Association of common genetic variants with lipid traits in the Indian population. PLoS One. 2014;9:e101688.
INdian DIabetes COnsortium. INDICO: The development of a resource for epigenomic study of Indians undergoing socioeconomic transition. Hugo J. 2011;5:65–69.
Tabassum R, Chauhan G, Dwivedi OP, Mahajan A, Jaiswal A, Kaur I, et al. Genome-wide association study for type 2 diabetes in indians identifies a new susceptibility locus at 2q21. Diabetes. 2013;62:977–86.
Giri AK, Banerjee P, Chakraborty S, Kauser Y, Undru A, Roy S, et al. Genome wide association study of uric acid in Indian population and interaction of identified variants with Type 2 diabetes. Sci Rep. 2016;6:21440.
Bandesh K, Prasad G, Giri AK, Voruganti VS, Butte NF, Cole SA, et al. Genome-wide association study of C-peptide surfaces key regulatory genes in Indians. J Genet. 2019;98:8.
Prasad G, Giri AK, INDICO, Basu A, Tandon N, Bharadwaj D. Genome-wide association study for C-reactive protein in Indians replicates known associations of common variants. J Genet. 2019;98:20.
Purcell S, Neale B, Todd-Brown K, Thomas L, Ferreira MAR, Bender D, et al. PLINK: A tool set for whole-genome association and population-based linkage analyses. Am J Hum Genet. 2007;81:559–75.
Willer CJ, Li Y, Abecasis GR. METAL: Fast and efficient meta-analysis of genomewide association scans. Bioinformatics. 2010;26:2190–1.
Delaneau JF, Zagury J, Marchini. Improved whole chromosome phasing for disease and population genetic studies. Nat Methods. 2013;10:5–6.
Howie BN, Donnelly P, Marchini J. A flexible and accurate genotype imputation method for the next generation of genome-wide association studies. PLoS Genet. 2009;5:e1000529.
Szklarczyk D, Morris JH, Cook H, Kuhn M, Wyder S, Simonovic M, et al. The STRING database in 2017: quality-controlled protein-protein association networks, made broadly accessible. Nucleic Acids Res. 2017;45:D362–68.
Fabregat A, Jupe S, Matthews L, Sidiropoulos K, Gillespie M, Garapati P, et al. The reactome pathway knowledgebase. Nucleic Acids Res. 2018;46(D1):D649–D655.
Mostafavi S, Ray D, Warde-Farley D, Grouios C, Morris Q. GeneMANIA: a real-time multiple association network integration algorithm for predicting gene function. Genome Biol. 2008;9(Suppl 1):S4.
der Sluis V, Dolan CV, Li J, Song Y, Sham P, Posthuma D, et al. MGAS: a powerful tool for multivariate gene-based genome-wide association analysis. Bioinformatics. 2015;31:1007–15.
Giri AK, Bharadwaj S, Banerjee P, Chakraborty S, Parekatt V, Rajashekar D, et al. DNA methylation profiling reveals the presence of population-specific signatures correlating with phenotypic characteristics. Mol Genet Genom. 2017;292:655–62.
GTEx Consortium. The genotype-tissue expression (GTEx) project. Nat Genet. 2013;45:580–5.
Davis CA, Hitz BC, Sloan CA, Chan ET, Davidson JM, Gabdank I, et al. The Encyclopedia of DNA elements (ENCODE): data portal update. Nucleic Acids Res. 2018;46:D794–D801.
Khan A, Fornes O, Stigliani A, Gheorghe M, Castro-Mondragon JA, van der Lee R, et al. JASPAR 2018: update of the open-access database of transcription factor binding profiles and its web framework. Nucleic Acids Res. 2018;46(D1):D260–D266.
Casper J, Zweig AS, Villarreal C, Tyner C, Speir ML, Rosenbloom KR, et al. The UCSC Genome Browser database: 2018 update. Nucleic Acids Res. 2018;46:D762–D769.
Surakka I, Horikoshi M, Mägi R, Sarin AP, Mahajan A, Lagou V, et al. The impact of low-frequency and rare variants on lipid levels. Nat Genet. 2015;47:589–97.
Below JE, Parra EJ, Gamazon ER, Torres J, Krithika S, Candille S, et al. Meta-analysis of lipid-traits in Hispanics identifies novel loci, population-specific effects, and tissue-specific enrichment of eQTLs. Sci Rep. 2016;6:e19429.
Kurano M, Tsukamoto K, Kamitsuji S, Kamatani N, Hara M, Ishikawa T, et al. Genome-wide association study of serum lipids confirms previously reported associations as well as new associations of common SNPs within PCSK7 gene with triglyceride. J Hum Genet. 2016;61:427–33.
Rebhan M, Chalifa-Caspi V, Prilusky J, Lancet D. GeneCards: integrating information about genes, proteins and diseases. Trends Genet. 1997;13:163.
