Abstract
Metachromatic leukodystrophy due to Arylsulfatase A enzyme deficiency is an autosomal recessive disorder caused by biallelic variations in ARSA gene. Till date 186 variations have been reported in ARSA gene worldwide, but the variation spectrum in India is not known. The aim of this study was to identify the variation profile in Indian patients presenting with features of Arylsulfatase A deficient metachromatic leukodystrophy. We sequenced the ARSA gene in 51 unrelated families and identified 36 variants out of which 16 were novel. The variations included 23 missense, 3 nonsense, and 6 frameshift variants (3 single-base deletions and 3 single-base duplications), 1 indel, one 3 bp deletion, and 2 splice site variations. The pathogenicity of the novel variations was inferred with the help of mutation prediction softwares like MutationTaster, SIFT, Polyphen-2, PROVEAN, and HANSA. The effects of the identified sequence variants on the protein structure were studied using in silico methods. The most common variation was c.931 C > T(p.Arg311*), found in 11.4% (14 out of 122 alleles) of the tested individuals. To the best of our knowledge, this study is the first of its kind in India with respect to the size of the cohort and the molecular diagnostic method used and one of the largest cohorts of metachromatic leukodystrophy studied till date.
Similar content being viewed by others
Introduction
Metachromatic leukodystrophy (MLD) (OMIM #250100) is an autosomal recessive lysosomal storage disorder. It is caused by the variations in ARSA gene (OMIM *607574), which results in the deficiency of Arylsulfatase A enzyme and accumulation of cerebroside-3-sulfate in the lysosomes of the tissues of central and peripheral nervous system. Intralysosomal storage of galactosylsulfatides leads to progressive demyelination and a variety of neurological symptoms like gait disturbance and cognitive regression. Though the disease occurs pan-ethnically with an estimated frequency of 1 in 40,000 [1] the incidence in India is not yet known.
Three different clinical forms of MLD have been characterized based on age at onset of symptoms: late infantile (onset before 30 months), juvenile (30 months–16 years) and adult (after 16 years). In the late infantile and juvenile forms, patients have gait disturbances, blindness, loss of speech, quadriparesis, peripheral neuropathy, and seizures. In the adult patients, the major signs are behavioral disturbances and dementia with a slow progression over several decades [2].
The ARSA gene located on chromosome 22q13 consists of eight exons and spans a genomic region of 3.2 kb. The ARSA cDNA hybridizes to three different mRNA species of 2.1, 3.7, and 4.8 kb and codes for a 507 aminoacids protein that contains three potential N-glycosylation sites [3]. About 186 variations causing MLD have been identified so far in ARSA gene [4]. In this study, we attempted to analyze the variation spectrum of ARSA gene in Indian patients with MLD.
Methods
Patient recruitment
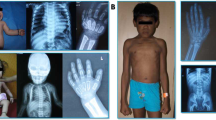
The patients described in this study were unrelated and belonged to different regions in India. The patients were recruited to the study after obtaining informed consent. In case of children, consent was obtained from parents or guardians. The Institute Ethics Committee approved the study. A total of 51 families with at least one affected individual with MLD were included in the study. In 41 families where an affected individual was available for evaluation, a detailed clinical examination was done by a Clinical Geneticist. Magnetic resonance imaging (MRI) of brain was done whenever possible and assay of Arylsulfatase enzyme was done in all patients with clinical and radiological features of MLD. A total of 4 ml blood was collected in ethylenediaminetetraacetic acid vacutainer for all these individuals. In families where previous affected individuals were not available for evaluation, from the available medical records, a diagnosis of MLD was made. Blood samples of parents were collected in such families for molecular genetic analysis.
Arylsulfatase A enzyme assay
Arylsulfatase A enzyme activity in patients was measured in leukocytes obtained from the whole blood sample, using the substrate 10 mM P-Nitrocatechol, in 0.5 M sodium acetate–acetic acid buffer with pH 5.2 and containing 0.023% sodium pyrophosphate and 9.94% sodium chloride. Protein concentration in the lysate was determined by the Lowry method [5] and the ideal concentration was 0.2–0.3 mg of protein. Totally, 200 μl substrate and 200 μl leukocyte homogenate were incubated for 1 h at 37 °C. Then the absorbance was read at 515 nm using Shimadzu UV 2450 spectrophotometer. The enzymatic activity was expressed as nmol of substrate hydrolyzed per mg of protein per hour. For standardization of reference range of enzymes, 20 normal control samples were used. For testing each sample, another control was used. The reference enzyme activity for leukocytes was in the range of 25–80 nmol/h/mg and for fibroblasts was in the range of 190–415 nmol/h/mg. Enzyme assay was performed in our center in 37 out of the 41 patients. In ten families, the index patient was no longer alive. In the remaining four patients, ARSA enzyme levels were not available, but molecular diagnosis was performed based on clinical and radiological features.
