Abstract
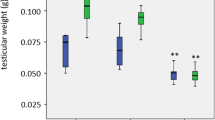
Peroxisomal biogenesis disorders include Zellweger syndrome and milder phenotypes, such as neonatal adrenoleukodystrophy (NALD). Our previous study of a NALD patient with a marked deterioration by a fever revealed a mutation (Ile326Thr) within a SH3 domain of PEX13 protein (Pex13p), showing a temperature-sensitive (TS) phenotype in peroxisomal biogenesis. Clinical TS phenotypes also have been reported in several genetic diseases, but the molecular mechanisms still remain to be clarified. The immunofluorescent staining with anti-Pex13p antibody also revealed TS phenotype of the I326T mutant protein itself in the patient cells. Protease digestion of the recombinant Pex13p-SH3 domain showed an increase of protease susceptibility, suggesting a problem of mutant protein fold. Conformational analyses against urea denaturation using urea gradient gel electrophoresis or fluorescence emission from tryptophan residue revealed that the mutant protein should be easily unfolded. Far-UV circular dichroism (CD) spectra demonstrated that both wild-type and the mutant protein have antiparallel beta-sheets as their secondary structure with slightly different extent. The thermal unfolding profiles measured by CD showed a marked lower melting temperature for I326T protein compared with that of wild-type protein. Analysis of the protein 3D-structure indicated that the Ile326 should be a core residue for folding kinetics and the substitution of Ile326 by threonine should directly alter the kinetic equilibrium, suggesting a marked increase of the unfolded molecules when the patient had a high fever. Structural analyses of the protein in the other genetic diseases could provide an avenue for better understanding of genotype-phenotype correlations.
Similar content being viewed by others
Main
Peroxisomal disorders represent an expanding group of genetic disorders in humans and comprise ~20 different disorders (1,2). Generally, peroxisomal disorders are divided into two groups; PBD (MIM#601539) and single peroxisomal enzyme deficiencies. PBD include ZS (MIM#214100), NALD (MIM#202370), infantile Refsum disease (MIM#266510), and rhizomelic chondrodysplasia punctata type 1 (MIM#215100). Our recent studies have shown that there is genetic heterogeneity among these patients, as concluded from complementation studies that have so far shown 13 different groups (3). The exact functional role of these different PEX gene products (peroxins) is largely unknown, but recent studies have revealed part of the importing mechanisms of peroxisomal proteins in association with peroxisome targeting signal receptors 1 and 2 (4).
A central role in the peroxisomal assembly is played by the PEX13 protein (Pex13p) together with Pex14p and Pex5p (5). Pex13p is a peroxisomal membrane protein containing an SH3, which typically requires the PxxP motif for its binding partners, as observed in Pex14p (6,7). Mutation analysis of PEX13 revealed that two patients belonging to complementation group H with the ZS and NALD phenotypes carried a nonsense and a missense mutation, respectively (8). The patient with the milder form, NALD, had a missense mutation with substitution of isoleucine (Ile) 326 by threonine (Thr), I326T, within a SH3 domain, and showed normal development just before a sudden deterioration with a high fever due to a respiratory infection. Additionally, the patient's fibroblasts showed a TS phenotype in peroxisome assembly and function (8).
TS phenotypes in human have been reported in several diseases, including anemia, thromboembolism, PBD, and trichothiodystrophy (8–11). The genotypes of the patients have been revealed, but the precise molecular mechanisms between genotypes and clinical phenotypes are still to be clarified. We report here a part of structural basis of the TS phenotype caused by the PEX13 mutation.
PATIENTS AND METHODS
Patient.
The index patient is a male of Caucasian parents with a nonconsanguineous marriage. Pregnancy, delivery, and developmental milestones during the first postnatal year were normal. At age 16 mo he became ill with a high fever due to an upper respiratory infection, followed by rapidly progressive hypotonia and muscle weakness leading to severe generalized paresis that made gavage feeding necessary. Loss of hearing and visual decline followed within 6 mo of onset. No external dysmorphia or hepatosplenomegaly was observed.
Skin fibroblasts of the patient showed no catalase-containing particle, and the clinical features and the cytologic appearance of the fibroblasts were compatible to the phenotypes of neonatal adrenoleukodystrophy. Biochemical analyses including fatty acids indicated peroxisomal abnormalities (C24:0/C16:0: 0.15; normal control: 0.58), but dihydroxyacetone phosphate acyltransferase activity was normal in the patient's fibroblasts (patient: 2.26; control: 1.55 nmol/mg/120 min) (8).
The patient was classified as new group H, with the genetic mutation being the homozygote of T997C of the PEX13 gene resulting in a missense mutation, I326T (8). The cell lines obtained from the patient showed a TS phenotype in the peroxisomal biogenesis detected by catalase import after incubation at 30°C instead of 37°C. It was demonstrated in a gene expression experiment using the PEX13-defective CHO mutant that the I326T was indeed the TS mutation (8). This study was approved by the Institutional Review Board of Gifu University. Informed consent was obtained from parents.
