Abstract
Cornelia de Lange syndrome (CdLS) is a rare neurodevelopmental syndrome for which mutations in five causative genes that encode (SMC1A, SMC3, RAD21) or regulate (NIPBL, HDAC8) the cohesin complex, account for ~70% of cases. Herein we report on four female Subjects who were found to carry novel intragenic deletions in HDAC8. In one case, the deletion was found in mosaic state and it was determined to be present in ~38% of blood lymphocytes and in nearly all cells of a buccal sample. All deletions, for which parental blood samples were available, were shown to have arisen de novo. X-chromosome inactivation studies demonstrated marked skewing, suggesting strong selection against the mutated HDAC8 allele. Based on an investigation of the deletion breakpoints, we hypothesize that microhomology-mediated replicative mechanisms may be implicated in the formation of some of these rearrangements. This study broadens the mutational spectrum of HDAC8, provides the first description of a causative HDAC8 somatic mutation and increases the knowledge on possible mutational mechanisms underlying copy number variations in HDAC8. Moreover our findings highlight the clinical utility of considering copy number analysis in HDAC8 as well as the analysis on DNA from more than one tissue as an indispensable part of the routine molecular diagnosis of individuals with CdLS or CdLS-overlapping features.
Similar content being viewed by others
Introduction
Cornelia de Lange syndrome (CdLS; OMIM# 122470,300590, 610759, 300882, and 614701) is a genetically heterogeneous disorder characterized by distinctive facial features, hirsutism, upper limb anomalies, growth retardation, and intellectual disabilities. Classic CdLS is associated with a severe phenotype with profound growth and neurodevelopmental delays, striking dysmorphic craniofacial features and reduction defects of the upper extremities ranging from complete absence of the forearms to oligodactyly [1]. Individuals with milder, atypical phenotype have less severe growth, cognitive and limb involvement, but often have facial features consistent with CdLS [2]. An estimated 60% of clinically well-defined CdLS cases harbor a de novo heterozygous mutation in the cohesin loader gene NIPBL [3]. About 10% of the patients carry a variant in the structural components of the chomatin-associated complex cohesin, SMC1A [4, 5], SMC3 [4], or RAD21 [6]. These patients present with a milder or less typical phenotype. A fifth CdLS gene, HDAC8, encodes a vertebrate SMC3 deacetylase that has roles in catalyzing the deacetylation of SMC3 as well as in recycling of cohesin for the next cell cycle [7].
Recent studies indicate that a significant proportion of CdLS individuals, in whom mutations in the five-known CdLS-associated genes were excluded by conventional Sanger sequencing performed on DNA isolated from blood, harbor somatic mutations in NIPBL in DNA derived from buccal mucosa [8,9,10]. Among the viable explanations are, the limited sensitivity of Sanger sequencing for detection of mosaicism and the loss of mutations in leukocytes due to reversion and leukocyte specific selection against mutant cells [8, 11]. Mosaicism has also been reported in SMC1A and SMC3 [8, 12]; however, somatic mosaicism for a mutation in HDAC8 resulting in a CdLS phenotype has not been previously reported.
To date, 41 different loss-of-function mutations in HDAC8 including six large deletions and one insertion have been reported [7, 13, 14], the majority of which were shown to have occurred de novo [13, 14].
In addition to typical CdLS features, HDAC8-patients can also exhibit some distinct features including hypertelorism, a broad or bulbous nasal tip, tooth anomalies, and mosaic patches of hyperpigmented skin [13, 14]. The phenotypic spectrum of HDAC8-related CdLS is consistent with HDAC8 localization on the X chromosome, with males being more severely affected than females. Females, who represent more than two-thirds of reported patients, display variable clinical severity that is heavily influenced by X-inactivation patterns. Most female patients demonstrate complete skewing towards full expression of the normal allele in blood, indicating a strong selection against the mutant allele [13, 14].
