Abstract
Newborns adjust to the extrauterine environment by developing intestinal immune homeostasis. Appropriate initial bacterial colonization is necessary for adequate intestinal immune development. An environmental determinant of adequate colonization is breast milk. Although the full-term infant is developmentally capable of mounting an immune response, the effector immune component requires bacterial stimulation. Breast milk stimulates the proliferation of a well-balanced and diverse microbiota, which initially influences a switch from an intrauterine TH2 predominant to a TH1/TH2 balanced response and with activation of T-regulatory cells by breast milk–stimulated specific organisms (Bifidobacteria, Lactobacillus, and Bacteroides). As an example of its effect, oligosaccharides in breast milk are fermented by colonic bacteria producing an acid milieu for bacterial proliferation. In addition, short-chain fatty acids in breast milk activate receptors on T-reg cells and bacterial genes, which preferentially mediate intestinal tight junction expression and anti-inflammation. Other components of breast milk (defensins, lactoferrin, etc.) inhibit pathogens and further contribute to microbiota composition. The breast milk influence on initial intestinal microbiota also prevents expression of immune-mediated diseases (asthma, inflammatory bowel disease, type 1 diabetes) later in life through a balanced initial immune response, underscoring the necessity of breastfeeding as the first source of nutrition.
Similar content being viewed by others
Main
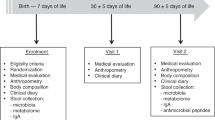
Although it is unclear to what extent mother’s microbiota during pregnancy influences the newborn infant, it is generally accepted that the newborn infant leaves an almost germ-free intrauterine environment to enter a highly contaminated extrauterine state, containing an abundance of potentially pathologic organisms which can evoke numerous infectious conditions. To prepare for the “shock” of birth, the fetus, during the last trimester of pregnancy, has swallowed numerous amniotic trophic factors which can help stimulate the maturation of immune defenses. Thus, at the time of birth of a full-term infant, the newborn has the capacity to protect itself in the external environment. However, for the efferent component of host defense, leading to intestinal homeostasis, to be generated initial bacterial intestinal colonization must be adequate. Colonizing bacteria communicate through a symbiotic relationship with intestinal epithelial and immunologic structures to develop the actual activation of host defense ( Figure 1 ) (1). A number of factors help to determine adequate initial colonization in the neonate, including a major determinant, the breastfed diet. In this review, the importance of initial colonization will be reviewed and the vital contribution made by exclusive breastfeeding underscored.
Diagram of mucosal immune defenses expressed in the human neonatal intestine as a function of gestational age. (a) All components of mucosal immune function are mature at birth in the term human infant. (b) However, the stimulus of initial colonization is needed before defenses become operational. Selected intestinal immune defenses include (1) a specialized epithelium (M cell) overlying, (2) Peyer’s patches containing aggregates of lymphoid elements, (3) interstitial lymphocytes, and (4) intraepithelial lymphocytes resulting in efferent immune responses. (Reprinted from: Robert M. Insoft, Ian R. Sanderson, W. Allan Walker. Development of immune function in the intestine and its role in neonatal diseases. Pediatr Clin 43:551–71; copyright © 1996, with permission from Elsevier.)
Initial Colonization
The full-term, vaginally delivered newborn infant leaves an almost germ-free intrauterine environment and on passing through the birth canal ingests maternal colonic and vaginal microbiota which acts as a nidus for initial colonization. During the next year, the introduction of oral feedings, e.g., breast vs. formula feeding, and weaning to solid foods completes the process resulting in 1014 bacteria colonizing principally the distal small intestine and colon with diverse microbiota (2). Normal colonization is dependent on many factors including a genetic contribution and the final signature microbiome is unique to the individual. The initial colonization process has an important stimulating effect on the developing intestine, particularly mucosal host defense.
Improper colonization, dysbiosis (3), delays the process and results in increased susceptibility to newborn and infant infections and altered immune homeostasis leading to the expression of immune-mediated disease (4,5) and metabolic diseases (6) later in life.
Infants who are born prematurely or by cesarean section have a disrupted phase I of colonization with the nidus of initial colonizing bacteria coming from other sources, e.g., maternal skin or the microbiota in the hospital NICU, and the balance of bacterial phyla and diversity of individual species are strikingly reduced (7). The ensuing dysbiosis has been associated with an increase in diseases such as asthma (8) and celiac disease (9). In addition, perinatal use of antibiotics, particularly in premature infants, can interfere with the balance of bacterial phyla and diversity of individual species and makes the newborn much more susceptible to infectious inflammatory diseases (10,11). Long-term follow up on the use of antibiotics during the first year of life has shown an association with an increased incidence of inflammatory bowel disease (12), asthma (13), and increased weight gain leading to adult obesity (14). These studies suggest that optimal initial colonization is critical in the adjustment of the newborn to the extrauterine environment.
Initial Colonization and Host Defense
Colonizing bacteria can act as an ancillary body organ. There are 10 times the number of bacteria in the intestine as there are eukaryotic cells in the body and 100-fold more bacterial genes than human genes in the body (15). Furthermore, colonizing bacteria collectively are more metabolically active than the human liver. Accordingly, this “ancillary organ” is essential for normal gastrointestinal development and function. This is particularly true for the neonate’s adjustment to the extrauterine environment through a symbiotic relationship (16).
