Abstract
Chromosome transfer technology, including chromosome modification, enables the introduction of Mb-sized or multiple genes to desired cells or animals. This technology has allowed innovative developments to be made for models of human disease and humanized animals, including Down syndrome model mice and humanized transchromosomic (Tc) immunoglobulin mice. Genome editing techniques are developing rapidly, and permit modifications such as gene knockout and knockin to be performed in various cell lines and animals. This review summarizes chromosome transfer-related technologies and the combined technologies of chromosome transfer and genome editing mainly for the production of cell/animal models of human disease and humanized animal models. Specifically, these include: (1) chromosome modification with genome editing in Chinese hamster ovary cells and mouse A9 cells for efficient transfer to desired cell types; (2) single-nucleotide polymorphism modification in humanized Tc mice with genome editing; and (3) generation of a disease model of Down syndrome-associated hematopoiesis abnormalities by the transfer of human chromosome 21 to normal human embryonic stem cells and the induction of mutation(s) in the endogenous gene(s) with genome editing. These combinations of chromosome transfer and genome editing open up new avenues for drug development and therapy as well as for basic research.
Similar content being viewed by others
Chromosome transfer technology
Chromosomes can be transferred from donor cells to recipient cells using several techniques of chromosome transfer. Different research groups are developing chromosome transfer techniques involving microcell-mediated chromosome transfer (MMCT) [1, 2], and the transfection of chromosomes with liposome carriers [3].
Chinese hamster ovary (CHO), A9, and DT40 cells are mainly used as donor cells in MMCT (Fig. 1a) because they are capable of micronucleation with elongated inhibition of microtubules [4] in response to colcemid [1] or TN16 and griseofulvin treatment [5]. Microcells are isolated by the centrifugation of cells forming micronuclei. Those microcells contains one or a few chromosomes and can be fused with recipient cells using polyethylene glycol (PEG) [1]. Because the efficiency of PEG-MMCT is low, virus protein-mediated MMCT such as MV-MMCT and Retro-MMCT was developed to increase the efficiency of chromosome transfer [6, 7].
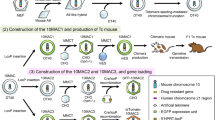
Chromosome transfer technology and the top-down HAC/MAC. a Chromosome transfer technology: MMCT. Donor CHO, A9, and DT40 cells are exposed to a microtubule inhibitor leading to micronucleation. Microcells are then purified and fused to target cells. Microcell hybrids, target cells containing the desired chromosome, can be obtained. b Construction of the top-down HAC/MAC. HACs and MACs are stable in humans and mice, respectively
Liposome carriers can also be used to transfer chromosomes. In this process, naked chromosomes are isolated from cells that have been induced to undergo mitotic arrest with colcemid treatment. The chromosomes collected by sucrose cushion centrifugation can be transfected to recipient cells using liposome carriers [3]. However, the efficiency of this method is low, so it is less commonly used for chromosome transfer than MMCT.
Applications of MMCT and the hurdles of conventional genome engineering technology
MMCT has been applied to various fields of molecular biology and biotechnology, e.g., chromosome mapping, functional assays associated with chromosome structure [2, 8], and the generation of transchromosomic (Tc) animals [2]. For example, an A9 human monochromosome library was used to transfer a target human chromosome via MMCT [9]. This library includes all human autosomes and X chromosome, which were deposited in the Japanese Collection of Research Bioresources Cell Bank [10].
Chromosome mapping is based on a complementation study [11, 12], and involves the introduction of a human chromosome via MMCT to rescue the phenotype of the model cells, such as cancer cell lines or disease model cells. This technique has been used for the identification of genes related to tumor suppression [13, 14], genomic imprinting [15, 16], DNA repair [17,18,19], metastasis and genomic instability [20], telomerase regulation [21,22,23], mitochondrial disorders [24], and lysosomal storage diseases [25]. MMCT has also been applied to investigate chromosomal functions such as kinetochore assembly, telomere function, and high-order chromosome architecture [8, 26, 27]. Tc animals harboring an extra copy of a human chromosome have also been generated via MMCT [28], including a Down syndrome (DS) mouse model [29, 30] and humanized immunoglobulin mice [31] and cattle models [32]. MMCT technologies and their applications are described in more detail in our previous review [2].
Although large DNA can be transferred using chromosome transfer technologies, difficulties remain using conventional genome engineering technologies for: (1) the efficient production of knockout (KO) animals for humanization and (2) the multiple and/or sophisticated genome manipulation required to improve existing human disease- and humanized animal-models, and to establish functional cell-based models. Therefore, the development of more efficient and convenient genome manipulation technology is urgently needed.
Genome editing techniques involving artificial nucleases mainly use zinc-finger nucleases [33], transcription activator-like effector nucleases [34], and clustered regularly interspaced short palindromic repeats/CRISPR-associated protein 9 (CRISPR/Cas9) [35]. These genome editing technologies allow KO or knock in cultures cells as well as animals to be produced. Details of the genome editing technologies are described in other reviews [36, 37]. Here, we describe the development of previously reported chromosome transfer-related technologies, and a new generation of chromosome transfer technology combined with genome editing techniques.
Combination of chromosome engineering and chromosome transfer
Chromosome modification in homologous recombination-proficient chicken DT40 cells
Two decades ago, chromosome modification relied on the occurrence of random events. Tc mice expressing the human immunoglobulin gene and genes on human chromosome 21 were generated with human chromosome fragments randomly obtained via MMCT [31] and X-ray irradiation-mediated MMCT [38], respectively. To perform required modifications without depending on random events, the homologous recombination (HR)-proficient chicken DT40 cell line, which has the capacity to undergo MMCT, was used for targeted gene insertion [15] and telomere-associated chromosome truncation to induce Mb-sized chromosomal deletions [39]. Such targeted chromosome modifications enable fine design of the chromosome and advanced applications to be made, e.g., the construction of mammalian artificial chromosome vectors including human artificial chromosomes (HACs) and mouse artificial chromosomes (MACs) (Fig. 1b), which can be used to generate humanized animals containing desired Mb-sized DNA.
Chromosome modification via genome editing in CHO and A9 cells
Although chromosome modification using HR-proficient chicken DT40 cells has enabled various unique developments to be made, it first requires the transfer of a targeted human chromosome to DT40 cells and then re-transfer of the modified chromosome to CHO cells before transferring it to the desired cell type (Fig. 2a). Additionally, the donor cell type determines the MMCT efficiency, which is low when using DT40 cells, except for transfer to recipient CHO cells. Moreover, repeated MMCT is labor-intensive and time-consuming (Fig. 2a). Chromosome modification using CHO or A9 cells, but not DT40 cells, can avoid these problems, because repeated MMCT is not required. CRISPR/Cas9-mediated HR and telomere-associated chromosome truncation have recently been achieved in CHO and A9 cells, and the modified chromosomes were shown to be transferrable (Fig. 2b) [40]. Although the HR efficiency in CHO and A9 cells is lower than that of DT40 cells and chromosome modification requires negative selection with suicide genes or fluorescent markers [41] to eliminate cells harboring unexpected insertions of the modification plasmid vector, the CHO/A9 cell method has the advantage that the modified chromosome can be directly transferred to desired cells.
