Abstract
Biliary atresia (BA) is a virus-induced autoimmune disease associated with abnormal DNA methylation patterns that contribute to disease presentation. This study examined DNA methylation patterns, changes to genes associated with methylation regulation, and changes to the autoimmune-related gene interferon gamma (IFN-γ) in CD4+ T cells from BA patients. We demonstrated that genomic DNA isolated from CD4+ T cells harvested from infants presenting with BA were hypomethylated relative to healthy controls. In addition, DNA methyltransferase (DNMT1) and DNMT3a mRNA levels were significantly lower in BA CD4+ T cells compared with controls and methyl-DNA-binding domain proteins (MBD1) mRNA expression (but not MBD4 detected at higher levels in BA patients), which was significantly lower in CD4+ T cells from BA infants than in controls. DNMT1 expression positively correlated with global DNA methylation in BA CD4+ T cells. IFN-γ mRNA expression levels in BA patients were also significantly increased, and the IFN-γ gene promoter region was hypomethylated in BA CD4+ T cells compared with controls and negatively correlated with DNA methylation. These data suggest that methylation changes in CD4+ cells may contribute to BA disease presentation and progression by affecting the expression of genes associated with autoimmunity.
Similar content being viewed by others
Main
Biliary atresia (BA) can develop into neonatal cholestasis via undefined mechanisms and is characterized by fibrosclerosing and inflammatory destruction of the extrahepatic and intrahepatic biliary system during the first few weeks of life (1,2). Neonatal cholestasis is a devastating disease that leads to cirrhosis, requiring liver transplantation as the only option for therapy in the majority of cases (3). The etiology and pathogenesis of bile duct obstruction in children with BA remains largely unknown. One theory suggests that it may result from a primary perinatal hepatobiliary viral infection that elicits autoimmune-mediated bile duct injury (4) associated primarily with CD4+ T helper 1 (Th-1) cell-mediated inflammatory processes (5), and genome-wide association studies have identified BA susceptibility loci on several chromosomes (6,7). However, studies carried out in twins demonstrated that nongenetic factors also play an important role in mediating generalized BA pathogenesis (8) even though specific causes of generalized BA remain obscure because no common (specific) environmental factors that trigger disease progression (either directly or via an autoimmune response) have yet been identified.
DNA methylation is the only genetically programmed DNA modification process in mammals involved in the regulation of several biological processes, including gene transcription, X-chromosome inactivation, genomic imprinting, and chromatin modification (9–11). DNA methylation plays a critical role in maintaining T-cell function, and a growing body evidence indicates that failure to maintain DNA methylation levels and patterns in mature T cells can result in T-cell-mediated autoimmune responses in vitro and autoimmunity in vivo (12). Defective maintenance of DNA methylation may result in the development of many autoimmune diseases including systemic lupus erythematosus, rheumatoid arthritis, and multiple sclerosis (13–15).
In general, methylation of CpG islands within promoter or enhancer regions suppresses target gene transcription (16) and DNA methylation patterns are established and maintained by DNA methyltransferases (DNMTs). In humans, three enzymes are known to have DNMT activity: DNMT1, DNMT3a, and DNMT3b. DNMT3a and DMMT3b are responsible for de novo methylation and modify unmethylated DNA. In contrast, DNMT1 acts on hemimethylated DNA and is thought to be required for maintaining methylation patterns (17). Methylated DNA is recognized by a conserved family of methyl-DNA-binding domain proteins (MBDs) consisting of five known members in mammals: MBD1, MBD2, MBD3, MBD4, and methyl-CpG-binding protein-2 (MeCP2). All MBDs play active roles in regulating DNA methylation, heterochromatin formation, and gene transcription (18).
In the present work, we investigated global DNA methylation levels (and mRNA expression patterns of DNMTs and MBDs genes) in CD4+ T cells harvested from infants presenting with BA and identified a pattern of aberrant genomic DNA methylation and DNMT and MBD expression. We also demonstrated increased gene expression of IFN-γ in CD4+ T cells collected from pediatric BA patients who correlated with the DNA methylation status of the IFN-γ gene promoter region. IFN-γ is critical to the development of protective innate and adaptive immune responses against viral and intracellular bacterial infections, and aberrant IFN-γ expression is associated with a number of autoinflammatory and autoimmune diseases. Our results provide novel insights into the pathogenesis of BA.