Several mechanisms have been proposed for the formation of disease-associated copy number variations (CNVs). Non-allelic homologous recombination (NAHR) is the main mechanism for the formation of recurrent genomic rearrangements by using low-copy repeats as a substrate for recombination [15]. Moreover, non-homologous end joining (NHEJ) and the DNA replication-based mechanisms of fork stalling and template switching (FoSTeS)/microhomology-mediated break-induced replication (MMBIR) [16] have been found to be play an important role in the generation of nonrecurrent and complex rearrangements in human diseases [17, 18]. While disease-causing CNVs have been reported in HDAC8, the mutational mechanism underlying these rearrangements has not been elucidated.
In this study, we describe the identification of four female Subjects with CdLS-overlapping phenotypes carrying novel intragenic deletions in HDAC8, including the first report of somatic mosaicism. Furthermore, we characterized the breakpoint junctions of the deletions to gain insight into the underlying molecular mechanisms of these rearrangements.
Materials and methods
Patients
Over the past 6 years, 126 samples were referred to our laboratory for HDAC8 deletion/duplication analysis. In all patients, previous testing performed using Sanger sequencing and exon-targeted oligonucleotide array comparative genomic hybridization (array-CGH), had excluded the presence of point mutations and copy number abnormalities in NIPBL, SMC1A, SMC3, and RAD21 as well as point mutations in HDAC8. Genomic DNA was isolated from blood leukocytes on the AutoGenFlex STAR robotic workstation (Autogen, Holliston, MA) following the manufacturer’s instructions.
Array comparative genomic hybridization
Deletion and duplication analysis of the HDAC8 gene was performed using a high resolution, custom-designed, exon-targeted 4 × 180 K array-CGH platform (Agilent Technologies, Santa Clara, CA). A total of 1811 probes spanned the HDAC8 gene, with an average resolution of ~1 probe/20 bp across the coding regions. Genomic DNA samples of the patients and gender-matched control were processed and co-hybridized onto microarray slides according to the manufacturer recommended procedures (Agilent Technologies). Microarray images were scanned at 2 µ resolution and the data was extracted using ImaGene (9.0) and analyzed using the Nexus software (7.1) (BioDiscovery, Hawthorne, CA). The genomic copy number was defined by analysis of the normalized log2 (Cy5/Cy3) ratio average of the CGH signal. Regions that reached a threshold of at least −0.32 were considered suspicious for copy number losses consistent with deletions.
Quantitative real-time PCR
Relative quantification of HDAC8 exon 3 in the DNA isolated from blood and buccal sample of Subject 4 was performed by quantitative real-time PCR with primers HDAC8 ex3 F (5′-TAAGCCTAAAGTGGCCTCCA-3′) and HDAC8 ex3 R (5′-CATATTCTATGGAGTCCGGATG’-3′). Analysis was performed using 20 ng of genomic DNA. Reactions were done in triplicate, wherein each target region was co-amplified with an internal control (PMP22: NM_000304.2) using the SYBR-Green detection chemistry and the ABI PRISM 7500 sequence detection system (Applied Biosystems, Grand Island, NY). Relative quantification of the target gene was determined by the comparative threshold cycle method (ΔΔCt) [19]. DNA samples from a male and female control with normal HDAC8 copy number were used to estimate the approximate proportion of mutant cells in the blood and buccal samples of Subject 4.
HDAC8 breakpoint junction sequence analysis
Breakpoint sequence analysis of the HDAC8 deletions was performed by PCR primer walking using HotStar Taq polymerase (Qiagen, Germantown, MD). PCR primers were designed from the HDAC8 reference sequence (NM_018486.2) across each deleted region derived from the array-CGH results assuming the most likely rearrangement (Supplementary Table 1). Sequencing products were compared to the HDAC8 reference sequence using Mutation Surveyor software version 3.01 (SoftGenetics, State College, PA) and breakpoint sequences were aligned manually or using MultAlin (http://multalin.toulouse.inra.fr/multalin/) [20].
