Abstract
The isobutyryl-CoA dehydrogenase (IBD) enzyme is involved in the degradation of valine. IBD deficiency was first reported in 1998 and subsequent genetic investigations identified acyl-CoA dehydrogenase (ACAD) 8, now IBD, as the gene responsible for IBD deficiency. Only three individuals homozygous or compound heterozygous for variations in the IBD gene have been reported. We present IBD deficiency in an additional four newborns with elevated C4-carnitine identified by tandem mass spectrometry (MS/MS) screening in Denmark and the United States. Three showed urinary excretions of isobutyryl-glycine, and in vitro probe analysis of fibroblasts from two newborns indicated enzymatic IBD defect. Molecular genetic analysis revealed seven new rare variations in the IBD gene (c.348C>A, c.400G>T, c.409G>A, c.455T>C, c.958G>A, c.1000C>T and c.1154G>A). Furthermore, sequence analysis of the short-chain acyl-CoA dehydrogenase (SCAD) gene revealed heterozygosity for the prevalent c.625G>A susceptibility variation in all newborns and in the first reported IBD patient. Functional studies in isolated mitochondria demonstrated that the IBD variations present in the Danish newborn (c.409G>A and c.958G>A) together with a previously published IBD variation (c.905G>A) disturbed protein folding and reduced the levels of correctly folded IBD tetramers. Accordingly, low/no IBD residual enzyme activity was detectable when the variant IBD proteins were overexpressed in Chang cells.
Similar content being viewed by others
Main
The acyl-CoA dehydrogenases (ACAD) (EC 1.3.99) are a family of nuclear-encoded mitochondrial enzymes involved in the metabolism of fatty acids and branched chain amino acids (1–3). Inherited deficiencies of the ACADs are important causes of metabolic disorders. During the catabolism of valine and isoleucine, the branched side chains are converted into isobutyryl-CoA and 2-methylbutyryl-CoA, respectively. These dehydrogenations were initially suggested to be carried out by one enzyme, the short branched chain acyl-CoA dehydrogenase (SBCAD) (2,4). However, identification of the ACAD8 gene, which encodes isobutyryl-CoA dehydrogenase (IBD), and subsequent functional analysis, demonstrated that IBD is responsible for the conversion of isobutyryl-CoA to methacrylyl-CoA in the valine metabolism (5,6). The first patient with IBD deficiency was reported in 1998 (7) and genetic analysis revealed homozygosity for the IBD c.905G>A (Ala302Gln) variation (3). Tandem mass spectrometry (MS/MS) based newborn screening is established in several countries world wide, and allows early detection and intervention of a wide range of metabolic disorders (8). With the addition of IBD deficiency to the list of metabolic disorders, the number of metabolites that must be traceable to identify specific inborn errors of metabolism has increased. Acyl-carnitine profiles representing elevated C4-carnitine may either indicate IBD deficiency or short-chain acyl-CoA dehydrogenase (SCAD) deficiency, as the routine MS/MS analysis does not allow discrimination between the C4-metabolites, isobutyryl-carnitine and butyryl-carnitine, which accumulate in these enzyme defects (7,9). SCAD (EC 1.3.99.2) is a mitochondrial fatty acid oxidation enzyme involved in the dehydrogenation of butyryl-CoA into crotonyl-CoA. To discriminate between SCAD deficiency and IBD deficiency, the MS/MS analysis of newborns with elevated C4-carnitine requires further evaluation for follow-up. Gas chromatography/mass spectrometry (GC/MS) analysis is used for quantification of the urine organic acids, ethylmalonic acid (EMA), methylsuccinic acid and n-butyrylglycine, all indicative of SCAD deficiency (9), while isobutyryl-glycine is suggestive of an IBD-related defect (10). However, the isobutyryl-glycine may not be elevated and genetic analysis should be performed to identify a possible defect in either SCAD or IBD. Three children have been reported with variations in the IBD gene; two newborns (11) and a clinically symptomatic patient (3,7). Yet another newborn was identified by MS/MS screening, but genetic analysis was not performed (9). The functional consequences of the identified IBD variations have not been evaluated, except for the Ala302Gln (c.905G>A) variant protein, which was inactive upon expression in E. coli and in mammalian cells (3). Protein structure and folding studies have demonstrated that disease-associated variations in the ACAD genes often cause protein misfolding and instability (12,13). The SCAD enzyme was shown to be very susceptible to misfolding due to a number of amino acid alterations (13), and variations in the medium-chain acyl-CoA dehydrogenase (MCAD) enzyme may also cause misfolding and/or increased thermal instability (12,14,15).