MATERIALS AND METHODS
Subjects.
Fifteen infants with BA with a mean age (mean ± SEM) of 59 ± 5.6 d were recruited from the Outpatient Pediatric Clinic and Inpatient ward at the Children's Hospital, Medical Center of Fudan University. Healthy controls (n = 12; mean age of 67 ± 7.4 d) were recruited from the Children's Hospital, Medical Centre of Fudan University. BA patients and controls were age- and gender matched. Pathological diagnosis of BA patients was confirmed independently by two pathologists. Experimental protocols were reviewed and approved by the human ethics committee of the Medical Center of Fudan University, and written informed consent was obtained from the parents of all subjects.
CD4+ T cell isolation.
Peripheral blood was collected in heparinized tubes from patients and controls. Peripheral blood mononuclear cells (PBMCs) were isolated by Ficoll-Hypaque density gradient centrifugation (Tianjin Haoyang Biological Manufacture, Co., Ltd, Tianjin, China), and CD4+ T cells were isolated by positive selection using magnetic beads as described by the manufacturer (Miltenyi Biotec, Bergisch Gladbach, Germany). The purity of enriched CD4+ T cells isolates was evaluated by flow cytometry and was higher than 94%.
RNA isolation and real-time quantitative RT-PCR (qRT-PCR).
Total RNA from CD4+ T cells was isolated using the DNA/RNA Isolation kit (Tiangen Biotech, Beijing, China). qRT-PCR was performed using an ABI 7500 instrument (Applied Biosystems 7500; ABI, Foster City, CA), and mRNA levels were quantified using the QuantiTect SYBR Green RT-PCR kit (TaKaRa Biotech, Co., Dalin, China). Serial dilutions of sample RNA were also included to generate a standard curve used to calculate the relative concentrations of transcript in each RNA sample examined. Negative controls (distilled water substituted for RNA) were also run for each sample, and β-actin was amplified and used as a loading control. Primers used are listed in Table 1.
Genomic DNA extraction and measurement of global DNA methylation.
Genomic DNA was isolated from CD4+ T cells using the DNA/RNA Isolation kit (Tiangen). Global DNA methylation was measured using the Methylamp Global DNA Methylation Quantification Kit as described by the manufacturer (Epigentek Group, Inc., New York, NY). This kit yields accurate measures of methylcytosine content expressed as a percentage of the total cytosine content. Briefly, DNA was immobilized on a strip well with high affinity for DNA and DNA methylation quantified by an ELISA-like reaction using an anti-5-methylcytosine antibody. The amount of methylated DNA is proportional to the OD, and the degree of DNA methylation determined was based on the OD readings obtained.
IFN-γ genomic DNA extraction and bisulfite sequencing.
IFN-γ genomic DNA was isolated from CD4+ T cells using the DNA/RNA Isolation kit (Tiangen). Determination of disulfite conversion was performed using the EpiTect Bisulfite Kit (Qiagen, Germany). The 5′-flanking sequences were identified by bisulfite treatment of purified DNA followed by PCR amplification of a 658 base pair (bp) fragment located immediately 5′ to the IFN-γ gene transcription start site (−579 to +79, containing five CG pairs). Fragments were cloned into the pMD19-T vector (TaKaRa Biotech), and five independent clones were sequenced for each of the amplified fragments (19,20). The following primers were used: forward, 5′-TTGTGTGGTTTGTATTGTATTTT-3′ and reverse, 5′- TCAAACTAATCAAATCCAAAAA-3′.
Statistical analysis.
Data are expressed as the mean ± SEM. The t test was used to determine statistically significant differences between groups, and p-values <0.05 were considered significant. All analyses were performed using the SPSS Version 13.0 (SPSS, Chicago, IL) software.
RESULTS
Global DNA methylation of CD4+ T cell DNA from BA infants and healthy controls.
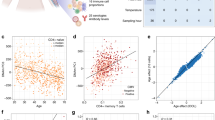
Global DNA methylation of DNA isolated from infants presenting with BA (n = 15) and from healthy controls (n = 12) was characterized and demonstrated that the mean genomic CD4+ T cell DNA methylation levels were significantly reduced in BA infants compared with healthy controls (37.37 ± 1.086% versus 55.2 ± 0.751%, p < 0.0001; Fig. 1).