Cell lines and immunofluorescence study.
Fibroblasts from patient H02, who showed Zellweger phenotype (W234ter in PEX13), and patient K01, who showed Zellweger phenotype (Q189ter in PEX14), were used as negative controls in immunofluorescent staining. Fibroblasts from patient H01 (I326T in PEX13), patient H02 (W234ter in PEX13), patient K01 (Q189ter in PEX14), and control were cultured at 37°C or 30°C for 7 d (3,8). The cells were fixed with 4% paraformaldehyde/0.1 M potassium phosphate, pH 7.4, permeabilized with 0.1% Triton X-100/PBS, and blocked with 4% FCS/0.1% Triton X-100/PBS. In these fixed fibroblasts, peroxisomes were immunohistochemically stained with rabbit anti-rat-peroxisomal AOX and rabbit anti-human DBP antibodies. For double immunofluorescence staining, rabbit anti-rat Pex14p and mouse anti-human Pex13p (Abcam Ltd., Cambridge, UK) antibodies, and FITC-conjugated goat anti-rabbit (Tago Biosource International, Camarillo, CA) and Cy3-conjugated goat anti-mouse (Chemicon International, Temecula, CA) secondary antibodies were used. For observation we used a confocal laser-scanning microscope (LSMS PASCAL, Carl Zeiss GmbH, Jena, Germany) (12).
Purification of recombinant proteins.
Protein expression and purification were performed as described previously with minor modification (13). Briefly, Escherichia coli BL21 cells were transformed with the vector, pGEX6P-3 coding the GST-fused Homo sapiens (Hs) Pex13p-SH3 domain, which was confirmed as a binding domain for Pex14p (14). The colony with the highest expression level was cultivated overnight in 200 mL of the LB medium with 100 μg/mL ampicillin. The culture was transferred into 2 L of the LB medium with 100 μg/mL ampicillin and incubated at 37°C until OD 600 = 0.45, and then was cooled to 18°C. IPTG (final concentration 0.1 mM) was added to the medium when OD 600 = 0.5. The culture was further incubated at 18°C for 7 h.
The bacterial cell pellet was resuspended in buffer-A (50 mM Tris-HCl, pH 8.0, 400 mM KCl, 10 mM 2-mercaptoethanol, and 1 mM EDTA) with 1 mM PefaBloc (Roche Molecular Biochemicals, Indianapolis, IN), lysed by sonication, and then centrifuged. The clear lysate was applied onto a GST affinity column (Amersham Pharmacia Biotech, Piscataway, NJ) and the column washed with buffer-A. The captured fusion protein was cleaved by PreScission protease (Amersham Pharmacia Biotech) at a ratio of 1% (wt/wt) at 4°C and HsPex13p-SH3(wild-type or I326T) protein was eluted by buffer-A.
Protease susceptibility assay.
Twenty microliters of GST-Pex13p(wild-type) or (I326T) (1 mg/mL) was incubated with 0.1% wt/wt of chymotrypsin at 20, 30, or 40°C for 1 h and analyzed by SDS-PAGE (15–25% gradient).
Conformational stability analyses of HsPex13-SH3.
Conformational stability of HsPex13-SH3(wild-type or I326T) was analyzed by TUG gel electrophoresis and fluorometry (10,15). TUG polyacrylamide gels [linear gradient 0–8 M urea in 7.5% (wt/vol) acrylamide] were prepared as previously described with minor modification using the DCode system (Bio-Rad, Hercules, CA) (10). Tris-glycine buffer (25 mM Tris,192 mM glycine) was used as a running buffer. One hundred micrograms of HsPex13-SH3(wild-type or I326T) was dissolved in sample buffer (62.5 mM Tris-HCl pH6.8, 40% glycerol, 0.01% BPB) and applied evenly across the top of the gel, and electrophoresis was performed at 25 mA constant current for 3 h, keeping the buffer and the gel at 20°C with the DCode temperature control system.
Conformational stability of HsPex13-SH3(wild-type or I326T) was also analyzed as previously described with minor modification using a F-2000 fluorimeter (Hitachi, Yokohama, Japan) (15). Tryptophan residues were selectively excited at 295 nm and the emitted signal was recorded from 309 to 550 nm. Measurements were performed at 20°C in 50 mM phosphate buffer (pH 7.5) containing 100 mM Na2SO4. Urea (1–8 M) was used as the denaturant and the protein concentration in each experiment was 15 μM.
Circular dichroism of HsPex13-SH3 at different temperatures.
All the circular dichroism (CD) measurements were acquired using an Aviv 202 stopped-flow circular dichroism spectrometer (Aviv Associated, Lakewood, NJ) (16,17). Far-UV CD spectra were recorded between 200 and 250 nm, using a 1-nm slit width and a 0.1-cm path length cell thermostated at 5°C. Typically, CD signals were collected for 10 s at each wavelength. Thermal unfolding of HsPex13-SH3 was monitored by measuring the CD signal at 218 nm containing 62 μM of HsPex13-SH3 in 50 mM phosphate buffer (pH 7.5) with 100 mM Na2SO4 in a temperature range from 5 to 85°C with a heating rate of 1°C/min.