Genetic analysis
The genomic DNA was isolated from whole blood using the phenol chloroform extraction method. Primers for the eight exons and the flanking intron–exon boundaries of ARSA were designed by Primer 3 software Input version 0.4.0 and NCBI primer BLAST software. Polymerase chain reaction (PCR) was carried out in the Bio-Rad DNA Engine Peltier Thermal Cycler (Bio-Rad Laboratories, Inc., Hercules, CA) for all 8 exons of ARSA with specific primers, for 25 cycles, with each cycle consisting of denaturation at 94 °C for 1 min, annealing temperature specific to the primers for 55 s and extension at 72 °C for 50 s, with an initial denaturation for 8 min at 96 °C and final extension for 7 min at 72 °C. Bidirectional sequencing was carried out on all the purified PCR products by capillary electrophoresis on ABI 3130 automated genetic analyzer (Applied Biosystems, Foster City, CA). The sequencing data were analyzed using the software EMBOSS [6] and Chromas Lite. The Human Genome Variation Society nomenclature guidelines were used for reporting the identified sequence variations [7]. Nucleotide numbering for ARSA variations is based on cDNA sequence of Ensembl gene transcript ID ENST00000547307. In 41 families, where the proband samples were available, sequencing was first done for the proband, followed by parents, if samples were available. In ten families where proband samples were not available, sequencing of the gene in parents was performed.
In silico characterization of identifies variants
Common polymorphisms and natural variants were filtered out using dbSNP, 1000 Genome browser, ExAC genome browser and in house database. By searching databases like ClinVar and Human Genome Mutation Database (HGMD), known pathogenic variants were identified. The disease causing potential of novel missense pathogenic variants was predicted using the mutation pathogenicity prediction softwareslike MutationTaster [8], HANSA [9], SIFT [10], PROVEAN [11], and PolyPhen-2 [12]. Evolutionary conservation of the mutated amino acidresidues was checked across different species. Mendelian segregation pattern of the identified novel sequence variants in the respective families was studied wherever feasible.
In silico characterization of the effects of novel variants on protein sequence and structure
Homologous sequences of P15289 (protein product of ARSA gene) [13] were obtained by running three rounds of PSIBLAST [14] against the non-redundant database. Multiple sequence alignments were generated using homologous sequences having sequence coverage of at least 95% of the query sequence with ClustalW-2.1 [15] running on default parameters. Gribskov’s score was calculated using the PROPHECY program of EMBOSS by incorporating the BLOSUM62 matrix [6]. The X-ray structure of the P15289 protein was also used for further analysis (PDB code 1AUK) [16]. The percent solvent accessible surface areas of the residues were calculated using PSA program of the JOY suite [17] TRANSEQ program of EMBOSS was used to obtain translated products of the deletion mutant c.576 del C and insertion mutants c.188_189 ins A, and c.445_446 ins T6. PyMOL [18] and Swiss-PdbViewer4.0.4 [19] were used for performing structural analysis and rendering figures. Prediction of the domains affected by the variations was performed using the Uniprot [13] and PROSITE databases [20].
Prenatal diagnosis
Chorion villus sampling (CVS) was done around 11–12 weeks gestation and enzyme assay and sequencing of ARSA were done on the fetal sample. Enzyme assay was done directly in the obtained CVS sample without any prior culturing. Targeted variation analysis was done in CVS DNA based on the variations identified in the proband or in the carrier parents.
Results
Clinical and biochemical evaluation
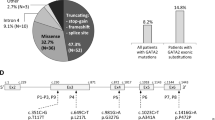
Of the 41 probands, 20 were males and 21 were females. The age of presentation ranged from 18 months to 26 years. A total of 80% (33/41) of patients had late infantile form of MLD, 14% (6/41) had juvenile type, and 4% (2/41) had adult onset type of MLD. All of them showed evidence of leukodystrophy on MRI brain. The clinical features and molecular spectrum are depicted in Table 1. Consanguinity in parents was present in 39 out of 51 families (76%). In all the consanguineous families, homozygous variations were found. In 13 families without consanguinity, homozygous variations were found in 7 families and compound heterozygous variations were found in 6 families.
