Abstract
Preterm infants are at risk of developing sepsis, necrotizing enterocolitis (NEC), chronic lung disease (CLD), and retinopathy of prematurity (ROP). We used high-throughput mass spectrometry to investigate whether cord blood proteins can be used to predict development of these morbidities. Cord blood plasma from 44 infants with a birth weight of <1500 g was analyzed by surface-enhanced laser desorption/ionization time-of-flight mass spectrometry (SELDI-TOF). Six infants developed ROP ≥stage II, 10 CLD, three sepsis, and one NEC. We detected 814 protein signals representing 330 distinct protein species. Nineteen biomarkers were associated with development of ≥stage II ROP [false-discovery rate (FDR) <5%] and none with CLD. Several proteins with molecular weight (Mr) 15–16 kD and pI 4–5 were detected with increased abundance in infants with ROP, while similar Mr proteins with pI 7–9 were less abundant in these patients. Sodium dodecylsulfate–polyacrylamide gel electrophoresis (SDS-PAGE) and sequence analysis identified these proteins as α-, β-, and γ-globin chains. Partial deamidation of Asn139 in β-globin chains was observed only in the pI 4–5 proteins. We conclude that there are several promising biomarkers for the risk of ROP. Deamidation of globin chains is especially promising and may indicate underlying prenatal pathologic mechanisms in ROP. Validation studies will be undertaken to determine their clinical utility.
Similar content being viewed by others
Main
Preterm infants are at risk of developing complications such as sepsis, NEC, CLD, and ROP (1–6). Approximately 0.3%–3.2% of low birth weight infants (501–1500 g) have more than one of these morbidities (3). The reason why some preterm infants of the same gestational age have a mild neonatal course while others go on to develop particular morbidities is of great interest to neonatologists. It has been hypothesized that the release of proinflammatory cytokines such as interleukin (IL)-6, IL-1β, IL-8, tumor necrosis factor α in amniotic fluid and other body fluids before birth may play a role in several adverse outcomes, including CLD, periventricular leukomalacia (PVL), and cerebral palsy (4,7–9).
Biomarker molecules that could identify the onset of or susceptibility for a particular disease would be valuable. Low molecular weight proteins present in plasma offer the potential to be such biomarkers. Several techniques can be used to identify these proteins or potential biomarkers. Antibody arrays can be useful in detection of proteins in disease states where there is a known panel of proteins implicated in the disease process. However, other methods for detection are needed in multifactorial diseases where the identity of the protein involved in the molecular pathogenesis is not yet known. Two-dimensional (2-D) electrophoresis is one such method that is useful as a screening technique for identification of proteins >20 kD. Another strategy is to screen and identify protein biomarkers using mass spectrometry (MS), an analytical technique that identifies unknown compounds by measuring the masses of individual molecules converted to ions (10). In this study, we used SELDI-TOF MS, a technique that is useful for high-throughput screening of several samples simultaneously and that is capable of determining the molecular mass of peptides and proteins in a complex mixture, without breaking apart the analytes (11,12). This technique is complementary to 2-D electrophoresis because it is most useful for the screening and identification of smaller peptides and proteins (2–30 kD). Analysis of plasma proteins using SELDI and bioinformatics, the technique used in this study, has been applied successfully for the detection of biomarkers for prostate and ovarian cancer, renal allograft rejection, liver disease, intra-amniotic infection, Alzheimer's disease, and Creutzfeldt-Jakob disease (13–19). In SELDI, plasma or serum proteins are adsorbed onto surfaces designed to bind proteins by different mechanisms. A matrix molecule dissolved in H2O/acetonitrile is typically added, and the matrix and adsorbed proteins allowed to cocrystallize on the SELDI surface. Upon irradiation with an ultraviolet laser beam, the matrix plus proteins are desorbed from the surface, the matrix becomes ionized, and transfers charge to the peptides/proteins. The molecular mass of the ions is then determined from the time of flight to a detector after acceleration in an electric field.
The goal of this study was to use SELDI-TOF to generate mass spectra and bioinformatics to analyze these spectra for the detection and identification of biomarkers in cord blood that would be predictive of future complications such as sepsis, NEC, CLD, PVL, and ROP in preterm infants. We hypothesized that intrapartum events such as hypoxia, stress, or infection would affect protein patterns in cord blood plasma and might influence the development of specific morbidities in preterm infants.
METHODS
Patient and sample collection.
