Abstract
Introduction:
To examine the immune-modulatory effects of probiotics during early infancy, Bifidobacterium breve M-16V (B. breve) was administered to rat pups during the newborn or weaning period, and the expression of inflammatory genes was investigated using a cDNA microarray and real-time PCR.
Results:
After B. breve administration, significant increases in the numbers of Bifidobacterium in both the cecum and colon were confirmed during the newborn period. The numbers of upregulated and downregulated genes were greater during the weaning period than in the newborn period and were greatest in the colon, with fewer genes altered in the small intestine and the fewest in the spleen. The expression of inflammation-related genes, including lipoprotein lipase (Lpl), glutathione peroxidase 2 (Gpx2), and lipopolysaccharide-binding protein (Lbp), was significantly reduced in the colon during the newborn period. In weaning rat pups, the expression of CD3d, a cell surface receptor–linked signaling molecule, was significantly enhanced in the colon; however, the expression of co-stimulatory molecules was not enhanced.
Discussion:
Our findings support a possible role for B. breve in mediating anti-inflammatory and antiallergic reactions by modulating the expression of inflammatory molecules during the newborn period and by regulating the expression of co-stimulatory molecules during the weaning period.
Methods:
Gene expression in the intestine was investigated after feeding 5×108 cfu of B. breve every day to the F344/Du rat from days 1 to 14 (newborn group) and from days 21 to 34 (weaning group). mRNA was extracted from intestine, and the expression of inflammatory gene was analyzed by microarray and real-time PCR.
Similar content being viewed by others
Main
As part of the delivery process, the neonate leaves the mostly germ-free intrauterine environment and enters a highly contaminated extrauterine world. It is thought that the process of intestinal bacterial colonization occurs within the first few hours to days after birth, which initiates the rapid immunological development of the intestine. However, the precise mechanisms of this development are not well understood.
Probiotics are defined as “live microorganisms, which when administered in adequate amounts confer a health benefit to the host” (1). Lactic acid bacteria, such as lactobacilli and bifidobacteria, are among the best-known candidates for probiotics. In numerous human trials, although the precise mechanisms have not been determined, intake of lactobacilli and bifidobacteria has been reported to be beneficial with respect to stool frequency and stool consistency (2,3). The potential benefit of probiotics in the modulation and regulation of the immune response has also been reported in both humans and animals (4,5,6,7,8).
Bifidobacterium breve M-16V (B. breve) is a probiotic strain that was isolated from the fecal sample of a healthy baby. Oral administration of B. breve to mice with IgE-mediated hypersensitivity to cow milk or ovalbumin promotes a significant improvement in allergic symptoms (9,10). Although B. breve influences bacterial colonization in the intestine and enhances TGF-β1 signaling by regulating Smad7 expression in preterm infants (11,12), the precise mechanisms of these interactions have not been well established.
In this study, we investigated the immune-modulatory effects of B. breve during infancy. Gene expression in the rat colon, small intestine, and spleen was examined by microarray and real-time PCR after feeding 5 × 108 colony-forming units/animal B. breve to rat pups during the newborn (days 1–14) or weaning (days 21–34) period.
Results
Intestinal Flora
Intestinal microflora analysis by real-time PCR revealed a significant increase in Bifidobacterium and a decrease in Bacteroides populations, which were confirmed in the cecum and colon, after B. breve administration to newborn rats as compared to controls (P < 0.01 each) ( Figure 1a,b ). In contrast, there was not a significant difference in the bacterial density of the cecum or colon in weaning rats administered B. breve ( Figure 1c,d ).
Bacterial colonization. Density of bacteria that colonized (a) the cecum and (b) the colon on (c) day 15 (newborn period) and (d) day 35 (weaning period). Bacterial density is expressed as 16S rDNA copies/g fecal sample for total bacteria, Bifidobacterium, Bacteroides, and Lactobacillus in the log scale. Filled bars represent the B. breve group, and open bars represent controls. Data are presented as the mean + SD; n = 14, newborn period; n = 9, weaning period; **P < 0.01.