In this study we identify seven new rare variations in the IBD gene in four newborns with elevated C4-carnitine detected in Danish and US MS/MS screening programs. We investigate the functional consequences of some of the newly identified IBD variations by overexpressing the variant IBD proteins in Chang cells. Protein folding abilities are evaluated using an established mitochondrial folding/processing assay.
METHODS
Patients.
Newborn A is a healthy Danish girl born at term after a normal pregnancy with birth weight 3,900 g, length 55 cm, and head circumference 38 cm. She was the only newborn identified with elevated C4-carnitine in the Danish MS/MS screening of 50,000 newborns. The blood spot C4-carnitine was 1.1 μM (cut-off level < 0.93 μM) on day 8, decreasing to 0.8 μM (cut-off level < 0.52 μM) in a confirmatory sample obtained on day 24. GC/MS analyses revealed slightly increased isobutyryl-glycine. Plasma amino acids were not elevated. Initial plasma carnitine was slightly reduced (total 29 μM; control 30–73 μM), while free carnitine was in the control range at follow-up at age 2.5 y. She receives no carnitine supplementation or other medications. Her psychomotor development, language development, and growth have been normal. She recovered normally from a number of intercurrent illnesses, but during these episodes she received extra carbohydrates.
Newborn B is a Native American girl born to non-consanguineous parents. She was small for a full-term infant. She was identified in the MS/MS screening of 9,198 babies in South Dakota, and all babies with elevated C4-carnitine were further evaluated by plasma acyl-carnitine and organic acid analysis. Newborn screening on day 1 showed elevated C4-carnitine (2.9 μM; cut-off level < 1.53 μM), followed by a repeated elevation on day 8 (2.6 μM). GC/MS analysis of urine revealed slightly increased EMA and unremarkable isobutyryl-glycine. Echocardiogram at age 1 y showed mild branch peripheral pulmonary stenosis, but no evidence of cardiomyopathy. At age 3 y, 10 mo (15.6 kg and 101.6 cm), she was completely asymptomatic with no significant developmental delays. She did have somewhat decreased muscle tone as an infant. She is currently receiving L-carnitine (32 mg/kg/d) therapy.
Newborns C and D were identified with elevated C4-carnitine in the MS/MS screening of 900,000 newborns in North Carolina, and all babies identified with elevated C4-carnitine and all were further evaluated by plasma acyl-carnitine profiles and urine organic acids (16).
Newborn C is an American-born girl of Caucasian ethnicity born after a normal pregnancy; no known consanguinity. Newborn screening on day 2 revealed elevated C4-carnitine (3.23 μM; cut-off level < 1.53 μM), and a repeated elevation (2.33 μM) was observed on day 18. A follow-up plasma acyl-carnitine profile revealed C4-carnitine of 1.52 μM (control < 0.46 μM), and urine organic acids showed elevated isobutyryl-glycine. Plasma C4-carnitine remained slightly elevated (median 0.54 μM, range 0.29-2.81 μM; control < 0.46 μM), and isobutyryl-glycine was infrequently present in urine. Plasma carnitine (total and free) was in the control range. Motor development and early speech development was normal; however, at age 5 y she received therapy for moderate expressive speech delay. Growth was normal, although height was at the 90–95th percentile, weight at the 75th percentile, and head circumference at the 50th percentile. The patient was treated with a valine-restricted diet, riboflavin (100 mg/d), and L-carnitine (300 mg/d) supplementation. During any febrile illness she received supplemental carbohydrates until symptoms had abated.
Newborn D is an Native American boy born after a normal pregnancy with birth weight 3,610 g and length 52.7 cm. He has two older, clinically unaffected siblings and there was no known consanguinity. Newborn screening on day 5 revealed elevated C4-carnitine (2.41 μM; cut-off level < 1.53 μM), and a repeated profile on day 37 showed 2.40 μM. A plasma acyl-carnitine profile revealed C4-carnitine of 1.84 μM (control < 0.46 μM), and urine organic acids revealed elevated isobutyryl-glycine. Plasma C4-carnitine level remained slightly elevated (median 1.07 μM, range 0.54–2.39 μM; control < 0.46 μM), and isobutyryl-glycine was infrequently present in urine. Plasma carnitine (total and free) was initially elevated (total 94 μM; control 25–66 μM), but dropped into the control range by 9 mo. Motor development and early speech development was normal; however, at 2 y he received therapy for speech delay. He has had recurrent ear infections with effusions. Growth was normal; although height was at the 75–90th percentile, weight at the 75th percentile, and head circumference at the 50th percentile. The patient was treated with a valine-restricted diet, riboflavin (100 mg/d), and L-carnitine (280–400 mg/d) supplementation. He had several episodes of lethargy during infancy, but evaluation by a pediatrician revealed no significant abnormalities. During any remarkable illnesses he received supplemental carbohydrates, until symptoms had abated.