X-inactivation analysis
X-inactivation (XCI) analysis was performed using a well-characterized assay that measures the methylation status in the 5′UTR of the Androgen Receptor gene of the inactivated X chromosome [21]. X-inactivation patterns were determined by comparing the ratios of amplified DNA products for each allele before versus after digestion. Amplification bias for the smaller alleles is accounted for using an empirically determined correction factor. The following XCI thresholds were used for interpretation: below 80% is considered random X-inactivation, 80–90% is considered moderately skewed, and above 90% is considered highly skewed.
Results
Intragenic deletions in HDAC8 were identified in four female Subjects (Table 1). Consent for photographs was obtained for Subjects 1 and 2 (Fig. 1). All Subjects have clinical features similar to those observed in typical CdLS, including non-specific features such short stature (4/4), microcephaly (4/4), poor weight gain (2/4), and more typical CdLS craniofacial features including synophrys (4/4), wide nasal bridge (2/4), smooth philtrum (2/4), and posteriorly rotated ears (3/4). Other findings suggestive of CdLS include cutis marmorata (1/4), hirsutism (1/4), small hands and feet (3/4), and brachydactyly (4/4). A bulbous nasal tip, a feature previously reported to be more typical in CdLS patients with HDAC8 mutations [13], was reported to be present in Subjects 1 and 2 (Fig. 1). All Subjects exhibited some degree of speech impairment, ranging from verbal apraxia to absent speech. Limb anomalies were not detected in any of the Subjects. In addition, several congenital defects were reported in our patients, including atrial septal defects, cleft palate, and diaphragmatic hernia.
Clinical findings in Subject 1 at age 6 years and Subject 2 at age 8 years. a, b Facial features of Subject 1, showing synophrys, broad nasal bridge, lowset ears, widely spaced teeth. c 2–3 toe syndacytly and d brachydactyly in Subject 1. e Facial features of Subject 2, showing mild synophrys, wide nasal bridge, bulbous nasal tip. f Tapered fingers and brachydactyly in Subject 2 (Color figure online)
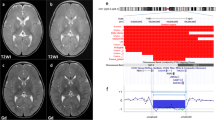
All deletions are novel, range in size from ~2.4 to 70 kb and encompass up to three exons (Fig. 2). One deletion encompassing exons 3 and 4 was found to be in an apparently mosaic state in blood (Subject 4) as the array-CGH data showed a low-level reduction in the Cy5/Cy3 fluorescence log2 ratio (average probes’ log2 ratio of approximately −0.3) of oligonucleotide probes interrogating these exons. Repeat analysis on DNA obtained from a buccal swab from this individual confirmed the presence of the deletion in a larger proportion of cells (average probes’ log2 ratio of approximately −0.5). Quantitative PCR analysis indicated that, compared to a normal female control, the relative copy number of HDAC8 exon 3 was 81% in the blood and 48% in the buccal sample (Fig. 3). This indicates that ~38% of blood lymphocytes contain a deletion of one allele of HDAC8 and nearly all of the cells in the buccal sample contain the deletion.
Array-CGH results for each Subject. HDAC8 deletions are highlighted in red and evidenced by the reduced log2 fluorescence ratios. The approximate positions of HDAC8 exons 1 to 7 are indicated as blue vertical bars above the array-CGH chart. The x and y axes represent genomic coordinates and the log2 hybridization signals, respectively (Color figure online)
The deletions occurred de novo in all cases for which parental testing was available (3 out of 4). X-inactivation studies on DNA from blood revealed significant skewing in Subjects 1 to 3. Subject 4, who carried the mosaic deletion, showed a random X-inactivation pattern in blood and moderate skewing in buccal cells (Table 1).
