Abstract
Background:
Chronic rhinosinusitis (CRS) is characterized by mucous overproduction and submucosal gland hyperplasia. The global protein profile of sinonasal secretions in pediatric CRS has not been studied. We hypothesized that MUC5B, a glandular mucin, would be relatively increased in CRS secretions compared to other mucins.
Methods:
Secretions were collected at Children’s National Health System (Children’s National) from CRS patients undergoing sinus surgery and from control patients without CRS undergoing craniofacial procedures. Proteins were extracted, digested to peptides, and analyzed by mass spectometry. Fold change significance was calculated using the QSpec algorithm. Western blot analysis was performed to validate proteomic findings.
Results:
In total, 294 proteins were identified. Although both MUC5B and MUC5AC were identified in a majority of samples, the relative abundance of MUC5B was found to be significantly higher (P < 0.05). Western blot data validated these findings. Other proteins with the highest significant positive-fold change in CRS samples were BP1 fold-containing family A member 1, chitinase-3-like protein 1, plastin-2, serpin 10, and BP1 fold-containing family B member 1.
Conclusion:
Overall, our data demonstrate an increase of MUC5B abundance in the sinus secretions of pediatric patients with CRS.
Similar content being viewed by others
Main
Chronic rhinosinusitis (CRS) is a common chronic upper respiratory illness that affects more than 16% of the US population (1) and accounts for more than 18 million office visits and direct healthcare costs of over 4 billion dollars a year (2). Although the pathogenesis of CRS has not yet been established at the molecular level, it is well accepted that the pathophysiology reflects a chronic inflammatory process of the paranasal and sinus mucosa, characterized by the overproduction of mucus lasting for 12 wk or more.
Mucin glycoproteins are the primary macromolecular component of mucus. Mucins, as well as other mucosal components, act as physiological barrier against contaminants and bacteria by forming a viscoelastic mucus gel. However, when overproduced, mucins and mucus contribute to the stagnation of secretions, obstruction of sinonasal clearance and chronic infection. To date, 20 mucin glycoproteins have been identified (3). These include the secretory mucins, of which MUC2, MUC5AC, and MUC5B are the best studied, and membrane-tethered mucins, of which MUC1 and MUC4 are presently the best investigated. In healthy lower airways, MUC5AC is typically expressed in the goblet cells, while MUC5B is restricted to the submucosal glands (3). In sinus mucosa, we have shown that MUC5AC is expressed only in goblet cells and that children with CRS do not exhibit goblet cell hyperplasia compared to controls (4) and that MUC5B mucin is expressed in goblet cells, both in control and CRS patients (5). Instead, submucosal glandular hyperplasia is the characteristic phenotype of pediatric CRS and MUC5B is the predominant glandular mucin (5).
To date, there are no studies on MUC5AC and MUC5B levels in sinonasal secretions from children, although MUC5B levels are increased in adult CRS secretions (6). There also appears to be differences in the pathophysiology of CRS in children compared to adults such that results from adult studies may not be applicable to pediatric CRS (7). Several proteomic studies analyzed nasal lavage fluid from adults (8,9,10,11) from individuals with and without various forms of sinusitis, but to our knowledge no reports in children. Identifying the exact macromolecular mucin content present in pediatric sinonasal secretions would not only provide clues as to the potential pathophysiology of this condition in children, but could also spark research into innovative ways to specifically target the viscous properties of pediatric sinonasal secretions.
This study was designed to interrogate the global differential expression of proteins, specifically in pediatric CRS secretions, compared to non-CRS secretions using semiquantitative and quantitative techniques in order to determine the correlation between protein abundance and disease state. Given that we have shown that pediatric CRS is histologically characterized by an increase in submucosal glands, a tissue in which MUC5B is well expressed (reviewed in ref. 3), we hypothesized that sinonasal secretions from CRS pediatric patients would contain an overabundance of MUC5B relative to other mucin glycoproteins.
Results
Table 1 lists the demographic information for the patient samples. Mean age of CRS patients was 9.8 y (range 2–14 y). Mean age of control patients was 9.2 y(2–18 y). All CRS subjects had radiographic confirmed evidence of CRS with symptoms lasting greater than 3 mo. Control patients did not have any symptoms of or treatment for CRS and were undergoing surgery for non-CRS indications.