Relevant informed consents have been obtained, and appropriate institutional review boards in Denmark and the US approved the use of control material.
In vitro probe analysis of cultured patient fibroblasts.
For in vitro probe analysis, skin fibroblasts from newborn C were incubated with 800 μM [U-13C]-valine (13C-labeling of all carbon atoms) and 400 μM L-carnitine according to (9). In two separate assays, fibroblasts from newborn B were incubated either with 800 μM [U-13C]-valine and 400 μM L-carnitine, or with 200 μM 16-2H3-palmitate (three deuterium labels on carbon-16 atom) and 400 μM L-carnitine as described in (7,17).
PCR amplification and sequencing.
DNA was extracted from whole blood or dried blood spots (Guthrie cards) using a kit (Gentra Systems). PCR amplification of IBD exons and surrounding introns was performed as described elsewhere (3). PCR fragments were sequenced on a 3100-Avant genetic analyzer using BigDye® Terminator v1.1 Cycle Sequencing kit (Applied Biosystems).
Allelic discrimination.
Samples were genotyped for the IBD c.409G>A and c.958G>A variations using unlabelled PCR primers and fluorescence labelled TaqMan probes for single nucleotide polymorphism (SNP) genotyping (Applied Biosystems). Details are available on request. The TaqMan™ PCR was performed on an ABI PRISM® 7700 (Applied Biosystems) according to the manufacturer's instructions.
Plasmid construction.
The coding region of human IBD cDNA was cloned between HindIII and ApaI sites of the pcDNA3.1+ expression vector (InVitrogen) (6). The variations, c.409G>A (Gly137Arg) and c.958G>A (Ala320Tha), were introduced into the pcDNA3.1-IBD wild type plasmid between PflmI and AscI, and BspEI and BlpI sites, respectively, using PCR-based mutagenesis. Correct orientation and sequence of the inserts were confirmed by sequencing. Construction of pcDNA3.1-IBD 905G>A (Arg302Gln) is published elsewhere (3).
Expression of wild type and variant IBD proteins.
Transfection of Chang cells was performed with 11.25-μg plasmid DNA using the FuGENE 6 reagent (Roche Molecular Biochemicals), and cells were harvested 48 h post-transfection. Western blot analysis was performed as previously described with an antibody raised in rabbits against a recombinant human IBD protein (3,6,13). The activity of overexpressed IBD enzymes were measured in lysates supplemented with isobutyryl-CoA (0.2 mM) with ferricenium ion as electron acceptor. The formation of methacrylyl-CoA was quantified using HPLC (HPLC) as described elsewhere (6).
Mitochondrial folding/processing assay.
IBD biogenesis/turnover was investigated in isolated rat liver mitochondria essentially as described in (13). In brief, in vitro transcription and translation of IBD cDNA in pcDNA3.1+ was performed in rabbit reticulocyte lysate (Promega) in the presence of [35S]-methionine (10μCi/μL, Amersham Biosciences). The reaction was stopped by adding cycloheximide at 0.15 μg/mL final concentration. Isolated rat liver mitochondria were incubated with the radiolabeled IBD protein for 30 min at 26°C (pulse), followed by 3 h at 37°C (chase). Aliquots were withdrawn at different time points (10, 30, 60, 120, and 180 min) and treated as described (13). The supernatant was analyzed by non-denaturing (native; 4–15% gel, BioRad) and denaturing (SDS; 12.5% gel, BioRad) PAGE. The pellet fraction was analyzed by denaturing PAGE (SDS; 12.5% gels, BioRad). Visualization of radiolabeled protein was performed on a Phosphoimager (STORM 804) and quantification was done using the ImageQuant software (Amersham Biosciences).
RESULTS
Genetic analysis of children with elevated c4-carnitine.