Analysis of the breakpoint regions revealed that for all four breakpoints the distal or proximal ends of the deletions harbored repetitive sequences, including short or long interspersed repeat elements. In addition, breakpoints from each of these individuals showed two to three base-pairs of shared microhomology between proximal and distal reference sequences (Fig. 4). The proximal breakpoint of Subject 1, which maps to a long interspersed repeat element (LINE; L1PB1), has a three base-pair microhomology (AAC) at the breakpoint junction (Fig. 4a). The deletion in Subject 2 lies within a short interspersed repetitive element (SINE, AluSz6) at the proximal breakpoint and has a deletion-insertion of three base-pairs (CAC) and a two base-pair microhomology (CT) at the breakpoint junction (Fig. 4b). The proximal breakpoint for Subject 3 lies within a long interspersed repeat element (LINE; LIPA3) and shows a two base-pair microhomology (GT) at the breakpoint junction (Fig. 4c). The distal deletion breakpoint of Subject 4 maps to a short interspersed repeat element (SINE; AluJr) and has a two base-pair microhomology (GC) at the breakpoint junction (Fig. 4d).
Breakpoint sequence analysis of the HDAC8 deletions. DNA sequences were aligned to the normal wild-type proximal and distal sequences. Sequence homology to the normal proximal and distal wild-type sequences are shown in green and red. Boxed sequences (blue) indicate regions of microhomology and dash red lines reveal the breakpoint junctions (Color figure online)
Discussion
We describe the identification of four novel intragenic deletions in the HDAC8 gene, uncovering a genetic etiology in ~3% of individuals in this cohort. All Subjects presented with clinical features consistent with CdLS/atypical CdLS (Table 1, Fig. 1). The deletions range in size from 2.4 to 70 kb, encompass up to three exons and appear to be novel to this study supporting the broad allelic heterogeneity of HDAC8 mutations in CdLS (Fig. 2).
To date, six gross deletions and one insertion in HDAC8 have been reported [13], however no insights have been provided into the developmental mechanisms of these rearrangements. This prompted us to characterize deletion breakpoints molecularly and examine the breakpoint regions. Interestingly, we uncovered the presence of two to three base-pairs of microhomology at the breakpoints in all cases (Fig. 4). Furthermore, analysis of the sequences surrounding the breakpoints revealed that in all cases the rearrangements occurred in, or in the proximity of, repetitive elements that are known to increase the genomic instability in certain regions [22]. The features observed at the breakpoint junctions for these cases suggest that a replication-based repair mechanism such as FoSTeS/MMBIR [16, 17] could underlie a proportion of these rearrangements in HDAC8, similar to what has been already proposed for intragenic copy number aberrations in the NIPBL gene [23]. The presence of microhomology at the breakpoints in all cases argues against a NHEJ mechanism. Furthermore, the insertion-deletion of the three base-pair CAC sequence in Subject 2 indicates that at least one additional template-switch occurred to acquire this unique sequence, and strongly supports a FoSTeS/MMBIR mechanism.
Somatic mosaicism in NIPBL has been described in multiple patients with CdLS. In a proportion of patients, the mutation is identifiable only in buccal samples and is undetectable in blood [8]. Mosaicism has also been reported in SMC1A and SMC3 [8, 12]. To our knowledge, the mosaic case reported herein is the first described patient with a mosaic HDAC8 mutation associated with clinical features of CdLS. Only one other case of somatic mosaicism for an HDAC8 variant has been reported in the apparently unaffected mother of two patients with HDAC8-related CdLS [14]. Quantitative PCR analysis indicated that in this Subject ~38% of blood lymphocytes and nearly all buccal cells contained the deletion. A comparison of the provided clinical features (Table 1) did not seem to suggest a significant difference in the severity of the phenotype of Subject 4 versus other patients possibly suggesting that the mosaicism cannot mitigate the phenotypic impact of a damaging mutation. In addition, because testing was only performed in peripheral blood and buccal cells, the results may not accurately reflect the mutation load in other tissues.
The HDAC8 mosaic deletion reported here again highlights the fact that the prevalence of somatic mosaicism in the CdLS-associated genes could be underestimated, as many of these cases may remain undetected. Given that mosaicism in other CdLS genes is not infrequent, it is likely that a proportion of patients included in this study have somatic mosaicism in any of the five-known CdLS genes but Sanger sequencing and array-CGH failed to identify a mutation in DNA isolated from blood. The use of more sensitive techniques, such as targeted analysis of the five CdLS genes by deep next generation sequencing of DNA isolated from blood and/or buccal cells is likely to be available in clinical diagnostic settings in the near future.