Proteomic Characterization of Secretions
Proteins were separated by Sodium dodecyl sulfate polyacrylamide gel electrophoresis (SDS-PAGE) and visualized using Bio-Safe Coomassie stain ( Figure 1 ). For proteomics analysis, the lanes from each sample were serially cut and processed as described in the Methods. A total of 242 proteins were identified by mass spectometry.
Protein content of clinical samples. Coomassie stained 4–12% Bis-Tris gels with desalted sinonasal secretion samples. The first lane of each gel contains molecular weight markers, shown on left (kDa). Each sample lane was loaded with 50 μg total protein. After electrophoresis each lane was serially sliced and in-gel digested for proteomic processing. C, control (nondiseased); CRS, chronic rhinosinusitis secretions.
To identify which of the 242 proteins demonstrated the most significant differences in abundance in the two cohorts, the differential expression of each protein was calculated using the QSpec algorithm (12). Proteins were considered to be increased in CRS samples or control samples based on P value ≤ 0.001 and absolute fold change ≥ 1.5. The 47 proteins that meet these criteria are listed in Table 2 based on the highest fold change in descending order. The protein with the highest significant positive-fold change in CRS samples was bactericidal/permeability-increasing fold containing family A, member 1 (BPIFA1) (log2 fold change = 3.93). The protein with the most significant negative-fold change (decreased abundance in CRS samples) was retinal dehydrogenase 1 (ALDH1A1, log2 fold change = −4.39).
In order to analyze the categories of proteins that were differentially expressed in CRS compared to control sinonasal secretions, the UniProt database was used to analyze the 47 proteins in Table 2 by Gene Ontology (GO) terms. The top seven most frequent biological processes for CRS and control samples are shown in Figure 2a , b respectively. The number of proteins in metabolic and cellular processes were similar for both cohorts. In some processes, e.g., developmental, proteins were more frequently present in control, but not CRS samples. The number of proteins in response to stimulus and biological regulation processes was almost double in the CRS samples. Of note, immune system process proteins were only identified in CRS samples.
Gene ontology representation of identified proteins in clinical specimens. Significant proteins (P ≤ 0.0001 and fold change ≥1.5) identified in chronic rhinosinusitis secretions (CRS) and control sinonasal secretion samples were compared to the human UniProt Database. The top seven protein enriched Gene Ontology (GO) terms (biological processes) in (a) CRS and (b) control samples are represented.
Mucin Protein Identification: MUC5B Was Significantly More Abundant Compared to MUC5AC in CRS Samples by Proteomic Analysis
The unique number of peptides and spectral counts for MUC5AC and MUC5B, which are similar in molecular weight and therefore comparable (13), are shown in Table 3 . MUC5B abundance by spectral count average was significantly higher than MUC5AC spectral count average (P = 0.043) in CRS secretions ( Figure 3 ). There was no significant difference between the spectral counts for MUC5B compared to MUC5AC in control samples (P = 0.180).
Spectral count of MUC5AC and MUC5B. Average normalized spectral count of MUC5B peptides compared to MUC5AC peptides in sinonasal secretions from pediatric chronic rhinosinusitis secretions (CRS) and control patients. Significantly more MUC5B was detected compared to MUC5AC in CRS samples (*P = 0.043, compared to MUC5AC in CRS), but not in control samples. White bar: nondiseased MUC5AC; black bar: nondiseased MUC5AB; gray bar: CRS MUC5AC; hatched bar: CRS MUC5B.
Western Blot Analysis of Secretory Mucins in Sinonasal Secretions
The secretory mucins MUC5AC and MUC5B were the only mucin glycoproteins identified in sinonasal secretions by mass spectrometry (MS). Mucins are typically underrepresented in proteomic analysis because of their large size and physical properties (14,15). Thus, we evaluated the abundance of MUC5B and MUC5AC using western blot techniques. Figure 4a , b show representative western blots for MUC5B and MUC5AC mucins in sinonasal samples. Due to limited sample availability, only one of the three control samples (sample C2) and four of the five CRS samples (Samples CRS 1–4) that were analyzed by proteomics were available for validation by western blot analysis. In general, there was good correlation between spectral counts and western blot band intensity. The densitometry signal for each protein was normalized to the positive biologic control signal for each blot. The normalized densitometry counts were then averaged per cohort. The average MUC5AC band intensity was lower in CRS than in control secretions, although this did not reach statistical significance (P = 0.06). On the other hand, parallel to the proteomics data, there was an increase (7.5-fold) in MUC5B intensity in CRS relative to control secretions, although this was not statistically significant (P = 0.11) ( Figure 4c ).