The arylsulfatase enzyme activity of the patients tested in our laboratory ranged from 0 to 19 nmol/h/mg. The reference enzyme activity was in the range of 25–80 nmol/h/mg. Enzyme activity was not measurable in eight patients.
ARSA variations
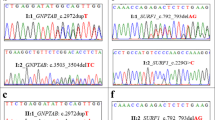
A total of 36 variations were identified out of which 16 were novel variants. The variations included 23 missense variants, 3 nonsense variants, and 6 frameshift variations (3 single base deletions and 3 single base duplications), 1 indel, one 3 bp deletion, and 2 splice site variations [Supplementary Table S1 (30−40)]. The exon wise distribution of the variations is shown in Fig. 1. Nine out of 36 variants were identified in exon 5 (25%). In the ten families, where the proband was not available, in two families, both spouses had known variations and in one family one spouse had a known variation and the other had a novel variant.
In silico characterization of novel variants
All the 16 novel variants were predicted to be disease causing by MutationTaster, SIFT, Polyphen-2, PROVEAN, and HANSA (Supplementary Table S2). None of these variants were present in population database 1000 Genome and pathogenic variant databases like Clinvar and HGMD. Two variants, c.1174 G > A and c.379 G > A, were present in one and two heterozygotes, respectively in ExAC database.
Characterization of the effects of novel variants on protein structure and function by in silico analysis
The human ARSA protein has 507 aminoacids and has a single sulfatase domain. The distribution of the variations in relation to the 8 exons and 507 aminoacids is shown in Fig. 1. Multiple sequence alignments indicate that all the residues at the affected sites are highly conserved and any changes to these positions are expected to cause deleterious effects (Supplementary Figure S1). A total of seven novel nonsynonymous missense variations viz.p.Gly34Glu(c.101 G > A), p.Gln139Lys(c.415 C > A), p.Gly127Arg(c.379 G > A), p.Pro180Gln(c.539 C > A), p.Arg299Trp (c.896 G > C), p.Gly392Arg(c.1174 G > A), and p.Phe399Ser(c.1196 T > C) were identified (Supplementary Table S2) and the effects of these novel variations were analysed on the ARSA protein structure (P15289) (Supplementary Figure S2). In the protein, the residues Gln at 139 and Pro at 180 are buried where as the residues Gly at 34 and Arg at 299 are exposed or solvent accessible (Supplementary Table S3).
In parallel to this approach, comparison of the hydrophobicity of normal and mutated protein sequences of variations Gly34Glu, Pro180Gln, and Arg299Trp showed a wide range of alterations in the overall hydrophobicity at the sites of variation, suggesting the protein structure instability (Supplementary Figure S3).
The deletion and insertion variations c.188_189 ins A, c.576 del C, and c.445_446 ins T induce frameshift variations and premature termination of the C-terminal domains (Supplementary Figure S4).
Pseudodeficiency alleles
In our study the following polymorphisms were identified in 21 patients: c.1049 A > G(p.Asn350Ser) [pseudodeficiency allele], c.1172 C > G(p.Thr391Ser), c.220 G > C(p.Ala74Pro), and p.Try190Cys in addition to the putative pathogenic variations.
Prenatal diagnosis
Prenatal diagnosis was done for 11 families based on proband diagnosis. Of the 11 fetuses, 3 were affected and 8 were unaffected (Table 1).
Discussion
Metachromatic leukodystrophy due to Arylsulfatase A deficiency should be suspected in any patient with progressive neurological deterioration, white matter changes on MRI of brain and a low level of arylsulfatase enzyme. However, decreased activity of arylsulfatase enzyme is not sufficient to diagnose MLD because of the presence of pseudodeficiency allele where the enzyme activity could be 5–20% of normal controls. Hence, for establishing the diagnosis of MLD molecular testing is essential. It is also essential in carrier status detection, especially in families with pseudodeficiency alleles.
In our study, majority of the patients had late infantile form of MLD (80%), which was comparable to the results from a previous study conducted in India [21] where 55% of the enrolled subjects (11/20) had late infantile type of disease.