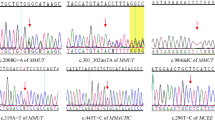
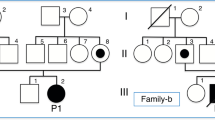
Newborn A was identified in the Danish MS/MS newborn screening with an acyl-carnitine profile showing C4-carnitine above the cut-off level (Table 1). GC/MS analysis of urine from newborn A did not show increased levels of EMA, a biochemical marker of SCAD deficiency, but documented small amounts of isobutyryl-glycine, pointing to an IBD-related defect. Accordingly, we performed sequence analysis of all exons and flanking introns in the IBD gene. We found heterozygosity for two variations, c.409G>A in exon 4 and c.958G>A in exon 9, causing the amino acid substitutions Gly137Arg and Ala320Thr, respectively (Fig. 1). Analysis of the IBD gene in both parents showed paternal inheritance of the c.409G>A variation, while c.958G>A was maternally inherited, thus confirming compound heterozygosity in newborn A. Carrier frequencies of these variations in the Danish population were examined by analysis of 415 control individuals using a specific assay (see Methods section). One carrier of the c.409G>A variation was identified and confirmed by sequence analysis, whereas no c.958G>A carriers were found. Data are summarized in Table 1.
Three newborns (B, C, and D) were identified with elevated blood spot C4-carnitine (Table 1) in US newborn screening programs. Follow-up GC/MS analysis of urine samples from newborns C and D revealed small amounts of isobutyryl-glycine, suggesting an IBD-related defect. Fibroblasts from newborns B and C were tested by in vitro probe analysis; fibroblasts were not obtained from newborns A and D. In two separate assays, fibroblasts from newborn B and controls were incubated with either 16-2H3-palmitate or [U-13C]-valine according to published methods (7). Incubation with 16-2H3-palmitate and L-carnitine revealed a three times higher level of unlabelled C4-carnitine compared with 2H3-butyryl-carnitine, and incubation with [U-13C]-valine and L-carnitine resulted in 20-fold higher accumulation of [U-13C]-isobutyryl-carnitine, but also a 2-fold higher level of unlabelled C4-carnitine. Fibroblasts from newborn C were incubated with [U-13C]-valine and L-carnitine, and increased accumulations of [U-13C]-isobutyryl-carnitine were detectable in the culture medium (0.93 μM; control 0.013 ± 0.004) (9). These results indicated decreased IBD enzyme activity in newborns B and C, but may also indicate a mild decrease of SCAD enzyme activity. We performed sequence analysis of all exons and flanking introns in the IBD gene from newborns B, C, and D. Newborn B was compound heterozygous for a c.455T>C (Met152Thr) variation in exon 4 and a c.1154A>G (Gln385Arg) variation in exon 10 (Fig. 1). Analysis of a healthy older half-sibling, who was heterozygous for the c.455T>C variation, indicated that c.455T>C and c.1154A>G segregate on separate alleles. In newborn C we found compound heterozygosity for a c.348C>A variation in exon 3, changing cysteine 116 to a stop codon (Cys116Stp), and a c.1000C>T variation (Arg334Cys) in exon 9 (Fig. 1). Analysis of the parents showed that these variations were located on different alleles. Newborn D was homozygous for a c.400G>T variation (Asp134Tyr) in exon 4 (Fig. 1), and analysis of the parents showed heterozygosity for this variation. To investigate if any of these variations are common in the US population, we analyzed exons 3, 4, 9 and 10 in 64 US control individuals. None of the variations were present. Data are summarized in Table 1.
Because the C4-carnitine elevations identified by the routine MS/MS analysis were not further separated and because fibroblasts from newborn B and from the first reported IBD patient (Patient E in Table 1; (3,7)) showed a two-fold increase of unlabelled C4-carnitine when incubated with [U-13C]-valine, a mild decrease of SCAD enzyme activity cannot be excluded. Accordingly, the promoter region and all exons in the SCAD gene were analyzed in newborns A, B, C, and D. We found heterozygosity for the prevalent c.625G>A SCAD susceptibility variation in all four newborns. No other variations were identified, except for heterozygosity for a silent c.669G>A (F55F) variation in newborn C. Also, Patient E showed heterozygosity for the c.625G>A variation. These findings demonstrated simultaneous presence of variations in the IBD gene and the prevalent c.625G>A SCAD susceptibility variation in all five individuals investigated.
Overexpression of variant IBD proteins in Chang cells.