Sequence alignment and homology modeling of the structures.
The sequence alignment of the Pex13p-SH3 domains of different species and the HsFyn-SH3 domain were performed by ClustalW with a BLOSUM matrix (18). Homology modeling of the HsPex13-SH3 domain was performed with a Geno3D system using the crystal structure of the Saccharomyces cerevisiae (Sc) Pex13-SH3 domain (PDB code: 1N5Z) as a template (19). Structural verification of the calculated models was performed with PROCHECK (20). Structural rendering was performed with RasMol software (21).
RESULTS
Immunofluorescence studies of patient fibroblasts.
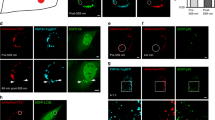
Fibroblasts of the control, patients H01 (I326T), H02 (W234ter), and K01 (Q189ter in PEX14) were incubated for 7 d at 37°C or 30°C and subjected to immunofluorescence staining. The punctuate staining pattern for intact peroxisomes and the intact import of AOX and DBP were observed in the control fibroblasts at 37°C and 30°C (Fig. 1), whereas no immunofluorescence stained particles with these antibodies could be observed in the H02 fibroblasts at either 37°C or 30°C (Fig. 1). In the H01 fibroblasts, only a few particles could be observed at 37°C (Fig. 1, A-a and B-a), whereas numerous particles could be observed at 30°C (Figure 1, A-d and B-d), indicating a TS phenotype.
Protein import into peroxisomes depends on temperature in fibroblasts harboring the mutation I326T in PEX13. Fibroblasts from control, patient H01 (I326T in PEX13), and patient H02 (W234ter in PEX13) were cultured at 37°C or 30°C. Cells were stained with (A) anti-human AOX and (B) anti-human DBP rabbit antibodies, and anti-rabbit IgG antibody with FITC. Import of peroxisomal enzymes is impaired at 37°C in H01 cells, but is restored at 30°C, indicating the temperature-sensitive phenotype of peroxisomal biogenesis with protein import via peroxisomal targeting signal receptors. In contrast, H02 cells show no restoration, even at 30°C.
In a double immunofluorescence staining study using mouse anti-Pex13p and rabbit anti-Pex14p antibodies, expression of Pex13p on peroxisomes was also impaired at 37°C in H01 cells (Fig. 2A-e); but it was restored at 30°C (Fig. 2B-e), indicating the temperature-sensitive nature of the protein with the I326T mutation. In addition, the Pex14p assembly on the peroxisomes was also impaired at 37°C in H01 cells (Fig. 2A-a), so that the assembly of Pex14p on peroxisomes was dependent on Pex13p expression, but not vice versa. Neither Pex13p nor Pex14p expression was detected in the H02 fibroblasts at 37°C or 30°C (Fig. 2). However, an amount of Pex14p equivalent to that of wild-type cells was confirmed in the H01 and H02 cell lysates by immunoblot analysis (authors' unpublished data). The findings suggested that Pex14p incorporation into the peroxisome in H01 cells was not due to its defect, but that the impairment was due to the lack of the HsPex13p binding function (22).
Pex13p expression and Pex14p assembly depend on temperature in fibroblasts harboring the mutation I326T in PEX13. Fibroblasts from control, patient H01 (I326T in PEX13), patient H02 (W234ter in PEX13), and patient K01 (Q189ter in PEX14) were cultured at (A) 37°C or (B) 30°C for 7 d. Cells were stained with mouse anti-human Pex13 and rabbit anti-rat Pex14p antibodies. Expression of Pex13p on peroxisomes is impaired at 37°C in H01 cells, but it is restored at 30°C, indicating the temperature-sensitive nature of protein with the I326T mutation. Pex14p assembly on the peroxisomes is also impaired at 37°C in H01 cells, but is restored at 30°C, suggesting that the loss of assembly of HsPex14p in H01 cells is due to the lack of the HsPex13p adaptor function.
Protease susceptibility assay.
Protease susceptibility of Pex13p was examined with chymotrypsin. HsPex13p-SH3(I326T) incubated with chymotrypsin showed marked susceptibility to protease cleavage at all temperatures compared with HsPex13p-SH3(wild-type) (Fig. 3). The finding suggested that the problem of SH3 domain fold in I326T mutant, which results in an increase of protease susceptibility.