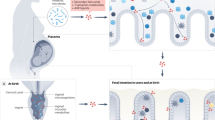
Several recent reviews have emphasized the vital importance of symbiosis to the development and maintenance of intestinal host defense (17,18). In a review in Nature Reviews Immunology (19), Artis underscores the important contributions of commensal bacteria to the intestinal epithelial cell barrier ( Figure 2 ). He suggests that not only do intestinal bacteria stimulate lymphoid elements, but they also influence components of the single layer of epithelium that makes up the mucosal barrier by stimulating the development of microvilli (19,20) and tight junctions (21). These bacteria also activate the release of mucin from epithelial goblet cells to form the glycocalyx as a physical and antibacterial barrier (19,20,21,22). In addition, he provides evidence that intestinal bacteria can stimulate epithelial cells and Paneth cells to release antibacterial peptides into the glycocalyx and lumen which helps to regulate the composition of colonizing intestinal bacteria (19,23). In a seminal study that underscores how colonizing bacteria influence intestinal protective function, germ-free mice were colonized with a single human intestinal commensal bacterium, Bacteroides thetaiodamicronm, and after colonization the intestinal epithelial RNA was analyzed for altered gene expression (24). Extensive activation of epithelial genes that influenced mucosal barrier function was noted including an upregulation of the polymeric IgA (pIgA) receptor. This study illustrated how initial bacterial colonization affects the small intestine of newborns at birth, since the newborn infant is like a germ-free rodent receiving its first colonization.
The intestinal epithelial-cell barrier. Simple columnar epithelial cells exhibit physical and biochemical adaptations to microbial colonization to maintain barrier integrity including actin-rich microvillar extensions (a), epithelial-cell tight junctions (b), apically attached and secreted mucins that form a glycocalyx (c), and the production of various antimicrobial peptides (d). Specialized intestinal epithelial cells known as M (microfold) cells overlie Peyer’s patches and lymphoid follicles to facilitate luminal sampling. M cells exhibit reduced mucin secretion and have modified apical and basolateral surfaces (e) to promote uptake and transport of luminal contents to professional antigen-presenting cells that inhabit the subepithelial dome (SED) of the Peyer’s patches and lymphoid follicles (f). Specialized dendritic cell (CD) subsets can also extend dendrites between the tight junctions of intestinal epithelial cells to sample luminal contents (g). (Reprinted with permission from Macmillan Publishers Ltd: Nat Rev Immunol (ref. 19), copyright © 2008.)
In a more recent review by Hooper et al. (25), a reciprocal relationship between the colonizing intestinal bacteria and the mucosal immune system is stressed. She emphasizes the contribution that mucosal immune defenses have made in defining the composition of intestinal bacteria, not only within the intestinal lumen, but also on the luminal surface. For example, an absence of/or reduction in pIgA levels results in a strikingly different microbiome in mice, which in some instances can contribute to the phenotypic expression of disease. For example, in a recent study the Toll-like receptor-5 (TLR-5) gene, located on the basolateral surface of intestinal enterocytes (ligand-bacterial flagella), was knocked out, resulting in an altered intestinal microbiota leading to the phenotypic expression of obesity and metabolic syndrome (26). The intestinal microbiota was implicated in this disease, because a transfer of the microbiome from TLR-5 knockout mouse to a genetically identical, but germ-free mouse, resulted in the same expression of disease. Accordingly, it has been proposed that the microbiome in numerous disease states (inflammatory bowel disease (IBD), type 1 diabetes, and necrotizing enterocolitis) may function as an intermediary determinant of the phenotypic disease expression (27).
Colonizing commensal bacteria can affect the development of both the innate and adaptive immune system. For example, pattern recognition receptors on enterocytes and mucosal immune cells interact with microbial surface-expressed or secreted ligands to evoke an innate, self-limited inflammatory response for pathogen mucosal penetration prevention (7,28). To prevent continuous inflammation of the intestine due to exposure of large numbers of colonizing bacteria, the absolute expression of these receptors, particularly the luminal surface expression of TLR-2 and 4, is reduced when the neonate leaves the intrauterine environment (29). In addition, in the mature intestine several mechanisms have evolved to distinguish an appropriate innate immune response to pathogens (in order to prevent penetration of the epithelium) from harmless commensal bacteria (30). Unfortunately, the immature enterocyte has not yet developed these inhibitory mechanisms and fetal/premature intestine responds to all colonizing bacteria with inflammation (31), undoubtedly a contributing factor to necrotizing enterocolitis. For example, a commensal bacterium (Salmonella typhimurium, nonpathogenic strain) interacting with the enterocyte’s luminal surface can interfere directly with the NFκB transcriptional inflammatory cytokine pathway by inhibiting the ubiquitination of phosphorylated IκB, a molecule which binds NFκB and prevents its entrance into the nucleus for transcription (32). In addition, B. thetaiodamicron interacting with the mature enterocyte luminal surface can evoke a nuclear reaction that disassociates NFκB from its transcription site on the chromosome and transports it out of the nucleus (33). In addition, several intracellular pattern recognition receptors, e.g., TLR-9, can also stimulate cellular proliferation of damaged epithelium due to pathogen penetration (34).