Cord blood samples from infants with a birth weight <1500 g delivered at participating institutions were collected immediately after birth in heparinized tubes. Samples were centrifuged at 5000 × g, and the plasma was stored at −80°C. Outcome data on all infants were obtained from the database maintained on these infants admitted to the neonatal intensive care unit. Gestational age was determined by the best obstetric estimate using the last menstrual period and results of an ultrasound examination. The definition of the various morbidities was that used for collection of data for the National Institute of Child Health and Human Development (NICHD) database. NEC was diagnosed if it was ≥stage II as described by the modified Bell's criteria (20); CLD was defined as oxygen need at 36 wk of gestation, with or without radiographic data (21); and early-onset sepsis was defined as sepsis at <7 d of age. The stage of ROP used for the analysis was the highest stage of ROP recorded during the hospitalization by retinal examination by a trained ophthalmologist. Samples were processed after data collection on neonatal outcomes was available. The Human Subjects Committee at both institutions approved the study.
SELDI-TOF MS.
All plasma samples were processed simultaneously, and each sample was examined in duplicate. Plasma (20 μL) was initially denatured with 30 μL 9 mol/L urea/2%CHAPS/50 mmol/L Tris (pH 9) and applied to a strong anion exchange bead at pH 9 (Q-ceramic HyperD F, Ciphergen Biosystems, Fremont, CA) in a 96-well Silent Screen filtration plate (Millipore, Bedford, MA). Using a pipetting robot (Biomek2000, Beckman-Coulter Inc.), the sample-loaded anion exchange beads were stepwise extracted (200 μL each) with five buffers of decreasing pH, from pH 9 to pH 3. A final sixth fraction was obtained by extracting the beads with 33.3% isopropanol/16.7% acentonitrile (ACN)/50% H2O/0.2% trifluoroacetic acid (TFA). An aliquot (10 μL) of each of the above six fractions was applied to each spot of three different SELDI ProteinChip arrays (Ciphergen), chosen to bind proteins by different mechanisms, using protocols essentially as described by the manufacturer. Samples were bound to (1) CM10 ProteinChips, a weak cation exchanger that binds proteins with net positive charge, in 0.1 mol/L sodium acetate buffer at pH 4; (2) H50 ProteinChips, a hydrophobic surface that functions as a reverse phase sorbent, in a buffer containing 10% ACN/0.1% TFA; and (3) IMAC30 ProteinChips, the immobilized metal affinity capture chip charged with Cu2+, a surface that binds histidine, tryptophan, and cysteine residues, in a buffer of 0.1 mol/L sodium phosphate/0.5 mol/L NaCl, pH 7. After binding for 1 h, the ProteinChips were washed three times with the appropriate binding buffer, followed by two brief washes with water, and allowed to dry. The energy-absorbing matrix molecule sinapinic acid (Sigma Chemical Co.) was then added to each ProteinChip surface (two applications of 0.5 μL of a saturated solution in 50% ACN/0.5% TFA). Mass spectra of proteins that bound to each type of protein chip surface were generated for each of the six fractions of each plasma sample using a Ciphergen PBSIIc mass spectrometer, with a total of 36 spectra generated for each sample (six fractions on three types of arrays, run in duplicate). Spectra were externally calibrated using spectra of a mixture of protein standards in the range of 7–30 kD.
Bioinformatics.
Spectra were analyzed by simultaneous spectrum analysis (SSA), an algorithm that identifies, quantifies, and processes protein peak information from SELDI-TOF MS spectra (22; http://med.stanford.edu/labs/harvey_cohen/software.html). The SSA algorithm facilitates identification of the location and boundaries of peaks in individual spectra and thus identification of all possible peaks across all the spectra. SSA improves the signal-to- noise ratio for detection of peaks by averaging spectra from all samples on a given ProteinChip surface and fraction. It then calculates the area of each peak in each spectrum. The spectra are normalized by a procedure in which peak magnitudes that are conserved across duplicate spectra are weighted more than highly variable peaks. Peaks that are noisy (poorly reproducible among replicates) are eliminated if they do not pass an F test with 99% confidence. Analysis parameters were minimum and maximum peak widths of 0.3% and 0.8% from 3000 mass-to-charge ratio (m/z) to 13,000 m/z, 0.2% and 0.8% from 13,000 m/z to 18,000 m/z, and 0.8% and 2.5% from 18,000 m/z to 75,000 m/z.