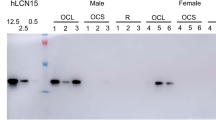
Western blots of mucin glycoproteins. (a) Representative western blot of MUC5AC mucin in sinonasal secretions from children with or without chronic rhinosinusitis secretions (CRS). Samples were electrophoresed on 1% agarose gels and immunostained at a dilution of 1:4,000 using an anti-MUC5AC rabbit polyclonal antibody (see Methods). 30 μg protein from sinonasal secretions was loaded. The sample numbers are listed above the lane. C, control (nondiseased). Saliva (30 μg) was used as a negative internal control and A549 cell lysates (30 μg) as a positive internal control for MUC5AC mucin. (b) Representative western blot analysis of MUC5B in sinonasal secretions from children with or without CRS. The membrane used in a was stripped and immunostained with an anti-MUC5B rabbit polyclonal antibody (see Methods) at a dilution of 1:300. 30 μg of protein for all samples was loaded. This time, A549 cell lysates were used as a negative internal control and saliva as a positive internal control for MUC5B mucin. (c) The average fold change of MUC5AC band intensity decreased in disease to a level 0.298 of control band intensity, although this did not reach statistical significance (P = 0.058). On the other hand, parallel to the proteomics data, there was a marked 7.47-fold change increase in MUC5B signal in disease relative to control, although this did not reach statistical significance either (P = 0.11).
Discussion
For this study, we aimed to gain a better understanding of the protein components in sinonasal secretions from CRS patients compared to sinonasal secretions from control patents. In contrast to other studies, we utilized sinonasal secretions obtained during surgery from CRS and control patients directly from the osteomeatal region, rather than nasal lavage fluid. Utilizing label-free proteomic techniques, we successfully created a protein profile showing the varying levels of expression of 294 proteins in sinonasal secretions from control and CRS pediatric patients. A large number of the proteins identified as having significant fold change in CRS sinonasal secretions were involved in innate immune response.
Most of the proteins with significant fold changes identified in our CRS sinonasal samples were assigned to ‘immune system processes’ by GO, supporting the concept that sinonasal mucosa in CRS patients is in a chronic inflammatory state. Several of the innate immunity proteins that show increased expression in CRS secretions in our study, e.g., BPIFA1, DMBT1, S100A8, S100A9, LTF, MYH9, and IGHA1, have been reported as being dysregulated in studies of CRS nasal lavage fluid or sinus tissues (6,11,16,17,18,19). These proteins are discussed below.
BPIFA1, which has the highest fold change in CRS sinonasal secretions, is a member of the BPI or PLUNC family of proteins (20) found in secretions from the upper airways, nose, and mouth. While the functions of BPI proteins have not yet been fully elucidated, they appear to have multifunctional roles in airway immune defense. Other proteomic studies identified high expression of BPIFA1 in CRS nasal mucosa tissue and verified this observation by immunohistochemistry (16). Since BPIFA1 is well expressed in submucosal glands in the respiratory tract (21,22), its overexpression in CRS sinonasal secretions further supports submucosal gland hyperplasia in CRS sinus mucosa.
DMBT1, also known as GP340, is a glycoprotein implicated in immune response (23), mucosal inflammation and epithelial regeneration (24). Its scavenger receptor cysteine rich domains (25) and glycan side chains (26) facilitate its binding to various pathogens (25), allowing it to function as part of the innate defense mucosal mechanism to remove bacteria, viruses or other infectious agents. Interestingly, DMBT1 is strongly associated with MUC5B in respiratory tract secretions (27).
The S100 proteins, specifically S100A8 and S100A9, are proinflammatory mediators that induce chemotaxis of neutrophils and monocytes. Each has been identified in CRS samples. S100A9 is overexpressed in nasal secretory cells from CRS patients with nasal polyps (28), as well as nasal mucus (29) and nasal lavage fluid from CRS patients (19). S100A8 is overexpressed in sinonasal mucosa tissue from patients with CRS with nasal polyps (30,31).
LTF, MYH9, and IGHA1 are also increased in CRS samples. Proteomic studies have shown overexpression of LTF, an antimicrobial protein, in the nasal lavage fluid of patients with acute sinusitis (19), asthmatic CRS (11), smokers (10), and seasonal allergic rhinitis patients (32), as well as nasal mucus and tissue from CRS patients (29,33). MYH9, which aids in cell motility and maintains cell shape (34), and IGHA1, which acts in immune system defense (35), also exhibit increased levels in proteomic studies on nasal lavage fluid from acute sinusitis (8) and asthmatic CRS (11) patients.