In like manner, intestinal commensals can influence the adaptive mucosal immune system. In an important observation published a decade ago (35,36), it has been shown that dendritic cells underlying the intestinal epithelium can extend appendages between enterocytes into the intestinal lumen. Commensal luminal bacteria interacting through pattern recognition receptors can stimulate these dendritic cells to release cytokines, which create a microenvironment that allows naïve T-helper cells (TH0) to create balanced subclasses of TH cells (TH1, TH2, TH17, and THreg) ( Figure 3 ). Newborn infants are born with a TH2 bias to protect them from intrauterine rejection (8,37). It takes appropriate colonization of the newborn to shift this bias to balanced TH cell subsets for effective adaptive immunity. This is underscored by a persistent TH2 response in newborn rat pups born to a germ-free mother (38). In like manner, Andrew Macpherson (39) has shown that the pIgA response to intestinal bacteria is contained within the mucosal compartment of the intestine. Commensal bacteria in the lumen or on the intestinal surface are taken up by dendritic cells that penetrate the epithelium into the lumen or underlie the epithelial surface of microfold cells, which in turn overlie Peyer’s patches. These dendritic cells laden with bacteria migrate to the mesenteric lymph node where components of the engulfed bacteria are presented to B lymphocytes resulting in maturation to antibody-specific activated pIgA plasma cells which release pIgA that is shuttled across the epithelium to help contain bacterial attachment and penetration. These two examples underscore the importance of bacterial colonization in the development and maintenance of mucosal immune function.
Cartoon depicting subepithelial dendritic cells extending their appendages between enterocytes into the intestinal lumen. These appendices, displaying Toll-like receptors, are activated by colonizing bacteria and secrete cytokines that activate naïve T-helper cells (TH0) to mature into a balanced T-helper cell response (TH1, TH, TH17, and Treg). (Reproduced with permission from ref. 8.)
We recognize that there is an increase in immune-mediated diseases (allergy and autoimmune diseases) and a decrease in infectious diseases over the last half century. This shifting disease paradigm explained by the “Hygiene Hypothesis” (40) has evolved over the last decade to principally implicate the Western lifestyle, particularly the Western diet (this will be discussed in detail later), in disrupted bacterial colonization. A development of a normal mucosal immune system, particularly tolerance, is paramount for the infant and the child in later life with regard to the absence of disease expression. In the absence of tolerance development or when tolerance is lost in later life due to the excessive use of antibiotics; for example, the individual is more likely to develop allergy or autoimmune disease (41). Sudo et al. (42) have reported that tolerance does not develop in germ-free animals but only with adequate colonization. Subsequent studies by Scandinavian investigators (43) demonstrated that newborn colonization was necessary for complete tolerance. Recently, neonatal colonization of rodents has been shown to contain the activation of natural killer T cells which is involved in gut inflammation and lung allergy (44). Accordingly, early colonization is necessary for appropriate development of mucosal immune defense, which in turn is required for prevention of immune-mediated diseases later in life. In addition, clinical and animal studies have shown the loss of tolerance with extensive antibiotic treatment in adults (18,45,46). These observations may help to explain late onset expression of autoimmune disease or allergy, now commonly seen by physicians.
In recent years, several studies have suggested that specific bacteria can contribute to the development of tolerance. These observations are relevant, since many of these specific bacteria are preferentially increased in the intestine during the early stages of colonization with breastfed infants (discussed in detail in section “Breastfeeding and Intestinal Colonization”). For example, Kasper’s group has reported that a carbohydrate on the surface of Bacteroides fragillis, polysaccharide A, can interact with the Toll receptor 2 (TLR-2) on intestinal dendritic cells to preferentially activate cytokines, which create a microenvironment that facilities the preferential proliferation of FOXP3 T cells (T regulatory cells) in the intestinal lamina propria (47), as shown by germ-free transfer studies of polysaccharide A positive and negative B. fragillis (15,48), and polysaccharide A ingestion alone (49). In like manner, certain species of Clostridia have been shown to increase T reg cells and to protect against IgE-mediated disease (50). These observations strongly suggest that not only is appropriate colonization in general necessary in the newborn period, but specific organisms seen in breastfed infants (51) may also be necessary to achieve early tolerance to innocuous antigens and bacteria, thereby preventing the expression of allergy and autoimmune diseases later in life.
Nutrition and Intestinal Colonization
An important environmental determinate of the composition of bacteria colonizing the intestine is nutrition. The human diet provides energy to increase the proliferation of bacteria colonizing the intestine, particularly anaerobic bacteria residing in the distal small intestine, cecum, and the colon. The most recent iteration of the so called “Hygiene Hypothesis” is that Western life style, particularly Western diet, has influenced changes in colonizing bacteria which are implicated in the striking increase in many chronic diseases (52). Alterations in diet can influence the proliferation of large families of bacteria (phyla) based on the capacity of prokaryotic cells to adapt quickly to their nutritional environment (53). A recent publication in Nature suggests that large families of microbiota, called enterotypes, evolve based on functional needs of the intestine, e.g., diet, drug metabolism, etc. (54). For example, this association has recently been demonstrated in a study of large numbers of adult volunteers who, under a clinical research center supervision, were given either an exclusive simple or complex carbohydrate diet or a diet rich in protein or fats (saturated vs. unsaturated) for prolonged periods of time (55). Over this long period of dietary change (approximately 60 d) but not over shorter periods (1–2 wk), the intestinal colonizing bacteria evolved into a specific enterotype. In addition, a basic study of 10 cloned commensal bacteria, grown in media with the energy source protein, carbohydrate, or fat (56), showed a striking impact on bacterial proliferation and bacterial gene expression due to the specific diet. Finally, a recent clinical study comparing the microbiome of two populations of children showed striking differences depending on their diet. Intestinal microbiomes of African children ingesting an all vegetarian, high fiber diet were compared with age-matched children consuming a Western diet in Florence, Italy. Their microbiomes were shown to be strikingly different (57). Although not specifically studied, the spectrum of disease in these two populations is remarkably different suggesting that dietary influence on the intestinal microbiome may affect the child’s health and disease burden. Finally, in an epidemiologic study of Japanese citizens, an increased expression of a classic Westernized diseases, e.g., Crohn disease, paralleled changes in the Japanese diet from a typical Eastern to a Westernized diet with increase ingestion of animal protein and saturated fat (58). These studies strongly suggest that diet can influence the intestinal microbiome and disease burden.