To investigate the functional consequences of the IBD variations c.409G>A (Gly137Arg) and c.958G>A (Arg320Thr) identified in newborn A, and of the previously published c.905G>A (Arg302Gln) variation in patient E, these variant proteins were overexpressed in Chang cells. Isobutyryl-CoA conversion to methacrylyl-CoA was measured in whole cell extracts (Fig. 2) and the steady state levels of IBD protein were examined by Western blot analysis (Fig. 3). No activity was detectable in cells overexpressing IBD Gly137Arg and IBD Arg302Gln variant proteins, while IBD Ala320Thr expressing cells showed approximately 20% activity compared with wild type. Western blot showed indistinguishable amounts of steady state IBD protein in cells expressing wild type, Ala320Thr, and Arg302Gln protein variants. No steady state protein was detectable in Gly137Arg-expressing cells.
IBD activity assay. The conversion of isobutyryl-CoA into methacrylyl-CoA by the IBD enzyme was measured for wild type, Gly137Arg, Arg302Gln, Ala320Thr variant proteins and for an empty vector. Results are means of three independent transfections measured in triplicates. Standard deviations represent variation of the three transfections.
Western blot analysis. Chang cell extracts overexpressing IBD wild type, Gly137Arg, Arg302Gln, and Ala320Thr were separated by SDS-PAGE (10 μg total cellular protein was loaded in each lane) and probed with anti-IBD antibodies. An empty expression vector was used as negative control. Results are representative of three independent experiments.
In vitro folding of variant IBD proteins in isolated mitochondria.
We investigated biogenesis and turnover of variant IBD proteins using a previously described mitochondrial folding/processing assay (13). In a pulse-chase experiment, isolated rat liver mitochondria were incubated with in vitro synthesized radiolabeled IBD proteins (wild type, Gly137Arg, Arg302Gln or Ala320Thr) for 30 min at 26°C (“pulse”) followed by 3 h at 37°C (“chase”), and aliquots were removed at different time points. Solubilized mitochondria were fractionated by centrifugation and soluble IBD protein remained in the supernatant, whereas the pellet fraction contained insoluble IBD protein. Folding abilities of the newly imported IBD proteins were examined by native PAGE analysis (Fig. 4A). The native gel showed that all IBD proteins were imported into the mitochondria and associated with mitochondrial chaperones (hsp60). The hsp60 chaperone association has been verified by western blotting previously (13). Wild type IBD protein was rapidly released from the hsp60 chaperone complexes and assembled into tetramers. Also the IBD Arg302Gln variant appeared to be capable of forming tetrameric protein, albeit in reduced amounts compared with the wild type. The IBD Gly137Arg variant, which was undetectable on the Western blot (Fig. 3), exhibited prolonged association with hsp60 and was very slowly released from the chaperone complexes without forming tetramers, indicating that this IBD variation confers inability of the variant protein to reach the functional tetrameric conformation. Also the IBD Ala320Thr protein variant demonstrated increased chaperone retention with a low production of IBD tetramers.
Mitochondrial folding/processing assay. Isolated mitochondria were incubated with in vitro synthesized radiolabeled IBD wild type, Gly137Arg, Arg302Gln, or Ala320Thr proteins and aliquots were withdrawn at different time points. (A) Native PAGE analysis. IBD-hsp60 complexes and IBD tetramers are marked by arrows. (B) SDS-PAGE analysis of soluble (upper panel) and insoluble (lower panel) mitochondrial fractions. IBD monomers as well as precursor (uncleaved) and mature IBD are marked by arrows. (C) Quantification of IBD protein. Total IBD protein was estimated by adding soluble (white+gray) and insoluble (black) protein from 4B and set to 100% at time 10 min. Based on the native gels in 4A, the soluble IBD protein was further divided into tetramers (white) and a non-tetrameric part (gray) comprising hsp60 complexes and other soluble folding intermediates. Results are representative of two separate experiments.
To examine turnover of the variant IBD proteins, total amounts of soluble (Fig. 4A, upper panel) and insoluble (Fig. 4B, upper panel) IBD protein was investigated by SDS-PAGE. These results demonstrated a reduction in soluble IBD Gly137Arg variant protein, whereas wild type and the other variant IBD proteins, Arg302Gln and Ala320Thr, remained soluble throughout the incubation time. According to the insoluble fraction, IBD Gly137Arg variant proteins appeared to precipitate rather than being degraded, thus demonstrating a severely impaired folding. Figure 4C represents a quantitative illustration of the native and SDS-PAGE analyses in Fig. 4A and B.