Mucins in Sinonasal Secretions
Mucin data from our proteomic and western blot analysis indicated a trend of overexpression of MUC5B mucin in pediatric CRS sinonasal secretions relative to MUC5AC. This result is not unexpected, based on studies showing an increase in submucosal glands in CRS patients (5,36) and expression of MUC5B in the mucosal cells in submucosal glands of the sinus mucosa (5). Proteomic analysis showed that MUC5B was significantly overexpressed in CRS samples, whereas MUC5AC did not reach statistical significance. In line with our results, MUC5B, but not MUC5AC, has been identified by proteomic analyses of nasal mucus from CRS patients with nasal polyps (29).
Viswanathan et al. evaluated MUC5B and MUC5AC mucin expression in sinus mucosa samples by ELISA to show a statistically significant increase in MUC5B levels in CRS compared to controls with no difference in MUC5AC levels between the cohorts (6); a finding that is also consistent with our proteomic data.
Conclusions
To our knowledge, this study represents the first global protein profile analysis of pediatric sinonasal secretions from patients with and without CRS, characterizing the presence of proteins in CRS samples which are to likely to play a role in the inflammatory nature of CRS. Furthermore, MUC5B has an increased abundance trend compared to MUC5AC in CRS patients, likely reflecting the increased glandular hyperplasia characteristic of the disease process. Many of the significantly increased proteins in CRS sinonasal secretions are implicated in airway inflammatory reaction and immune response and some might be useful to further investigate as potential biomarkers for CRS.
Methods
Sample Collection and Preparation
Sinonasal secretions were collected from pediatric patients according to the institutional guidelines at Children’s National Medical Center (Children’s National). Studies were approved by the Institutional Review Board of Children’s National. All subjects/families signed informed consent for participation in this study. Control secretions were obtained from patients without known CRS disease undergoing craniofacial and/or neurosurgical surgeries. CRS secretions were collected from patients undergoing functional endoscopic sinus surgery for CRS. Exclusion criteria for CRS patients included: Patients with CRS who do not require surgery. Exclusion criteria for controls included: a history of sinonasal surgery, current sinonasal infection, sinonasal or allergic symptoms within the previous 3 mo, and/or treatment with topical nasal steroids within 1 mo or antihistamines within 3 mo before surgery. Computed tomography and or magnetic resonance imaging for controls all had normal paranasal sinuses, these images had all been obtained for clinically indicated reasons and never for sole purposes of this study. Using endoscopic guidance, samples were collected directly from the osteomeatal complex, put on ice, and transferred to the lab. Endoscopically directed sampling of secretions from the osteomeatal complex have been shown to be an accurate representation for the properties of secretions from inside the sinuses, at least in terms of bacterial presence (37,38). Secretions from the left and right osteomeatal complexes were combined and centrifuged at 1,100 rpm at 4 °C for 5 min. The supernatant was transferred to a clean tube and 1 ml of ice cold phosphate-buffered saline was added. The same supernatant was centrifuged at 1,500 rpm for 10 min at 4 °C. Samples were then desalted and concentrated on an Amicon Ultra-4 3K filter units (Millipore, Billerica, MA). Protein content was analyzed by a bicinchoninic acid (BCA) protein assay (Thermo Scientific, Rockford, IL). Eight secretion samples were used for proteomic analysis (three control, five CRS). Four additional samples (three control and one CRS), were used for Western analyses.
Proteomic Sample Preparation
A 100 μg of total protein for samples were resuspended (NuPAGE LDS with reducing agent) and subjected to one-dimensional SDS gel electrophoresis using a 4–12% Bis-Tris polyacrylamide gel (Life Technologies, Carlsbad, CA) at 200 V for 50 min. The gel was fixed with a 50% methanol, 45% ddh2O, and 5% acetic acid solution and stained with Bio-Safe Coomassie for protein visualization. Each gel lane was serially sliced into 33 segments and in-gel digested with trypsin as previously (39).
