Abstract
Long-read sequencing technology is now capable of reading single-molecule DNA with an average read length of more than 10 kb, fully enabling the coverage of large structural variations (SVs). This advantage may pave the way for the detection of unprecedented SVs as well as repeat expansions. Pathogenic SVs of only known genes used to be selectively analyzed based on prior knowledge of target DNA sequence. The unbiased application of long-read whole-genome sequencing (WGS) for the detection of pathogenic SVs has just begun. Here, we apply PacBio SMRT sequencing in a Japanese family with benign adult familial myoclonus epilepsy (BAFME). Our SV selection of low-coverage WGS data (7×) narrowed down the candidates to only six SVs in a 7.16-Mb region of the BAFME1 locus and correctly determined an approximately 4.6-kb SAMD12 intronic repeat insertion, which is causal of BAFME1. These results indicate that long-read WGS is potentially useful for evaluating all of the known SVs in a genome and identifying new disease-causing SVs in combination with other genetic methods to resolve the genetic causes of currently unexplained diseases.
Similar content being viewed by others
Introduction
Short-read next-generation sequencing (NGS) is now widely applied in medical research and genetic testing for the detection of pathogenic single-nucleotide variants, and small insertions and deletions (indels). This short-read technology has achieved tremendous success in the discovery of many genes causative of human disease. However, many patients with conditions for which the genetic cause is unknown are still encountered, suggesting that certain types of pathogenic variation evade detection by the currently available short-read technology [1, 2]. Structural variations (SVs) spanning more than hundreds to tens of thousands of base pairs should be far beyond the reach of short reads (of ~150 bp in length). Many algorithms using depth of coverage, split reads, and paired reads have been developed to detect SVs using short-read WES and WGS data [3, 4]. However, the implementation of these algorithms to routine WES/WGS analysis for the detection of SVs is a challenge. Moreover, in these algorithms, it is difficult to accurately detect intermediate-size SVs (50 bp to several kilobases in size) and there is also the problem of numerous false-positive calls. As such, long-read sequencing technology is an attractive option for reliably detecting novel SVs [5, 6].
BAFME is an autosomal-dominant adult-onset neurological disease characterized by tremulous myoclonus (cortical tremor), and infrequent generalized epileptic seizure. The major electrophysiological findings are generalized epileptiform discharges and photosensitivity in electroencephalogram (EEG), and cortical reflex myoclonus (giant somatosensory evoked potentials, C-reflex, and spikes preceding myoclonus on EEG jerk-locked back-averaging) [7,8,9,10]. Four loci associated with BAFME have been mapped by linkage analysis [11]. Among them, BAFME1 at 8q24 has been documented in Japanese and Chinese families. Clinical anticipation of BAFME was also described in these two Asian populations [12, 13], suggesting that repeat expansion is associated with BAFME1. Recently, Ishiura et al. discovered the intronic pentanucleotide repeat expansions in the sterile alpha motif domain containing 12 gene (SAMD12) with an expanded repeat length in the range of 2.2–18.4 kb in Japanese families [14]. This new finding was also followed by Chinese BAFME1 families [15]. The currently available Pacbio SMRT sequencing using the Sequel system is now capable of reading >10-kb DNA [16]. Thus, we reasoned that it should have the potential to fully cover the SAMD12 repeat expansion and reliably prove the expanded repeat.
Here, we applied long-read WGS using PacBio together with a conventional method to detect the repeat expansion of SAMD12 in a family affected by BAFME. Even low-coverage long-read WGS may be useful to detect known and novel pathogenic SVs.
Materials and methods
Subjects
Five members of a single family suffered from BAFME (I-1, II-1, II-7, III-1, and III-2), which was inherited in an autosomal-dominant fashion (Fig. 1a). Patients showed cortical tremor and epilepsy with clinical anticipation. III-2 developed cortical tremor earlier than II-7 (Supplementary Table S1). IV-1, a 16-year-old girl, was asymptomatic at the time of study entry. III-4 and III-7 were not clinically evaluated. Four individuals (II-2, II-7, III-2, and III-6) participated in this study. II-2 is a nonconsanguineous (unrelated) spouse of II-1. She suffered from idiopathic generalized epilepsy but not BAFME. Written informed consent for inclusion in the study was obtained from all participants. This study was approved by the Institutional Review Board of Yokohama City University School of Medicine and University of Occupational and Environmental Health, Japan.