In a variation update of ARSA and PSAP genes causing MLD by Cesani et al. [22], 200 distinct ARSA-MLD alleles were identified. A total of 66.5% of those variants were missense, 7.5% were nonsense, 6.5% were splice site variations and 12% were frameshift variations due to deletions, inversions, and duplications. In our study, the most common variations were missense (63.8%; 23/36), followed by frameshift variations (16.6%; 6/36), nonsense variations (8.3%; 3/36), and splicing variations (5.5%; 2/36).
The most common variant identified in our cohort was c.931 C > T (p.Arg311*), which is a known disease causing variant. This variant was identified in 14/122 alleles, which were checked, accounting for 11.4% of the disease causing variants, followed by c.459 + 1 G > A, which was seen in 9.8% of all alleles tested. The most common disease causing variation in patients with late infantile form from European ancestry is c.459 + 1 G > A [23]. In a previous study done in India, c.459 + 1 G > A was the most common disease causing variant identified [21], but the sample size of that study was limited. In adult onset or juvenile phenotype, the most common reported variants were the missense variants c.1277 C > T and c.536 T > G [22]. Only one patient in our cohort had the variant c.1277 C > T in a compound heterozygous state. He had adult onset disease and the other variant he had, was a known disease causing variant, c.1288 G > T. The variation profile identified in our study was distinct from the variation spectrum in the previous Indian study [22] and studies from other populations [24, 25, 26, 27]. In our study, 44% (16/36) were novel variants, implying the uniqueness of variation spectrum in Indian population. Identification of a common variation in Indian population would help in designing a low-cost targeted variation testing in patients with suspected MLD before resorting to costlier methods for molecular diagnosis.
Previous studies have shown that majority of ARSA-MLD alleles clustered around exon 2 and exon 5 [22]. In our study 25% of variants were in exon 5, but only 11% of variants were in exon 2. Hence, exon 5 can be considered as a “common mutated exon” for variations in Indian patients and thus this information can be used to screen for exon 5 in Indian patients followed by sequencing of all exons.
In our cohort, 76% of families were consanguineous. But in the 13 nonconsanguineous families, more than 50% had homozygous variations. This is not an uncommon phenomenon in India where inbreeding is prevalent and can result in autosomal recessive diseases in offspring and this has been documented in previous molecular studies from India [28, 29].
The c.459 + 1 G > A is a null allele and makes no protein and hence causes late infantile type of MLD. In our study, the three probands who had homozygous c.459 + 1 G > A had no detectable arylsulfatase enzyme and had a late infantile type of MLD. However, we were unable to derive any correlation between the phenotype, genotype, and biochemical parameters.
Treatment for MLD is mainly supportive even though hematopeotic stem cell transplantation (HSCT) has been suggested as a curative therapy especially in presymptomatic or early symptomatic juvenile or adult onset disease. HSCT is not recommended for patients with late infantile MLD. Majority of the patients in our cohort had late infantile MLD and were managed by supportive care. None of the patients in with juvenile or adult onset MLD in our cohort underwent hematopeotic stem cell transplant (HSCT).
To conclude, this study helped in identifying the variation spectrum of ARSA gene in Indian patients with MLD and aided in identifying the most common type of variation seen in Indian patients. This study stresses the need for molecular diagnosis in all patients with MLD, so that carrier testing, genetic counseling, and prenatal diagnosis can be provided for families with MLD.
References
Heinisch U, Zlotogora J, Kafert S, Gieselmann V. Multiple mutations are responsible for the high frequency of metachromatic leukodystrophy in a small geographic area. Am J Hum Genet. 1995;56:51–7.
Von Figura K, Gieselmann V, Jacken J. Metachromatic leukodystrophy. In: Scriver CR, Beaudet AL, Sly WS, Valle D, editors. The Metabolic and Molecular Bases of Inherited Disease. New York, NY: McGraw-Hill; 2001:3695–724.
Gort L, Coll MJ, Chabas A. Identification of 12 novel mutations and two new polymorphisms in the arylsulfatase A gene: haplotype and genotype–phenotype correlation studies in Spanish metachromatic leukodystrophypatients. Hum Mutat. 1999;14:240–8.
Hgmd.cf.ac.uk. (2018). [online] Available at: http://www.hgmd.cf.ac.uk/
Lowry OH, Rosebrough NJ, Farr AL, Randall RJ. Protein measurement with the folin phenol reagent. J Biol Chem. 1951;193:265–75.
Rice P, Longden I, Bleasby A. EMBOSS: The European Molecular Biology Open Software Suite. Trends Genet. 2000;16:276–7.