MS and Protein Identification
MS analysis and data processing was carried out using methods as previously reported by our group (39). Briefly, tryptic peptides were injected via an autosampler and loaded onto a C18 trap column followed by separation on a C18 reverse-phase column using an Eksigent nano-HPLC system (Dublin, CA). The Linear Ion Trap Mass Spectrometer (LTQ) (ThermoFisher Scientific, Waltham, MA) was operated in data-dependent mode with dynamic exclusion in which one cycle of experiments consisted of a full-MS (300–2,000 m/z) survey scan and five subsequent MS/MS scans of the most intense peaks. Each survey scan file was searched for protein identification using the Sequest algorithm in the Bioworks Browser software (ThermoFisher Scientific, version 3.3.1) against the Uniprot database indexed for human, fully tryptic peptides, two missed cleavages, and potential modification of oxidized methionine (15.9949 Da). Data generation parameters were Peptide Tolerance of 1.5 Da and Fragment Ion Tolerance of 1 Da. Search results were uploaded into ProteoIQ software (version 2.3.07, Premier Biosoft, Palo Alto, CA) and filtered based on the following criteria: XCorr > 1.9, peptides > 6 amino acids, 2 spectra per peptide, 0.98 peptide probability and 0.95 protein probability. Proteomics data were filtered using QSpec’s statistical algorithm to determine which proteins had significant fold-change values. Data were submitted through the QSpec web submission page (12) (Ann Arbor, MI). The QSpec algorithm, is tailored for the analysis of differential proteins expression using label-free spectra count data and small sample numbers (12). To account for blood and other contamination, data were normalized to total scan counts after subtraction of albumin, hemoglobin, and keratins. This was done in order to remove the likely confounding contribution of these overabundant proteins to the overall proteinacious content of the biospecimen. Dealing with blood contamination in proteomic analyses of biospecimens remains a standard problem.
Western Blotting
Western blot analysis was performed using established protocols in our laboratories (39,40). For mucin protein blotting, samples containing 40 µg total protein were separated by electrophoresis in 1.0% agarose gels and transferred to polyvinylidene fluoride (PVDF) membranes (Millipore, Billerica, MA). The positive control for MUC5B was human saliva to which protease inhibitor cocktail (Sigma, St Louis, MO) was added at a 1:100 dilution on collection and for MUC5AC was A549 lung carcinoma cell lysates. For MUC5AC immunostaining, the membrane was probed using LUM5-1 (41), a rabbit polyclonal anti-MUC5AC antibody generously provided to us by Dr Mehmet Kesimer, University of North Carolina at Chapel Hill at a 1:4,000 dilution in a 2.5% milk solution in phosphate-buffered saline for 1 h at room temperature followed by incubation at 4 °C overnight. The antibody (LUM5-1 epitope) was raised against the sequence RNQDQQGPFKMC in the cysteine rich nontandem repeat domains of MUC5AC. MUC5B expression was detected with a rabbit polyclonal α-MUC5B H-300 (Santa Cruz Biotechnology, Dallas, TX) at 1:300 dilution. The respective intensities of positive bands relative to the positive control were densitometrically semiquantified with Quantity One 4.3.1 image processing software (BioRad, Hercules, CA) using equal sized box-shaped markers.
Statistical Analysis
Protein fold change was considered to be significant if P value (or false discovery rate) was less than or equal to 0.0001. Additionally, only proteins with an absolute fold change greater than or equal to 1.5 were taken into account. Mann–Whitney tests for nonparametric data were used for densitometry analyses of western blots.
Statement of Financial Support
This work was supported by a Flight Attendant Medical Research Institute Clinical Innovator Award, Miami, FL.
Disclosure
The authors declare no conflict of interest.
References
Benson V, Marano MA . Current estimates from the National Health Interview Survey, 1992. Vital Health Stat 1994;10:1–269.
Gliklich RE, Metson R . The health impact of chronic sinusitis in patients seeking otolaryngologic care. Otolaryngol Head Neck Surg 1995;113:104–9.
Rose MC, Voynow JA . Respiratory tract mucin genes and mucin glycoproteins in health and disease. Physiol Rev 2006;86:245–78.
Peña MT, Aujla PK, Patel KM, Zalzal GH, Rose MC . Immunohistochemical analyses of MUC5AC mucin expression in sinus mucosa of children with sinusitis and controls. Ann Otol Rhinol Laryngol 2005;114:958–65.
Peñia MT, Aujla PK, Zudaire E, et al. Localization and expression of MUC5B and MUC7 mucins in pediatric sinus mucosa. Ann Otol Rhinol Laryngol 2007;116:389–97.