Pedigree of a family with pathogenic structural variation of SAMD12. a Pedigree of BAFME and segregation of SAMD12 variant. Black and white indicate affected and unaffected statuses with respect to the BAFME phenotype, respectively. Symbols with gray outlines represent participants subjected to genetic analysis. Asterisk indicates an individual whose genomic DNA was analyzed by long-read WGS using the PacBio Sequel system. b Validation of SAMD12 repeat expansion by Southern blotting. SacI-digested genomic DNA was run on a 1.6% agarose gel (w/v) in 1.0× TBE and probed with DIG-labelled SAMD12 probe. Control and unaffected individuals showed a signal corresponding to a size of 2239 bp. Arrow indicates the allele with expanded repeat
SMRTbell library preparation
Genomic DNA was extracted from peripheral blood leukocytes using QuickGene DNA whole blood kit (Kurabo) for three controls. Genomic DNA for the BAFME-affected family was extracted from peripheral blood leukocytes by standard phenol-chloroform DNA extraction. The size and integrity of genomic DNA were assessed by pulse-field agarose gel electrophoresis and the DNA concentration was measured by a Qubit fluorometer (Life Technologies). Seven micrograms of genomic DNA in a 150-µl volume was fragmented using g-TUBE (Covaris) by centrifugation at 1500 × g for 2 min twice. Recovered DNA was purified and concentrated using AMpure PB magnetic beads (Beckman Coulter).
SMRTbell Template Prep Kit 1.0 SPv3, Sequel Binding Kit 2.0, SMRTbell Clean Up Column v2 Kit, and MagBead Kit v2 (Pacific Biosciences) were used for SMRTbell library construction. SMRTbell template DNA/polymerase complex was used for sequencing on the PacBio Sequel system.
Five micrograms of fragmented DNA was subjected to SMRTbell library preparation in accordance with the manufacturer’s instructions (Procedure & Checklist >20 kb Template Preparation Using BluePippin Size-Selection System for Sequel Systems, Pacific Biosciences).
The resulting SMRTbell template was size-selected by BluePippin (Sage Science) and enriched for DNA fragments of >10 kb in size. Extraction conditions were set as follows: 0.75% DF Marker S1 high-pass 6–10 kb vs3 with a base-pair threshold start value (BP start) of 10,000. The size-selected library was purified by AMpure PB and then subjected to a DNA damage repair reaction. SMRTbell template DNA was annealed with Sequencing Primer v3 at 20 °C for 1 h. For polymerase binding, primer-annealed SMRTbell template DNA was incubated at 30 °C for 4 h with Sequel Polymerase 2.0. SMRTbell template DNA/polymerase complex was then purified using SMRTbell Clean Up Column. The purified complex was diluted to achieve an on-plate loading concentration of 20 pM, and then mixed and incubated with MagBead at 4 °C for 1 h to prepare MagBead-bound SMRTbell complex. This complex was loaded onto Sequel SMRT Cell 1M v2 and sequenced using Sequel Sequencing Kit 2.0. Data were collected for 6 h for each SMRT cell.
Data analysis using SMRT analysis module provided by SMRT link
Four SMRT cells were used for III-2, control 2, and control 3, which generated mean genome-wide coverage of 7×, 8×, and 6×, respectively. For control 1, mean coverage of 13× was obtained by using 10 SMRT cells. Raw statistics on the sequencing performance is described in Supplementary Table S2.