DISCUSSION
MS/MS based newborn screening for abnormal acyl-carnitine profiles is now being used in many countries worldwide. In most cases the acyl-carnitine profiles pinpoint specific diseases; however, not all acyl-carnitines can be distinguished routinely. For instance, the accumulation of C4-carnitine may represent butyryl-carnitine or isobutyryl-carnitine, or a mixture. Accumulation of butyryl-carnitine is suggestive of SCAD deficiency, an enzymatic defect that may cause severe clinical symptoms, while accumulation of isobutyryl-carnitine may indicate IBD deficiency, an enzymatic defect of yet unclear clinical relevance. It is therefore crucial that the mechanisms underlying abnormal acyl-carnitine profiles are investigated in detail. We examined four newborns with elevated C4-carnitine identified by newborn screening. Three newborns (A, C, and D) showed urinary excretions of isobutyryl-glycine, an indicator of IBD deficiency, and fibroblasts from two newborns (B and C) revealed an enzymatic IBD defect using in vitro probe analysis. These biochemical findings strongly suggested an IBD-related defect rather than SCAD deficiency, and genetic analysis demonstrated variations on both alleles of the IBD gene in all newborns. We identified seven new variations in the IBD gene, thus more than doubling the total number of published IBD variations to 13 (3,11,18). So far, little is known about the prevalence of IBD deficiency, except that it has been detected in very few individuals (Table 1). In this study, we investigated 830 Danish control alleles for the two variations, c.409G>A and c.958G>A, identified in newborn A, and found only one carrier of the c.409G>A variation. We also estimated carrier frequencies of the IBD variations, c.348C>A, c.455T>C, c.400G>T, c.1000C>T, and c.1154A>G, present in the US newborns (B, C, and D) and found none of these in 128 control alleles. Some of the recently reported IBD variations (Met128Ile in exon 4 (11); Pro148Leu in exon 4, and Arg330Trp in exon 9 (18)) are also located in the sequenced exons, and we found none of these to be present in the US control group. These findings indicate a diverse spectrum of IBD gene variations with no prevalent variations.
Functional studies of the IBD variations from newborn A demonstrated reduced activity (< 20% of wild type) of the IBD Ala320Thr enzyme, whereas no residual activity was detectable for the IBD Gly137Arg variant when overexpressed in Chang cells. This was supported by experiments in isolated mitochondria which showed reduced amounts of IBD Ala320Thr tetramers, suggesting that the Ala320Thr variation disturbed, but not destroyed, the protein folding. The Gly137Arg variant demonstrated a severely impaired folding without production of functional IBD tetramers, but with increased tendency to precipitate. As in agreement with previous reports, the IBD Arg302Gln variant identified in patient E was found to have approximately same solubility as the wild type, but with higher levels of non-tetrameric IBD protein (3). Altogether, the investigated IBD variations, Gly137Arg (c.409G>A), Arg302Gln (c.905G>A) and Ala320Thr (c.958G>A), affected the IBD protein folding with consequently low or no production of functional IBD enzyme. These results add to the notion that enzymatic defects associated with variations in the ACADs usually affect folding and stability of the variant proteins (12–15).
Surprisingly, SCAD gene analysis revealed heterozygosity for the prevalent c.625G>A susceptibility variation in five (newborns A, B, C, D, and patient E) of five individuals investigated. Footnote 1 The c.625G>A variant of the SCAD gene is frequent in the European population and is considered to confer increased susceptibility to clinical SCAD deficiency in combination with other genetic or non-genetic factors (20). Although, previous reports have indicated that newborn blood spots from c.625G>A heterozygotes may not have significantly higher C4-carnitine concentrations than c.625G homozygotes (21,22), in vitro probe analysis of newborn B and patient E suggests that c.625G>A heterozygosity may contribute to an elevation of C4-carnitine in cultured fibroblasts as reported in (23). However, more cases should be investigated to clarify if there is an association between IBD deficiency and the SCAD c.625G>A variation.
Including the four newborns presented in this paper, IBD deficiency has now been reported in nine individuals worldwide (7,9,11,18). Although significant clinical symptoms were observed in the first patient reported, the IBD deficient newborns identified by screening have either remained asymptomatic or presented with milder clinical abnormalities. However, until more knowledge about IBD deficiency has accumulated, newborns with elevated C4-carnitine should be carefully monitored, as they may be at risk of developing clinical symptoms.
Notes
With a c.625G>A carrier frequency of 34.8 (19), the estimated probability that five random selected individuals will have this variation is very small: (0.348)5 = 0.0051 0.5.