Den Dunnen JT, Antonarakis SE. Mutation nomenclature extensions and suggestions to describe complex mutations: a discussion. Hum Mutat. 2000;15:7–12.
Schwarz JM, Cooper DN, Schuelke M, Seelow D. MutationTaster2: mutation prediction for the deep-sequencing age. Nat Methods. 2014;11:361–2.
Acharya V, Nagarajaram HA. Hansa: an automated method for discriminating disease and neutral human nsSNPs. Human Mutat. 2011;33:332–7.
Kumar P, Henikoff S, Ng PC. Predicting the effects of coding non-synonymous variants on protein function using the SIFT algorithm. Nat Protoc. 2009;4:1073–81.
Choi, Y, Sims, G, Murphy, S, Miller, J, Chan, A. Predicting the functional effect of amino acid substitutions and indels. PLoS ONE. 2012;7:e46688.
Adzhubei I, Schmidt S, Peshkin L, Ramensky V, Gerasimova A, Bork P, et al. A method and server for predicting damaging missense mutations. Nat Methods. 2010;7:248–9.
ARSA - Arylsulfatase A precursor—Homo sapiens (Human)—ARSA gene & protein. Uniprot.org. http://www.uniprot.org/uniprot/P15289. Published 2018. Accessed August 10, 2018.
BLAST: Basic Local Alignment Search Tool. Blast.ncbi.nlm.nih.gov. http://blast.ncbi.nlm.nih.gov/Blast.cgi. Published 2018. Accessed August 10, 2018.
Chenna R, Sugawara H, Koike T, Lopez R, Gibson TJ, Higgins DG, et al. Multiple sequence alignment with the Clustal series of programs. Nucleic Acids Res. 2003;31:3497–3500.
Bond CS, Clements PR, Ashby SJ, Collyer CA, Harrop SJ, Hopwood JJ, et al. Structure of a human lysosomalsulfatase. Structure. 1997;5:277–89.
Mizuguchi K, Deane CM, Blundell TL, Johnson MS, Overington JP. JOY: protein sequence–structure representation and analysis. Bioinformatics. 1998;14:617–23.
DeLano WL. Unraveling hot spots in binding interfaces: progress and challenges. Curr Opin Struct Biol. 2002;12:14–20.
Guex N, Peitsch MC. SWISS-MODEL and the Swiss-PdbViewer: an environment for comparative protein modeling. Electrophoresis. 1997;18:2714–23.
ExPASy—PROSITE. Prosite.expasy.org. https://prosite.expasy.org. Published 2018. Accessed August 10, 2018.
Shukla P, Vasisht S, Srivastava R, Gupta N, Ghosh M, Kumar M, et al. Molecular and structural analysis of metachromatic leukodystrophy patients in Indian population. J Neurol Sci. 2011;301:38–45.
Cesani M, Lorioli L, Grossi S, Amico G, Fumagalli F, Spiga I, et al. Mutation update of ARSA and PSAP genes causing metachromatic leukodystrophy. Hum Mutat. 2015;37:16–27.
Polten A, Fluharty A, Fluharty C, Kappler J, von Figura K, Gieselmann V. Molecular basis of different forms of metachromatic leukodystrophy. New Engl J Med. 1991;324:18–22.
Chen L, Yan H, Cao B, Wu Y, Gu Q, Xiao J, et al. Identification of novel ARSA mutations in Chinese patients with metachromatic leukodystrophy. Int J Genom. 2018;2018:1–9.
Grossi S, Regis S, Rosano C, Corsolini F, Uziel G, Sessa M, et al. Molecular analysis of ARSA and PSAP genes in twenty-one Italian patients with metachromatic leukodystrophy: identification and functional characterization of 11 novelARSAalleles. Hum Mutat. 2008;29:E220–E230.
Ługowska A, Berger J, Tylki-Szymańska A, Löschl B, Molzer B, Zobel M, et al. Molecular and phenotypic characteristics of metachromatic leukodystrophy patients from Poland. Clin Genet. 2005;68:48–54.
Berger J, Löschl B, Bernheimer H, Lugowska A, Tylki‐Szymanska A, Gieselmann V, et al. Occurrence, distribution, and phenotype of arylsulfatase A mutations in patients with metachromatic leukodystrophy. Am J Med Genet. 1997;69:335–40.
Dalal A, Bhavani GS, Togarrati P, Bierhals T, Nandineni M, Danda S, et al. Analysis of theWISP3gene in Indian families with progressive pseudorheumatoid dysplasia. Am J Med Genet A. 2012;158A:2820–8.