Breastfeeding and Intestinal Colonization
As stated above, nutrition is an important environmental determinate of gut microbiota. This observation is most apparent at birth with the first time exposure to oral feedings. Furthermore, several recent publications suggest that breastfeeding protection is primarily mediated through bacterial colonization (59). Since newborn intestinal microbiota is in a formative stage of development, modifications may have an important and timely impact on development of intestinal immune function. An appropriate maturation of both mucosal immune function and initial bacterial colonization is very important to immune homeostasis and short-/long-term health of the infant (18,601). Furthermore, an initial transient proliferation of health promoting bacteria, e.g., “pioneer” bacteria, in a timely manner in the early neonatal period may be very important to intestinal host defense. In subsequent sections, the evidence to support the importance of breastfeeding in initial intestinal colonization will be systematically reviewed (60).
The Bifidogenic Effect of Breastfeeding
For over 40 y, it has been known that breastfeeding, particularly in the first weeks of neonatal life, can stimulate the active proliferation of various strains of Bifidobacteria, e.g., the bifidogenic effect. A study reported in Pediatrics in 1983 (61), using standard culture techniques, compared microbiota from infants fed breast milk vs. infant formula. Over the first 3 wk, the breastfed infants have a striking increase in Bifidobacteria and Lactobacillus species compared with formula-fed infants who had a more mature microbiota pattern containing organisms such as enterobacteria and enterococci. The authors suggested that breastfeeding created an intestinal milieu conducive to the growth of these bifidogenetic organisms. Breastfed infant’s intestinal content was noted to be acidic (pH 5.0) and contain an abundance of short-chain fatty acids, particularly lactic acid, whereas with formula-fed infants intestinal content was more alkaline (pH 7.1) and had lesser levels of short-chain fatty acids. In subsequent studies, the high content of oligosaccharides (approximately 8% of total breast milk nutrients) was identified as the principal breast milk factor affecting the intestinal milieu and therefore stimulating Bifidobacteria activation (62). These oligosaccharides pass undigested through the small intestine and enter the colon where they are fermented by endogenous microbiota (63). Extensive observations over the last two decades have established the impact of breast milk, particularly oligosaccharides, on the microbial composition of the intestine and its effect on gut development, particularly immune homeostasis (37,62). The term prebiotic has been used to describe the endogenous bacterial stimulatory effect of breast milk oligosaccharides (4).
The Breastfeeding Effect on Other Organisms
Breast milk factors, also effect other health promoting microorganisms such as Lactobacillus, Bacteroides, and Clostridia, which have been shown to be important during the neonatal period in activating immunologic functions such as tolerance (64), mucus production (65), tight junction expression (19,66), and TH cell balance (17,67), necessary in the immature intestine to provide mucosal barrier homeostasis. Furthermore, the short-chain fatty acids (acetate, proprionic, and butyrate) produced by bacterial fermentation of complex carbohydrates have also been shown to provide important gut functions. In addition to being absorbed by colonic cells as a source of energy, these short-chain fatty acids have also been shown to provide important immune protective function. For example, a recent publication has shown that the short-chain fatty acid, acetate, produced by fermentation of “pioneer” bacteria interacts with a receptor on T reg cells in the lamina propria to stimulate proliferation (68). Several functions for butyrate have been reported, including activation of immunologic molecules through epigenetic mechanisms (69).
Metagenomic analysis of colonizing bacteria in breastfed infants identified several health promoting, anaerobic bacteria earlier in the neonatal period than previously reported (70) including Bacteroides, Lactobacillus, and Clostridia. Fortuitously, these individual species have been shown to have important, unique functions in the developing mucosal immune system as discussed above, which appear to affect the newborn’s immune system in the very important postpartum period. Furthermore, this early activation of the mucosal immune system appears to provide lifelong protection against expression of disease (8,18,37).
Recent studies have expanded the importance of breast milk oligosaccharides in bacterial stimulation. When breast milk oligosaccharides, compared with commercial oligosaccharides, are used as the only source of carbon in the culture of Bifidobacterium longum subsp. infantis, not only is there bacterial proliferation, but bacterial genes are stimulated, which have a greater effect on enterocyte protection (71,72). In studies from this laboratory (73), we have identified a secreted product from the same B. infantis which on partial characterization is likely a glycan of less than 30 kD size. This factor has an anti-inflammatory effect in a fetal small intestinal cell line (H4) and functions by causing a maturation of the innate inflammatory immune response to an inflammatory stimulus with a reduction in the expression of TLR-4, its signaling molecules and its transcription factor NFκB genes and an increase in negative regulators of this response (TOLLIP, SIGIIR, and A-20) as seen in mature enterocytes (73,74). Finally, a recent study of the molecular response of intestinal bacteria to breast vs. infant formula feeding has shown that the products of breastfed intestinal bacteria have a greater effect on protective genes in enterocytes than products of formula-fed infants (75). Collectively, these recent studies suggest that a major protective impact of breastfeeding on newborn intestinal immune function may be mediated through intestinal bacteria.