Percent global DNA methylation. Methylation levels in CD4+ T cells harvested from infants presenting with BA (▪) or from healthy controls (□) were determined. Methylcytosine levels are expressed as a percent of total cytosine in CD4+ T cells determined by ELISA. Relative to controls, the mean global methylcytosine levels were significantly decreased in CD4+ T cells of BA infants. *p < 0.05.
Expression of DNMTs and MBDs in CD4+ T cells of BA infants and healthy controls.
To determine the cause of DNA hypomethylation in infants presenting with BA, the transcription levels of DNMTs and MBDs in CD4+ T cells were assessed using qRT-PCR. DNMT1 and DNMT3a mRNA levels were significantly lower in BA CD4+ T cells (0.0048 ± 0.0005 versus 0.0072 ± 0.0003, p = 0.0012 and 0.0012 ± 0.0002 versus 0.0024 ± 0.0003, p = 0.0210, respectively; Fig. 2A). However, no significant differences in DNMT3b expression were observed between BA infants and controls (0.0005 ± 0.0001 versus 0.0007 ± 0.0001, p = 0.2283). Of the five MBD family members, only the MBD1 mRNA expression levels were significantly diminished in BA infant CD4+ T cells (0.0056 ± 0.0007 versus 0.0106 ± 0.0009, p = 0.0029) compared with controls. By contrast, MBD4 mRNA expression levels were significantly increased in CD4+ T cells (0.0536 ± 0.005 versus 0.0285 ± 0.006; p = 0.0199). No significant differences in the expression levels of MBD2, MBD3, or MeCP2 in CD4+ T cells (MBD2: 0.0203 ± 0.0040 versus 0.0275 ± 0.0020; p = 0.1992; MBD3: 0.0019 ± 0.00008 versus 0.0020 ± 0.0003; p = 0.7645; and MeCP2: 0.0074 ± 0.0020 versus 0.0094 ± 0.0020; p = 0.4686) were observed between BA infants and controls (Fig. 2B).
Relative mRNA levels of DNMTs (A) and MBDs (B) in CD4+ T cells harvested from BA infants (▪) and healthy controls (□) measured by qRT-PCR. Data represent the mean expression levels normalized to β-actin ± SEM. Relative to the controls, DNMT1, DNMT3a, and MBD1 expression levels were significantly decreased in BA CD4+ T cells and the MBD4 expression levels were increased. *p < 0.05.
Correlation between global hypomethylation with DNMTs and expression of MBDs.
We next analyzed the relationship between global DNA methylation status and expression of DNMTs or MBDs in CD4+ T cells harvested from BA infants. This analysis demonstrated that DNMT1 expression positively correlated with the overall methylation levels in CD4+ T cells (r = 0.6290, p = 0.0120; Fig. 3). However, there was no significant correlation between DNMT3a, DNMT3b, or MBD expression levels and global methylation status in BA CD4+ T cells compared with controls.
IFN-γ expression and promoter DNA methylation in BA CD4+ T cells.
To investigate whether specific genes were hypomethylated in BA CD4+ T cells, we examined the expression levels of IFN-γ mRNA and the methylation status of a known promoter fragment of the IFN-γ gene. We chose to focus on this gene because others have shown that the IFN-γ gene is particularly sensitive to alterations in methylation status in addition to being associated with autoimmunity (21–23). As shown in Figure 4A, INF-γ mRNA levels in BA CD4+ T cells were significantly higher than levels observed in healthy controls (0.0008 ± 0.0002 versus 0.0003 ± 0.0001; p = 0.0376). To define the methylation status of the IFN-γ gene promoter of CD4+ T cells from BA infants and healthy controls, a 658-bp fragment located immediately 5′ to the IFN-γ gene transcription start site (−579 to +79) was analyzed using bisulfite genomic DNA sequencing. Figure 4B demonstrates that the average methylation level of the IFN-γ gene promoter in CD4+ T cells. The mean methylation level of the five CG pairs within the amplified fragment (positions −458, −372, −297, −187, and −55) was significantly lower in BA infants than in healthy controls (47.1 ± 1.899% versus 60.7 ± 4.280%; p = 0.0035; Fig. 4C). Moreover, we found that the average methylation status of five CG pairs present in the IFN-γ gene correlated with the mRNA expression levels in CD4+ T cells examined from BA infants and healthy controls (r = −0.5720, p = 0.0260; Fig. 4D).