Viswanathan H, Brownlee IA, Pearson JP, Carrie S . MUC5B secretion is up-regulated in sinusitis compared with controls. Am J Rhinol 2006;20:554–7.
Zang HR, Wang T, Li YC, et al. [A histopathological study: chronic rhinosinusitis in adolescents versus adults]. Zhonghua Yi Xue Za Zhi 2009;89:1975–8.
Casado B, Pannell LK, Viglio S, Iadarola P, Baraniuk JN . Analysis of the sinusitis nasal lavage fluid proteome using capillary liquid chromatography interfaced to electrospray ionization-quadrupole time of flight- tandem mass spectrometry. Electrophoresis 2004;25:1386–93.
Casado B, Pannell LK, Iadarola P, Baraniuk JN . Identification of human nasal mucous proteins using proteomics. Proteomics 2005;5:2949–59.
Ghafouri B, Ståhlbom B, Tagesson C, Lindahl M . Newly identified proteins in human nasal lavage fluid from non-smokers and smokers using two-dimensional gel electrophoresis and peptide mass fingerprinting. Proteomics 2002;2:112–20.
Benson LM, Mason CJ, Friedman O, Kita H, Bergen HR 3rd, Plager DA . Extensive fractionation and identification of proteins within nasal lavage fluids from allergic rhinitis and asthmatic chronic rhinosinusitis patients. J Sep Sci 2009;32:44–56.
Choi H, Fermin D, Nesvizhskii AI . Significance analysis of spectral count data in label-free shotgun proteomics. Mol Cell Proteomics 2008;7:2373–85.
Cao R, Wang TT, DeMaria G, Sheehan JK, Kesimer M . Mapping the protein domain structures of the respiratory mucins: a mucin proteome coverage study. J Proteome Res 2012;11:4013–23.
Kesimer M, Kirkham S, Pickles RJ, et al. Tracheobronchial air-liquid interface cell culture: a model for innate mucosal defense of the upper airways? Am J Physiol Lung Cell Mol Physiol 2009;296:L92–L100.
Ali M, Lillehoj EP, Park Y, Kyo Y, Kim KC . Analysis of the proteome of human airway epithelial secretions. Proteome Sci 2011;9:4.
Min-man W, Hong S, Zhi-qiang X, Xue-ping F, Chang-qi L, Dan L . Differential proteomic analysis of nasal polyps, chronic sinusitis, and normal nasal mucosa tissues. Otolaryngol Head Neck Surg 2009;141:364–8.
Kim JK, Kim SS, Rha KW, et al. Expression and localization of surfactant proteins in human nasal epithelium. Am J Physiol Lung Cell Mol Physiol 2007;292:L879–84.
Chen Y, Thai P, Zhao YH, Ho YS, DeSouza MM, Wu R . Stimulation of airway mucin gene expression by interleukin (IL)-17 through IL-6 paracrine/autocrine loop. J Biol Chem 2003;278:17036–43.
Casado B . Proteomics for nasal secretion analysis. Curr Allergy Asthma Rep 2004;4:224–9.
Bingle CD, Craven CJ . PLUNC: a novel family of candidate host defence proteins expressed in the upper airways and nasopharynx. Hum Mol Genet 2002;11:937–43.
Campos MA, Abreu AR, Nlend MC, Cobas MA, Conner GE, Whitney PL . Purification and characterization of PLUNC from human tracheobronchial secretions. Am J Respir Cell Mol Biol 2004;30:184–92.
Bingle L, Cross SS, High AS, et al. SPLUNC1 (PLUNC) is expressed in glandular tissues of the respiratory tract and in lung tumours with a glandular phenotype. J Pathol 2005;205:491–7.
Hartshorn KL, White MR, Mogues T, Ligtenberg T, Crouch E, Holmskov U . Lung and salivary scavenger receptor glycoprotein-340 contribute to the host defense against influenza A viruses. Am J Physiol Lung Cell Mol Physiol 2003;285:L1066–76.
Kang W, Reid KB . DMBT1, a regulator of mucosal homeostasis through the linking of mucosal defense and regeneration? FEBS Lett 2003;540:21–5.
Bikker FJ, Ligtenberg AJ, Nazmi K, et al. Identification of the bacteria-binding peptide domain on salivary agglutinin (gp-340/DMBT1), a member of the scavenger receptor cysteine-rich superfamily. J Biol Chem 2002;277:32109–15.