SELDI-TOF MS peaks in the spectra of fractionated plasma samples are highly redundant, e.g. the same protein is often detected on different protein chip surfaces or with two different ionizations. Therefore, a clustering algorithm was used to create composite features by averaging protein peaks likely to represent the same protein species or highly correlated fragments or modifications (manuscript in preparation). These features were tested using the Wilcoxon rank-sum test to determine differences between infants who developed a particular complication and those who did not. Multiple-hypothesis testing was controlled by the FDR method (23,24). FDR controls for multiple hypotheses testing by setting the threshold for statistical significance such that there are 20 times more significant results than expected by chance (fewer than 5% of reported results are expected to be type I false-positive errors).
SDS-PAGE and MALDI-TOF/TOF MS.
Identification of biomarkers by SELDI was followed by SDS-PAGE and MALDI-TOF/TOF-MS to determine the identify of the biomarkers. Of the 19 biomarkers identified by SELDI in ROP infants, SDS-PAGE and MALDI-TOF/TOF-MS was used to further identify one 15-kD protein peak or biomarker that was present in fraction 4 on CM10 surfaces at a high magnitude in all six infants with ≥stage II ROP (see Results section). To identify this protein peak, samples from all six anion exchange fractions from one patient with ROP were run on a one-dimensional (1-D) SDS gel. A band at approximately 15 kD from fractions 2 (pH 7 extract of anion exchange beads) and 4 (pH 4 extract) was isolated, and the proteins were extracted from the gel piece in 50% formic acid/25% acetonitrile/15% isopropyl alcohol/10% H2O. The extracted proteins were reanalyzed on the SELDI ProteinChip CM10 arrays to verify that they were the same biomarker that had been identified by SELDI and then digested with trypsin. Digests of protein extracted from both fractions were spotted to matrix-assisted laser desorption/ionization (MALDI) target plates and analyzed by MALDI-TOF MS (Applied Biosystems 4700), a MS method in which a solid matrix is used to obtain a peptide mass fingerprint (25). The 10 most abundant ions were analyzed by tandem mass spectrometry (MS/MS) to obtain the sequence of these peptides. These sequences were analyzed by the Mascot search engine to identify the specific protein. Liquid chromatography (LC)-MALDI analysis of tryptic peptides was performed with a Michrom C18 reverse-phase column (0.1 mm × 15 cm) eluted with a gradient of acetonitrile in water containing 0.1% formic acid and 0.005% heptafluorobutyric acid. Samples were collected directly onto stainless steel MALDI target plates. Peptides were sequenced by MS/MS to identify differences in posttranslational protein modifications between fractions 2 and 4.
RESULTS
Patient demographics.
A total of 44 patient samples were analyzed. The median birth weight was 1001 g (range, 482–1489) and the median gestational age was 28 wk (range, 24–32). There were 22 male and 22 female infants, 12 African American, 14 white, 7 Asian, and 11 Hispanic. The complications that were present in this group of infants were CLD (10 positive, 30 negative, four unknown), ROP >stage II (six positive, 38 negative), sepsis (three positive, 39 negative, two unknown), and NEC (one positive, 41 negative, two unknown). The number of patients who developed sepsis and NEC were not sufficient to perform statistical tests. The analysis was therefore limited to comparison of infants with and without ROP >stage II and with and without CLD. Details of the ROP status in the six patients are presented in Table 1.
Analysis of protein spectra.
We detected 814 protein peaks with high confidence. Since the same protein is often detected on different chips or with two different ionizations, the clustering algorithm was applied to create composite features by averaging correlated peaks. There were 329 signals likely to represent distinct proteins. These were reduced to 249 composites containing a single protein peak, 68 containing two, 34 containing between three and 14, and eight containing 15 or more. These eight appear to represent highly abundant proteins (e.g. albumin) that appear at several ionization states and in several chromatographic fractions.
CLD.
No differences were seen between patients who did or did not develop CLD. The lowest p value by the Wilcoxon rank-sum test was 0.0025 (pI 7 fraction, H50 hydrophobic surface, m/z 5498), which is expected to occur by chance 0.82 times in this data set, and the lowest FDR was 68%.
ROP.