Abbreviations
- ACAD:
-
acyl-CoA dehydrogenase
- EMA:
-
ethylmalonic acid
- GC/MS:
-
gas chromatography/mass spectrometry
- IBD:
-
isobutyryl-CoA dehydrogenase
- MS/MS:
-
tandem mass spectrometry
- SCAD:
-
short-chain acyl-CoAdehydrogenase
References
Ikeda Y, Dabrowski C, Tanaka K 1983 Separation and properties of five distinct acyl-CoA dehydrogenases from rat liver mitochondria. Identification of a new 2-methyl branched chain acyl-CoA dehydrogenase. J Biol Chem 258: 1066–1076
Ikeda Y, Tanaka K 1983 Purification and characterization of 2-methyl-branched chain acyl coenzyme A dehydrogenase, an enzyme involved in the isoleucine and valine metabolism, from rat liver mitochondria. J Biol Chem 258: 9477–9487
Nguyen TV, Andresen BS, Corydon TJ, Ghisla S, Abd-El Razik N, Mohsen AW, Cederbaum SD, Roe DS, Roe CR, Lench NJ, Vockley J 2002 Identification of isobutyryl-CoA dehydrogenase and its deficiency in humans. Mol Genet Metab 77: 68–79
Rozen R, Vockley J, Zhou L, Milos R, Willard J, Fu K, Vicanek C, Low-Nang L, Torban E, Fournier B 1994 Isolation and expression of a cDNA encoding the precursor for a novel member (ACADSB) of the acyl-CoA dehydrogenase gene family. Genomics 24: 280–287
Telford EA, Moynihan LM, Markham AF, Lench NJ 1999 Isolation and characterisation of a cDNA encoding the precursor for a novel member of the acyl-CoA dehydrogenase gene family. Biochim Biophys Acta 1446: 371–376
Andresen BS, Christensen E, Corydon TJ, Bross P, Pilgaard B, Wanders RJ, Ruiter JP, Simonsen H, Winter V, Knudsen I, Schroeder LD, Gregersen N, Skovby F 2000 Isolated 2-methylbutyrylglycinuria caused by short/branched-chain acyl-CoA dehydrogenase deficiency: identification of a new enzyme defect, resolution of its molecular basis, and evidence for distinct acyl-CoA dehydrogenases in isoleucine and valine metabolism. Am J Hum Genet 67: 1095–1103
Roe CR, Cederbaum SD, Roe DS, Mardach R, Galindo A, Sweetman L 1998 Isolated isobutyryl-CoA dehydrogenase deficiency: an unrecognized defect in human valine metabolism. Mol Genet Metab 65: 264–271
Rinaldo P, Tortorelli S, Matern D 2004 Recent developments and new applications of tandem mass spectrometry in newborn screening. Curr Opin Pediatr 16: 427–433
Koeberl DD, Young SP, Gregersen NS, Vockley J, Smith WE, Benjamin DK, An Y, Weavil SD, Chaing SH, Bali D, McDonald MT, Kishnani PS, Chen YT, Millington DS 2003 Rare disorders of metabolism with elevated butyryl- and isobutyryl-carnitine detected by tandem mass spectrometry newborn screening. Pediatr Res 54: 219–223
Lehnert W 1994 Long-term results of selective screening for inborn errors of metabolism. Eur J Pediatr 153: S9–S13
Sass JO, Sander S, Zschocke J 2004 Isobutyryl-CoA dehydrogenase deficiency: isobutyrylglycinuria and ACAD8 gene mutations in two infants. J Inherit Metab Dis 27: 741–745
Bross P, Corydon TJ, Andresen BS, Jorgensen MM, Bolund L, Gregersen N 1999 Protein misfolding and degradation in genetic diseases. Hum Mutat 14: 186–198
Pedersen CB, Bross P, Winter VS, Corydon TJ, Bolund L, Bartlett K, Vockley J, Gregersen N 2003 Misfolding, degradation, and aggregation of variant proteins. The molecular pathogenesis of short chain acyl-CoA dehydrogenase (SCAD) deficiency. J Biol Chem 278: 47449–47458
O'Reilly L, Bross P, Corydon TJ, Olpin SE, Hansen J, Kenney JM, McCandless SE, Frazier DM, Winter V, Gregersen N, Engel PC, Andresen BS 2004 The Y42H mutation in medium-chain acyl-CoA dehydrogenase, which is prevalent in babies identified by MS/MS-based newborn screening, is temperature sensitive. Eur J Biochem 271: 4053–4063
Andresen BS, Dobrowolski SF, O'Reilly L, Muenzer J, McCandless SE, Frazier DM, Udvari S, Bross P, Knudsen I, Banas R, Chace DH, Engel P, Naylor EW, Gregersen N 2001 Medium-chain acyl-CoA dehydrogenase (MCAD) mutations identified by MS/MS-based prospective screening of newborns differ from those observed in patients with clinical symptoms: identification and characterization of a new, prevalent mutation that results in mild MCAD deficiency. Am J Hum Genet 68: 1408–1418