Eriksson C, Frängsmyr L, Danielsson Niemi L, et al. Variant size- and glycoforms of the scavenger receptor cysteine-rich protein gp-340 with differential bacterial aggregation. Glycoconj J 2007;24:131–42.
Thornton DJ, Davies JR, Kirkham S, et al. Identification of a nonmucin glycoprotein (gp-340) from a purified respiratory mucin preparation: evidence for an association involving the MUC5B mucin. Glycobiology 2001;11:969–77.
Al Badaai Y, DiFalco MR, Tewfik MA, Samaha M . Quantitative proteomics of nasal mucus in chronic sinusitis with nasal polyposis. J Otolaryngol Head Neck Surg 2009;38:381–9.
Tewfik MA, Latterich M, DiFalco MR, Samaha M . Proteomics of nasal mucus in chronic rhinosinusitis. Am J Rhinol 2007;21:680–5.
Upton DC, Welham NV, Kuo JS, Walker JW, Pasic TR . Chronic rhinosinusitis with nasal polyps: a proteomic analysis. Ann Otol Rhinol Laryngol 2011;120:780–6.
Wu X, Ghimbovschi S, Aujla PK, Rose MC, Peña MT . Expression profiling of inflammatory mediators in pediatric sinus mucosa. Arch Otolaryngol Head Neck Surg 2009;135:65–72.
Ghafouri B, Irander K, Lindbom J, Tagesson C, Lindahl M . Comparative proteomics of nasal fluid in seasonal allergic rhinitis. J Proteome Res 2006;5:330–8.
Fukami M, Stierna P, Veress B, Carlsöö B . Lysozyme and lactoferrin in human maxillary sinus mucosa during chronic sinusitis. An immunohistochemical study. Eur Arch Otorhinolaryngol 1993;250:133–9.
Johnstone DB, Zhang J, George B, et al. Podocyte-specific deletion of Myh9 encoding nonmuscle myosin heavy chain 2A predisposes mice to glomerulopathy. Mol Cell Biol 2011;31:2162–70.
Kang HK, Cox DW . Tandem repeats 3’ of the IGHA genes in the human immunoglobulin heavy chain gene cluster. Genomics 1996;35:189–95.
Tos M, Mogensen C . Mucus production in chronic maxillary sinusitis. A quantitative histopathological study. Acta Otolaryngol 1984;97:151–9.
Nadel DM, Lanza DC, Kennedy DW . Endoscopically guided sinus cultures in normal subjects. Am J Rhinol 1999;13:87–90.
Talbot GH, Kennedy DW, Scheld WM, Granito K ; Endoscopy Study Group. Rigid nasal endoscopy versus sinus puncture and aspiration for microbiologic documentation of acute bacterial maxillary sinusitis. Clin Infect Dis 2001;33:1668–75.
Preciado D, Goyal S, Rahimi M, et al. MUC5B Is the predominant mucin glycoprotein in chronic otitis media fluid. Pediatr Res 2010;68:231–6.
Berger JT, Voynow JA, Peters KW, Rose MC . Respiratory carcinoma cell lines. MUC genes and glycoconjugates. Am J Respir Cell Mol Biol 1999;20:500–10.
Thornton DJ, Carlstedt I, Howard M, Devine PL, Price MR, Sheehan JK . Respiratory mucins: identification of core proteins and glycoforms. Biochem J 1996;316 (Pt 3):967–75.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Saieg, A., Brown, K., Pena, M. et al. Proteomic analysis of pediatric sinonasal secretions shows increased MUC5B mucin in CRS. Pediatr Res 77, 356–362 (2015). https://doi.org/10.1038/pr.2014.187
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/pr.2014.187
This article is cited by
-
Lacticaseibacillus casei AMBR2 modulates the epithelial barrier function and immune response in a donor-derived nasal microbiota manner
Scientific Reports (2020)
-
Modelling upper respiratory tract diseases: getting grips on host-microbe interactions in chronic rhinosinusitis using in vitro technologies
Microbiome (2018)
-
Untargeted analysis of the airway proteomes of children with respiratory infections using mass spectrometry based proteomics
Scientific Reports (2018)
-
In vivo imaging of the Muc5b gel-forming mucin
Scientific Reports (2017)
-
Purification and characterization of microRNAs within middle ear fluid exosomes: implication in otitis media pathophysiology
Pediatric Research (2017)