CD4+ IFN-γ mRNA and methylation levels. Measurement of INF-γ mRNA by qRT-PCR in CD4+ T cells from BA infants (▪) was significantly increased compared with healthy controls (□) (A). Results represent expression levels normalized to β-actin ± SEM. The mean methylation status of each of the five CG pairs of the IFN-γ gene in CD4+ T cells harvested from BA infants (▪) and healthy controls (□) is shown (B). The average methylation status at positions −458, −372, −297, −187, and −55 was lower in patient samples (•) compared with controls (○) (C). Average DNA methylation levels of IFN-γ negatively correlated with the relative IFN-γ mRNA levels in CD4+ T cells from BA infants compared with controls (D) (r = −0.572; p = 0.026). *p < 0.05.
DISCUSSION
In this study, we assessed DNA methylation levels in CD4+ T cells harvested from infants presenting with BA using a global DNA methylation quantification kit to detect genomic DNA methylation levels. Our results demonstrated that CD4+ T cells exhibited significantly diminished global DNA methylation levels in infants presenting with BA compared with control subjects. Furthermore, decreased global methylation of BA CD4+ T cells correlated with decreased expression of DNMT1 and increased expression of IFN-γ.
In humans, DNA methylation is carried out by three enzymes, DNMT1, DNMT3a, and DNMT3b. DNMT3a and DNMT3b are primarily responsible for de novo methylation that occurs primarily during early development. Maintenance of genomic methylation patterns is mediated primarily by DNMT1. Both de novo DNA methylation and maintenance of methylation patterns are mediated by DNMTs (24,25), and the abundance of DNMT1 has been correlated with the status of DNA methylation (26). In CD4+ T cells from BA infants, both DNMT1 and DNMT3a mRNA levels were reduced compared with control subjects. Furthermore, DNMT1 expression levels statistically correlated with the degree of hypomethylation in CD4+ T cells of BA infants. These data suggested that it is likely that reduced DNMT1 expression in CD4+ T cells contributed to BA development. Because BA is associated with CD4+ Th-1 cell-mediated immunity, it suggests that a specific immune response is involved with BA pathogenesis (5). Our present results showed that the mean methylation patterns of CD4+ T cells from BA infants were significantly decreased compared with control subjects suggesting that hypomethylation of CD4+ T cell DNA may have affected development or resulted in the development of BA.
Data presented in this report demonstrated that CD4+ T cells from BA infants presented with reduced levels of MBD1 mRNA and higher levels of MBD4 mRNA compared with levels observed in controls. In addition, MBD2, MBD3, and MeCP2 levels were decreased in BA infants, although the differences did not reach statistical significance. Despite expression irregularities, we did not find a significant correlation between global methylation levels and mRNA levels for any of the five MBD proteins in BA CD4+ T cells. On one hand, MBD1, MBD2, MBD3, and MeCP2 have been linked to transcriptional repression and defects that can affect DNA methylation (27); on the other hand, MBD4 can demethylate DNA by means of its 5′-mC (5-methylcytosine) DNA glycosylate activity (28) suggesting that MBD family members may play different roles in the DNA methylation process taking place in BA CD4+ T cells. However, given the relatively small sample size examined in the present study, it is possible that relationships between T-cell methylation levels and expression of either DNMT3b or MBD could be identified if a larger BA cohort were analyzed.
IFN-γ over-expression is sufficient to elicit BA autoimmune responses in mouse and human lymphocytes. In a mouse model of rotavirus-induced BA, rotavirus infection triggered hepatobiliary inflammation mediated by IFN-γ produced by CD4+ and CD8 + T cells, resulting in progressive jaundice and growth failure. The genetic loss of IFN-γ significantly suppressed tissue-specific targeting of T lymphocytes and completely prevented the inflammation and fibrosing obstruction of the extrahepatic bile ducts, a key pathogenic component associated with the BA progression (29). In children presenting with BA, most of the genes examined showed differential lymphocyte function with activation of osteopontin, a regulator of cell-mediated immunity in T-helper lymphocytes and suppression of immunoglobulin genes during the early stages of disease associated with IFN-γ production (30). The overexpression of IFN-γ suggested that Th-1-associated cytokines were important to the pathogenesis of BA (30) and that IFN-γ gene-mediated immunity (through CD4+ T cells) could play an important role in mediating BA pathogenesis.