Secondary analysis using base-called data was performed on SMRT analysis v5.1.0. Structural variants were called using PBSV with the default settings, an application provided by SMRT analysis. PBSV (https://github.com/PacificBiosciences/pbsv) is a mapping-based structural variant caller for PacBio SMRT reads. PacBio reads are mapped to a reference human genome (GRCh37/hg19) using the long read mapper NGMLR. The CIGAR strings (Compact Idiosyncratic Gapped Alignment Report), which are a compressed representation of the aligned reads to the reference genome, are scanned to find deletions and insertions ≥50 bp. Nearby events are clustered and summarized into a SV call. Minimum SV length, minimum reads that support SV, and minimum percentage of variant reads were set to 50 bp, two reads, and 20%, respectively. PBSV called two types of SV, insertion and deletion. Each SV call was classified by the sequence pattern and assigned to one of the following categories: Alu, L1, SVA (SINE-VNTR-Alu class of retrotransposons), tandem repeat, and unannotated. When comparing the insertion calls among different individuals, regions up to 50 bp in length might be misaligned due to high sequence error rates of long-read sequencing; such inaccuracies were thus ignored and grouped into a single unit with the same/similar SVs. The resequencing application provided by SMRT analysis was used to summarize the mapping statistics in order to evaluate the data quality because PBSV does not generate such metrics (Supplementary Table S2).
Southern blot analysis
Six micrograms of genomic DNA was digested with SacI. Digested DNA was run on a 0.8% (Supplementary Fig. S1b) or 1.6% (Fig. 1b) agarose gel (w/v) in 1.0× TBE and transferred to a positively charged nylon membrane using capillary transfer. The DNA probe for studying intronic repeat expansion of SAMD12 was prepared as previously described [14]. Digoxigenin-labeled probe, DIG-(TGAAA)9 and DIG-(AGAAA)9 were purchased from Integrated DNA Technologies. The same membrane was stripped and reused for hybridization, according to the manufacturer’s instructions (Merck).
Dot plot analysis
Dot plots for the DNA sequence were created using Gepard [17]. By manual inspection, subread 3 and the 3′ end of subread 1 showed large discrepancies from the reference sequence. Subread 3 covered the repeat expansion at SAMD12, but the genomic position and sequence of the repeat were inconsistent with those of subreads 1 and 2. These discrepancies might have arisen from sequencing errors when using Sequel Sequencing Chemistry 2.0 and/or base-calling software. These errors might have occurred because of loss of fidelity of the polymerase or miscalibration of the detection system. Since subreads 1 and 2 are consistent, we excluded subread 3 and the 3′ end of subread 1 (subread 1: 11,840–13,651) from further analysis.
Results
We encountered a four-generation Japanese family affected by BAFME (Fig. 1a). The clinical manifestations of all of the participants in this study are summarized in Supplementary Table S1. Previous studies suggested that a major cause of BAFME in affected families in Japan is the presence of a common ancestor in which the SAMD12 variant and repeat expansion in intron 4 of SAMD12 occurred [14, 15]. Consistent with this, Southern blot analysis showed a heterozygous SV at the SAMD12 intronic repeat region in the affected individual (III-2). This SV cosegregated with the BAFME phenotype (Fig. 1b). It should be noted that III-2 and II-7 had similar repeat expansion sizes after the paternally germline passage (Supplementary Fig. S1).
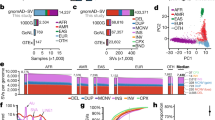
Based on Southern blot analysis, III-2 had a repeat length of approximately 4 kb, which could be fully covered by Pacbio SMRT sequencing in view of its current capacity. To characterize the SAMD12 variant with respect to repeat size and genomic position, genomic DNA from III-2 was analyzed by long-read WGS using the Pacbio Sequel system (mean genome-wide coverage of 7×). Genomic DNA from three control individuals was also sequenced for comparison (mean genome-wide coverage of 13×, 8×, and 6×) (Supplementary Table S2). WGS data were analyzed using PBSV, which is a structural variant caller for PacBio reads. A total of 9138 insertions and 6498 deletions were called in III-2 (Fig. 2a). Among them, 2420 insertions and 1086 deletions were found to be specific to III-2 (lacking in the controls), including six SVs (four insertion and two deletion calls) in the BAFME1-linked region (Fig. 2b, c). PBSV suggested the presence of a 4661-bp insertion at chr8: 119,379,051 (GRCh37/hg19), which was supported by two subreads (subreads 1 and 2) (Supplementary Table S3). This 4661-bp insertion was mapped between two repetitive sequences, AluSq2 (chr8: 119,378,770–119,379,051) and (TAAAA)n (chr8: 119,379,052–119,379,172) (Supplementary Figs. S2a and S3), which is consistent with previous reports [14, 15]. We created a dot plot of subreads 1 and 2 against the corresponding human reference genome sequence. The dot plot showed that the insertion was a novel sequence, rather than a tandem duplication (Fig. 2d). Then, we compared subreads 1 and 2 with each other. The created dot plot showed that these subreads were consistently similar in a region corresponding to the 4661-bp insertion of repetitive sequences (Supplementary Fig. S2b). In fact, 99.33% (4630 of 4661 bp) was masked by the RepeatMasker Open-4.0 program (http://www.repeatmasker.org). A total of 95.41% was found to be a low-complexity sequence, composed of GA or A-rich repeats (Supplementary Table S4).