Other Breast Milk Factors that Influence Intestinal Colonization
As stated previously, intestinal immune function can influence the composition of colonizing bacteria. The newborn’s mucosal immune system host defenses are not entirely operational at birth or in the neonatal period. Breast milk, a dynamic fluid, provides passive protective functions to the deficient newborn. For example, in early studies from this laboratory, we have shown that pIgA as a function of total breast milk protein decreased as the infant’s own endogenous pIgA increased over the first month of life (8,16). At various stages of mucosal immune development, breast milk provides passive protective factors such as pIgA, antibacterial peptides (defensins), and components of the innate immune response (soluble TLR-2 and 4, CD14 and MD2). These passive factors allow the newborn to protect itself from invading pathogens. These factors also contribute to the composition of luminal and enterocyte surface bacteria. For example, a decrease in pIgA levels in breast milk has been shown to result in a different intestinal microbiota than with adequate pIgA levels (59,76). This difference in bacterial composition has also been shown after reduction in soluble components of the innate immune response (77). Other protective factors in breast milk, e.g., lactoferrin, lysozyme, defensins, shown to be protective in young breastfed infants also cause protection through their influence on intestinal microbiota (23,78). Thus, as we understand the importance of initial intestinal colonization in the newborn and older child’s health, we realized that much of this effect is mediated through breast milk’s influence on intestinal microbiota, the quintessential dietary environmental factor in colonization.
The Breast Milk Microbiome
It has long been known that breast milk contains microorganisms, originally thought to be due to contamination by skin microbes from mother’s areola or from bacteria present in the infant’s mouth. Over the last few years with the introduction of molecular analysis of the microbiome in breast milk, microbiota from mother’s intestine are also shown in breast milk (79). Human and animal studies suggest that microorganisms from the maternal gut can be taken up by an increasingly permeable intestine due to hormonal effects on enterocyte tight junctions in the later stages of pregnancy and into early lactation. These migrant intestinal bacteria are engulfed by lymphoid cells which home to the engorged breast and are then secreted into breast milk where bacteria are released. Evidence in human lactating mothers shows large quantities of circulating lymphoid cells with engulfed bacteria. In animal studies, labeled bacteria placed into the maternal gut of lactating rodents appear in the breast milk. These studies suggest the presence of a microbial enteromammary circulation during lactation. Although these maternal gut microbes represent a small number of bacteria (103/cc breast milk) compared colonizing bacteria that exist, they may represent another inoculum for the intestinal colonization process, particularly in the presence of breast milk oligosaccharides acting as prebiotics. The actual role for breast milk microbes, however, remains to be established.
Dysbiosis and Disease States
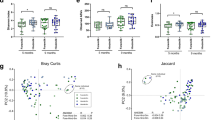
As stated above, there are many circumstances in which a disrupted environment can influence initial colonization (mode of delivery, prematurity, and antibiotics). Such a disruption in appropriate colonizing bacteria leads to an alteration in phyla and less diverse individual bacterial species resulting in dysbiosis (3). A recent review article has underscored the difference in microbiota in the microbiome of children with diseases and their age-matched controls ( Figure 4 ) (4,5,6). At this point, we cannot evoke the dysbiosis as a cause or as an effect of the disease. However, as previously stated, fecal microbiota transplants from disease phenotypes of either animal models or humans to germ-free animals have resulted in a phenotypic expression of the disease. As stated, this observation implicates the dysbiotic microbiome in disease expression, but does not provide evidence for cause. Furthermore, breastfeeding has been repeatedly shown to influence initial colonization and breastfed infants have a lower incidence of immune-mediated disease suggesting an association, but not necessarily an effect. Although an important environmental factor in initial colonization, breastfeeding is not the only environmental factor involved and cannot, for example, account for inherent genetic defects. However, a published meta-analysis has strongly suggested that early exclusive breastfeeding can prevent diseases expressed later in life (asthma, celiac disease, obesity, etc.) ( Figure 5 ) (80).
Disease states reveal phylum-level differences compared with healthy controls. Comparisons of the relative abundances of predominant bacterial phyla in (a) inflammatory bowel disease (IBD), (b) type 2 diabetes, and (c) necrotizing enterocolitis (NEC) are matched with healthy controls are displayed in two columns. Fecal samples from infants with NEC and patients with type 2 diabetes were compared with healthy controls revealing a predominance of Proteobacteria in patients with NEC. Cecal samples from patients with IBD were compared with healthy controls, and relative abundances were assessed. Firmicutes—maroon; Actinobacteria—dark blue; Bacteroidetes—yellow; Verrucomicrobia—green; Fusobacteria—purple; Proteobacteria—light blue. (Reprinted with permission from Macmillan Publishers Ltd: Nat Rev Microbiol (ref. 5), copyright © 2011.)
Exclusive breastfeeding (≥4 mo) and asthma during the first 8 y of life among children in a birth cohort from Stockholm (n = 3,825). Adjusted for sex, birth weight, parental history of allergic disease, maternal smoking during pregnancy and/or at birth, and maternal age. Odds ratios and 95% confidence intervals (CIs) were estimated by using general estimated equation (GEEs), after exclusion of 474 children with symptoms of wheeze or eczema during the breastfeeding period. (Reprinted from: Inger Kull, Erik Melen, Johan Alm, et al. Breast-feeding in relation to asthma, lung function, and sensitization in young schoolchildren. J Allergy Clin Immunol 125;1013–19; copyright © 2010, with permission from Elsevier.)