To determine whether DNA methylation changes contributed to INF-γ overexpression in BA CD4+ T cells, the methylation status of the methylation-sensitive region within the proximal promoter of the IFN-γ gene (located −73 to −48 bp upstream of the transcription start site and sufficient to induce IFN-γ expression by activated T cells) (21) was assessed because methylation of the CpG motif at position −55 represents a major epigenetic regulatory mechanism (22,23). Also affecting INF-γ expression is the cAMP response element-binding protein (CREB), activating transcription factor (ATF), and activator protein-1 (AP-1) that bind to the proximal promoter (31,32). In this study, we found that the proximal INF-γ promoter region containing five CG pairs was hypomethylated in BA CD4+ T cells and that the average methylation status of this region was negatively correlated with IFN-γ mRNA expression. These observations suggested that DNA hypomethylation may have contributed to increased IFN-γ expression in BA cases consistent with previously reported observations describing that DNA hypomethylation lead to biliary defects and INF-γ-mediated signaling in Zebrafish (33).
The causes of hypomethylation associated with different diseases has been associated with altered DNMT activity, histone modifications (loss of trimethylation and increased acetylation), exogenous insults (diet, environment, and infection), noncoding RNA, or with defective DNA repair (34,35). In addition, folate plays an important role in human embryonic development because it serves as a single carbon group donor (the main methyl group donor of the cell) (36). Folate deficiencies are associated with a variety of birth defects including neural tube defects, Hirschsprung's disease, cleft lip, and cleft palate (36–38). Furthermore, exposure to toxic insults (such as arsenic) can have long-term effects in mice resulting in genome-wide DNA hypomethylation that enhances genetic instability (39). Interestingly, causes of hypomethylation are consistent with the causes of BA (genetic, infective, inflammatory, and toxic insults might contribute to BA development) suggesting that hypomethylation might be a contributing factor associated with BA development.
In summary, our findings demonstrated that global genomic DNA methylation was compromised in CD4+ T cells examined from infants presenting with BA and that these cells also had abnormal DNMTs and MBDs expression levels that likely contributed to BA pathogenesis. Our results also provided basic epigenetic data that contributed to a better understanding of the mechanisms by which autoimmune-related genes (such as the IFN-γ gene) become up-regulated in infants presenting with BA that ultimately results in hepatobiliary inflammation and fibrosing obstruction of bile ducts.
Abbreviations
- BA:
-
biliary atresia
- DNMT :
-
DNA methyltransferase
- MBD :
-
methyl-DNA-binding domain protein
- MeCP2 :
-
methyl-CpG-binding protein-2
- qRT-PCR:
-
real-time quantitative reverse transcriptase-polymerase chain reaction
- Th-1:
-
T helper 1
References
Hartley J, Harnden A, Kelly D 2010 Biliary atresia. BMJ 340: c2383
Santos JL, Choquette M, Bezerra JA 2010 Cholestatic liver disease in children. Curr Gastroenterol Rep 12: 30–39
Shen C, Zheng S, Wang W, Xiao XM 2008 Relationship between prognosis of biliary atresia and infection of cytomegalovirus. World J Pediatr 4: 123–126
Mack CL 2007 The pathogenesis of biliary atresia: evidence for a virus-induced autoimmune disease. Semin Liver Dis 27: 233–242
Mack CL, Tucker RM, Sokol RJ, Karrer FM, Kotzin BL, Whitington PF, Miller SD 2004 Biliary atresia is associated with CD4+ Th-1 cell-mediated portal tract inflammation. Pediatr Res 56: 79–87