Microarray Analysis of Total Gene Expression
Gene expression was examined by microarray using RNA isolated from whole-thickness colon, small intestine, or spleen samples and compared in rats with and without B. breve administration during the newborn and weaning periods. The number of upregulated or downregulated genes with a fold change >1.3 and a P value < 0.05 after B. breve administration was relatively small. The numbers of upregulated and downregulated genes were greater during the weaning period than in the newborn period. Furthermore, the number of genes with significantly changed expression was greatest in the colon, next greatest in the small intestine, and least in the spleen ( Figure 2 ).
Number of genes with fold changes >1.3 and P values < 0.05, representing upregulated and downregulated genes in the colon, small intestine, and spleen during the newborn (open bars) or weaning period (filled bars) after B. breve administration. Microarray analysis was performed on pooled RNA from three rat pups for each group.
Functional Categories of Upregulated and Downregulated Genes
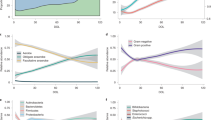
The genes with significantly altered expression were divided into seven immunological, functional categories based on the biological processes determined by Ingenuity Pathways Analysis. Histograms of the numbers of upregulated and downregulated genes in each functional category in colon, small intestine, and spleen during the newborn and weaning periods are shown in Figure 3 . The number of changed genes after B. breve administration was greater during the weaning period than in the newborn period. Furthermore, the greatest number of changed genes was observed in the colon, with fewer changed genes in the small intestine, and the fewest changed genes in the spleen. There were almost no changes in gene expression in the spleen after B. breve administration in either the newborn or weaning period.
Histograms presenting the numbers of genes upregulated and downregulated >1.3-fold in each functional category in the colon (open bars), small intestine (filled bars), and spleen (gray bars) after B. breve administration during the (a) newborn and (b) weaning periods. The functional category of each gene in the microarray was separated as shown in the lower part of the figure based on the Ingenuity Pathways Analysis. Microarray analysis was performed on pooled RNA from three rat pups per group.
Microarray Analysis of Typical Gene Expression
The comparison of microarray results in newborn rats with and without B. breve administration showed that 679, 168, and 23 genes in the colon had fold changes >1.2, 1.3, and 1.5, respectively. In addition, 1,071, 214, and 103 genes were significantly different between B. breve–fed rats and controls, with P values < 0.05, 0.01, and 0.005, respectively.
As a result, we identified four genes, Lpl, Gpx2, ClCa4, and Lbp, that had fold changes >1.5 and P values < 0.05 after B. breve administration in the newborn colon ( Table 1 ). With regard to these genes, a significant difference was observed only in the colon and not in the small intestine or spleen ( Figure 4a ). During the weaning period, no significant changes were observed in the expression of these four genes in the colon, small intestine, or spleen ( Figure 4b ).
Microarray analysis of gene expression in the colon, small intestine, and spleen after B. breve administration during the (a) newborn and (b) weaning periods. Filled bars represent the B. breve group, and open bars represent controls. The raw signal intensity of each group is shown as the mean + SD (n = 3). *P < 0.05. ClCa4, chloride channel, calcium-activated family member 4; Gpx2, glutathione peroxidase 2; LPS, lipopolysaccharide.
During the weaning period in rats administered B. breve, genes in the colon with fold changes >1.5 and P values < 0.01 were identified ( Table 2 ). The expression of most of these genes, as listed in Table 2 , was enhanced after B. breve administration rather than decreased as observed in newborn rats. Although the expression of CD3d was significantly increased in the colon, there were no accompanying significant changes in the expression of co-stimulatory molecules ( Figure 5b ). These gene changes were observed only in the colon and not in the small intestine or spleen. These changes were not observed in the colon, small intestine, or spleen during the newborn period ( Figure 5a ).