Summary and Conclusions
This review of the association of breastfeeding with initial colonization and its importance in the infant/child’s health suggests that a critical new protective function for breastfeeding exists and opens up many new areas for investigation for both clinical and research studies. At the very least, it provides strong support for infants being breastfed, if at all possible, for the first 4–6 mo of life.
Statement of Financial Support
Supported by National Institutes of Health (NIH) grants: National Institute of Digestive, Diabetes and Kidney Diseases P30 DK040561 (W.A.W. and R.S.I.), P01 DK033506; and National Institute of Child Health and Human Development R01 HD012437, R01 HD059126, Bethesda, MD, United States.
Disclosure
There is no conflict of interest to disclose.
References
Insoft RM, Sanderson IS, Walker WA . The development of immune function within the human intestine and its role in neonatal diseases. Pediatr Clin North Am 1996;43:551–71.
Palmer C, Bik EM, DiGiulio DB, Relman DA, Brown PO . Development of the human infant intestinal microbiota. Plos Biology 2010;5:e1566–73.
Penders J, Thijs C, Vink C, et al. Factors influencing the composition of the intestinal microbiota in early infancy. Pediatrics 2006;118:511–21.
Gareau MG, Sherman PM, Walker WA . Probiotics and the gut microbiota in intestinal health and disease. Nat Rev Gastroenterol Hepatol 2010;7:503–14.
Johnson CL, Versalovic J . The human microbiome and its potential importance to pediatrics. Pediatrics 2012;129:950–60.
Ridaura VK, Faith JJ, Rey FE, et al. Gut microbiota from twins discordant for obesity modulate metabolism of mice. Science 2013;341;1069–70.
Round JL, Mazmanian SK . The gut microbiota shapes intestinal responses during health and disease. Nat Rev Immunol 2009;9:313–23.
Kaplan JL, Shi H, Walker WA . The role of microbes in developmental immunologic programming: convergence of the Barker and hygiene hypotheses. Pediatr Res 2011;9:465–72.
Sellitto M, Bai G, Serena G, et al. Proof of concept of microbiome-metabolome analysis and delayed gluten exposure on celiac disease autoimmunity in genetically at-risk infants. PLoS One 2012;7:e33387.
Cho I, Yamanishi S, Cox L, et al. Antibiotics in early life alter the murine colonic microbiome and adiposity. Nature 2012;488:621–7.
Cotton CM, Taylor S, Stoll B, Goldberg RN, Hansen NI, Sanchez PJ . Prolonged duration of initial empirical antibiotic treatment is associated with increased rates of necrotizing enterocolitis and death for extremely low birth weight infants. Pediatrics 2009;123:59–66.
Kronman MP, Zaoutis TE, Haynes K, Feng R, Coffin SE . Antibiotic exposure and IBD development among children: a population-based cohort study. Pediatrics 2012;130:e794–803.
Marra F, Marra CA, Richardson K, et al. Antibiotic use in children is associated with increased risk of asthma. Pediatrics 2009;123:1003–10.
Kaplan JL, Walker WA . Early gut colonization and subsequent obesity risk. Curr Opin Clin Nutr 2012;15:278–84.
Neish AS . Reviews in basic and clinical gastroenterology: microbes in gastrointestinal health and disease. Gastroenterology 2009;136:65–80.
Walker WA . Bacterial colonization, probiotics and the development of intestinal host defense. Funct Food Rev 2009;1:13–19.
Hill DA, Artis D . Intestinal bacteria and the regulation of immune cell homeostasis. Annu Rev Immunol 2010;28:623–67.
Rautava S, Luoto R, Salminen S, Isolauri A . Microbial contact during pregnancy, intestinal colonization and human disease. Nat Rev Gastroenterol Hepatol 2011;9:565–76.
Artis D . Epithelial-cell recognition of commensal bacteria and maintenance of immune homeostasis in the gut. Nat Rev Immunol 2008;8:411–20.
McAuley JL, Linden SK, Png CW, et al. MUC1 cell surface mucin is a critical element of the mucosal barrier to infection. J Clin Invest 2007;117:2313–24.
Shen L, Turner JR . Role of epithelial cells in initiation and propagation of intestinal inflammation. Eliminating the static: tight junction dynamics exposed. Am J Physiol Gastrointest Liver Physiol 2006;290:G577–82.
Frey A, Giannasca KT, Weltzin R, et al. Role of the glycocalyx in regulating access of microparticles to apical plasma membranes of intestinal epithelial cells: implications for microbial attachment and oral vaccine targeting. J Exp Med 1996;184:1045–59.
Salzman NH, Ghosh D, Huttner KM, Paterson Y, Bevins CL . Protection against enteric salmonellosis in transgenic mice expressing a human intestinal defensin. Nature 2003;422:522–6.
Hooper LV, Wong MH, Thelin A, Hansson L, Falk PG, Gordon JI . Molecular analysis of commensal host-microbial relationships in the intestine. Science 2001;291:881–4.
Hooper LV, Littman DR, Macpherson AJ . Interactions between the microbiota and the immune system. Science 2012;336:1268–73.
Vijay-Kumar M, Aitken JD, Carvalho FA, et al. Metabolic syndrome and altered gut microbiota in mice lacking toll-like receptor 5. Science 2010;328:228–31.