Chromosome modification: gene insertion and telomere-associated chromosome truncation using CRISPR/Cas9 in CHO and A9 cells. a Diagram of conventional chromosome modification and the MMCT strategy. Three MMCT experiments are necessary to transfer the targeted chromosome to DT40 cells, re-transfer it to limited cell lines including CHO cells, and transfer it again to the targeted cells This multistep of chromosome modification and MMCT is a detour. b Diagram of a new chromosome modification and MMCT strategy. The CRISPR/Cas9-mediated chromosome modification can be completed in CHO and A9 cells without the need for homologous recombination in DT40 cells, so the targeted chromosome can be directly transferred from CHO and A9 cells to the targeted cells
On the other hand, genome editing technologies are associated with two potential problems caused by the off-target effect [42]. The first occurs on the target chromosome although CHO and A9 cells only contain a single targeted human chromosome. It is therefore important to ensure that an off-target sequence is located on the targeted chromosome. The second occurs on the host chromosome of CHO or A9 cells. In this case, it is not necessary to identify an off-target effect because the modified chromosome will be transferred to other cells from the host cells. Thus, targeted cells transferred the modified chromosome avoid the risk of off-target effects by CRISPR/Cas9. This is a particularly valuable advantage of chromosome modification via genome editing in CHO and A9 cells than conventional genome editing in targeted cells without MMCT.
DT40-mediated chromosome modification techniques require ~6 months for three MMCT procedures to be completed, including 1 month for MMCT and cell cloning, and another month for PCR and chromosome analyses including fluorescence in situ hybridization analysis. In contrast, CHO- or A9-mediated chromosome modification techniques can be completed more quickly (~2 months), so have been able to reduce the start-up hurdles for chromosome engineering, including the transfer of Mb-sized chromosome loci.
HAC/MAC technologies
Characteristics of HACs/MACs
HACs have mainly been constructed in two ways, the bottom-up or top-down method. The bottom-up approach generates de novo HAC formation by component assembly in host cells [43,44,45]. One such HAC vector (tet-O HAC) with an artificial centromere sequence is conditionally removable from host cells, so is particularly useful for gene function analysis [46]. The top-down approach generates HACs from a native chromosome by chromosome engineering technology, including gene-targeting and telomere-associated chromosome truncation in DT40 cells [47,48,49] (Fig. 1b). It is also possible to construct an artificial chromosome from any mammalian chromosome or cell type using the genome editing techniques described above. Most bottom-up HACs are circular and their construction relies on spontaneous assembly. In contrast, all top-down HACs are linear, precisely engineered, and stably maintained in host cells. The bi-HAC system utilizing both top-down and bottom-up HACs, and benefitting from the advantages of both, was reported previously [50].
Regarding Tc animal production, the generation of Tc mice carrying HACs with gene(s) of interest [51,52,53] has been achieved by using MMCT to establish mouse embryonic stem (ES) cells containing the HACs. If germline-transmissible chimeric animals cannot be developed using ES cells, cloning technologies can instead be applied to generate Tc animals via MMCT to fibroblasts. This technology was used to generate Tc calves producing human immunoglobulin [32].
Although HACs are stable in human cell lines and some cells and tissues of other species, they have shown variable retention rates in mouse tissues, being especially low in mouse hematopoietic cells with high turnovers [54]. To overcome this problem, MACs were generated by the top-down approach to have the same capacity as HAC vectors with the additional advantage of highly stable maintenance in mouse tissues including hematopoietic cells. Therefore, MACs are more valuable gene delivery vectors for the production of model mice than HACs [54, 55]. Although several groups have developed MACs using different methods, to our knowledge those derived from native mouse chromosomes via the top-down approach are the most stable in mice [56, 57]. In summary, HACs/MACs as gene delivery vectors can deliver large genomes, and are stably and independently maintained with defined copy numbers in host cells, as well as being transferrable to desired cell lines via MMCT [2].
Gene-loading techniques for HACs/MACs
Five types of gene cloning system can be utilized with top-down HACs/MACs: (a) HR-mediated gene insertion; (b) the Cre/loxP-mediated insertion system; (c) the Cre/loxP-mediated translocation system; (d) the multi-integrase (MI) system; and (e) the simultaneous or sequential integration of multiple gene-loading vectors (GLVs) (SIM) system. In system (a), gene loading with unique sequences in HACs/MACs via HR in DT40 cells has previously been performed [49, 54] (Fig. 3a). The same gene-loading system involving HR in other cells such as CHO and A9, but not DT40 cells, was succeeded by genome editing technologies [40].
Gene-loading methods applicable to HACs/MACs. a Homologous recombination-mediated gene insertion for gene of interest (GOI). b Cre/loxP-mediated circular DNA insertion. c Cre/loxP-mediated large genomic DNA fragment loading by reciprocal translocation. d The MI system. Five gene-loading vectors can theoretically be inserted into the MI system. e The SIM system. Three gene-loading vectors can be inserted simultaneously. The sequential method inserts gene-loading vectors one by one indefinitely. Both methods enable the theoretical indefinite integration of gene-loading vectors
HACs/MACs generally harbor loxP sites for gene loading. In system (b), circular vectors such as plasmids, P1 artificial chromosomes (PACs), and bacterial artificial chromosomes (BACs) were successfully loaded onto HACs/MACs by the Cre/loxP system for various purposes including functional analysis, monitoring system development, and gene therapy [58,59,60,61,62,63,64] (Fig. 3b). This loading system is particularly suited to when the gene of interest is covered by one of these circular vectors. In system (c), Mb-sized chromosome fragments such as the human CYP3A cluster and human dystrophin gene (~2.4 Mb) can be loaded onto top-down HACs/MACs [53, 65] (Fig. 3c). This system is suitable when the gene(s) of interest exceeds the delivery size limit of PACs and BACs. It involves manipulation of the chromosome containing the genomic region of interest by telomere-associated truncation and loxP targeting. The genomic region is eventually loaded on the HACs/MACs by Cre/loxP-mediated reciprocal translocation cloning in CHO cells.
Previously, the HAC/MAC system has mainly employed the Cre/loxP system in CHO cells for gene loading as mentioned above. However, two multiple gene-loading systems, the MI and SIM systems, have recently been developed for application in HACs/MACs with loxP sites [66, 67].
In system (d) (the MI system), four serine integrases and one recombinase (PhiC31, Bxb1, R4, TP901-1, and FLP) are utilized. Using all recombination sites, five GLVs can theoretically be inserted into the MI-HAC (Fig. 3d). For the efficient and convenient production of Tc animals carrying gene(s) of interest, a MAC carrying multi-integration sites (MI-MAC) was also constructed [54]. The MI-MAC has been transferred into mouse ES cells to enable gene(s) of interest to be directly loaded onto it [68]. The MI-MAC has also been transferred into HepG2 cells to verify cytotoxicity levels via luciferase reporter assay [69]. The MI-HAC/MAC system can be used in various cell types and has successfully achieved the insertion of large GLVs such as PACs and BACs and multiple genes into the MI-HAC/MAC [70,71,72]. Although limitations exist regarding the available drug-resistant genes for five GLVs loading onto the MI-HAC/MAC vector, this was solved by reusing the same drug-resistant genes by disrupting them with genome editing technology (Honma et al., 2017, unpublished data). This system is particularly valuable for gene loading when required number of gene loading is within five times and if the MI-HAC/MAC has already been transferred to the desired cell type.