Evaluation of long-read WGS. a Narrowing down SVs in III-2. PBSV called 9,138 insertions and 6,498 deletions. SVs fulfilling the following criteria were considered as candidates: (1) not present in three controls, (2) overlap with RefSeq genes, and (3) mapped around the BAFME1 locus. b III-2-specific insertion calls are plotted. The size of each insertion call (kb) is plotted against chromosomes (x-axis). PBSV classified each SV into one of five categories based on its sequence pattern, namely, Alu1 SINE repeat, L1 LINE repeat, SVA element, tandem repeat, and unannotated. c Visualizing the SV calls using the PBSV and SMRT sequencing results at the BAFME1 locus (chr8: 116,462,116–124,864,982). The sites of insertions and deletions are shown by vertical black and gray lines, respectively. Four insertion and two deletion calls remained after prioritization. PacBio subreads are shown at the bottom. Forward and reverse complement strands are shown by gray and open thick lines, respectively. Insertion calls are highlighted in black. d Dot plots of subreads 1 and 2 against the corresponding reference sequence. An unknown sequence of 4661 bp is inserted adjacent to the (TAAAA)n repeat sequence. The (TAAAA)n repeat is highlighted in gray
Discussion
Repetitive sequences are thought to be a major source of genomic instability [18, 19]. A total of 962,714 are described as simple tandem repeats in the RepeatMasker track of the UCSC genome browser. Such simple tandem repeats constitute polymorphic variation, but in some cases they become pathogenic and cause human genetic disorders. As gold standard methods for testing these pathogenic repeats, Southern blot analysis and/or repeat primed PCR are used. Recently, several algorithms using short-read NGS data were developed for SV detection [20,21,22,23]. However, these methods might require prior knowledge of the target repeat sequence and involve a computational burden when performing studies at the genome-wide level. It is highly anticipated that a long-read WGS approach can overcome these limitations.
In this study, we applied long-read WGS to detect SAMD12 intronic repeat expansion using the Pacbio Sequel system. An approximately 4.6-kb insertion at SAMD12 was correctly called by PBSV, a structural variant-calling application in SMRT Link v5.1.0. The size of the insertion in the PacBio data was 4661 bp, which was in good agreement with the size as estimated by the Southern blot analysis (Supplementary Fig. S1b). This indicates that this approach has the potential to increase the diagnostic yield of known repeat expansion diseases. However, the inserted sequence is suggested to be (TTTCT), rather than (TTTCA) as reported previously [14, 15]. Owing to the high sequence error rate of long-read sequencing technology (13–15% for PacBio SMRT sequencing), two subreads were insufficient to build a consensus on the actual sequence [16]. Indeed, Southern blot analysis using oligonucleotide probe (TGAAA)9 but not (AGAAA)9 identified the band corresponding to the mutated allele, indicating the presence of (TTTCA) repeat insertion in SAMD12 (Supplementary Fig. S4). Hence low-coverage Pacbio data can provide reliable size estimates for repeat length, but additional validation is required using higher-coverage Pacbio sequencing, perhaps with a targeted amplicon sequence or CRISPR-Cas9 targeted enrichment [24,25,26]. Moreover, accurate validation of the insertion sequence of each individual can also provide additional insights. For example, larger SAMD12 repeat expansion was suggested to be prone to occur through the maternal germline passage in BAFME [14]. In agreement with this, III-2 and II-7 had similar repeat expansion sizes when the SAMD12 variant was paternally transmitted (Supplementary Fig. S1). From this perspective, comparison of the repeat sequence with different repeat length within a pedigree might be valuable to provide insight into the mechanism behind repeat expansion and genotype–phenotype correlation including anticipation [27, 28].