There were 19 biomarkers that correlated strongly with eventual appearance of ROP ≥stage II with FDR <0.05. FDR of 0.05 corresponded to selecting biomarkers with p < 0.0028 (Table 2). The available data are not yet sufficient to develop and validate these proteins as clinical biomarkers for ROP. However, we wanted to determine whether any of these proteins or some combination thereof has the potential to serve as an early clinical biomarker for ROP. We evaluated each of the 19 biomarkers as a classifier that would maximize the fraction of correct classifications with each group (ROP versus no-ROP) given equal total weight.
Classification results using leave-one-out cross-validation are shown in Table 2. The table lists the 19 biomarkers arranged by effect size (i.e. the biomarkers most likely to be able to separate the ROP from no-ROP group), listing the relevant protein signals and the magnitude between-group differences. Sixteen of the 19 biomarkers were lower in the ROP patients, while three biomarkers, a 15-kD peak (see below) in the fourth chromatographic fraction (pH 4 elution) (biomarker 2), a molecule at 34.4 kD in the fifth chromatographic fraction (pH 3 elution) (biomarker 3), and a molecule at 7.6 kD in the sixth chromatographic fraction (organic fraction) on the H50 surface (hydrophobic) (biomarker 4) were higher in ROP patients.
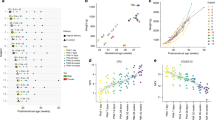
Several of the 19 biomarkers performed well as classifiers (i.e. able to separate the ROP from no-ROP), particularly biomarkers with molecular weight (Mr) from 15–16 kD in several chromatographic fractions. One of these 15- to 16-kD proteins was present in increased abundance in ROP patients in the pH 4 fraction (biomarker 2) (Fig. 1A). We realize that this does not constitute a validated test because potential biomarkers were preselected using all the data.
(A) Protein peak magnitude of the deamidated globin peaks (15–16 kD, pH fraction 4–5, weak-cation exchange surface). Units are arbitrary and shifted to have a mean of 0. The biomarker is notably higher in infants who developed ROP ≥stage II (p = 0.0001, FDR <0.05). ROP < stage II (diamonds); ROP ≥stage II (squares). (B) Deamidated globin peak magnitude vs gestational ages at birth shows that the biomarker is not related to chronological age.
Analysis and identification of an ROP biomarker.
We investigated the 15-kD biomarkers in greater detail by isolating the relevant proteins in all six fractions from one patient with ROP using SDS-PAGE 1-D gel electrophoresis and peptide fingerprinting and sequencing for identification (see Methods section). Detailed analysis by MALDI-TOF/TOF of proteins isolated from fractions 2 (pH 7) and 4 (pH 4) identified α- (six peptides, 58% sequence coverage), β- (10 peptides, 73% sequence coverage) and γ-(nine peptides, 78% sequence coverage) globin chains in both fractions. The presence of globin chains in the pH 4 fraction was surprising because these proteins are expected in higher pH fractions (pI for α-, β-, and γ-globins are 8.7, 6.8, and 6.7, respectively). We therefore searched for posttranslational modifications that could have affected the pI of the protein. A peptide with a singly charged monoisotopic mass of 1150.63 with the amino acid sequence VVAGVADALAHK (Fig. 2A) was observed in the pH 4 fraction but not the higher pH 7 fraction. This peptide corresponds to amino acids 133–144 of β-globin with deamidation of Asn 139. The corresponding peptide with monoisotopic mass of 1149.63 and the predicted sequence of β-globin residues 133–144 (VVAGVANALAHK, Fig. 2B) was observed to elute from the reverse phase column approximately 3.6 min later than the deamidated peptide. In summary, MALDI-TOF/TOF analysis identified the biomarker in the pH 4 fraction (biomarker 2) as a combination of α-, β-, and γ-globin chains with partial deamidation on Asn139 of the β-globin chain.
Figure 3 shows examples of mass spectra of the pH 4 globin biomarker in the six ROP ≥stage II infants next to six gestational age-matched infants without ROP. Figure 1A shows how the biomarker is distributed in the two groups (arbitrary units, shifted to overall mean of zero).
Protein mass spectra showing the deamidated globin biomarker peaks with molecular weights from 3 to 25 kD (top) and in detail from 15 to 16 kD (bottom). Biomarker peaks are highlighted in gray. Shown are spectra of the pH 4–5 fraction from all six patients with ROP ≥stage II on the CM10 surface against similar spectra from six gestational age–matched patients ROP <stage II. These biomarkers were isolated using a 1-D SDS gel and identified using MALDI-TOF MS to map peptides produced by trypsin digestion.