Microarray analysis of gene expression in the colon, small intestine, and spleen after B. breve administration during the (a) newborn and (b) weaning periods. Filled bars represent the B. breve group, and open bars represent controls. The raw signal intensity of each group is shown as the mean + SD (n = 3). **P < 0.01.
Real-Time PCR
Real-time PCR was performed to confirm the changes observed by microarray analysis with samples collected from 14 newborn and 9 weaning rat pups. In the newborn rat pups, the expression of Lpl, Gpx2, ClCa4, and Lbp was examined in the colon and compared to the expression of β-actin. The expression of Lpl, Gpx2, and ClCa4, but not Lbp, was significantly downregulated (P < 0.05; Figure 6 ).
Real-time PCR analysis of (a) lipoprotein lipase, (b) chloride channel, calcium-activated family member 4, (c) lipopolysaccharide-binding protein, and (d) glutathione peroxidase 2 gene expression in the colon after B. breve administration during the newborn period. Filled bars represent the B. breve group, and open bars represent controls. The relative gene expression in each group is shown as the mean + SD (n = 14). *P < 0.05.
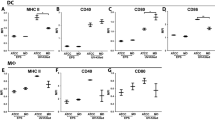
In the weaning rat pup colon, because the expression of CD3d was significantly enhanced according to microarray analysis, the expression of CD3d and its co-stimulatory molecules CD80, CD86, CD28, and CTLA-4 was examined by real-time PCR to confirm the effect of B. breve on signaling through these molecules. The expression of CD3d was significantly enhanced as compared to controls (P < 0.05), whereas there was no significant difference observed in the expression of co-stimulatory molecules in the colon ( Figure 7 ).
Real-time PCR analysis of (a) CD3d, (b) CD80, (c) CD86, (d) CD28, and (e) CTLA-4 gene expression in the colon after B. breve administration during the weaning period. Filled bars represent the B. breve group, and open bars represent controls. The relative gene expression in each group is shown as the mean + SD (n = 9). *P < 0.05.
Immunohistochemical Analysis
To confirm the effect of B. breve on signaling through CD3 and other co-stimulatory molecules, lymphoid follicles in the colon were stained with anti-CD3 and anti-CD86 antibodies ( Figure 8 ). CD3-positive cells were confirmed in both control and B. breve-administered weaning rat pups, and accumulation of staining with anti-CD3 antibody was confirmed in the B. breve group ( Figure 8c,d ) as compared with their controls ( Figure 8a,b ). The staining with anti-CD86 antibody was also confirmed, but it was light in both groups, and we found no significant difference in staining between the control ( Figure 8e,f ) and B. breve groups ( Figure 8g,h ).
Immunohistochemical analysis of two representative weaning rat intestines after receiving B. breve (c,d,g,h) and their controls (a,b,e,f). Intestinal lymphoid follicles were stained with (a–d) anti-CD3 and (e–h) anti-CD86 antibodies. Accumulation of staining with anti-CD3 antibody was confirmed in the B. breve group (c,d) as compared with controls (a,b). The staining with anti-CD86 antibody was light in both groups and showed no significant difference between the control (e,f) and B. breve groups (g,h). Original magnification ×400.
Discussion
In this study, we examined the immune-modulatory effects of probiotics during early infancy. B. breve was administered to rat pups during the newborn or weaning period, and the expression of inflammatory genes was investigated in the colon, small intestine, and spleen. Administration of B. breve during the newborn period was associated with a significant increase in the proportion of bifidobacteria and a decrease in the proportion of aerobic bacteria in both the cecum and the colon (P < 0.01 each). Furthermore, these changes were associated with altered expression of Lpl, Gpx2, and ClCa4 during the newborn period (fold change > 1.5, P < 0.05). Enhanced expression of Lpl has been observed in dextran sulfate sodium–induced colitis in hamsters (18). ClCa4 is considered to be an important molecule for the induction of diarrhea in cystic fibrosis (19). Gpx2 is a marker for the colonic inflammation that occurs in experimental colitis and inflammatory bowel diseases (20,21,22). It has been suggested that all of these molecules are involved in the pathogenesis of inflammatory changes in the intestine. Therefore, downregulating the expression of these molecules by the administration of B. breve may be beneficial for avoiding inflammation, e.g., necrotizing enterocolitis, in newborns.