In system (e) (the SIM system), two integrases and two recombinases (PhiC31, Bxb1, Cre, and FLP) are used. This system is applicable to HACs/MACs with loxP sites, and either simultaneous or sequential integration can be performed (Fig. 3e) [67]. The simultaneous integration system enables the simultaneous integration of three GLVs into HACs/MACs and further sequential insertions. The sequential integration system enables GLV loading one by one. Both methods theoretically allow the indefinite loading of GLVs onto HACs/MACs. Selection for gene insertion is performed by switching on/off hypoxanthine-guanine phosphoribosyl transferase (HPRT) reconstitution or neomycin resistance. Thus, gene loading onto HACs/MACs has to be performed in HPRT-deficient cells. Although gene loading with the SIM system is restricted in HPRT-deficient cells, HPRT KO in cells of interest can readily be performed by genome editing technology leading to direct gene loading onto HACs/MACs. The SIM system is particularly suitable when efficient simultaneous multi-gene loading or unlimited gene loading is required. HACs/MACs with MI or SIM systems are expected to be employed in a variety of fields and can be chosen depending on the purpose. Taken together, the combination of five cloning methods onto top-down HACs/MACs and genome editing will be useful for the cloning of desired genes and gene clusters. Regarding bottom-up HACs, systems (a, b, d, and e) are applicable to bottom-up HACs with loxP sites. However, bottom-up HACs cannot be used for reciprocal translocation-mediated gene cloning in system (c) because they are mostly circular and contain multiple loxP sites.
Humanized animal model: the CYP3A model mouse
The HAC/MAC system is a powerful tool for producing humanized animal models harboring multiple or large genes with physiological expression [2]. As an example, the humanized CYP3A mouse model was generated using HAC vectors [53]. CYP3A genes encode enzymes involved in drug metabolism and are associated with the metabolism of approximately 50% of commercially available drugs. However, CYP3A enzymes differ between species, so animal models such as the mouse and rat do not reflect human CYP3A-related pharmacokinetics. Therefore, humanized CYP3A model animals are useful for predicting CYP3A-related pharmacokinetics and toxicities of new drugs in humans.
CYP3A genes includingCYP3A4, CYP3A5, CYP3A7, and CYP3A43 form a cluster on human chromosome 7. It is difficult to introduce large genomic regions into mice by conventional technologies, so to produce a humanized CYP3A mouse model, human chromosome 7 was modified in DT40 cells by chromosome engineering and the CYP3A cluster was cloned into a HAC vector (CYP3A-HAC). The CYP3A-HAC was transferred into mouse ES cells to produce chimeric mice that transmitted the CYP3A-HAC through the germline. Further crossing with Cyp3a-KO mice produced fully humanized CYP3A mice that showed gender-, tissue-, and developmental stage-specific CYP3A expression, similar to humans [53, 73]. The humanized CYP3A mice also recapitulated the CYP3A metabolic activity observed in humans. Furthermore, whole-embryo cultures of the humanized CYP3A mouse embryo showed teratogenic effects of thalidomide that are not seen in rodents [74]. Thus, humanized CYP3A mice are expected to be useful for drug screening by predicting CYP3A-related drug metabolism.
Such drug metabolism-related genes generally form large clusters, so chromosome engineering including HAC/MAC technologies and chromosome transfer are powerful technologies for producing humanized mice carrying these genes. Furthermore, other humanized models of the human immune system or particular genetic diseases are likely to be produced in the future.
Previously, KO of endogenous orthologous genes was laborious, especially in ES cells for fully humanized model production [53]. Gene cluster KO requires the deletion of a large targeted region, mainly using the Cre/loxP system, so is labor- and time-intensive as well as expensive because targeting of two loxP sites and Cre/loxP-mediated chromosomal deletion must be performed in mouse ES cells, and chimeric mice formation assays are required at each step. To overcome the problem of ES cell-based technologies, genome editing technologies can be utilized to induce mutations, deletions of single genes, and large genomic deletions for orthologous gene KO [75]. Now, the combination of chromosome transfer and genome editing technology is crucial for the efficient production of fully humanized animal models (Fig. 4) [76]. If genome editing of KO endogenous genes can be applied to germline-transmitted Tc animals, both time and cost for mating Tc and KO animals will be saved. Additionally, the combination of HACs/MACs and genome editing can be adapted to generate humanized model rats which offer several advantages over mouse models [77, 78].
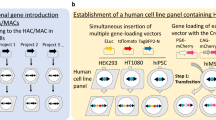
Graphical summary for the generation of humanized model mice. The development of genome editing techniques has enabled the rapid modification of chromosomes in ES cells and fertilized eggs. To generate a humanized model, genome editing techniques permit the knockout (KO) of orthologous gene loci in the mouse genome. Crossing KO mice with transchromosomic (Tc) mice generate fully humanized mice. Genome editing techniques realize the further modification of human genomic loci on the MAC to produce humanized mice carrying specific single-nucleotide polymorphisms or disease-associated mutations, deletions, or amplifications for a personalized model
Replacement of single-nucleotide polymorphisms in humanized animal models
Genome editing technologies enable the further modification of genomic sequences in pre-constructed HACs/MACs. Numerous single-nucleotide polymorphisms (SNPs) of human drug metabolism-related genes are reported to affect drug pharmacokinetics [79,80,81]. For example, CYP3A5 SNPs have been reported in intron 3: CYP3A5*1(6986A) and CYP3A5*3(g.6986A>G) [82,83,84]. CYP3A5*1 allele carriers express CYP3A5 mainly in the liver and intestine, while homozygotes of the CYP3A5*3 allele lack almost all CYP3A5 protein expression [85].
Recently, an MAC was utilized to produce humanized CYP3A mice (CYP3A-MAC mice), and human-like CYP3A-related drug metabolism was observed in the CYP3A-MAC mice as well as in the CYP3A-HAC mice (Kazuki et al., 2017, unpublished data) [53]. However, the CYP3A5 SNP genotype of humanized CYP3A mice was CYP3A5*3, and the mice lacked CYP3A5 protein expression. To produce humanized CYP3A mice that can be utilized to predict the CYP3A5 contribution to drug metabolism, the CYP3A5 SNP on the CYP3A-MAC was modified from CYP3A5*3 to CYP3A*1 by CRISPR/Cas9 in both mouse ES cells carrying the CYP3A-MAC and fertilized eggs carrying the CYP3A-MAC [86, 87, 106]. Modification with CRISPR/Cas9 was achieved by the transfection of mouse ES cells carrying the CYP3A-MAC using double-stranded circular DNA containing a 1 kb region of homology with the SNP, and pronuclear injection with 135 bp single-stranded DNA with the SNP as the donor. The resulting CYP3A5*1 mice showed higher CYP3A5 protein expression in their liver and intestine compared with CYP3A5*3 mice, and CYP3A5 was found to contribute more to metabolism in CYP3A5*1 mice. These results suggested that the CYP3A5 SNP had been functionally modified, and that the SNP effect could be recapitulated in mice because CYP3A5*1 mice showed the CYP3A5*1 carrier phenotype.