All Subjects exhibited skewed X-inactivation, consistent with previous reports [13, 14]. Subjects 1, 2 and 3 exhibited highly skewed X-inactivation in blood. Subject 4, who is mosaic for a deletion of exons 3 and 4, exhibited an X-inactivation ratio consistent with random inactivation in her blood sample and moderately skewed X-inactivation in buccal cells. While this patient’s deletion was detectable in blood, the presence of the deletion was more clearly demonstrated in buccal cells, suggesting a higher proportion of cells harboring the mutated allele in this cell lineage. Although X-inactivation studies are not able to distinguish which allele is preferentially inactivated, the observation that the sample with a higher proportion of deleted alleles has increased X-inactivation skewing suggests that the X chromosome with the HDAC8 deletion is likely to be preferentially inactivated.
In conclusion, this study identifies additional underlying causes of CdLS, describes the first instance of a somatic HDAC8 mutation in an individual with CdLS features and provides insight into the molecular bases of HDAC8 deletions. The frequency of the deletions identified in this cohort highlights the importance of utilizing multiple techniques including copy number analysis of HDAC8 in individuals with a suspected diagnosis of CdLS who are negative for mutations in other CdLS-associated genes. Finally, the identification of a somatic mutation in HDAC8 suggests that the analysis of DNA derived from buccal cells should be considered to investigate whether a patient may have a somatic mosaicism if lymphocyte analysis has failed to identify a mutation.
References
Kline AD, Krantz ID, Sommer A, Kliewer M, Jackson LG, FitzPatrick DR, et al. Cornelia de Lange syndrome: clinical review, diagnostic and scoring systems, and anticipatory guidance. Am J Med Genet A. 2007;143A:1287–96.
Rohatgi S, Clark D, Kline AD, Jackson LG, Pie J, Siu V, et al. Facial diagnosis of mild and variant CdLS: Insights from a dysmorphologist survey. Am J Med Genet A. 2010;152A:1641–53.
Krantz ID, McCallum J, DeScipio C, Kaur M, Gillis LA, Yaeger D, et al. Cornelia de Lange syndrome is caused by mutations in NIPBL, the human homolog of Drosophila melanogaster Nipped-B. Nat Genet. 2004;36:631–35.
Deardorff MA, Kaur M, Yaeger D, Rampuria A, Korolev S, Pie J, et al. Mutations in cohesin complex members SMC3 and SMC1A cause a mild variant of cornelia de Lange syndrome with predominant mental retardation. Am J Hum Genet. 2007;80:485–94.
Musio A, Selicorni A, Focarelli ML, Gervasini C, Milani D, Russo S, et al. X-linked Cornelia de Lange syndrome owing to SMC1L1 mutations. Nat Genet. 2006;38:528–30.
Deardorff MA, Wilde JJ, Albrecht M, Dickinson E, Tennstedt S, Braunholz D, et al. RAD21 mutations cause a human cohesinopathy. Am J Hum Genet. 2012;90:1014–27.
Deardorff MA, Bando M, Nakato R, Watrin E, Itoh T, Minamino M, et al. HDAC8 mutations in Cornelia de Lange syndrome affect the cohesin acetylation cycle. Nature. 2012;489:313–17.
Huisman SA, Redeker EJ, Maas SM, Mannens MM, Hennekam RC. High rate of mosaicism in individuals with Cornelia de Lange syndrome. J Med Genet. 2013;50:339–44.
Nizon M, Henry M, Michot C, Baumann C, Bazin A, Bessieres B, et al. A series of 38 novel germline and somatic mutations of NIPBL in Cornelia de Lange syndrome. Clin Genet. 2016;89:584–89.