Protease susceptibility of Pex13p(I326T). HsPex13p-SH3(wild-type or I326T) was incubated with 0.1% wt/wt of chymotrypsin at 20, 30, or 40°C for 1 h and analyzed by SDS-PAGE. HsPex13p-SH3(I326T) incubated with chymotrypsin shows marked susceptibility against protease cleavage at all temperatures compared with HsPex13p-SH3(wild-type). These findings indicate that the protein structure of HsPex13p-SH3(I326T) itself also has a temperature-sensitive character. Large arrow, intact Pex13p-SH3; small arrows, cleaved fragments; pre, preincubation; 20,30,40, incubation temperature.
Conformational stability analyses of HsPex13-SH3.
TUG gel analysis of HsPex13p-SH3 protein showed a significant migration between HsPex13-SH3(wild-type) and HsPex13-SH3(I326T), both showing a sigmoidal transition curve (Fig. 4A). Co-electrophoresis of mixed sample of HsPex13-SH3(wild-type) and HsPex13-SH3(I326T) clearly showed the difference of migration between them (Fig. 4B). HsPex13-SH3(I326T) migrated similarly to the HsPex13-SH3(wild-type) at around the 0 M and 8 M urea region, suggesting similar native and denatured conformations. However, the half point of the conformational transition in HsPex13-SH3(I326T) was observed at 2 M, whereas that in HsPex13-SH3(wild-type) was observed at 4 M. These results indicate that the conformational stability of HsPex13-SH3(I326T) is markedly lower than that of HsPex13-SH3(wild-type).
TUG gel analysis of HsPex13p-SH3 protein. (A) Electrophoresis of (left) HsPex13-SH3(wild-type) or (right) HsPex13-SH3(I326T) was performed on TUG polyacrylamide gels [linear gradient 0–8 M urea in 7.5% (wt/vol) acrylamide] using the DCode system (Bio-Rad), keeping the temperature at 20°C. Migration of HsPex13-SH3(I326T) showed a significantly different pattern compared with that of HsPex13-SH3(wild-type), having a sigmoidal curve. (B) Co-electrophoresis of mixed sample of HsPex13-SH3(wild-type) and HsPex13-SH3(I326T). Co-electrophoresis clearly showed the difference of migration between HsPex13-SH3(wild-type) and HsPex13-SH3(I326T). HsPex13-SH3(I326T) migrated similar to the HsPex13-SH3(wild-type) at 0 M or 8 M urea region, suggesting similar native or denatured conformations of them. However, the half point of the conformational transition in HsPex13-SH3(I326T) was observed at around 2 M, whereas that in HsPex13-SH3(wild-type) was observed at around 4 M. These results indicate that the conformational stability of HsPex13-SH3(I326T) is markedly lower than that of HsPex13-SH3(wild-type).
Conformational changes of HsPex13-SH3(wild-type or I326T) were examined using fluorescence from tryptophan (excitation, 295; and emission, above 309 nm). Fluorescence spectra were obtained in the same buffer with or without 8 M urea at 20°C (Fig. 5A). HsPex13-SH3(wild-type or I326T) in the absence of urea showed a different peak emission wavelength (wild-type, 336 nm; I326T, 342 nm) and intensity, whereas those in the presence of 8 M urea showed the same peak emission wavelength with a significant red-shift at 350 nm and the same intensity, which were significantly higher than those of protein without urea. These results indicate that the peak wavelength and intensity increases with the degree of exposure of the tryptophan residue in the SH3 domain (residue number Trp313 of HsPex13) to the solvent.
Conformational changes in denaturation by urea. (A) Conformational changes of HsPex13-SH3(wild-type or I326T) was measured using fluorescence from tryptophan (excitation, 295; and emission, above 309 nm). Measurements were performed at 20°C in 50 mM phosphate buffer (pH 7.5) containing 100 mM Na2SO4 with or without 8 M urea. HsPex13-SH3(wild-type or I326T) in the absence of urea showed a different emission peak and intensity (wild-type, 336 nm; I326T, 342 nm), whereas in the presence of 8 M urea showed the same emission peak at 350 nm and the same intensity significantly higher than those of protein in native state. These results indicate that the peak wavelength and intensity will increase with the degree of exposure of the tryptophan residue in the SH3 domain (residue number Trp313) and the native state protein of HsPex13-SH3(I326T) should have a different tertiary structure around Trp313. (B) Monitoring of the peak wavelength in emissions with a different urea concentration (0–8 M). HsPex13-SH3(wild-type) showed a lesser sigmoidal transition of the peak changing from 336 to 350 nm, whereas HsPex13-SH3(I326T) showed a nonsigmoidal transition of the peak changing from 342 to 350 nm. These findings indicate that HsPex13-SH3(I326T) has a structure with more exposed Trp313 even at native condition, and the population of the mutant protein would be skewed for the unfolded state even at 20°C.
Transitions of the peak wavelength were monitored with a different urea concentration (0–8 M). HsPex13-SH3(wild-type) showed a sigmoidal transition of the peak emission wavelength changing from 336 to 350 nm, whereas HsPex13-SH3(I326T) showed a lesser sigmoidal transition of the peak changing from 342 to 350 nm (Fig. 5B). These findings indicate that HsPex13-SH3(I326T) has a different population of native conformation even in a native condition at 20°C.