The immune system of the gut is continually exposed to benign foreign proteins, including foods, and yet, at the same time, the immune system must be able to react to pathogenic foreign proteins. To address this dichotomy, the gut immune system has acquired mechanisms to avoid excessive reactions to foods, known as tolerance. Previous reports on the induction of tolerance through administration of probiotics have mainly been focused on lactobacilli (23). Therefore, we used B. breve to examine its effect on tolerance. Although a number of genes were significantly upregulated in the colon, including CD3d, the group of changed genes did not include those for co-stimulatory molecules. To further validate the effect of B. breve on CD3 and co-stimulatory molecules, lymphoid follicles in the colon were stained with anti-CD3 and anti-CD86 antibodies. Accumulation of staining with anti-CD3 antibody but not with CD86 antibody was confirmed in B. breve–administered weaning rat pups ( Figure 8 ).
Because administration of B. breve enhanced the expression of CD3 but not co-stimulatory molecules, this bacterium would be beneficial for introducing tolerance to benign foreign peptides during the weaning period.
The administration of B. breve influenced gene expression during the newborn period and the weaning period in distinct ways. The total number of genes upregulated or downregulated (fold change > 1.5, P < 0.05) was ~10 times larger in the weaning group than in the newborn group; moreover, the changed genes were mostly observed in the colon, with few present in the small intestine and even fewer present in the spleen. One possible explanation for these observations is related to the differences in the total amounts of antigens encountered in various regions of the intestine. Because a larger number of food proteins enter the intestinal lumen during the weaning period than in the newborn period, and because B. breve tends to stay longer in the colon than in the small intestine, B. breve may primarily influence local immune reactions in the colon but not systemic reactions. Furthermore, B. breve may not be a strong, systemic immune regulator but rather may be capable of mildly modulating immune reactions by introducing tolerance. However, there is an additional possibility that larger amounts of B. breve could be administered to introduce significant changes in systemic immune reactions.
There were no significant differences in the mRNA levels of cytokines after B. breve administration in this study. O’Mahony et al. (24) reported a model of cytokine production under stimulation with anti-CD3 and anti-CD28 antibodies using splenocytes. Because our study simply examined the mRNA levels of cytokines and other signaling molecules without any stimulation, we cannot directly compare our results to theirs. However, O’Mahony et al. also reported that TNF-α and IFN-γ production from splenocytes of mice fed B. breve was reduced after stimulation with anti-CD3 and anti-CD28 compared with controls. These results suggest that B. breve regulates cytokine synthesis directly and indirectly.
Probiotics are useful for regulating inflammation as well as allergic reactions (25). Although lactobacilli and bifidobacteria are often used as probiotics, their functions seem to be different. Nonaka et al. suggested that lactobacilli modulate the Th1/Th2 immune balance toward Th1 reactions (8). Shima et al. reported that the Lactobacillus casei strain Shirota enhances the expression of genes involved in defense/immune functions and in lipid metabolism, whereas the B. breve strain Yakult downregulates the expression of many genes (26). Hoarau et al. suggested that B. breve can induce dendritic cell maturation through TLR2, with production of IL-10 regulating excessive Th1 responses as well as Th2 polarization (27). Our findings support a possible mechanism for B. breve action in which it modulates inflammation by downregulating the expression of inflammatory molecules during the newborn period. It would also promote tolerance by upregulating the expression of CD3 but not co-stimulatory molecules during the weaning period. Further study is needed to determine the precise effects of B. breve in immune reactions during early infancy.