HACs/MACs can deliver entire gene regions with elements for physiological expression, and can maintain defined copy numbers, especially single copies, in model animals [53,54,55]. Therefore, modified SNPs can be more efficiently identified in Tc animals than in conventional genome-edited animals with diploid genomes. The combination of HAC/MAC systems and genome editing technologies is valuable for validation of the SNP effect in humans by producing humanized model animals with corresponding SNPs. Because genome editing technologies enable human genes to be efficiently modified on HACs/MACs, disease-associated mutations-, deletions-, and amplification-harboring models that reflect individual patient phenotypes can be generated from humanized animals (Fig. 4).
In vitro models for aneuploidy and cancer
Chromosome transfer is a powerful technology for producing aneuploidy syndrome cell models such as trisomy 21, 18, and 13, and cancer-initiating cell models to reflect the contribution of specific chromosome amplifications to cancer progression. One of the aneuploidy syndromes, DS, also known as trisomy 21 (Ts21), is the most common genetic chromosomal abnormality. Infants with DS suffer from transient abnormal myelopoiesis (DS-TAM) at a high frequency, and approximately 20–30% of cases subsequently develop DS-related acute megakaryoblastic leukemia (DS-AMKL) [88,89,90]. Mutations in GATA1, encoding the megakaryocyte transcription factor, resulting in the production of N-terminal truncated GATA1 (GATA1s) is associated with DS-TAM and DS-AMKL [91,92,93]. The combination of GATA1 mutation(s) and constitutive Ts21 is the most likely cause of DS-TAM, and additional mutations are thought to result in DS-AMKL [94].
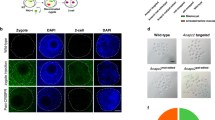
To understand the mechanism of the progression to TAM in DS patients, Ts21, GATA1s, and GATA1s/Ts21 human ES cells were generated by combining chromosome transfer and genome editing technologies [95] (Fig. 5i, ii). Notably, all ES cell lines generated were isogenic and genetically defined with the advantage of chromosome transfer. Zinc-finger nucleases were used for the targeted mutation of GATA1 exon 2 in human ES cells (WT-ES), and human ES cells in which both alleles were mutated (GATA1s-ES) were successfully obtained. Human chromosome 21 was then transferred via MMCT to WT-ES and GATA1s-ES cells to generate Ts21-ES and GATA1s/Ts21-ES, respectively. ES-sac-mediated in vitro hematopoietic differentiation analyses using the DS models revealed that Ts21 and the GATA1 mutations synergistically contributed to hematopoietic abnormalities. Although another group established isogenic cell lines via the spontaneous loss of chromosome 21 from induced pluripotent stem (iPS) cells derived from a DS patient, they reported the synergistic interaction of Ts21 and GATA1 mutations mainly from an assessment of non-isogenic cell lines with a different genotype [96]. It is crucial to establish isogenic cell lines for disease models harboring chromosome abnormalities.
Graphical summary of a disease model of aneuploidy syndrome. Chromosome transfer techniques enable the generation of a disease model associated with aneuploidy via the following methods: i Transfer of an additional chromosome in the same genome background; ii the reproduction of advanced mutations on the host genome for research into associated gene(s) using genome editing; iii, iv transfer of chromosomes with mutations or truncated chromosomes generated by genome editing in CHO or DT40 cells; and v identification of responsible gene(s) or loci using HACs/MACs with Mb-regions or gene(s)
The DS models generated by the combination of chromosome transfer and genome editing technology are useful for studying how GATA1 mutations associate with the onset of TAM in DS patients. Recently, genomic profiling suggested that several mutations contribute to the progression from DS-TAM to DS-AMKL [94]. Thus, GATA1s/Ts21-ES may be used to screen factors involved in DS-AMKL progression through introducing additional mutations into GATA1s/Ts21-ES cells by genome editing technologies (Fig. 5ii). The introduction of chromosome 21 with a partial deletion or specific gene mutation generated by genome editing technology into normal ES cells, corresponding to partial trisomy 21, will help us to screen the region or gene(s) responsible for the disease phenotype (Fig. 5iii, iv). Moreover, the transfer of HACs/MACs carrying specific loci or genes on hChr.21 will enable the identification of the gene (Fig. 5v).
Genome editing studies validating the contribution of recurrent mutations to colorectal carcinogenesis suggested that aneuploidy, as the gain of chromosomes such as 7, 13, and 20, may be associated with aggressive tumor phenotypes [97, 98]. Direct confirmation of colorectal cancer exacerbation can be achieved by generating aneuploidy models by MMCT of such human chromosomes. The combination of the two technologies will be useful for the generation of in vitro chromosomal abnormality models with multiple genetic alterations for the functional analysis of cancer development.
Conclusions and future perspectives
Chromosome transfer using MMCT is a unique and powerful technology for the generation of genetically modified animals for humanization and disease models. The introduction of Mb-sized genomic loci is technically difficult and takes several years using conventional BAC/transgenic-based genetic modification in mouse ES cells [99, 100].Genome editing techniques simplify the genetic modification and expand the applications. Moreover, the combination of chromosome transfer and genome editing offers a new generation of chromosome engineering.
Future prospects based on this combination are summarized in Fig. 6. Sophisticated chromosome engineering enables specific chromosomes to be monitored, leading to efficient chromosome sorting and transfer, and complex chromosome modification for various purposes. For example, CRISPR/Cas9 may be applicable to improve the efficiency of MMCT. Isolated microcells are a mixed population containing targeted and/or host chromosomes. The chromosome tagging system involving dCas9 [101] fused with fluorescence proteins may be useful in purifying microcells containing a targeted chromosome.
Considering the use of HACs in gene and cell therapy, to avoid the potential contamination of animal components and unknown viruses derived from fused microcells, HACs should be constructed in normal human cells such as iPS cells, and transferred without using cell lines of other species such as CHO, A9, or DT40 cells. Additionally, it will be beneficial to develop novel techniques to perform MMCT from human cells to targeted human cells such as patient-derived iPS cells.
The humanization of mice and larger animals such as cattle, pigs, and cats will be streamlined by efficient orthologous gene(s) KO using genome editing, and the generation of various humanized Tc animal models may promote drug development and therapy [102,103,104]. Once HACs/MACs with gene(s) of interest are constructed, they can be transferred to various mammalian cells and maintained independently without integration into the host genome. The generation of disease model animals such as Duchenne muscular dystrophy (DMD) with various deletions or mutations of the human DMD genome recapitulating individual human patients supports our understanding of detailed disease mechanisms resulting in different phenotypes.