Ranganath P, Matta D, Bhavani G, Wangnekar S, Jain J, Verma I, et al. Spectrum of mutations in the SMPD1 gene in Asian Indian patients with acid sphingomyelinase deficient Niemann–Pick disease. Am J Med Genet A. 2017;173:829–829.
Ługowska A, Wlodarski P, Płoski R, Mierzewska H, Dudzińska M, Matheisel A, et al. Molecular and clinical consequences of novel mutations in the arylsulfatase A gene. Clin Genet. 2009;75:57–64.
Berná L, Gieselmann V, Poupětová H, Hřebíček M, Elleder M, Ledvinová J. Novel mutations associated with metachromatic leukodystrophy: phenotype and expression studies in nine Czech and Slovak patients. Am J Med Genet A. 2004;129A:277–81.
Olkhovich N. Novel mutations in arylsulfatase A gene in three Ukrainian families with metachromatic leukodystrophy. Mol Genet Metab. 2003;80:360–3.
Harvey J, Nelson P, Carey W, Robertson E, Morris C. An arylsulfatase A (ARSA) missense mutation (T274M) causing late-infantile metachromatic leukodystrophy. Hum Mutat. 1993;2:261–7.
Hasegawa Y, Kawame H, Eto Y. Mutations in the Arylsulfatase A gene of Japanese patients with metachromatic leukodystrophy. DNA Cell Biol. 1993;12:493–8.
Draghia R, Letourneur F, Drugan C, Manicom J, Blanchot C, Kahn A, et al. Metachromatic leukodystrophy: identification of the first deletion in exon 1 and nine novel point mutations in the arylsulfatase A gene. Hum Mutat. 1997;9:234–42.
Rafi M. Disease-causing mutations in cis with the common arylsulfatase A pseudodeficiency allele compound the difficulties in accurately identifying patients and carriers of metachromatic leukodystrophy. Mol Genet Metab. 2003;79:83–90.
Barth M, Fensom A, Harris A. Identification of seven novel mutations associated with metachromatic leukodystrophy. Hum Mutat. 1995;6:170–6.
Biffi A, Cesani M, Fumagalli F, Carro U, Baldoli C, Canale S, et al. Metachromatic leukodystrophy—mutation analysis provides further evidence of genotype–phenotype correlation. Clin Genet. 2008;74:349–57.
Kreysing J, Bohne W, Bösenberg C, Marchesini S, Turpin JC, Baumann N, et al. High residual arylsulfatase A (ARSA) activity in a patient with late-infantile metachromatic leukodystrophy. Am J Hum Genet. 1993 Aug;53:339–46.
Gieselmann V, Zlotogora J, Harris A, Wenger DA, Morris CP. Molecular genetics of metachromatic leukodystrophy. Hum Mutat. 1994;4:233–42.
Acknowledgments
The authors are thankful to the patients and their families who participated in the study, for their kind cooperation. The authors are glad to acknowledge funding by Indian Council of Medical Research (54/5/2008-BMS) and the core support provided by the Centre for DNA Fingerprinting and Diagnostics, Hyderabad.
Funding
Indian Council of Medical Research (54/5/2008-BMS) and the core support provided by the Centre for DNA Fingerprinting and Diagnostics, Hyderabad.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Narayanan, D.L., Matta, D., Gupta, N. et al. Spectrum of ARSA variations in Asian Indian patients with Arylsulfatase A deficient metachromatic leukodystrophy. J Hum Genet 64, 323–331 (2019). https://doi.org/10.1038/s10038-019-0560-1
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s10038-019-0560-1
This article is cited by
-
The natural history and burden of illness of metachromatic leukodystrophy: a systematic literature review
European Journal of Medical Research (2024)
-
A systematic review on the birth prevalence of metachromatic leukodystrophy
Orphanet Journal of Rare Diseases (2024)
-
Identification and structural characterization of a pathogenic ARSA missense variant in two consanguineous families from Jammu and Kashmir (India) with late infantile metachromatic leukodystrophy
Molecular Biology Reports (2024)
-
Predicting disease severity in metachromatic leukodystrophy using protein activity and a patient phenotype matrix
Genome Biology (2023)
-
Chinese Cases of Metachromatic Leukodystrophy with the Novel Missense Mutations in ARSA Gene
Journal of Molecular Neuroscience (2021)