Besides biomarker 2, we were able to identify biomarkers 18 and 6. Biomarker 18 with the same Mr in the pH 7 fraction was similarly identified as the three globin chains, but no deamidation was observed. Of note, the deamidated globin chains were in increased abundance in samples from patients who developed ROP ≥stage II, while the other globin chains were in decreased abundance in those samples (fractions 2 and 6).
Using similar procedures, biomarker 6 (13.9 kD) in the pH 4 and pH 3 fractions was identified as transthyretin. This is a composite feature including transthyretin in two fractions and on two surfaces (weak cation exchanger and metal binding) as well as its second ionizations. This biomarker performed very well in classification (five of six ROP and 35 of 38 non-ROP correct).
We initially suspected that the 7.6-kD biomarker in the organic fraction on the hydrophobic surface (biomarker 4) was a second ionization of globin chains. However, it is negatively correlated to globin chains in the same fraction (R = −0.39, p < 0.02), which indicates that it is not a higher ionization of the globin chains.
Confounding variables: gestational age, birth weight, and collection center.
Confounding variables are a constant risk in this type of biomarker discovery, especially with small sample sizes. To check that we are not identifying biomarkers for low gestational age, we used Spearman's correlation to determine whether there was any correlation between each biomarker and gestational age. This is a nonparametric statistic that does not assume a linear relationship. Three biomarkers were correlated with gestational age with p ≤ 0.01 (biomarkers 9, 15, 17) and one with p = 0.04 (biomarker 3) (data available on request). None of the other 15 had significant correlation with gestational age (the next lowest had p = 0.10), and the deamidated globin biomarker did not correlate with gestational age (Fig. 1B).
Similar analyses with respect to birth weight found the same set of three biomarkers (9,15,17), as well as biomarker 19, correlated with birth weight with p < 0.02, and biomarker 5 with p = 0.04. None of the globin biomarkers or transthyretin correlated with gestational age or birth weight.
We also compared each biomarker in patient samples collected at the two institutions to check for systematic bias in collection. The patients with ROP were all at one center and thus excluded from this test to avoid having the ROP/non-ROP difference appear as a difference between institutions. Using the Wilcoxon rank-sum test on the non-ROP patients, the smallest p value was 0.024 for biomarker 3. With 19 biomarkers, a p value this low has a 37% probability of occurring by chance. No other p values suggested any systematic bias.
DISCUSSION
Our preliminary observation of the presence of higher levels of deamidated globin chains in the cord blood of preterm infants who later developed ≥stage II ROP suggests that factors present before birth may influence development of ROP in these infants. Deamidation is a nonenzymatic modification of asparaginyl and glutaminyl residues in peptides and proteins, resulting in aspartate and glutamate residues, respectively. It therefore confers a negative charge that can change the folding and conformation of the protein (26). This process is among the most commonly occurring posttranslational modifications, although its role is not completely understood. Some investigators have considered deamidation to be an undesirable form of protein damage, and it has been implicated in several pathologic conditions, including cataracts (27,28) and Alzheimer's disease (29). Another possibility is that deamidation is simply a dysregulation of normally occurring processes in a cell (30,31).
We did not find any correlation of this biomarker with CLD, thus arguing in favor of the biomarker being specific for stage 2 or greater ROP. Also, the marker was not found to be significantly correlated with gestational age or birth weight, thus confirming that it was not simply a marker of preterm birth. We were unable to examine other morbidities such as sepsis or NEC in this population and thus cannot comment on the role of this biomarker in other disease states in preterm infants.
We found biomarkers in the plasma of preterm infants for a presumptive localized disease that affects the retina. This raises the possibility that proteomics of low molecular weight plasma proteins can reveal molecules either involved in the pathogenesis of various disease states or indicative of pathogenesis.
Also, the detection of posttranslational modifications as potential biomarkers confirms the importance of multidimensional high-resolution separations in proteomic investigations. For example, examining the total content of globin chains would not have revealed deamidation, and there was no a priori reason to suspect that this protein modification had occurred.
One of the difficulties in conducting these analyses in preterm infants is obtaining a sufficient number of samples and in collecting adequate amounts of plasma. The results of this study, although novel and of high statistical significance, will need to be validated in a separate data set of preterm infants with and without ROP. Such validation will require another data set, as well as confirmation of the results using complementary techniques such as enzyme-linked immunosorbent assay or Western blotting wherever possible.