In addition to generating synergistic driver mutations by genome editing, aneuploidy models generated by chromosome transfer provide an important insight into the hallmarks of cancer. For basic research, the screening of potential tumor repressor genes or gene clusters including noncoding RNAs with chromosome mapping by chromosome transfer will be promoted by the precise manipulation of chromosomes through genome editing. Chromosome mapping technology also ensures physiological gene expression following single chromosome transfer to identify reliable candidates.
Taken together, the benefits of the combination of chromosome transfer and genome editing not only aid efficiency and the acceleration of research, but also promote novel approaches and new perspectives in the field, such as synthetic biology involving genome writing [105].
References
Fournier RE, Ruddle FH. Microcell-mediated transfer of murine chromosomes into mouse, Chinese hamster, and human somatic cells. Proc Natl Acad Sci USA. 1977;74:319–23
Oshimura M, Uno N, Kazuki Y, Katoh M, Inoue T. A pathway from chromosome transfer to engineering resulting in human and mouse artificial chromosomes for a variety of applications to bio-medical challenges. Chromosome Res. 2014;23:111–33.
Suzuki N, Itou T, Hasegawa Y, Okazaki T, Ikeno M. Cell to cell transfer of the chromatin-packaged human beta-globin gene cluster. Nucleic Acids Res. 2010;38:e33.
Nakayama Y, Uno N, Uno K, Mizoguchi Y, Komoto S, Kazuki Y, et al. Recurrent micronucleation through cell cycle progression in the presence of microtubule inhibitors. Cell Struct Funct. 2015;40:51–9.
Liskovykh M, Lee NC, Larionov V, Kouprina N. Moving toward a higher efficiency of microcell-mediated chromosome transfer. Mol Ther Methods Clin Dev. 2016;3:16043
Katoh M, Kazuki Y, Kazuki K, Kajitani N, Takiguchi M, Nakayama Y, et al. Exploitation of the interaction of measles virus fusogenic envelope proteins with the surface receptor CD46 on human cells for microcell-mediated chromosome transfer. BMC Biotechnol. 2010;10:37.
Suzuki T, Kazuki Y, Oshimura M, Hara T. Highly efficient transfer of chromosomes to a broad range of target cells using Chinese Hamster ovary cells expressing murine leukemia virus-derived envelope proteins. PLoS ONE. 2016;11:e0157187.
Kouprina N, Tomilin AN, Masumoto H, Earnshaw WC, Larionov V. Human artificial chromosome-based gene delivery vectors for biomedicine and biotechnology. Expert Opin Drug Deliv. 2014;11:517–35.
Kugoh H, Mitsuya K, Meguro M, Shigenami K, Schulz TC, Oshimura M. Mouse A9 cells containing single human chromosomes for analysis of genomic imprinting. DNA Res. 1999;6:165–72.
Tanabe H, Nakagawa Y, Minegishi D, Hashimoto K, Tanaka N, Oshimura M, et al. Human monochromosome hybrid cell panel characterized by FISH in the JCRB/HSRRB. Chromosome Res. 2000;8:319–34.
Meaburn KJ, Parris CN, Bridger JM, The manipulation of chromosomes by mankind: the uses of microcell-mediated chromosome transfer. Chromosoma. 2005;114:263–74
Doherty AM, Fisher EM. Microcell-mediated chromosome transfer (MMCT): small cells with huge potential. Mamm Genome. 2003;14:583–92.
Oshimura M, Barrett JC. Multiple pathways to cellular senescence: role of telomerase repressors. Eur J Cancer. 1997;33:710–5.
Doherty AMO, Fisher EMC. Microcell-mediated chromosome transfer (MMCT): small cells with huge potential. Mamm Genome. 2003;14:583–92.
Horike S, Mitsuya K, Meguro M, Kotobuki N, Kashiwagi A, Notsu T, et al. Targeted disruption of the human LIT1 locus defines a putative imprinting control element playing an essential role in Beckwith-Wiedemann syndrome. Hum Mol Genet. 2000;9:2075–83.
Mitsuya K, Meguro M, Lee MP, Katoh M, Schulz TC, Kugoh H, et al. LIT1, an imprinted antisense RNA in the human KvLQT1 locus identified by screening for differentially expressed transcripts using monochromosomal hybrids. Hum Mol Genet. 1999;8:1209–17.
Matsuura S, Weemaes C, Smeets D, Takami H, Kondo N, Sakamoto S, et al. Genetic mapping using microcell-mediated chromosome transfer suggests a locus for Nijmegen breakage syndrome at chromosome 8q21-24. Am J Hum Genet. 1997;60:1487–94.
Matsuura S, Tauchi H, Nakamura A, Kondo N, Sakamoto S, Endo S, et al. Positional cloning of the gene for Nijmegen breakage syndrome. Nat Genet. 1998;19:179–81.
Horibata K, Iwamoto Y, Kuraoka I, Jaspers NG, Kurimasa A, Oshimura M, et al. Complete absence of Cockayne syndrome group B gene product gives rise to UV-sensitive syndrome but not Cockayne syndrome. Proc Natl Acad Sci USA. 2004;101:15410–5.
Matsuura S, Matsumoto Y, Morishima K, Izumi H, Matsumoto H, Ito E, et al. Monoallelic BUB1B mutations and defective mitotic-spindle checkpoint in seven families with premature chromatid separation (PCS) syndrome. Am J Med Genet A. 2006;140:358–67.
Abe S, Tanaka H, Notsu T, Horike S, Fujisaki C, Qi DL, et al. Localization of an hTERT repressor region on human chromosome 3p21.3 using chromosome engineering. Genome Integr. 2010;1:6.
Qi DL, Ohhira T, Oshimura M, Kugoh H. Human chromosome 5 carries a transcriptional regulator of human telomerase reverse transcriptase (hTERT). Biochem Biophys Res Commun. 2010;398:695–701.
Qi DL, Ohhira T, Fujisaki C, Inoue T, Ohta T, Osaki M, et al. Identification of PITX1 as a TERT suppressor gene located on human chromosome 5. Mol Cell Biol. 2011;31:1624–36.
Seyda A, Newbold RF, Hudson TJ, Verner A, MacKay N, Winter S, et al. A novel syndrome affecting multiple mitochondrial functions, located by microcell-mediated transfer to chromosome 2p14-2p13. Am J Hum Genet. 2001;68:386–96.
Kurimasa A, Ohno K, Oshimura M. Restoration of the cholesterol metabolism in 3T3 cell lines derived from the sphingomyelinosis mouse (spm/spm) by transfer of a human chromosome 18. Hum Genet. 1993;92:157–62.
Wakai M, Abe S, Kazuki Y, Oshimura M, Ishikawa F. A human artificial chromosome recapitulates the metabolism of native telomeres in mammalian cells. PLoS ONE. 2014;9:e88530.
Fukagawa T, Nogami M, Yoshikawa M, Ikeno M, Okazaki T, Takami Y, et al. Dicer is essential for formation of the heterochromatin structure in vertebrate cells. Nat Cell Biol. 2004;6:784–91.