Garcia-Barceló MM, Yeung MY, Miao XP, Tang CS, Cheng G, So MT, Ngan ES, Lui VC, Chen Y, Liu XL, Hui KJ, Li L, Guo WH, Sun XB, Tou JF, Chan KW, Wu XZ, Song YQ, Chan D, Cheung K, Chung PH, Wong KK, Sham PC, Cherny SS, Tam PK 2010 Genome-wide association study identifies a susceptibility locus for biliary atresia on 10q24.2. Hum Mol Genet 19: 2917–2925
Leyva-Vega M, Gerfen J, Thiel BD, Jurkiewicz D, Rand EB, Pawlowska J, Kaminska D, Russo P, Gai X, Krantz ID, Kamath BM, Hakonarson H, Haber BA, Spinner NB 2010 Genomic alterations in biliary atresia suggest region of potential disease susceptibility in 2q37.3. Am J Med Genet A 152A: 886–895
Smith BM, Laberge JM, Schreiber R, Weber AM, Blanchard H 1991 Familial biliary atresia in three siblings including twins. J Pediatr Surg 26: 1331–1333
Geiman TM, Robertson KD 2002 Chromatin remodeling, histone modifications, and DNA methylation-how does it all fit together?. J Cell Biochem 87: 117–125
Reik W, Walter J 2001 Genomic imprinting: parental influence on the genome. Nat Rev Genet 2: 21–32
Chow J, Heard E 2009 X inactivation and the complexities of silencing a sex chromosome. Curr Opin Cell Biol 21: 359–366
Richardson B 2003 DNA methylation and autoimmune disease. Clin Immunol 109: 72–79
Richardson B, Scheinbart L, Strahler J, Gross L, Hanash S, Johnson M 1990 Evidence for impaired T cell DNA methylation in systemic lupus erythematosus and rheumatoid arthritis. Arthritis Rheum 33: 1665–1673
Karouzakis E, Gay RE, Michel BA, Gay S, Neidhart M 2009 DNA hypomethylation in rheumatoid arthritis synovial fibroblasts. Arthritis Rheum 60: 3613–3622
Baranzini SE, Mudge J, van Velkinburgh JC, Khankhanian P, Khrebtukova I, Miller NA, Zhang L, Farmer AD, Bell CJ, Kim RW, May GD, Woodward JE, Caillier SJ, McElroy JP, Gomez R, Pando MJ, Clendenen LE, Ganusova EE, Schilkey FD, Ramaraj T, Khan OA, Huntley JJ, Luo S, Kwok PY, Wu TD, Schroth GP, Oksenberg JR, Hauser SL, Kingsmore SF 2010 Genome, epigenome and RNA sequences of monozygotic twins discordant for multiple sclerosis. Nature 464: 1351–1356
Attwood JT, Yung RL, Richardson BC 2002 DNA methylation and the regulation of gene transcription. Cell Mol Life Sci 59: 241–257
Rountree MR, Bachman KE, Herman JG, Baylin SB 2001 DNA methylation, chromatin inheritance, and cancer. Oncogene 20: 3156–3165
Hendrich B, Bird A 1998 Identification and characterization of a family of mammalian methyl-CpG binding proteins. Mol Cell Biol 18: 6538–6547
Lu Q, Ray D, Gutsch D, Richardson B 2002 Effect of DNA methylation and chromatin structure on ITGAL expression. Blood 99: 4503–4508
Lu Q, Kaplan M, Ray D, Ray D, Zacharek S, Gutsch D, Richardson B 2002 Demethylation of ITGAL (CD11a) regulatory sequences in systemic lupus erythematosus. Arthritis Rheum 46: 1282–1291
Penix L, Weaver WM, Pang Y, Young HA, Wilson CB 1993 Two essential regulatory elements in the human interferon gamma promoter confer activation specific expression in T cells. J Exp Med 178: 1483–1496
Jones B, Chen J 2006 Inhibition of IFN-gamma transcription by site-specific methylation during T helper cell development. EMBO J 25: 2443–2452
Yano S, Ghosh P, Kusaba H, Buchholz M, Longo DL 2003 Effect of promoter methylation on the regulation of IFN-gamma gene during in vitro differentiation of human peripheral blood T cells into a Th2 population. J Immunol 171: 2510–2516
Okano M, Bell DW, Haber DA, Li E 1999 DNA methyltransferases Dnmt3a and Dnmt3b are essential for de novo methylation and mammalian development. Cell 99: 247–257
Pradhan S, Bacolla A, Wells RD, Roberts RJ 1999 Recombinant human DNA (cytosine-5) methyltransferase. I. Expression, purification, and comparison of de novo and maintenance methylation. J Biol Chem 274: 33002–33010
Song J, Rechkoblit O, Bestor TH, Patel DJ 2011 Structure of DNMT1-DNA complex reveals a role for autoinhibition in maintenance DNA methylation. Science 331: 1036–1040
Wade PA 2001 Methyl CpG-binding proteins and transcriptional repression. Bioessays 23: 1131–1137