Baquero-Montoya C, Gil-Rodriguez MC, Braunholz D, Teresa-Rodrigo ME, Obieglo C, Gener B, et al. Somatic mosaicism in a Cornelia de Lange syndrome patient with NIPBL mutation identified by different next generation sequencing approaches. Clin Genet. 2014;86:595–97.
Rohlin A, Wernersson J, Engwall Y, Wiklund L, Bjork J, Nordling M. Parallel sequencing used in detection of mosaic mutations: comparison with four diagnostic DNA screening techniques. Hum Mutat. 2009;30:1012–20.
Ansari M, Poke G, Ferry Q, Williamson K, Aldridge R, Meynert AM, et al. Genetic heterogeneity in Cornelia de Lange syndrome (CdLS) and CdLS-like phenotypes with observed and predicted levels of mosaicism. J Med Genet. 2014;51:659–68.
Kaiser FJ, Ansari M, Braunholz D, Concepción Gil-Rodríguez M, Decroos C, Wilde JJ, et al. Loss-of-function HDAC8 mutations cause a phenotypic spectrum of Cornelia de Lange syndrome-like features, ocular hypertelorism, large fontanelle and X-linked inheritance. Hum Mol Genet. 2014;23:2888–900.
Parenti I, Gervasini C, Pozojevic J, Wendt KS, Watrin E, Azzollini J, et al. Expanding the clinical spectrum of the ‘HDAC8-phenotype’ - implications for molecular diagnostics, counseling and risk prediction. Clin Genet. 2016;89:564–73.
Inoue K, Lupski JR. Molecular mechanisms for genomic disorders. Annu Rev Genomics Hum Genet. 2002;3:199–42.
Zhang F, Gu W, Hurles ME, Lupski JR. Copy number variation in human health, disease, and evolution. Annu Rev Genomics Hum Genet. 2009;10:451–81.
Lee JA, Carvalho CM, Lupski JRA. DNA replication mechanism for generating nonrecurrent rearrangements associated with genomic disorders. Cell. 2007;131:1235–47.
Hastings PJ, Ira G, Lupski JR. A microhomology-mediated break-induced replication model for the origin of human copy number variation. PLoS Genet. 2009;5:e1000327.
Livak KJ, Schmittgen TD. Analysis of relative gene expression data using real-time quantitative PCR and the 2(-Delta Delta C(T)) method. Methods. 2001;25:402–8.
Corpet F. Multiple sequence alignment with hierarchical clustering. Nucleic Acids Res. 1988;16:10881–90.
Allen RC, Zoghbi HY, Moseley AB, Rosenblatt HM, Belmont JW. Methylation of HpaII and HhaI sites near the polymorphic CAG repeat in the human androgen-receptor gene correlates with X chromosome inactivation. Am J Hum Genet. 1992;51:1229–39.
Argueso JL, Westmoreland J, Mieczkowski PA, Gawel M, Petes TD, Resnick MA. Double-strand breaks associated with repetitive DNA can reshape the genome. Proc Natl Acad Sci USA. 2008;105:11845–50.
Pehlivan D, Hullings M, Carvalho CM, Gonzaga-Jauregui CG, Loy E, Jackson LG, et al NIPBL rearrangements in Cornelia de Lange syndrome: evidence for replicative mechanism and genotype-phenotype correlation. Genet Med. 2012;14:313–22.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Electronic supplementary material
Rights and permissions
About this article
Cite this article
Helgeson, M., Keller-Ramey, J., Knight Johnson, A. et al. Molecular characterization of HDAC8 deletions in individuals with atypical Cornelia de Lange syndrome. J Hum Genet 63, 349–356 (2018). https://doi.org/10.1038/s10038-017-0387-6
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s10038-017-0387-6
This article is cited by
-
The emerging role of chromatin remodelers in neurodevelopmental disorders: a developmental perspective
Cellular and Molecular Life Sciences (2021)
-
Understanding the Landscape of X-linked Variants Causing Intellectual Disability in Females Through Extreme X Chromosome Inactivation Skewing
Molecular Neurobiology (2020)