Devoy A, Bunton-Stasyshyn RK, Tybulewicz VL, Smith AJ, Fisher EM. Genomically humanized mice: technologies and promises. Nat Rev Genet. 2012;13:14–20.
Shinohara T, Tomizuka K, Miyabara S, Takehara S, Kazuki Y, Inoue J, et al. Mice containing a human chromosome 21 model behavioral impairment and cardiac anomalies of Down’s syndrome. Hum Mol Genet. 2001;10:1163–75.
O’Doherty A, Ruf S, Mulligan C, Hildreth V, Errington ML, Cooke S, et al. An aneuploid mouse strain carrying human chromosome 21 with Down syndrome phenotypes. Science. 2005;309:2033–7.
Tomizuka K, Yoshida H, Uejima H, Kugoh H, Sato K, Ohguma A, et al. Functional expression and germline transmission of a human chromosome fragment in chimaeric mice. Nat Genet. 1997;16:133–43.
Kuroiwa Y, Kasinathan P, Choi YJ, Naeem R, Tomizuka K, Sullivan EJ, et al. Cloned transchromosomic calves producing human immunoglobulin. Nat Biotechnol. 2002;20:889–94.
Urnov FD, Miller JC, Lee YL, Beausejour CM, Rock JM, Augustus S, et al. Highly efficient endogenous human gene correction using designed zinc-finger nucleases. Nature. 2005;435:646–51.
Mahfouz MM, Li L, Shamimuzzaman M, Wibowo A, Fang X, Zhu JK. De novo-engineered transcription activator-like effector (TALE) hybrid nuclease with novel DNA binding specificity creates double-strand breaks. Proc Natl Acad Sci USA. 2011;108:2623–8.
Jinek M, Chylinski K, Fonfara I, Hauer M, Doudna JA, Charpentier E. A programmable dual-RNA-guided DNA endonuclease in adaptive bacterial immunity. Science. 2012;337:816–21.
Hsu PD, Lander ES, Zhang F. Development and applications of CRISPR-Cas9 for genome engineering. Cell. 2014;157:1262–78.
Wang H, La Russa M, Qi LS. CRISPR/Cas9 in genome editing and beyond. Annu Rev Biochem. 2016;85:227–64.
Hernandez D, Mee PJ, Martin JE, Tybulewicz VL, Fisher EM. Transchromosomal mouse embryonic stem cell lines and chimeric mice that contain freely segregating segments of human chromosome 21. Hum Mol Genet. 1999;8:923–33.
Kuroiwa Y, Shinohara T, Notsu T, Tomizuka K, Yoshida H, Takeda S, et al. Efficient modification of a human chromosome by telomere-directed truncation in high homologous recombination-proficient chicken DT40 cells. Nucleic Acids Res. 1998;26:3447–8.
Uno N, Hiramatsu K, Uno K, Komoto S, Oshimura M. CRISPR/Cas9-induced transgene insertion and telomere-associated truncation of a single human chromosome for chromosome engineering in CHO and A9 cells. Sci Rep. 2017;7:12739.
Lee JS, Kallehauge TB, Pedersen LE, Kildegaard HF. Site-specific integration in CHO cells mediated by CRISPR/Cas9 and homology-directed DNA repair pathway. Sci Rep. 2015;5:8572.
Cho SW, Kim S, Kim Y, Kweon J, Kim HS, Bae S, et al. Analysis of off-target effects of CRISPR/Cas-derived RNA-guided endonucleases and nickases. Genome Res. 2014;24:132–41.
Grimes BR, Schindelhauer D, McGill NI, Ross A, Ebersole TA, Cooke HJ. Stable gene expression from a mammalian artificial chromosome. EMBO Rep. 2001;2:910–4.
Mandegar MA, Moralli D, Khoja S, Cowley S, Chan DY, Yusuf M, et al. Functional human artificial chromosomes are generated and stably maintained in human embryonic stem cells. Hum Mol Genet. 2011;20:2905–13.
Ikeno M, Grimes B, Okazaki T, Nakano M, Saitoh K, Hoshino H, et al. Construction of YAC-based mammalian artificial chromosomes. Nat Biotechnol. 1998;16:431–9.
Iida Y, Kim JH, Kazuki Y, Hoshiya H, Takiguchi M, Hayashi M, et al. Human artificial chromosome with a conditional centromere for gene delivery and gene expression. DNA Res. 2010;17:293–301.
Katoh M, Ayabe F, Norikane S, Okada T, Masumoto H, Horike S, et al. Construction of a novel human artificial chromosome vector for gene delivery. Biochem Biophys Res Commun. 2004;321:280–90.
Kakeda M, Nagata K, Osawa K, Matsuno H, Hiratsuka M, Sano A, et al. A new chromosome 14-based human artificial chromosome (HAC) vector system for efficient transgene expression in human primary cells. Biochem Biophys Res Commun. 2011;415:439–44.
Kazuki Y, Hoshiya H, Takiguchi M, Abe S, Iida Y, Osaki M, et al. Refined human artificial chromosome vectors for gene therapy and animal transgenesis. Gene Ther. 2011;18:384–93.
Iida Y, Kazuki Y, Hayashi M, Ueda Y, Hasegawa M, Kouprina N, et al. Bi-HAC vector system toward gene and cell therapy. ACS Synth Biol. 2014;3:83–90.
Hasegawa Y, Ishikura T, Hasegawa T, Watanabe T, Suzuki J, Nakayama M, et al. Generating a transgenic mouse line stably expressing human MHC surface antigen from a HAC carrying multiple genomic BACs. Chromosoma. 2014;124:107–18.
Miyamoto K, Suzuki N, Sakai K, Asakawa S, Okazaki T, Kudoh J, et al. A novel mouse model for Down syndrome that harbor a single copy of human artificial chromosome (HAC) carrying a limited number of genes from human chromosome 21. Transgenic Res. 2014;23:317–29.
Kazuki Y, Kobayashi K, Aueviriyavit S, Oshima T, Kuroiwa Y, Tsukazaki Y, et al. Trans-chromosomic mice containing a human CYP3A cluster for prediction of xenobiotic metabolism in humans. Hum Mol Genet. 2013;22:578–92.
Takiguchi M, Kazuki Y, Hiramatsu K, Abe S, Iida Y, Takehara S, et al. A novel and stable mouse artificial chromosome vector. ACS Synth Biol. 2012;3:903–14.
Kazuki K, Takehara S, Uno N, Imaoka N, Abe S, Takiguchi M, et al. Highly stable maintenance of a mouse artificial chromosome in human cells and mice. Biochem Biophys Res Commun. 2013;442:44–50.
Shen MH, Mee PJ, Nichols J, Yang J, Brook F, Gardner RL, et al. A structurally defined mini-chromosome vector for the mouse germ line. Curr Biol. 2000;10:31–4.
Co DO, Borowski AH, Leung JD, van der Kaa J, Hengst S, Platenburg GJ, et al. Generation of transgenic mice and germline transmission of a mammalian artificial chromosome introduced into embryos by pronuclear microinjection. Chromosome Res. 2000;8:183–91.