Smith MI, Yatsuneji T, Manary MJ, et al. Gut microbiomes of Malawian twin pairs discordant for Kwashiorkor. Science 2013;339:548–55.
O’Hara AM, Shanahan F . Gut microbiota: mining for therapeutic potential. Clin Gastroenterol Hepatol 2007;5:274–84.
Abreu MT, Fukata M, Arditi M . TLR signaling in the gut in health and disease. J Immunol 2005;174:4453–60.
Otte JM, Cario E, Podolsky DK . Mechanisms of cross hyporesponsiveness to toll-like receptor bacterial ligands in intestinal epithelial cells. Gastroenterology 2004;126:1054–70.
Claud EC, Lu L, Anton PM, Savidge T, Walker WA, Cherayil BJ . Developmentally-regulated IκB expression in intestinal epithelium and susceptibility to flagellin-induced inflammation. Proc Natl Acad Sci USA 2004;101:7404–8.
Neish AS, Gerwitz AT, Zeng H, et al. Prokaryotic regulation of epithelial response by inhibition of IκB-α ubiquitination. Science 2000;289:1560–3.
Kelly D, Campbell JI, King TP, et al. Commensal anaerobic gut bacteria attenuate inflammation by regulating nuclear-cytoplasmic shuttling of PPAR-γ and ReIA. Nat Immunol 2004;5:104–11.
Katakura K, Lee J, Rachmilewitz D, Li G, Eckmann L, Raz E . Toll-like receptor 9-induced type I IFN protects mice from experimental colitis. J Clin Invest 2005;115:695–702.
Cobbold SP, Adams E, Nolan KF, Regateiro FS, Waldmann H . Connecting the mechanisms of T-cell regulation: dendritic cells as the missing link. Immunol Rev 2010;236:203–18.
Niess JH, Brand S, Gu X, et al. CX3CR1-mediated dendritic cell access to the intestinal lumen and bacterial clearance. Science 2005;307:254–8.
Weng M, Walker WA . The role of gut microbiota in programming the immune phenotype. J Dev Orig Health Dis 2013;4:203–14.
Conroy ME, Shi HN, Walker WA . The long-term health effects of neonatal microbial flora. Curr Opin Allergy Clin Immunol 2009;9:197–201.
Macpherson AJ, Uhr T . Induction of protective IgA by intestinal dendritic cells carrying commensal bacteria. Science 2004;303:1662–5.
Guarner F, Bourdet-Sicard R, Brandtzaeg P, et al. Mechanisms of disease: the hygiene hypothesis revisited. Nat Clin Pract Gastroenterol Hepatol 2006;3:275–84.
Nicholson JK, Holmes E, Wilson ID . Gut microorganisms, mammalian metabolism and personalized health care. Nat Rev Microbiol 2005;3:431–8.
Sudo N, Sawamura S, Tanaka K, Aiba Y, Kubo C, Koga Y . The requirement of intestinal bacterial flora for the development of an IgE production system fully susceptible to oral tolerance induction. J Immunol 1997;159:1739–45.
Karlsson MR, Kahu H, Hanson LA, Telemo E, Dahlgren UI . Neonatal colonization of rats induces immunological tolerance to bacterial antigens. Eur J Immunol 1999;29:109–18.
Olszak T, An D, Zeissig S, et al. Microbial exposure during early life has persistent effects on natural killer T cell function. Science 2012;336:489–93.
Bashir ME, Louie S, Shi HN, Nagler-Anderson C . Toll-like receptor 4 signaling by intestinal microbes influences susceptibility to food allergy. J Immunol 2004;172:6978–87.
Pabst O, Mowat AM . Oral tolerance to food protein. Nat Rev Immunol 2012;5:232–9.
Mazmanian SK, Kasper DL . The love–hate relationship between bacterial polysaccharides and the host immune system. Nat Rev Immunol 2006;6:849–58.
Mazmanian SK, Round JL, Kasper DL . A microbial symbiosis factor prevents intestinal inflammatory disease. Nature 2008;453:620–5.
O’Mahoney C, Scully P, O’Mahoney D, et al. Commensal-induced regulatory T cells mediate protection against pathogen-stimulated NF-κB activation. Pathology 2008;4:e1000112.
Atarashi K, Tanoue T, Shima T, et al. Induction of colonic regulatory T cells by indigenous Clostridium species. Science 2011;331:337–41.
Hooper LV . Bacterial contributions to mammalian gut development. Trends Microbiol 2004;12:129–34.
Maslowski KM, Mackay CR . Diet, gut microbiota and immune responses. Nat Immunol 2011;12:5–9.
Hooper LV, Xu J, Falk PG, Midtvedt T, Gordon JI . A molecular sensor that allows a gut commensal to control its nutrient foundation in a competitive ecosystem. Proc Natl Acad Sci USA 1999;96:9833–8.
Arumugam M, Raes J, Pelletier E, et al. Enterotypes of the human gut microbiome. Nature 2011;473:174–80.
Wu GD, Chen J, Hoffmann C, et al. Linking long-term dietary patterns with gut microbial enterotypes. Science 2011;334:105–8.
Faith JJ, McNulty NP, Rey FE, Gordon JI . Predicting a human gut microbiota’s response to diet in gnotobiotic mice. Science 2011;333:101–4.
DeFillippo C, Cavalieri D, DiPaola M, et al. Impact of diet in shaping gut microbiota revealed by a comparative study in children from Europe and rural Africa. Prac Natl Acad Sci USA 2010;107:14691–6.