Zhu B, Zheng Y, Angliker H, Schwarz S, Thiry S, Siegmann M, Jost JP 2000 5-Methylcytosine DNA glycosylase activity is also present in the human MBD4 (G/T mismatch glycosylase) and in a related avian sequence. Nucleic Acids Res 28: 4157–4165
Shivakumar P, Campbell KM, Sabla GE, Miethke A, Tiao G, McNeal MM, Ward RL, Bezerra JA 2004 Obstruction of extrahepatic bile ducts by lymphocytes is regulated by IFN-gamma in experimental biliary atresia. J Clin Invest 114: 322–329
Bezerra JA, Tiao G, Ryckman FC, Alonso M, Sabla GE, Shneider B, Sokol RJ, Aronow BJ 2002 Genetic induction of proinflammatory immunity in children with biliary atresia. Lancet 360: 1653–1659
Samten B, Howard ST, Weis SE, Wu S, Shams H, Townsend JC, Safi H, Barnes PF 2005 Cyclic AMP response element-binding protein positively regulates production of IFN-gamma by T cells in response to a microbial pathogen. J Immunol 174: 6357–6363
Cippitelli M, Sica A, Viggiano V, Ye J, Ghosh P, Birrer MJ, Young HA 1995 Negative transcriptional regulation of the interferon-gamma promoter by glucocorticoids and dominant negative mutants of c-Jun. J Biol Chem 270: 12548–12556
Matthews RP, Eauclaire SF, Mugnier M, Lorent K, Cui S, Ross MM, Zhang Z, Russo P, Pack M 2011 DNA hypomethylation causes bile duct defects in zebrafish and is a distinguishing feature of infantile biliary atresia. Hepatology 53: 905–914
Wilson AS, Power BE, Molloy PL 2007 DNA hypomethylation and human diseases. Biochim Biophys Acta 1775: 138–162
Pogribny IP, Beland FA 2009 DNA hypomethylation in the origin and pathogenesis of human diseases. Cell Mol Life Sci 66: 2249–2261
Bliek BJ, Steegers-Theunissen RP, Blok LJ, Santegoets LA, Lindemans J, Oostra BA, Steegers EA, de Klein A 2008 Genome-wide pathway analysis of folate-responsive genes to unravel the pathogenesis of orofacial clefting in man. Birth Defects Res A Clin Mol Teratol 82: 627–635
van der Linden IJ, Heil SG, van Egmont PM, van Straaten HW, den Heijer M, Blom HJ 2008 Inhibition of methylation and changes in gene expression in relation to neural tube defects. Birth Defects Res A Clin Mol Teratol 82: 676–683
Munnes M, Patrone G, Schmitz B, Romeo G, Doerfler W 1998 A 5′-CG-3′-rich region in the promoter of the transcriptionally frequently silenced RET protooncogene lacks methylated cytidine residues. Oncogene 17: 2573–2583
Sciandrello G, Caradonna F, Mauro M, Barbata G 2004 Arsenic-induced DNA hypomethylation affects chromosomal instability in mammalian cells. Carcinogenesis 25: 413–417
Acknowledgements
We thank Yijie Zheng, Shudao Xiong and Prof. Yiwei Chu at Department of Immunology, Shanghai Medical College, Fudan University, China, for providing technical assistance.
Author information
Authors and Affiliations
Corresponding author
Additional information
Supported by the National Natural Science Foundation of China (Grant 30973139) and the Science Foundation of Shanghai (Grant 09JC1402800).
The authors report no conflicts of interest.
Rights and permissions
About this article
Cite this article
Dong, R., Zhao, R. & Zheng, S. Changes in Epigenetic Regulation of CD4+ T Lymphocytesin Biliary Atresia. Pediatr Res 70, 555–559 (2011). https://doi.org/10.1203/PDR.0b013e318232a949
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1203/PDR.0b013e318232a949
This article is cited by
-
Biliary atresia-specific deciduous pulp stem cells feature biliary deficiency
Stem Cell Research & Therapy (2021)
-
MicroRNA-29b/142-5p contribute to the pathogenesis of biliary atresia by regulating the IFN-γ gene
Cell Death & Disease (2018)
-
Update on investigations pertaining to the pathogenesis of biliary atresia
Pediatric Surgery International (2017)
-
Global methylation, oxidative stress and relative telomere length in biliary atresia patients
Scientific Reports (2016)