Abbreviations
- CLD:
-
chronic lung disease
- FDR:
-
false-discovery rate
- LC-MS/MS:
-
liquid chromatography–tandem mass spectrometry
- MALDI-TOF:
-
matrix-assisted laser desorption/ionization time-of-flight mass spectrometry
- NEC:
-
necrotizing enterocolitis
- PVL:
-
periventricular leukomalacia
- ROP:
-
retinopathy of prematurity
- SELDI-TOF:
-
surface-enhanced laser desorption/ionization time-of-flight mass spectrometry
References
Wilson-Costello D, Friedman H, Minich N, Fanaroff AA, Hack M 2005 Improved survival rates with increased neurodevelopmental disability for extremely low birth weight infants in the 1990s. Pediatrics 115: 997–1003
Fanaroff AA, Hack M, Walsh MC 2003 The NICHD neonatal research network: changes in practice and outcomes during the first 15 years. Semin Perinatol 27: 281–287
Lemons JA, Bauer CR, Oh W, Korones SB, Papile LA, Stoll BJ, Verter J, Temprosa M, Wright LL, Ehrenkranz RA, Fanaroff AA, Stark A, Carlo W, Tyson JE, Donovan EF, Shankaran S, Stevenson DK 2001 Very low birth weight outcomes of the National Institute of Child health and human development neonatal research network, January 1995 through December 1996. NICHD Neonatal Research Network. Pediatrics 107: E1
Jobe AH, Bancalari E 2001 Bronchopulmonary dysplasia. Am J Respir Crit Care Med 163: 1723–1729
Cryotherapy for Retinopathy of Prematurity Cooperative Group 1988 Multicenter trial of cryotherapy for retinopathy of prematurity: preliminary results. Pediatrics 81: 697–706
Stoll BJ, Hansen NI, Adams-Chapman I, Fanaroff AA, Hintz SR, Vohr B, Higgins RD, National Institute of Child Health and Human Development Neonatal Research Network 2004 Neurodevelopmental and growth impairment among extremely low-birth-weight infants with neonatal infection. JAMA 292: 2357–2365
Dammann O, Leviton A 1998 Infection remote from the brain, neonatal white matter damage, and cerebral palsy in the preterm infant. Semin Pediatr Neurol 5: 190–201
Viscardi RM, Muhumuza CK, Rodriguez A, Fairchild KD, Sun CC, Gross GW, Campbell AB, Wilson PD, Hester L, Hasday JD 2004 Inflammatory markers in intrauterine and fetal blood and cerebrospinal fluid compartments are associated with adverse pulmonary and neurologic outcomes in preterm infants. Pediatr Res 55: 1009–1017
Vigneswaran R 2000 Infection and preterm birth: evidence of a common causal relationship with bronchopulmonary dysplasia and cerebral palsy. J Paediatr Child Health 36: 293–296
Domon B, Aebersold R 2006 Mass spectrometry and protein analysis. Science 312: 212–217
Petricoin EF, Zoon KC, Kohn EC, Barrett JC, Liotta LA 2002 Clinical proteomics: translating benchside promise into bedside reality. Nat Rev Drug Discov 1: 683–695
Conrads TP, Fusaro VA, Ross S, Johann D, Rajapakse V, Hitt BA, Steinberg SM, Kohn EC, Fishman DA, Whitely G, Barrett JC, Liotta LA, Petricoin EF 3rd, Veenstra TD 2004 High-resolution serum proteomic features for ovarian cancer detection. Endocr Relat Cancer 11: 163–178
Rubin RB, Merchant M 2000 A rapid protein profiling system that speeds study of cancer and other diseases. Am Clin Lab 19: 28–29
Wagner M, Naik DN, Pothen A, Kasukurti S, Devineni RR, Adam BL, Semmes OJ, Wright GL Jr 2004 Computational protein biomarker prediction: a case study for prostate cancer. BMC Bioinformatics 5: 26
Clarke W, Silverman BC, Zhang Z, Chan DW, Klein AS, Molmenti EP 2003 Characterization of renal allograft rejection by urinary proteomic analysis. Ann Surg 237: 660–664
Poon TC, Hui AY, Chan HL, Ang IL, Chow SM, Wong N, Sung JJ 2005 Prediction of liver fibrosis and cirrhosis in chronic hepatitis B infection by serum proteomic fingerprinting: a pilot study. Clin Chem 51: 328–335