Circular dichroism of PEX13-SH3 at different temperatures.
Far-UV CD spectra at 5°C in Figure 6A demonstrates that both HsPex13-SH3(wild-type) and HsPex13-SH3(I326T) have a minima around 218 nm, which corresponds to the typical antiparallel beta-sheets, but with different intensities. Antiparallel beta sheets are characteristic to the SH3 domain structures (17). On the other hand, thermally unfolded CD spectra at 85°C are almost the same for both proteins, suggesting similar unfolded conformations (Fig. 6A). A decrease of intensities at 218 nm upon unfolding may be due to an increase of the intermolecular interaction (intermolecular beta-sheet formation) and/or the change of contribution from the aromatic side-chain absorption. However, to address the differences in thermal stability between the two proteins, we measured their thermal denaturation profiles.
CD spectra of Pex13p-SH3. (A) CD spectra of Pex13p-SH3 in native and denatured states. The values of wild-type and I326T protein at 5°C have a similar minima at around 218 nm, which corresponds to the typical anti-parallel beta-sheets, but with different intensities. On the other hand, thermally unfolded CD spectra at 85°C are almost the same for both proteins, suggesting similar unfolded conformations. The measured temperatures are indicated in parenthesis. (B) Transition of HsPex13p-SH3(I326T) at a lower temperature. Melting temperature (Tm) of HsPex13-SH3 (wild-type) is ~58°C. However, Tm of HsPex13-SH3 (I326T) is significantly low, i.e. around 30°C. The significantly low thermal stability of HsPex13-SH3 (I326T) results in a low population of the native conformation even at 30°C. When the temperature increases over 40°C, most of the population may lose their native folds.
Thermal unfolding profiles for both proteins measured by CD intensities at 218 nm is displayed in Fig. 6B. Melting temperature (Tm) of HsPex13-SH3(wild-type) is ~58°C. However, melting temperature of HsPex13-SH3(I326T) is significantly lower, i.e. around 30°C. The significantly low thermal stability of HsPex13-SH3(I326T) results in a low population of the native conformation even at 30°C. When the temperature increases over 40°C, most of the population may lose their native folds (Fig. 6B). Thermal denaturation processes of HsPex13-SH3(wild-type) and HsPex13-SH3(I326T) were fully reversible between 5 and 85°C (data not shown).
Sequence alignment and homology modeling of the structures.
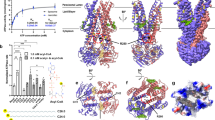
The sequence alignment of the Pex13p-SH3 domains of different species showed striking similarities, especially in mammalian species (>95%) (Fig. 7A). The amino acid identities among yeasts and mammalians were not so high (about 30%), but the substitutions of the amino acids were mostly conservative. Ile at 326 of HsPex13p is well conserved, except for Pichia pastoris Pex13p, but has a conservative amino acid, valine (Fig. 7A). The crystal structure showed that the Ile362 of ScPex13p, which corresponds to Ile326 of HsPex13, is one of the hydrophobic core residues of the beta-barrel structure of the SH3 domain, and that the residue also consists of a recognition site for the PxxP motif of ScPex14p, suggesting that the substitution of the residue affects structure and binding activity (Fig. 7B).
Structural analyses of Pex13p-SH3. (A) Sequence alignment of the Pex13-SH3 domains among different species (Hs: Homo sapiens, Mm: Mus musculus, Rn: Rattus norvegicus, Cg: Cricetulus griseus, Pp: Pichia pastoris, Sc: Saccharomyces cerevisiae), showing striking similarities, especially in mammalian species (>95%). Amino acid identities among yeasts and mammalians are relatively lower (about 30%), but the substitutions are mostly conservative. Isoleucine at 326 of HsPex13p is well conserved, except for PpPex13p but has a conservative amino acid, valine. (B) Crystal structure of ScPex13p. Ile362 of ScPex13p, corresponding to Ile326 of HsPex13, is one of the hydrophobic core residues of the beta-barrel structure of the SH3 domain, and the residue also consists of a recognition site for the PxxP motif of ScPex14p, suggesting that the substitution of the residue affects the structure and the binding activity. The overall structures are shown as a ribbon model and the residues of Ile362 of ScPex13p and the peptide with the PxxP motif of ScPex14p are indicated in spacefill representations—carbon in gray, nitrogen in blue, and oxygen in red.
Alignment of the HsPex13-SH3 and HsFyn-SH3 domains showed high similarity, and that the core “nucleus” hydrophobic residues, Ile28, Ala39, and Ile50, of HsFyn-SH3 determined by protein folding kinetic analyses (15,17) were also conserved in HsPex13-SH3 (Fig. 8A). Homology modeling of the HsPex13-SH3 domain confirmed that these three core nuclei residues were also structurally conserved (Fig. 8B, left and center).