The long-read WGS approach could be used to uncover pathogenic variants that remain undetected by the currently available short-read NGS approach. More than 15,000 intermediate-size SVs were called in III-2. As is the case with short-read NGS analysis, variant filtering is beneficial for prioritizing pathogenic SVs. In fact, we could effectively narrow down the candidate SVs for BAFME using low-coverage Pacbio data. A total of 15,636 SVs (9138 insertion and 6498 deletion calls) were initially called in III-2. Notably, 73.5% (6718 of 9138 insertion calls) and 83.3% (5412 of 6498 deletion calls) were present in at least one of the three control individuals, indicating the nonpathogenic nature of the majority of these SVs. We further focused on SVs overlapping with RefSeq genes (exons or introns) because known pathogenic repeat expansions were reported not only in the coding region, but also in the 5′ UTR, 3′ UTR, and introns of different genes associated with repeat expansion diseases [29]. After this filtering step, 1058 insertions and 482 deletions remained. Considering previous linkage mapping of BAFME, genes in four linked regions, 8q24 (BAFME1), 2p11.2–q11.2 (BAFME2), 5p15.31–p15.1 (BAFME3), and 3q26.32–q28 (BAFME4), should be prioritized [11, 30]. From this perspective, six (four insertions and two deletion), eight (four deletions and four insertions), and ten (eight insertions and two deletion) SVs remained at 8q24, 5p15.31–p15.1, and 3q26.32–q28, respectively. No SVs survived at 2p11.2–q11.2 (Supplementary Table S5). SAMD12 insertion (4.6 kb) is the only outlier with a size of more than 2 kb. Our results suggest that long-read WGS in combination with linkage mapping can be useful to identify novel pathogenic repeat expansions. Furthermore, pathogenicity might be suspected if certain SVs that are outliers in terms of their size are found in diseases exhibiting anticipation.
Currently, long-read sequencing technologies are expensive and have a high sequencing error rate, so they are not yet ready for clinical use. However, the ability to cover repetitive sequences should provide invaluable input for analyzing pathogenic SVs, even at low coverage. Such long read input will be aided by linkage analysis, multiple sample analysis within or without a pedigree, and comparison with the catalog of polymorphic SVs in human populations, as suggested in this study. The current PBSV version is only able to call two types of SV: insertions and deletions. Other types of SV including inversions, duplications, and even complex rearrangements should thus be the next targets of upcoming software [31,32,33]. We showed how to apply this technique to diseases for which the causative genetic factors remain unresolved and believe that it will enable the discovery of SVs for which the pathogenic effects have not been determined and even novel pathogenic SVs. In a revision process of this manuscript, Zheng et al. have also reported intronic repeat insertion in SAMD12 using long-read WGS, further proving useful of this challenging approach [34].
References
Huddleston J, Chaisson MJ, Meltz Steinberg K, Warren W, Hoekzema K, Gordon DS, et al. Discovery and genotyping of structural variation from long-read haploid genome sequence data. Genome Res. 2016;27:677–85.
Seo JS, Rhie A, Kim J, Lee S, Sohn MH, Kim CU, et al. De novo assembly and phasing of a Korean human genome. Nature. 2016;538:243–7.
Miyatake S, Koshimizu E, Fujita A, Fukai R, Imagawa E, Ohba C, et al. Detecting copy-number variations in whole-exome sequencing data using the eXome Hidden Markov Model: an 'exome-first' approach. J Hum Genet. 2015;60:175–82.
Pirooznia M, Goes FS, Zandi PP. Whole-genome CNV analysis: advances in computational approaches. Front Genet. 2015;6:138.
Merker JD, Wenger AM, Sneddon T, Grove M, Zappala Z, Fresard L, et al. Long-read genome sequencing identifies causal structural variation in a Mendelian disease. Genet Med. 2018;20:159–63.
Reiner J, Pisani L, Qiao W, Singh R, Yang Y, Shi L, et al. Cytogenomic identification and long-read single molecule real-time (SMRT) sequencing of a Bardet-Biedl Syndrome 9 (BBS9) deletion. NPJ Genom Med. 2018;3:3.