Otsuki A, Tahimic CG, Tomimatsu N, Katoh M, Chen DJ, Kurimasa A, et al. Construction of a novel expression system on a human artificial chromosome. Biochem Biophys Res Commun. 2005;329:1018–25.
Suda T, Katoh M, Hiratsuka M, Takiguchi M, Kazuki Y, Inoue T, et al. Heat-regulated production and secretion of insulin from a human artificial chromosome vector. Biochem Biophys Res Commun. 2006;340:1053–61.
Kurosaki H, Hiratsuka M, Imaoka N, Iida Y, Uno N, Kazuki Y, et al. Integration-free and stable expression of FVIII using a human artificial chromosome. J Hum Genet. 2011;56:727–33.
Hiratsuka M, Uno N, Ueda K, Kurosaki H, Imaoka N, Kazuki K, et al. Integration-free iPS cells engineered using human artificial chromosome vectors. PLos ONE. 2011;6:e25961.
Yamada H, Li YC, Nishikawa M, Oshimura M, Inoue T. Introduction of a CD40L genomic fragment via a human artificial chromosome vector permits cell-type-specific gene expression and induces immunoglobulin secretion. J Hum Genet. 2008;53:447–53.
Ren X, Katoh M, Hoshiya H, Kurimasa A, Inoue T, Ayabe F, et al. A novel human artificial chromosome vector provides effective cell lineage-specific transgene expression in human mesenchymal stem cells. Stem Cells. 2005;23:1608–16.
Uno N, Uno K, Komoto S, Suzuki T, Hiratsuka M, Osaki M, et al. Development of a safeguard system using an episomal mammalian artificial chromosome for gene and cell therapy. Mol Ther Nucleic Acids. 2015;4:e272.
Hoshiya H, Kazuki Y, Abe S, Takiguchi M, Kajitani N, Watanabe Y, et al. A highly stable and nonintegrated human artificial chromosome (HAC) containing the 2.4 Mb entire human dystrophin gene. Mol Ther. 2009;17:309–17.
Yamaguchi S, Kazuki Y, Nakayama Y, Nanba E, Oshimura M, Ohbayashi T. A method for producing transgenic cells using a multi-integrase system on a human artificial chromosome vector. PLoS ONE. 2011;6:e17267.
Suzuki T, Kazuki Y, Oshimura M, Hara T. A novel system for simultaneous or sequential integration of multiple gene-loading vectors into a defined site of a human artificial chromosome. PLoS ONE. 2014;9:e110404.
Yoshimura Y, Nakamura K, Endo T, Kajitani N, Kazuki K, Kazuki Y, et al. Mouse embryonic stem cells with a multi-integrase mouse artificial chromosome for transchromosomic mouse generation. Transgenic Res. 2015;24:717–27.
Wakuri S, Yamakage K, Kazuki Y, Kazuki K, Oshimura M, Aburatani S, et al. Correlation between luminescence intensity and cytotoxicity in cell-based cytotoxicity assay using luciferase. Anal Biochem. 2017;522:18–29.
Endo T, Noda N, Kuromi Y, Kokura K, Kazuki Y, Oshimura M, et al. Evaluation of an Hprt-Luciferase reporter gene on a mammalian artificial chromosome in response to cytotoxicity. Yonago Acta Med. 2016;59:174–82.
Kokura K, Kuromi Y, Endo T, Anzai N, Kazuki Y, Oshimura M, et al. A kidney injury molecule-1 (Kim-1) gene reporter in a mouse artificial chromosome: the responsiveness to cisplatin toxicity in immortalized mouse kidney S3 cells. J Gene Med. 2016;18:273–81.
Yasunaga M, Fujita Y, Saito R, Oshimura M, Nakajima Y. Continuous long-term cytotoxicity monitoring in 3D spheroids of beetle luciferase-expressing hepatocytes by nondestructive bioluminescence measurement. BMC Biotechnol. 2017;17:54.
Kobayashi K, Abe C, Endo M, Kazuki Y, Oshimura M, Chiba K. Gender difference of hepatic and intestinal CYP3A4 in CYP3A-humanized mice generated by a human chromosome-engineering technique. Drug Metab Lett. 2017;11.
Kazuki Y, Akita M, Kobayashi K, Osaki M, Satoh D, Ohta R, et al. Thalidomide-induced limb abnormalities in a humanized CYP3A mouse model. Sci Rep. 2016;6:21419.
Wang L, Shao Y, Guan Y, Li L, Wu L, Chen F, et al. Large genomic fragment deletion and functional gene cassette knock-in via Cas9 protein mediated genome editing in one-cell rodent embryos. Sci Rep. 2015;5:17517.
Boroviak K, Doe B, Banerjee R, Yang F, Bradley A. Chromosome engineering in zygotes with CRISPR/Cas9. Genesis. 2016;54:78–85.
Abbott A. Laboratory animals: the Renaissance rat. Nature. 2004;428:464–6.
Jacob HJ, Kwitek AE. Rat genetics: attaching physiology and pharmacology to the genome. Nat Rev Genet. 2002;3:33–42.
Werk AN, Cascorbi I. Functional gene variants of CYP3A4. Clin Pharmacol Ther. 2014;96:340–8.
Pinto N, Dolan ME. Clinically relevant genetic variations in drug metabolizing enzymes. Curr Drug Metab. 2011;12:487–97.
Sissung TM, Troutman SM, Campbell TJ, Pressler HM, Sung H, Bates SE, et al. Transporter pharmacogenetics: transporter polymorphisms affect normal physiology, diseases, and pharmacotherapy. Discov Med. 2012;13:19–34.
Kuehl P, Zhang J, Lin Y, Lamba J, Assem M, Schuetz J, et al. Sequence diversity in CYP3A promoters and characterization of the genetic basis of polymorphic CYP3A5 expression. Nat Genet. 2001;27:383–91.
Fukuen S, Fukuda T, Maune H, Ikenaga Y, Yamamoto I, Inaba T, et al. Novel detection assay by PCR-RFLP and frequency of the CYP3A5 SNPs, CYP3A5*3 and *6, in a Japanese population. Pharmacogenetics. 2002;12:331–4.
Lamba J, Hebert JM, Schuetz EG, Klein TE, Altman RB. PharmGKB summary: very important pharmacogene information for CYP3A5. Pharmacogenet Genomics. 2012;22:555–8.
Lin YS, Dowling AL, Quigley SD, Farin FM, Zhang J, Lamba J, et al. Co-regulation of CYP3A4 and CYP3A5 and contribution to hepatic and intestinal midazolam metabolism. Mol Pharmacol. 2002;62:162–72.
Oji A, Noda T, Fujihara Y, Miyata H, Kim YJ, Muto M, et al. CRISPR/Cas9 mediated genome editing in ES cells and its application for chimeric analysis in mice. Sci Rep. 2016;6:31666.