Conservation of hydrophobic core residues in Pex13p-SH3. (A) Alignment of the HsPex13-SH3 and HsFyn-SH3 domains. The HsFyn-SH3 domain is numbered according to Northey et al. (15) The high similarity indicates that conservation of the core “nucleus” hydrophobic residues, Ile28, Ala39, and Ile50, of HsFyn-SH3 is important for protein folding. (B) Structural comparison between HsFyn-SH3 and HsPex13-SH3s. Homology modeling of the HsPex13-SH3 domain confirms the structural conservation of the core “nucleus” hydrophobic residues. However, the mutant SH3 domain with I326T shows a different main chain topology especially around the interface for peptide binding, and the core nucleus residues can be only loosely formed due to the negatively charged side chain of Thr. The overall structures are shown as the main chain backbone, and the critical nucleus residues are indicated as spacefill representations, as in Figure 7.
However, the mutant SH3 domain with I326T has a different main chain topology, especially around the interface for peptide binding. Moreover, the core nucleus residues could be only loosely formed due to the negatively charged side chain of Thr (Fig. 8B, right). These findings suggest that the overall structure of the SH3(I326T) mutant would be fragile and be susceptible to protease digestion, which results in loss of interaction between SH3(I326T) and Pex14p.
DISCUSSION
Pex13p has a central role in the assembly of peroxisomal proteins, and interaction with Pex14p is a critical step for peroxisomal biogenesis (5,8). Pex13p has an SH3 domain in the C-terminal region that is known as an adaptor domain for the PxxP motif peptide, one that Pex14p also has (6,7). Our observations of the patient's fibroblasts revealed the temperature-sensitive nature of biogenesis of peroxisomes and import of peroxisomal enzymes (Fig. 1) (8). In addition, assembly of Pex14p also showed a temperature-sensitive character that depended on Pex13p expression (Fig. 2). The temperature-sensitive nature of the expression of HsPex13p(I326T) on peroxisomes suggests that the produced protein, especially the SH3 domain, is fragile and was degraded at 37°C, but that the protein structure could be maintained preserving its binding activity with Pex14p at 30°C (Fig. 2).
The principal stabilizing element in globular proteins such as the SH3 domain is the hydrophobic core, a subset of hydrophobic residues that is shielded from solvents in the native state structure. The role of hydrophobic core residues in maintaining the thermodynamic stability of proteins has been demonstrated; and protein folding kinetics studies have indicated a key role for hydrophobic core residues not only in stabilizing the native state but also in driving the folding reaction itself (15). Compaction of the hydrophobic core occurs in the early stages of folding and resulted in significant reductions in the folding rate caused by substitutions at a distinct region of the hydrophobic core (23,24).
In the HsFyn-SH3 domain, the stability and folding kinetics of protein have been characterized using >40 mutant proteins (15). The results showed that three core nucleus residues, Ile28, Ala39, and Ile 50 (Fig. 8B, left), were tightly packed even in the transition state of folding, and that substitutions at these positions cause the largest changes in the stability in the native state, and also in the folding rate; for example, the alanine mutant of Ile50 showed significant reduction in Tm (wild-type: 80.1°C; I50A: 44.2°C). Introducing the mutation also altered the folding and the unfolding kinetics at 25°C (folding versus unfolding: wild-type, 30.2/s versus 0.51/s; I50A, 4.2/s versus 13.8/s), indicating that slower folding and faster unfolding resulted in an equilibrium with the larger unfolded population of the proteins even at a temperature lower than its melting temperature (15). Similar characteristics of core nucleus residues have been investigated in other SH3 domains showing similar results (17,23–25).
Ile326 of HsPex13p-SH3 is a hydrophobic core residue of the barrel structure of the domain and also should act as a key core nucleus residue for protein folding with Leu301 and Ala316 (Fig. 8B, center). The substitution of Ile50 by serine of HsFyn-SH3 showed a drastic reduction in peptide binding activity with marked protein instability, suggesting similar effects as by the I326T mutation, introducing a charged side chain of threonine into the hydrophobic core of HsPex13p-SH3 (Fig. 8B, right). Protein stability assays performed here have shown the fragility of the SH3 domain conformation (Figs. 3–5), and that the melting temperature of I326T estimated by CD was markedly lower than 37°C (Tm: wild-type, 58°C; I326T, 30°C). These findings indicate that a large population of the expressed protein is not correctly folded even at a normal body temperature (Fig. 6B). The temperature-sensitive nature of the mutation I326T in HsPex13p should be recognized, not only as simple fragility, such as susceptibility to the proteases of the native protein, but also as an imbalance of folding and unfolding kinetics, which results in an increase of unfolded proteins even at a lower temperature, something similarly observed in other SH3 mutants (15,24,25).