Methods
Study Design
All animal experiments were approved by the Juntendo University Animal Experimental Ethics Committee and complied with the National Institutes of Health guidelines for animal care.
B. breve Supplementation Study
B. breve powder was commercially prepared from Morinaga Milk Industry (Zama, Japan), and this strain was deposited into the Belgian Co-Ordinated Collections of Micro-Organisms as strain LMG23729 (13). B. breve was cultured for 16 h at 37 °C in M17 broth (Difco, Detroit, MI), collected by centrifugation, and washed twice with phosphate-buffered saline. F344/Du rat pups were used in this study (Japan SLC, Shizuoka, Japan). The pups were randomly divided into four groups soon after birth. One group received 5 × 108 colony-forming units/animal live B. breve plus starch dissolved in phosphate-buffered saline daily from days 1 to 14 after delivery (newborn group, n = 14) via an orogastric catheter. A second group received an equal amount of B. breve on days 21–34 (weaning group, n = 9). The remaining two groups were control groups that received starch alone during these periods (n = 14 and 9, respectively, for the newborn and weaning period control groups). During the newborn period, all rat pups received breast milk from their mothers from days 0 to 21. All rats were killed 14 days after B. breve administration, and their colons, small intestines, and spleens were removed and washed with phosphate-buffered saline. mRNA was extracted from a colon sample 1 cm proximal to the anus, from an ileal sample 1 cm proximal to the ileocecal valve, and from the spleen. The rest of each colon sample was fixed with formalin and embedded in paraffin for histological analysis.
DNA Extraction From Fecal Samples
DNA was extracted from fecal samples as previously described (14). Briefly, fecal samples (20 mg) were washed three times in 1.0 ml phosphate-buffered saline and centrifuged at 13,000 rpm for 5 min. Fecal pellets were resuspended in 450 μl of extraction buffer (100 mM Tris/HCl, 40 mM EDTA, pH 9.0) and 50 μl of 10% sodium dodecyl sulfate. Glass beads (0.1 mm diameter, 300 mg) and 500 μl of buffer-saturated phenol were added to the suspension, and the mixture was vigorously vortexed for 30 s using a FastPrep FP 100A instrument (Funakoshi, Tokyo, Japan). After centrifugation at 13,000 rpm for 5 min, 400 μl of supernatant was extracted with phenol–chloroform, and 250 μl of supernatant was precipitated with isopropanol. Inhibitors were removed using a High Pure PCR Template Preparation Kit (Roche, Basel, Switzerland). Purified DNA was suspended in 200 μl of Tris-EDTA buffer (pH 8.0).
Real-Time PCR for Quantitative Determination of Bacteria
Real-time PCR analysis of microflora was performed using an ABI 7500 Fast Real-time PCR System (Applied Biosystems, Foster City, CA), SYBR Premix Ex Taq (Takara Bio, Shiga, Japan), and ROX reference dye II (Takara Bio) as an internal standard. Primer sets were previously reported: total bacteria and Bacteroides/Prevotella (15), Bifidobacterium (16), and Lactobacillus (17). The amplification program consisted of 1 cycle at 95 °C for 20 s followed by 40 cycles at 95 °C for 3 s, 60 °C for 30 s, and 72 °C for 30 s. Fluorescent products were detected at the last step of each cycle. Melting curves were obtained by heating the reaction products from 60 °C to 95 °C in 0.2 °C/s increments with continuous fluorescence collection.
RNA Extraction From Tissue Samples
Whole-thickness samples from colon, small intestine, and spleen obtained from rat pups were preserved in RNAlater solution (Applied Biosystems) at −30 °C. In preparation for microarray and real-time PCR analyses, the samples were minced and homogenized in Buffer RLT and then extracted using RNeasy Mini Kit spin columns (Qiagen, Germantown, MD). The quantity and purity of the RNA samples were determined with a NanoDrop ND-1000 Spectrophotometer (Thermo Scientific, Wilmington, MA) and the Experion RNA StdSens Analysis Kit (Bio-Rad Laboratories, Hercules, CA).