Gravett MG, Novy MJ, Rosenfeld RG, Reddy AP, Jacob T, Turner M, McCormack A, Lapidus JA, Hitti J, Eschenbach DA, Roberts CT Jr, Nagalla SR 2004 Diagnosis of intra-amniotic infection by proteomic profiling and identification of novel biomarkers. JAMA 292: 462–469
Lewczuk P, Esselmann H, Groemer TW, Bibl M, Maler JM, Steinacker P, Otto M, Kornhuber J, Wiltfang J 2004 Amyloid beta peptides in cerebrospinal fluid as profiled with surface enhanced laser desorption/ionization time-of-flight mass spectrometry: evidence of novel biomarkers in Alzheimer's disease. Biol Psychiatry 55: 524–530
Sanchez JC, Guillaume E, Lescuyer P, Allard L, Carrette O, Scherl A, Burgess J, Corthals GL, Burkhard PR, Hochstrasser DF 2004 Cystatin C as a potential cerebrospinal fluid marker for the diagnosis of Creutzfeldt-Jakob disease. Proteomics 48: 2229–2233
Walsh MC, Kliegman RM 1986 Necrotizing enterocolitis: treatment based on staging criteria. Pediatr Clin North Am 33: 179–201
Shennan AT, Dunn MS, Ohlsson A, Lennox K, Hoskins EM 1988 Abnormal pulmonary outcomes in premature infants: prediction from oxygen requirement in the neonatal period. Pediatrics 82: 527–532
Carlson SM, Najmi A, Whitin JC, Cohen HJ 2005 Improving feature detection and analysis of surface-enhanced laser desorption/ionization-time of flight mass spectra. Proteomics 5: 2778–2788
Benjamini Y, Hochberg Y 1995 Controlling the false discovery rate—a practical and powerful approach to multiple testing. J R Statist Soc Ser B 57: 289–300
Storey JD, Tibshirani R 2003 Statistical significance for genomewide studies. Proc Natl Acad Sci U S A 100: 9440–9445
Yates JR 2000 Mass spectrometry—from genomics to proteomics. Trends Genet 16: 5–8
Robinson NE, Robinson ZW, Robinson BR, Robinson AL, Robinson JA, Robinson ML, Robinson AB 2004 Structure-dependent nonenzymatic deamidation of glutaminyl and asparaginyl pentapeptides. J Pept Res 63: 426–436
Hoenders HJ, Bloemendal H 1983 Lens proteins and aging. J Gerontol 38: 278–286
Takemoto L, Emmons T, Granstrom D 1990 The sequences of two peptides from cataract lenses suggest they arise by deamidation. Curr Eye Res 9: 793–797
Watanabe A, Takio K, Ihara Y 1999 Deamidation and isoaspartate formation in smeared tau in paired helical filaments. Unusual properties of the microtubule-binding domain of tau. J Biol Chem 274: 7368–7378
Deverman BE, Cook BL, Manson SR, Niederhoff RA, Langer EM, Rosova I, Kulans LA, Fu X, Weinberg JS, Heinecke JW, Roth KA, Weintraub SJ 2002 Bcl-xL deamidation is a critical switch in the regulation of the response to DNA damage. Cell 111: 51–62
Robinson NE 2002 Protein deamidation. Proc Natl Acad Sci U S A 99: 5283–5288
Acknowledgements
The authors thank the neonatology postdoctoral fellows and staff, Barbara Piane and the General Clinical Research Center nurses without whom this work would not be possible. They also acknowledge the support of the NICHD Neonatal Research Network for funding the Generic Database.
Author information
Authors and Affiliations
Corresponding author
Additional information
This work was supported in part by grant 5 M01 RR000070 from the National Institutes of Health and by the Pediatric Research Fund, Packard Hospital, Palo Alto, CA (A.M., H.J.C.).
Rights and permissions
About this article
Cite this article
Madan, A., El-Ferzli, G., Carlson, S. et al. A Potential Biomarker in the Cord Blood of Preterm Infants Who Develop Retinopathy of Prematurity. Pediatr Res 61, 215–221 (2007). https://doi.org/10.1203/pdr.0b013e31802d776d
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1203/pdr.0b013e31802d776d
This article is cited by
-
Cord blood transforming growth factor-β-induced as predictive biomarker of retinopathy of prematurity in preterm infants
Graefe's Archive for Clinical and Experimental Ophthalmology (2023)
-
Identifying technical aliases in SELDI mass spectra of complex mixtures of proteins
BMC Research Notes (2013)