Ikeda A, Kakigi R, Funai N, Neshige R, Kuroda Y, Shibasaki H. Cortical tremor: a variant of cortical reflex myoclonus. Neurology. 1990;40:1561–5.
Inazuki G, Naito H, Ohama E, Kawase Y, Honma Y, Tokiguchi S, et al. [A clinical study and neuropathological findings of a familial disease with myoclonus and epilepsy—the nosological place of familial essential myoclonus and epilepsy (FEME)]. Seishin Shinkeigaku Zasshi—Psychiatr Et Neurol Jpn. 1990;92:1–21.
Mikami M, Yasuda T, Terao A, Nakamura M, Ueno S, Tanabe H, et al. Localization of a gene for benign adult familial myoclonic epilepsy to chromosome 8q23.3-q24.1. Am J Hum Genet. 1999;65:745–51.
van Rootselaar AF, van Schaik IN, van den Maagdenberg AM, Koelman JH, Callenbach PM, Tijssen MA. Familial cortical myoclonic tremor with epilepsy: a single syndromic classification for a group of pedigrees bearing common features. Mov Disord. 2005;20:665–73.
Cen ZD, Xie F, Xiao JF, Luo W. Rational search for genes in familial cortical myoclonic tremor with epilepsy, clues from recent advances. Seizure. 2016;34:83–89.
Hitomi T, Kondo T, Kobayashi K, Matsumoto R, Takahashi R, Ikeda A. Clinical anticipation in Japanese families of benign adult familial myoclonus epilepsy. Epilepsia. 2012;53:e33–6.
Cen Z, Huang C, Yin H, Ding X, Xie F, Lu X, et al. Clinical and neurophysiological features of familial cortical myoclonic tremor with epilepsy. Mov Disord. 2016;31:1704–10.
Ishiura H, Doi K, Mitsui J, Yoshimura J, Matsukawa MK, Fujiyama A, et al. Expansions of intronic TTTCA and TTTTA repeats in benign adult familial myoclonic epilepsy. Nat Genet. 2018;50:581–90.
Cen Z, Jiang Z, Chen Y, Zheng X, Xie F, Yang X, et al. Intronic pentanucleotide TTTCA repeat insertion in the SAMD12 gene causes familial cortical myoclonic tremor with epilepsy type 1. Brain. 2018;141:2280–8.
Ardui S, Ameur A, Vermeesch JR, Hestand MS. Single molecule real-time (SMRT) sequencing comes of age: applications and utilities for medical diagnostics. Nucleic Acids Res. 2018;46:2159–68.
Krumsiek J, Arnold R, Rattei T. Gepard: a rapid and sensitive tool for creating dotplots on genome scale. Bioinformatics. 2007;23:1026–8.
Carvalho CM, Lupski JR. Mechanisms underlying structural variant formation in genomic disorders. Nat Rev Genet. 2016;17:224–38.
Hannan AJ. Tandem repeats mediating genetic plasticity in health and disease. Nat Rev Genet. 2018;19:286–98.
Doi K, Monjo T, Hoang PH, Yoshimura J, Yurino H, Mitsui J, et al. Rapid detection of expanded short tandem repeats in personal genomics using hybrid sequencing. Bioinformatics. 2014;30:815–22.
Dolzhenko E, van Vugt J, Shaw RJ, Bekritsky MA, van Blitterswijk M, Narzisi G, et al. Detection of long repeat expansions from PCR-free whole-genome sequence data. Genome Res. 2017;27:1895–903.
Tang H, Kirkness EF, Lippert C, Biggs WH, Fabani M, Guzman E, et al. Profiling of short-tandem-repeat disease alleles in 12,632 human whole genomes. Am J Hum Genet. 2017;101:700–15.
Dashnow H, Lek M, Phipson B, Halman A, Sadedin S, Lonsdale A, et al. STRetch: detecting and discovering pathogenic short tandem repeat expansions. Genome Biol. 2018;19:121.