Microarray Hybridization and Data Analysis
To analyze gene expression, a microarray assay was performed according to the manufacturer’s instructions for the Ambion WT Expression Kit (Applied Biosystems) and GeneChip WT Terminal Labeling and Controls Kit (Affymetrix, Santa Clara, CA). Briefly, total RNA was reverse-transcribed into cDNA with random primers, and then cRNA was synthesized from this cDNA by in vitro transcription. The resulting cRNA was used as a template for the second cycle of cDNA synthesis. This cDNA was then fragmented, labeled, and hybridized to a GeneChip Rat Gene 1.0 ST Array (Affymetrix) at 45 °C for 17 h. This array encompasses ~29,000 genes. The chips were washed, stained with streptavidin-phycoerythrin, and scanned using a GeneChip Scanner 3000 (Affymetrix).
Scanned images were converted to CEL files using the GeneChip Command Console Software (AGCC) (Affymetrix), and data were analyzed with GeneSpring GX v11 software (Agilent Technologies, Santa Clara, CA). Raw intensity values from each chip were normalized using the RMA16 algorithm. Genes showing a greater than 1.3-fold difference in signal intensity compared to controls and a significant difference in expression between rats administered B. breve and those not (t-test, P < 0.05) were considered upregulated or downregulated genes. Data were also functionally analyzed through the use of Ingenuity Pathways Analysis (Ingenuity Systems, http://www.ingenuity.com).
Real-Time PCR
The expression of specific signaling molecules in rat colon was examined by real-time PCR. TaqMan probe–based real-time quantitative reverse transcription PCR was performed using cDNA synthesized from total rat colon RNA preparations (High Capacity cDNA Reverse Transcription Kit; Applied Biosystems) and analyzed with a 7500 Real-Time PCR System (Applied Biosystems) using the default protocols. The expression of each gene was normalized to the expression of β-actin using the standard curve method. Primers for lipoprotein lipase (Lpl; Rn00561482_m1); glutathione peroxidase 2 (Gpx2; Rn00822100_gH); chloride channel, calcium-activated family member 4 (ClCa4; Rn01748633_m1); lipopolysaccharide-binding protein (Lbp; Rn00567985_m1); CD3d (Rn00565890_m1); CD80 (Rn00709368_m1); CD86 (Rn00571654_m1); CD28 (Rn00565469_m1); cytotoxic T-lymphocyte antigen 4 (CTLA-4; Rn00581545_m1); and β-actin (Rn00667869_m1) were prepared using TaqMan Gene Expression Assays (Applied Biosystems).
Immunohistochemical Analysis
Paraffin-embedded sections were used for immunohistochemical analysis. Deparaffinized sections were incubated with either polyclonal rabbit anti-CD3 antibody (Invitrogen, Camarillo, CA) or monoclonal mouse anti-CD86 antibody (Santa Cruz Biotechnology, Santa Cruz, CA). After washing, the sections were incubated with biotin-conjugated secondary antibody. The sections were then incubated with avidin peroxidase (Sigma-Aldrich, St. Louis, MO). Peroxidase activity was detected with 3,3′-diaminobenzenzidine-tetra-hydrochloride (Sigma-Aldrich) in Tris–HCl containing 0.01% H2O2. Each section was counterstained with hematoxylin before examination by light microscopy. Nonspecific staining was evaluated on sections stained without the primary antibody.
Statistical Analysis
Differences in mRNA expression measured by real-time PCR and in microflora determinations were analyzed with Student’s t-test, and P < 0.05 was considered statistically significant.
References
Report of a Joint FAO/WHO Expert Consultation on Evaluation of Health and Nutritional Properties of Probiotics in Food Including Powder Milk with Live Lactic Acid Bacteria, 2001. (http://www.who.int/foodsafety/publications/fs_management/en/probiotics.pdf).