Noda T, Oji A, Ikawa M. Genome editing in mouse zygotes and embryonic stem cells by introducing SgRNA/Cas9 expressing plasmids. Methods Mol Biol. 2017;1630:67–80.
Massey GV, Zipursky A, Chang MN, Doyle JJ, Nasim S, Taub JW, et al. A prospective study of the natural history of transient leukemia (TL) in neonates with Down syndrome (DS): Children’s Oncology Group (COG) study POG-9481. Blood. 2006;107:4606–13.
Malinge S, Izraeli S, Crispino JD. Insights into the manifestations, outcomes, and mechanisms of leukemogenesis in Down syndrome. Blood. 2009;113:2619–28.
Gamis AS, Alonzo TA, Gerbing RB, Hilden JM, Sorrell AD, Sharma M, et al. Natural history of transient myeloproliferative disorder clinically diagnosed in Down syndrome neonates: a report from the Children’s Oncology Group Study A2971. Blood. 2011;118:6752–9. quiz 6996
Wechsler J, Greene M, McDevitt MA, Anastasi J, Karp JE, Le Beau MM, et al. Acquired mutations in GATA1 in the megakaryoblastic leukemia of Down syndrome. Nat Genet. 2002;32:148–52.
Xu G, Nagano M, Kanezaki R, Toki T, Hayashi Y, Taketani T, et al. Frequent mutations in the GATA-1 gene in the transient myeloproliferative disorder of Down syndrome. Blood. 2003;102:2960–8.
Ahmed M, Sternberg A, Hall G, Thomas A, Smith O, O’Marcaigh A, et al. Natural history of GATA1 mutations in Down syndrome. Blood. 2004;103:2480–9.
Yoshida K, Toki T, Okuno Y, Kanezaki R, Shiraishi Y, Sato-Otsubo A, et al. The landscape of somatic mutations in Down syndrome-related myeloid disorders. Nat Genet. 2013;45:1293–9.
Kazuki Y, Yakura Y, Abe S, Osaki M, Kajitani N, Kazuki K, et al. Down syndrome-associated haematopoiesis abnormalities created by chromosome transfer and genome editing technologies. Sci Rep. 2014;4:6136.
Banno K, Omori S, Hirata K, Nawa N, Nakagawa N, Nishimura K, et al. Systematic Cellular Disease Models Reveal Synergistic Interaction of Trisomy 21 and GATA1 Mutations in Hematopoietic Abnormalities. Cell Rep. 2016;15:1228–41.
Matano M, Date S, Shimokawa M, Takano A, Fujii M, Ohta Y, et al. Modeling colorectal cancer using CRISPR-Cas9-mediated engineering of human intestinal organoids. Nat Med. 2015;21:256–62.
Drost J, van Jaarsveld RH, Ponsioen B, Zimberlin C, van Boxtel R, Buijs A, et al. Sequential cancer mutations in cultured human intestinal stem cells. Nature. 2015;521:43–47.
Murphy AJ, Macdonald LE, Stevens S, Karow M, Dore AT, Pobursky K, et al. Mice with megabase humanization of their immunoglobulin genes generate antibodies as efficiently as normal mice. Proc Natl Acad Sci USA. 2014;111:5153–8.
Lee EC, Liang Q, Ali H, Bayliss L, Beasley A, Bloomfield-Gerdes T, et al. Complete humanization of the mouse immunoglobulin loci enables efficient therapeutic antibody discovery. Nat Biotechnol. 2014;32:356–63.
Chen B, Gilbert LA, Cimini BA, Schnitzbauer J, Zhang W, Li GW, et al. Dynamic imaging of genomic loci in living human cells by an optimized CRISPR/Cas system. Cell. 2013;155:1479–91.
Do L, Wittayarat M, Terazono T, Sato Y, Taniguchi M, Tanihara F, et al. Effects of duration of electric pulse on in vitro development of cloned cat embryos with human artificial chromosome vector. Reprod Domest Anim. 2016;51:1039–43.
Kuroiwa Y, Kasinathan P, Matsushita H, Sathiyaselan J, Sullivan EJ, Kakitani M, et al. Sequential targeting of the genes encoding immunoglobulin-mu and prion protein in cattle. Nat Genet. 2004;36:775–80.
Yang L, Guell M, Niu D, George H, Lesha E, Grishin D, et al. Genome-wide inactivation of porcine endogenous retroviruses (PERVs). Science. 2015;350:1101–4.
Boeke JD, Church G, Hessel A, Kelley NJ, Arkin A, Cai Y, et al. GENOME ENGINEERING. The Genome Project-Write. Science. 2016;353:126–7.
Abe S, Kobayashi K, Oji A, Sakuma T, Kazuki K, Takehara S, et al. Modification of single-nucleotide polymorphism in a fully humanized CYP3A mouse by genome editing technology. Sci Rep. 2017;7:15189.
Acknowledgments
We thank the graduate students and technical staff at Tottori University for their technical assistance, and Drs H. Kugoh, M. Hiratsuka, and T. Ohbayashi for critical discussions. Our studies in this review were supported in part by The Mochida Memorial Foundation for Medical and Pharmaceutical Research (to Y.K.), The Naito Foundation (to Y.K.), The Takeda Science Foundation (to Y.K.), the Regional Innovation Strategy Support Program from the Ministry of Education, Culture, Sports, Science and Technology of Japan (to Y.K. and M.O.), the Funding Program for Next Generation World-Leading Researchers (NEXT Program) from the Japan Society for the Promotion of Science (JSPS) (to Y.K.), the Basic Science and Platform Technology Program for Innovative Biological Medicine from Japan Agency for Medical Research and Development, AMED (to Y.K.), and JSPS KAKENHI Grant Number 25221308 (to M.O.), Grant Number 15H04285 (to Y.K.), and Grant Number 15K19615 (to N.U.).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
Dr. M.O. is CEO of, and a shareholder in, Trans Chromosomics Inc. The remaining authors declare that they have no competing interests.
Additional information
Narumi Uno and Satoshi Abe contributed equally to this work.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-ShareAlike 4.0 International License, which permits any non-commercial use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. If you remix, transform, or build upon this article or a part thereof, you must distribute your contributions under the same license as the original. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-sa/4.0/.
About this article
Cite this article
Uno, N., Abe, S., Oshimura, M. et al. Combinations of chromosome transfer and genome editing for the development of cell/animal models of human disease and humanized animal models. J Hum Genet 63, 145–156 (2018). https://doi.org/10.1038/s10038-017-0378-7
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s10038-017-0378-7
This article is cited by
-
Human artificial chromosome carrying 3p21.3-p22.2 region suppresses hTERT transcription in oral cancer cells
Chromosome Research (2023)
-
Panel of human cell lines with human/mouse artificial chromosomes
Scientific Reports (2022)
-
Blastocyst complementation using Prdm14-deficient rats enables efficient germline transmission and generation of functional mouse spermatids in rats
Nature Communications (2021)
-
Construction of stable mouse artificial chromosome from native mouse chromosome 10 for generation of transchromosomic mice
Scientific Reports (2021)
-
Y chromosome in health and diseases
Cell & Bioscience (2020)