McFarland KN, Liu J, Landrian I, Godiska R, Shanker S, Yu F, et al. SMRT sequencing of long tandem nucleotide repeats in SCA10 reveals unique insight of repeat expansion structure. PLoS ONE 2015;10:e0135906.
Schule B, McFarland KN, Lee K, Tsai YC, Nguyen KD, Sun C, et al. Parkinson’s disease associated with pure ATXN10 repeat expansion. NPJ Park Dis. 2017;3:27.
Höijer I, Tsai YC, Clark TA, Kotturi P, Dahl N, Stattin EL, et al. Detailed analysis of HTT repeat elements in human blood using targeted amplification-free long-read sequencing. Hum Mutat. 2018;39:1262–72.
Mirkin SM. DNA structures, repeat expansions and human hereditary disorders. Curr Opin Struct Biol. 2006;16:351–8.
Landrian I, McFarland KN, Liu J, Mulligan CJ, Rasmussen A, Ashizawa T. Inheritance patterns of ATCCT repeat interruptions in spinocerebellar ataxia type 10 (SCA10) expansions. PLoS ONE 2017;12:e0175958.
Usdin K, House NC, Freudenreich CH. Repeat instability during DNA repair: Insights from model systems. Crit Rev Biochem Mol Biol. 2015;50:142–67.
Mori S, Nakamura M, Yasuda T, Ueno S, Kaneko S, Sano A. Remapping and mutation analysis of benign adult familial myoclonic epilepsy in a Japanese pedigree. J Hum Genet. 2011;56:742–7.
English AC, Salerno WJ, Reid JG. PBHoney: identifying genomic variants via long-read discordance and interrupted mapping. BMC Bioinform. 2014;15:180.
Fang L, Hu J, Wang D, Wang K. NextSV: a meta-caller for structural variants from low-coverage long-read sequencing data. BMC Bioinform. 2018;19:180.
Sedlazeck FJ, Rescheneder P, Smolka M, Fang H, Nattestad M, von Haeseler A, et al. Accurate detection of complex structural variations using single-molecule sequencing. Nat Methods. 2018;15:461–8.
Zeng S, Zhang MY, Wang XJ, Hu ZM, Li JC, Li N, et al. Long-read sequencing identified intronic repeat expansions in SAMD12 from Chinese pedigrees affected with familial cortical myoclonic tremor with epilepsy. Journal of medical genetics 2018. https://doi.org/10.1136/jmedgenet-2018-105484.
Acknowledgements
We would like to thank all of the subjects for participating in this study. We also thank N. Watanabe, T. Miyama, M. Sato, and K. Takabe for their technical assistance and A. Wenger (Pacific Biosciences) for helpful comments. We are also grateful to Edanz Group (www.edanzediting.com/ac) for editing a draft of this manuscript. This work was supported by AMED under grant numbers JP18ek0109280, JP18dm0107090, JP18ek0109301, JP18ek0109348, and JP18kk020500; by JSPS KAKENHI under grant numbers JP17K15630, JP17H01539, JP17K10080, and JP17K15630; by JST under the Creation of Innovation Centers for Advanced Interdisciplinary Research Areas Program in the Project for Developing Innovation Systems; the Ministry of Health, Labor, and Welfare; and Takeda Science Foundation.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Supplementary information
Rights and permissions
About this article
Cite this article
Mizuguchi, T., Toyota, T., Adachi, H. et al. Detecting a long insertion variant in SAMD12 by SMRT sequencing: implications of long-read whole-genome sequencing for repeat expansion diseases. J Hum Genet 64, 191–197 (2019). https://doi.org/10.1038/s10038-018-0551-7
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s10038-018-0551-7
This article is cited by
-
Sequencing and characterizing short tandem repeats in the human genome
Nature Reviews Genetics (2024)
-
DeepRepeat: direct quantification of short tandem repeats on signal data from nanopore sequencing
Genome Biology (2022)
-
A guide for the diagnosis of rare and undiagnosed disease: beyond the exome
Genome Medicine (2022)
-
An update on the neurological short tandem repeat expansion disorders and the emergence of long-read sequencing diagnostics
Acta Neuropathologica Communications (2021)
-
Founder effect of the TTTCA repeat insertions in SAMD12 causing BAFME1
European Journal of Human Genetics (2021)