Shoda R, Matsueda K, Yamato S, Umeda N . Epidemiologic analysis of Crohn disease in Japan: increased dietary intake of n-6 polyunsaturated fatty acids and animal protein relates to the increased incidence of Crohn disease in Japan. Am J Clin Nutr 1996;63:741–5.
Rogier EW, Frantz AL, Bruno ME, et al. Secretory antibodies in breast milk promote long-term intestinal homeostasis by regulating the gut microbiota and host gene expression. Proc Natl Acad Sci USA 2014;111:3074–9.
Rautava S, Walker WA . Breastfeeding—an extrauterine link between mother and child. Breastfeed Med 2009;4:3–10.
Yoshioka H, Iseki K, Fujita K . Development and differences of intestinal flora in the neonatal period in breast-fed and bottle-fed infants. Pediatrics 1983;72:317–21.
Bode L . Human milk oligosaccharides: every baby needs a sugar mama. Glycobiology 2012;22:1147–62.
Sela DA, Mills DA . Nursing our microbiota: molecular linkages between bifodobacteria and milk oligosaccharides. Trends Microbiol 2010;76:7373–81.
Barnes MJ, Powrie F . The guts’ Clostridium cocktail. Science 2011;331:289–90.
Frey A, Giannasca KT, Weltzin R, et al. Role of the glycocalyx in regulating access of microparticles to apical plasma membranes of intestinal epithelial cells: implications for microbial attachment and oral vaccine targeting. J Exp Med 1996;184:1045–59.
Duerr CU, Hornef MW . The mammalian intestinal epithelium as integral player in the establishment and maintenance of host-microbial homeostasis. Semin Immunol 2012;24:25–35.
Geuking MB, Cahenzli J, Lawson MA, et al. Intestinal bacterial colonization induces mutualistic regulatory T cell responses. Immunity 2011;34:794–806.
Smith PM, Howitt MR, Panikov N, et al. The microbial metabolites, short-chain fatty acids, regulate colonic Treg cell homeostasis. Science 2013;341:569–73.
Weng M, Walker WA, Sanderson IR . Butyrate regulates the expression of pathogen-triggered IL-8 in intestinal epithelia. Pediatr Res 2007;62:542–6.
Jost T, Lacroix C, Braegger C, Chassard C . New insights into gut microbiota establishment in healthy breast fed neonates. PLoS One 2012;7:e44595–603.
Garrido D, Kim JH, German JB, Raybould HE, Mills DA . Oligosaccharide binding proteins from Bifidobacterium longum subsp. infantis reveal a preference for host glycans. PLoS One 2011;6:e17315.
Chichlowski M, De Lartigue G, German JB, Raybould HE, Mills DA . Bifidobacteria isolated from infants and cultured on human milk oligosaccharides affect intestinal epithelial function. J Pediatr Gastroenterol Nutr 2012;55:321–7.
Ganguli K, Meng D, Rautava S, Lu L, Walker WA, Nanthakumar N . Probiotics prevent necrotizing enterocolitis by modulating enterocyte genes that regulate innate immune-mediated inflammation. Am J Physiol Gastrointest Liver Physiol 2013;304:G132–41.
Weng M, Ganguli K, Zhu W, Shi HN, Walker WA . Conditioned media from Bifidobacteria infantis protects against Cronobacter seakzakii-induced intestinal inflammation in newborn mice. Am J Physiol Gastrointest Liver Physiol 2014; 306:G779–87.
Schwartz S, Friedberg I, Ivanov IV, et al. A metagenomic study of diet-dependent interaction between gut microbiota and host in infants reveals differences in immune response. Genome Biol 2012;13:r32.
Macpherson AJ, Geuking MB, Slack E, Hepfelmeier S, McCor KD . The habitat, double like, citizenship and forgetfulness of IgA. Immunol Rev 2012;245:132–46.
LeBouder E, Rey-Nores JE, Raby AC, et al. Modulation of neonatal microbial recognition: TLR-mediated innate immune responses are specifically and differentially modulated by human milk. J Immunol 2006;176:3742–52.
Newburg DS, Walker WA . Protection of the neonate by the immune system of developing gut and of human milk. Pediatr Res 2007;61:2–8.
Jeurink PV, van Bergenhenegouwen J, Jiménez E, et al. Human milk: a source of more life than we imagine. Benef Microbes 2013;4:17–30.
Kull I, Melen E, Alm J, et al. Breast-feeding in relation to asthma, lung function, and sensitization in young schoolchildren. J Allergy Clin Immunol 2010;125:1013–9.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Walker, W., Iyengar, R. Breast milk, microbiota, and intestinal immune homeostasis. Pediatr Res 77, 220–228 (2015). https://doi.org/10.1038/pr.2014.160
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/pr.2014.160
This article is cited by
-
Comparing microbiotas of foals and their mares’ milk in the first two weeks after birth
BMC Veterinary Research (2024)
-
Factors associated with stunting: gut inflammation and child and maternal-related contributors among under-five children in Hawassa City, Sidama Region, Ethiopia
BMC Nutrition (2023)
-
Feeding practice during infancy is associated with attention-deficit/hyperactivity disorder and autism spectrum disorder: a population-based study in South Korea
European Journal of Pediatrics (2023)
-
The gut microbiome of obese postpartum women with and without previous gestational diabetes mellitus and the gut microbiota of their babies
Diabetology & Metabolic Syndrome (2022)
-
Preterm birth and Kawasaki disease: a nationwide Japanese population-based study
Pediatric Research (2022)