Frazier DM, Millington DS, McCandless SE, Koeberl DD, Weavil SD, Chaing SH, Muenzer J The tandem mass spectrometry newborn screening experience in North Carolina: 1997-2005. J Inherit Metab Dis 29: 76–85
Yang BZ, Ding JH, Roe D, Dewese T, Day DW, Roe CR 1998 A novel mutation identified in carnitine palmitoyltransferase II deficiency. Mol Genet Metab 63: 110–115
Battaile KP, Nguyen TV, Vockley J, Kim JJ 2004 Structures of isobutyryl-CoA dehydrogenase and enzyme-product complex: comparison with isovaleryl- and short-chain acyl-CoA dehydrogenases. J Biol Chem 279: 16526–16534
Corydon MJ, Gregersen N, Lehnert W, Ribes A, Rinaldo P, Kmoch S, Christensen E, Kristensen TJ, Andresen BS, Bross P, Winter V, Martinez G, Neve S, Jensen TG, Bolund L, Kolvraa S 1996 Ethylmalonic aciduria is associated with an amino acid variant of short chain acyl-coenzyme A dehydrogenase. Pediatr Res 39: 1059–1066
Corydon MJ, Vockley J, Rinaldo P, Rhead WJ, Kjeldsen M, Winter V, Riggs C, Babovic-Vuksanovic D, Smeitink J, de Jong J, Levy H, Sewell AC, Roe C, Matern D, Dasouki M, Gregersen N 2001 Role of common gene variations in the molecular pathogenesis of short-chain acyl-CoA dehydrogenase deficiency. Pediatr Res 49: 18–23
Nagan N, Kruckeberg KE, Tauscher AL, Bailey KS, Rinaldo P, Matern D 2003 The frequency of short-chain acyl-CoA dehydrogenase gene variants in the US population and correlation with the C(4)-acylcarnitine concentration in newborn blood spots. Mol Genet Metab 78: 239–246
van Maldegem BT, Waterham HR, Duran M, van d V, van Woerden CS, Bobu LL, Wanders RJ, Wijburg FA 2005 The 625G>A SCAD gene variant is common but not associated with increased C4-carnitine in newborn blood spots. J Inherit Metab Dis 28: 557–562
Young SP, Matern D, Gregersen N, Stevens RD, Bali D, Liu HM, Koeberl DD, Millington DS 2003 A comparison of in vitro acylcarnitine profiling methods for the diagnosis of classical and variant short chain acyl-CoA dehydrogenase deficiency. Clin Chim Acta 337: 103–113
Acknowledgements
The authors thank Hans Eiberg for providing DNA from Danish controls.
Author information
Authors and Affiliations
Corresponding author
Additional information
Supported by the March of Dimes Foundation (Grant 1-FY-2003-688), Læge Sofus Carl Emils Friis og Hustru Olga Doris Friis's Legat, Aase og Ejnar Danielsens Fond, Oda og Hans Svenningsens Fond, Fonden af 1870, Vanførefonden, Aarhus University Hospital's Research Initiative, The Danish Medical Research Council, and Institute of Experimental Clinical Research, Aarhus University.
Rights and permissions
About this article
Cite this article
Pedersen, C., Bischoff, C., Christensen, E. et al. Variations in IBD (ACAD8) in Children with Elevated C4-Carnitine Detected by Tandem Mass Spectrometry Newborn Screening. Pediatr Res 60, 315–320 (2006). https://doi.org/10.1203/01.pdr.0000233085.72522.04
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1203/01.pdr.0000233085.72522.04
This article is cited by
-
Phenotype, genotype and long-term prognosis of 40 Chinese patients with isobutyryl-CoA dehydrogenase deficiency and a review of variant spectra in ACAD8
Orphanet Journal of Rare Diseases (2021)
-
DDIEM: drug database for inborn errors of metabolism
Orphanet Journal of Rare Diseases (2020)
-
Metabolic heritability at birth: implications for chronic disease research
Human Genetics (2014)
-
Enzymology of the branched‐chain amino acid oxidation disorders: the valine pathway
Journal of Inherited Metabolic Disease (2012)
-
The ACADS gene variation spectrum in 114 patients with short-chain acyl-CoA dehydrogenase (SCAD) deficiency is dominated by missense variations leading to protein misfolding at the cellular level
Human Genetics (2008)