From the findings above, it can be assumed that when a patient with the I326T mutation in Pex13p has a high body temperature, such as 40°C, correctly folded protein would be almost lost, and the loss of Pex13p leads to the loss of assembly of the peroxisomal targeting receptors including Pex14p; then total peroxisomal biogenesis would be abolished, with the dysfunction leading to the deterioration of the patient. Moreover, a large proportion of Pex13p is unfolded even at a normal body temperature, but the residual adaptor activity by correctly folded proteins can somehow maintain peroxisomal functions as shown in the residual activity of the fatty acid metabolism (4,8). In PBD, TS phenotypes in Pex1, Pex2, and Pex6 with milder forms have been reported (26,27) and therapeutic effects have been reported in cells from patients with milder phenotypes, but not those with the severe phenotype, ZS (28). Treatment of PBD fibroblasts with 4-phenylbutyrate showed an induction of peroxisomes having a 2-fold increase in transcription of related genes. If the absolute amount of correctly folded protein increases even with the same ratio of the unfolded protein, it could provide a protective treatment for TS phenotype patients.
Mutation effects of noncatalytic residues in genetic diseases have been implicated simply as protein degradation by structural instability. However, as shown in this study, which has directly revealed the folding state of protein in the TS phenotype mutation (I326T), more precise investigations utilizing structural analyses could reveal the true nature of the mutations; showing a part of pathogenesis of TS phenotype in peroxisomal disease. The structural approaches in the other genetic diseases also can contribute to better understanding of genotype-phenotype correlation and assist the development of new therapeutic approaches including the use of chemical chaperones (29).
Abbreviations
- AOX:
-
peroxisomal acyl-CoA oxidase
- CD:
-
circular dichroism
- DBP:
-
D-bifunctional protein
- GST:
-
glutathione-S-transferase
- NALD:
-
neonatal adrenoleukodystrophy
- PBD:
-
peroxisomal biogenesis disorder
- Pex:
-
Peroxin
- SH3:
-
src-homology 3
- TS:
-
temperature-sensitive
- TUG:
-
transverse urea gradient
- ZS:
-
Zellweger syndrome
References
Moser HW 1999 Genotype-phenotype correlations in disorders of peroxisome biogenesis. Mol Genet Metab 68: 316–327
Fujiki Y 2000 Peroxisome biogenesis and peroxisome biogenesis disorders. FEBS Lett 476: 42–46
Shimozawa N, Tsukamoto T, Nagase T, Takemoto Y, Koyama N, Suzuki Y, Komori M, Osumi T, Jeannette G, Wanders RJ, Kondo N 2004 Identification of a new complementation group of the peroxisome biogenesis disorders and PEX14 as the mutated gene. Hum Mutat 23: 552–558
Suzuki Y, Shimozawa N, Orii T, Tsukamoto T, Osumi T, Fujiki Y, Kondo N 2001 Genetic and molecular bases of peroxisome biogenesis disorders. Genet Med 3: 372–376
Urquhart AJ, Kennedy D, Gould SJ, Crane DI 2000 Interaction of Pex5p, the type 1 peroxisome targeting signal receptor, with the peroxisomal membrane proteins Pex14p and Pex13p. J Biol Chem 275: 4127–4136
Douangamath A, Filipp FV, Klein AT, Barnett P, Zou P, Voorn-Brouwer T, Vega MC, Mayans OM, Sattler M, Distel B, Wilmanns M 2002 Topography for independent binding of α-helical ligands to a peroxisomal SH3 domain. Mol Cell 10: 1007–1017
Kami K, Takeya R, Sumimoto H, Kohda D 2002 Diverse recognition of non-PxxP peptide ligands by the SH3 domains from p67(phox), Grb2 and Pex13p. EMBO J 21: 4268–4276
Shimozawa N, Suzuki Y, Zhang Z, Imamura A, Toyama R, Mukai S, Fujiki Y, Tsukamoto T, Osumi T, Orii T, Wanders RJ, Kondo N 1999 Nonsense and temperature-sensitive mutations in PEX13 are the cause of complementation group H of peroxisome biogenesis disorders. Hum Mol Genet 8: 1077–1083
Zinkham WH, Liljestrand JD, Dixon SM, Hutchison JL 1979 Observations on the rate and mechanism of hemolysis in individuals with Hb Zurich [His E7(63) leads to Arg]: II. Thermal denaturation of hemoglobin as a cause of anemia during fever. Johns Hopkins Med J 144: 109–116
Bruce D, Perry DJ, Borg JY, Carrell RW, Wardell MR 1994 Thromboembolic disease due to thermolabile conformational changes of antithrombin Rouen-VI (187 Asn-Asp). J Clin Invest 94: 2265–2274
Vermeulen W, Rademakers S, Jaspers NG, Appeldoorn E, Raams A, Klein B, Kleijer WJ, Hansen LK, Hoeijmakers JH 2001 A temperature-sensitive disorder in basal transcription and DNA repair in humans. Nat Genet 27: 299–303