Yang YX, He M, Hu G, et al. Effect of a fermented milk containing Bifidobacterium lactis DN-173010 on Chinese constipated women. World J Gastroenterol 2008;14: 6237–43.
Coccorullo P, Strisciuglio C, Martinelli M, Miele E, Greco L, Staiano A . Lactobacillus reuteri (DSM 17938) in infants with functional chronic constipation: a double-blind, randomized, placebo-controlled study. J Pediatr 2010;157: 598–602.
Kalliomäki M, Salminen S, Arvilommi H, Kero P, Koskinen P, Isolauri E . Probiotics in primary prevention of atopic disease: a randomised placebo-controlled trial. Lancet 2001;357: 1076–9.
Xiao JZ, Kondo S, Yanagisawa N, et al. Probiotics in the treatment of Japanese cedar pollinosis: a double-blind placebo-controlled trial. Clin Exp Allergy 2006;36: 1425–35.
Inoue Y, Iwabuchi N, Xiao JZ, Yaeshima T, Iwatsuki K . Suppressive effects of bifidobacterium breve strain M-16V on T-helper type 2 immune responses in a murine model. Biol Pharm Bull 2009;32: 760–3.
Torii A, Torii S, Fujiwara S, Tanaka H, Inagaki N, Nagai H . Lactobacillus Acidophilus strain L-92 regulates the production of Th1 cytokine as well as Th2 cytokines. Allergol Int 2007;56: 293–301.
Nonaka Y, Izumo T, Izumi F, et al. Antiallergic effects of Lactobacillus pentosus strain S-PT84 mediated by modulation of Th1/Th2 immunobalance and induction of IL-10 production. Int Arch Allergy Immunol 2008;145: 249–57.
Schouten B, van Esch BC, Hofman GA, et al. Cow milk allergy symptoms are reduced in mice fed dietary synbiotics during oral sensitization with whey. J Nutr 2009;139: 1398–403.
Hougee S, Vriesema AJ, Wijering SC, et al. Oral treatment with probiotics reduces allergic symptoms in ovalbumin-sensitized mice: a bacterial strain comparative study. Int Arch Allergy Immunol 2010;151: 107–17.
Li Y, Shimizu T, Hosaka A, Kaneko N, Ohtsuka Y, Yamashiro Y . Effects of Bifidobacterium breve supplementation on intestinal flora of low birth weight infants. Pediatr Int 2004;46: 509–15.
Fujii T, Ohtsuka Y, Lee T, et al. Bifidobacterium breve enhances transforming growth factor beta1 signaling by regulating Smad7 expression in preterm infants. J Pediatr Gastroenterol Nutr 2006;43: 83–8.
Abe F, Miyauchi H, Uchijima A, Yaeshima T, Iwatsuki K Stability of bifidobacteria in powdered formula. Int J Food Sci Tech 2009;44: 718–24.
Odamaki T, Xiao JZ, Iwabuchi N, et al. Fluctuation of fecal microbiota in individuals with Japanese cedar pollinosis during the pollen season and influence of probiotic intake. J Investig Allergol Clin Immunol 2007;17: 92–100.
Bartosch S, Fite A, Macfarlane GT, McMurdo ME . Characterization of bacterial communities in feces from healthy elderly volunteers and hospitalized elderly patients by using real-time PCR and effects of antibiotic treatment on the fecal microbiota. Appl Environ Microbiol 2004;70: 3575–81.
Matsuki T, Watanabe K, Fujimoto J, et al. Quantitative PCR with 16S rRNA-gene-targeted species-specific primers for analysis of human intestinal bifidobacteria. Appl Environ Microbiol 2004;70: 167–73.