Yang J, Benyamin B, McEvoy BP, Gordon S, Henders AK, Nyholt DR, et al. Common SNPs explain a large proportion of heritability for human height. Nat Genet. 2010;42:565–9.
Type 2 Diabetes Knowledge Portal. Accessed on: 10 October 2018. Available at: http://www.type2diabetesgenetics.org/
De Bruin RG, Shiue L, Prins J, De Boer HC, Singh A, Fagg WS, et al. Quaking promotes monocyte differentiation into pro-atherogenic macrophages by controlling pre-mRNA splicing and gene expression. Nat Commun. 2016;7:10846.
Fernandez-Ruiz I, Puchalska P, Narasimhulu CA, Sengupta B, Parthasarathy S. Differential lipid metabolism in monocytes and macrophages: influence of cholesterol loading. J Lipid Res. 2016;57:574–86.
Aouadi M, Vangala P, Yawe JC, Tencerova M, Nicoloro SM, Cohen JL, et al. Lipid storage by adipose tissue macrophages regulates systemic glucose tolerance. AJP Endocrinol Metab. 2014;307:E374–E383.
Li Y, Ma Z, Jiang S, Hu W, Li T, Di S, et al. A global perspective on FOXO1 in lipid metabolism and lipid-related diseases. Prog Lipid Res. 2017;66:42–49.
Golson MJ, Kaestner KH. Fox transcription factors: from development to disease. Development. 2016;143:4558–70.
He L, Gomes AP, Wang X, Yoon SO, Lee G, Nagiec MJ, et al. mTORC1 Promotes Metabolic Reprogramming by the Suppression of GSK3-Dependent Foxk1 Phosphorylation. Mol Cell. 2018;70:P949–960.
Hashemolhosseini S, Wegner M. Impacts of a new transcription factor family -mammalian GCM proteins in health and disease. J Cell Biol. 2004;166:765–8.
Akmal M, Kasim SE, Soliman AR, Massry SG. Excess parathyroid hormone adversely affects lipid metabolism in chronic renal failure. Kidney Int. 1990;37:854–8.
Psychiatric GWAS Consortium Bipolar Disorder Working Group. et al. Large-scale genome-wide association analysis of bipolar disorder identifies a new susceptibility locus near ODZ4. Nat Genet. 2011;43:977–83.
Huang YJ, Tsai SY, Chung KH, Chen PH, Huang SH, Kuo CJ. State-dependent alterations of lipid profiles in patients with bipolar disorder. Int J Psychiatry Med. 2018;53:273–81.
Guay SP, Brisson D, Lamarche B, Marceau P, Vohl MC, Gaudet D, et al. DNA methylation variations at CETP and LPL gene promoter loci: New molecular biomarkers associated with blood lipid profile variability. Atherosclerosis. 2013;228:413–20.
Acknowledgements
The authors thank all study participants. We acknowledge the support and participation of members of the INDICO consortium in data generation. We also thank GLGC study for summary statistics for meta-analysis. KB acknowledges CSIR for Senior Research Fellowship (CSIR-SRF). GP and AKG acknowledge UGC for Senior Research Fellowship (UGC-SRF).
Funding
This work was supported by the Council of Scientific and Industrial Research [CSIR], Government of India through Centre for Cardiovascular and Metabolic Disease Research [CARDIOMED] project [Grant No: BSC0122-(8)]. This work was also funded by the Department of Science and Technology-PURSE-II (DST/SR/PURSE Phase II/11) given to Jawaharlal Nehru University, New Delhi, India.
Author information
Authors and Affiliations
Consortia
Contributions
KB wrote the final version of manuscript. KB and GP researched the data, interpreted the results and drafted the manuscript. KB, GP, AKG, YK, and MU generated the data. AKG performed whole genome DNA methylation study in Indians. AB supervised the statistical analysis and critically reviewed the manuscript. NT provided intellectual inputs. DB is guarantor of work who conceived, supervised, obtained financial support, and oversaw the entire study.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
About this article
Cite this article
Bandesh, K., Prasad, G., Giri, A.K. et al. Genome-wide association study of blood lipids in Indians confirms universality of established variants. J Hum Genet 64, 573–587 (2019). https://doi.org/10.1038/s10038-019-0591-7
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s10038-019-0591-7
This article is cited by
-
Genetics and epigenetics of diabetes and its complications in India
Human Genetics (2024)
-
Genome-wide analysis identifies novel loci influencing plasma apolipoprotein E concentration and Alzheimer’s disease risk
Molecular Psychiatry (2023)
-
Multi-trait discovery and fine-mapping of lipid loci in 125,000 individuals of African ancestry
Nature Communications (2023)
-
A Nutrigenetic Update on CETP Gene–Diet Interactions on Lipid-Related Outcomes
Current Atherosclerosis Reports (2022)
-
Meta-analysis of sub-Saharan African studies provides insights into genetic architecture of lipid traits
Nature Communications (2022)