Shimozawa N, Tsukamoto T, Suzuki Y, Orii T, Shirayoshi Y, Mori T, Fujiki Y 1992 A human gene responsible for Zellweger syndrome that affects peroxisome assembly. Science 255: 1132–1134
Li A, Kato Z, Ohnishi H, Hashimoto K, Matsukuma E, Omoya K, Yamamoto Y, Kondo N 2003 Optimized gene synthesis and high expression of human interleukin-18. Protein Expr Purif 32: 110–118
Otera H, Setoguchi K, Hamasaki M, Kumashiro T, Shimizu N, Fujiki Y 2002 Peroxisomal targeting signal receptor Pex5p interacts with cargoes and import machinery components in a spatiotemporally differentiated manner: conserved Pex5p WXXXF/Y motifs are critical for matrix protein import. Mol Cell Biol 22: 1639–1655
Northey JG, Di Nardo AA, Davidson AR 2002 Hydrophobic core packing in the SH3 domain folding transition state. Nat Struct Biol 9: 126–130
Kuwata K, Matumoto T, Cheng H, Nagayama K, James TL, Roder H 2003 NMR-detected hydrogen exchange and molecular dynamics simulations provide structural insight into fibril formation of prion protein fragment 106–126. Proc Natl Acad Sci U S A 100: 14790–14795
Mok YK, Elisseeva EL, Davidson AR, Forman-Kay JD 2001 Dramatic stabilization of an SH3 domain by a single substitution: roles of the folded and unfolded states. J Mol Biol 307: 913–928
Rastogi PA 2000 MacVector. Integrated sequence analysis for the Macintosh. Methods Mol Biol 132: 47–69
Combet C, Jambon M, Deleage G, Geourjon C 2002 Geno3D: automatic comparative molecular modelling of protein. Bioinformatics 18: 213–214
Laskowski RA, Rullmannn JA, MacArthur MW, Kaptein R, Thornton JM 1996 AQUA and PROCHECK-NMR: programs for checking the quality of protein structures solved by NMR. J Biomol NMR 8: 477–486
Sayle RA, Milner-White EJ 1995 RASMOL: biomolecular graphics for all. Trends Biochem Sci 20: 374
Fransen M, Vastiau I, Brees C, Brys V, Mannaerts GP, Van Veldhoven PP 2004 Potential role for Pex19p in assembly of PTS-receptor docking complexes. J Biol Chem 279: 12615–12624
Martinez JC, Serrano L 1999 The folding transition state between SH3 domains is conformationally restricted and evolutionarily conserved. Nat Struct Biol 6: 1010–1016
Riddle DS, Grantcharova VP, Santiago JV, Alm E, Ruczinski I, Baker D 1999 Experiment and theory highlight role of native state topology in SH3 folding. Nat Struct Biol 6: 1016–1024
Di Nardo AA, Larson SM, Davidson AR 2003 The relationship between conservation, thermodynamic stability, and function in the SH3 domain hydrophobic core. J Mol Biol 333: 641–655
Imamura A, Shimozawa N, Suzuki Y, Zhang Z, Tsukamoto T, Fujiki Y, Orii T, Osumi T, Wanders RJ, Kondo N 2000 Temperature-sensitive mutation of PEX6 in peroxisome biogenesis disorders in complementation group C (CG-C): comparative study of PEX6 and PEX1. Pediatr Res 48: 541–545
Walter C, Gootjes J, Mooijer PA, Portsteffen H, Klein C, Waterham HR, Barth PG, Epplen JT, Kunau WH, Wanders RJ, Dodt G 2001 Disorders of peroxisome biogenesis due to mutations in PEX1: phenotypes and PEX1 protein levels. Am J Hum Genet 69: 35–48
Wei H, Kemp S, McGuinness MC, Moser AB, Smith KD 2000 Pharmacological induction of peroxisomes in peroxisome biogenesis disorders. Ann Neurol 47: 286–296
Perlmutter DH 2002 Chemical chaperones: a pharmacological strategy for disorders of protein folding and trafficking. Pediatr Res 52: 832–836
Acknowledgements
The authors thank M. Komori for providing anti-Pex13p antibody and N. Usuda for anti-AOX and DBP antibodies. We also thank Y. Yatsuda, A. Seki, S. Shimizu, Y. Kondo, M. Kojima, and K. Kasahara for their assistance.
Author information
Authors and Affiliations
Corresponding author
Additional information
Supported, in part, by a grant-in-aid for Scientific Research (13670791) from the Japan Society for the Promotion of Science, by a grant for Child Health and Development (14C-3) from the Ministry of Health, Labour and Welfare, by the Uehara Memorial Foundation, and by the Naito Foundation.
Rights and permissions
About this article
Cite this article
Hashimoto, K., Kato, Z., Nagase, T. et al. Molecular Mechanism of a Temperature-Sensitive Phenotype in Peroxisomal Biogenesis Disorder. Pediatr Res 58, 263–269 (2005). https://doi.org/10.1203/01.PDR.0000169984.89199.69
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1203/01.PDR.0000169984.89199.69