Delroisse JM, Boulvin AL, Parmentier I, Dauphin RD, Vandenbol M, Portetelle D . Quantification of Bifidobacterium spp. and Lactobacillus spp. in rat fecal samples by real-time PCR. Microbiol Res 2008;163: 663–70.
Karlsson A, Jägervall A, Pettersson M, Andersson AK, Gillberg PG, Melgar S . Dextran sulphate sodium induces acute colitis and alters hepatic function in hamsters. Int Immunopharmacol 2008;8: 20–27.
Ritzka M, Stanke F, Jansen S, et al. The CLCA gene locus as a modulator of the gastrointestinal basic defect in cystic fibrosis. Hum Genet 2004;115: 483–91.
Te Velde AA, Pronk I, de Kort F, Stokkers PC . Glutathione peroxidase 2 and aquaporin 8 as new markers for colonic inflammation in experimental colitis and inflammatory bowel diseases: an important role for H2O2? Eur J Gastroenterol Hepatol 2008;20: 555–60.
Dincer Y, Erzin Y, Himmetoglu S, Gunes KN, Bal K, Akcay T . Oxidative DNA damage and antioxidant activity in patients with inflammatory bowel disease. Dig Dis Sci 2007;52: 1636–41.
Kruidenier L, Kuiper I, Van Duijn W, et al. Imbalanced secondary mucosal antioxidant response in inflammatory bowel disease. J Pathol 2003;201: 17–27.
Van Overtvelt L, Moussu H, Horiot S, et al. Lactic acid bacteria as adjuvants for sublingual allergy vaccines. Vaccine 2010;28: 2986–92.
Wall R, Ross RP, Shanahan F, et al. Metabolic activity of the enteric microbiota influences the fatty acid composition of murine and porcine liver and adipose tissues. Am J Clin Nutr 2009;89: 1393–401.
Ruemmele FM, Bier D, Marteau P, et al. Clinical evidence for immunomodulatory effects of probiotic bacteria. J Pediatr Gastroenterol Nutr 2009;48: 126–41.
Shima T, Fukushima K, Setoyama H, et al. Differential effects of two probiotic strains with different bacteriological properties on intestinal gene expression, with special reference to indigenous bacteria. FEMS Immunol Med Microbiol 2008;52: 69–77.
Hoarau C, Lagaraine C, Martin L, Velge-Roussel F, Lebranchu Y . Supernatant of Bifidobacterium breve induces dendritic cell maturation, activation, and survival through a Toll-like receptor 2 pathway. J Allergy Clin Immunol 2006;117: 696–702.
Acknowledgements
The authors are grateful to Yumiko Sakurai for her extended technical support.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
H.I. and M.N. are employed by Morinaga Milk Industry Co., Ltd.
Rights and permissions
About this article
Cite this article
Ohtsuka, Y., Ikegami, T., Izumi, H. et al. Effects of Bifidobacterium breve on inflammatory gene expression in neonatal and weaning rat intestine. Pediatr Res 71, 46–53 (2012). https://doi.org/10.1038/pr.2011.11
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/pr.2011.11
This article is cited by
-
Microbial and immune faecal determinants in infants hospitalized with COVID-19 reflect bifidobacterial dysbiosis and immature intestinal immunity
European Journal of Pediatrics (2023)
-
Evaluation of Porcine Intestinal Epitheliocytes as an In vitro Immunoassay System for the Selection of Probiotic Bifidobacteria to Alleviate Inflammatory Bowel Disease
Probiotics and Antimicrobial Proteins (2021)
-
Effect of probiotic Bifidobacterium bifidum G9-1 on the relationship between gut microbiota profile and stress sensitivity in maternally separated rats
Scientific Reports (2018)
-
Gene expression profiling in the intestinal mucosa of obese rats administered probiotic bacteria
Scientific Data (2017)
-
Adamdec1, Ednrb and Ptgs1/Cox1, inflammation genes upregulated in the intestinal mucosa of obese rats, are downregulated by three probiotic strains
Scientific Reports (2017)