Abstract
The molecular background of an intermediate type of dystrophinopathy [Duchenne and Becker muscular dystrophy (DMD/BMD)] remains to be clarified, and out-of -frame and in-frame mutations of the dystrophin gene are shown to be causes of DMD and BMD, respectively. In a boy with this disease, dystrophin mRNA extracted from lymphocytes and muscle were analyzed both qualitatively and quantitatively using reverse transcription PCR. Three different dystrophin mRNA were found to be produced via the use of three cryptic splice acceptor sites resulting from a novel point mutation of 2831-2A>G at the conserved splice acceptor site of intron 20. One of three mRNA showed an insertion of six nucleotides of intron 20 between exons 20 and 21 (dys+6) that encoded two novel amino acids in the rod domain of dystrophin. Two other mRNA species showed an insertion of seven nucleotides of intron 20 between exons 20 and 21 (dys+7) or a seven-nucleotide deletion in exon 21 (dys−7). Quantitative analysis of each dystrophin mRNA expressed in the boy's skeletal muscle disclosed that around 95% and 5% of dystrophin mRNAs were dys−7 and dys+6, respectively, whereas these two mRNA were almost equally expressed in lymphocytes. It is suggested that production of a small fraction of in-frame mRNA in muscle explains the molecular background of the intermediate type of dystrophinopathy in the index case. This finding underlines the potential of genetic therapeutic strategies aimed to modify mRNA in DMD to generate a much milder disease.
Similar content being viewed by others
Main
X-linked dystrophinopathy resulting from mutations in the dystrophin gene is the most common cause of inherited myopathy affecting approximately one in 3500 males. It shows varying degrees of severity, ranging from the severe DMD to the mild BMD (1). DMD is a rapidly progressive disease occurring during childhood in which an affected individual loses the ability to walk before the age of 12 y. BMD has a slower rate of progression; those affected remain ambulatory beyond the age of 16 y and may even lead near-normal lives. In BMD, the mean age of symptoms, such as muscle weakness and difficulty in walking, is reported to be 11 y (2). Patients who lose the ability to walk between age 12 and 16 y are classified as having an intermediate form of the dystrophinopathy, DMD/BMD (3).
In approximately 60% of DMD/BMD cases, a partial deletion or duplication of the dystrophin gene can be detected (4). The severity of the disease depends on the effect of the mutation on the translational reading frame of the dystrophin mRNA. According to the reading frame rule, deletions or duplications that create a shift in the reading frame lead to the more severe DMD phenotype, whereas the milder BMD phenotype occurs if the open reading frame is maintained after the deletion or duplication (5). More than 90% of deletion or duplication mutations identified in DMD/BMD are consistent with the reading frame rule (6). Exceptions to the rule have been reported, however, and the production of an alternative splicing product maintaining the reading frame is considered a key to modifying the clinical phenotype to the milder BMD (7–9). Recently, the analysis of dystrophin mRNA expressed in lymphocytes has become commonly used in identifying small mutations of the dystrophin gene (10–12) and led to the identification of not only small mutations but also aberrant dystrophin mRNA. As a result, a nearly 100% correlation has been obtained between the clinical phenotype and the reading frame rule (13). However, the reading frame rule does not explain the molecular background of the intermediate form DMD/BMD.
Splicing is a process to remove introns from pre-mRNA and splice sites are determined by so-called splicing consensus sequences that are located at intron/exon boundaries. Shapiro's splicing probability scores (14) describe the degree to which individual splice junctions match with the consensus sequence. Mutations at splicing consensus sequences result in exon skipping, activation of cryptic splice sites, or both, producing aberrant mRNA (15). Because the dystrophin gene consists of 79 exons spread over 3000 kb and encodes only a 14-kb long mRNA (4, 16, 17), the splicing of an extremely large intron (up to 200 kb) seems to be regulated in a more sophisticated way than in other genes. Mutations at the splicing consensus sequence of the dystrophin gene have been reported to result in either exon skipping or the activation of cryptic splice sites (18–20). However, examples of the activation of multiple cryptic splice sites have yet to be reported for the dystrophin gene, although many examples have been found in other genes (21, 22). The absence of reports of multiple splice activation is considered to reflect the highly regulated process for splicing of dystrophin pre-mRNA.
In this article, three dystrophin mRNA were found to be produced by the activation of three cryptic splice sites through inactivation of the authentic splice acceptor site. Furthermore, quantification of each dystrophin mRNA showed that a small fraction of in-frame dystrophin mRNA was present in intermediate DMD/BMD.
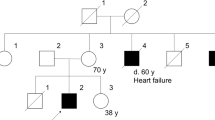
A 14-y-old boy was referred to Kobe University Hospital for a genetic diagnosis of his muscle weakness. As an infant, he started to crawl at age 9 mo but did not begin to walk until the age of 2 y. A slowness to climb stairs was noticed at age 6 y. At age 9 y, he showed a waddling gait and was clinically diagnosed with DMD based on a highly elevated serum creatine kinase (CK) level. His muscle weakness progressed but he could still walk independently and climb stairs using a stair-holder at age 12 y. He had a bone fracture in his right ankle and was wheelchair-bound at the age of 13 y, leading to a diagnosis of intermediate DMD/BMD (3). There was no family history of neuromuscular disease. Laboratory examination disclosed an elevated level of serum CK (8642 IU/L; control, <169 IU/L). He is mentally retarded and his IQ is 59. An electromyogram showed a typical myogenic pattern, but no abnormalities were detected by either x-ray or ECG examinations. To diagnose the myopathy, a muscle biopsy was performed on his biceps. Consent for this study was obtained from his parents. The ethical committee of Kobe University Graduate School of Medicine approved the study.
METHODS
Immunohistochemical analysis
The muscle biopsy sample was examined histologically. An indirect immunofluorescence analysis was performed using three dystrophin antibodies recognizing the N-terminal (NCL-Dys3), the rod (NCL-Dys1), and the C-terminal (NCL-Dys2) domains of dystrophin (Novocastra Laboratories, Newcastle, U.K.) as previously described (23). Normal skeletal muscle tissue was simultaneously stained with the panel of antibodies.
Mutation analysis.
For mutation analysis of the dystrophin gene, DNA was isolated from peripheral blood of the index case and members of his family by standard phenol-chloroform extraction methods. Southern blot analysis was performed using Hin dIII restriction enzyme-digested DNA as a template and dystrophin cDNA fragments as probes (4). This technique allows for the full extent of any deletions or duplications to be recognized. To characterize the genomic mutations, a region encompassing 59, 181, and 79 bp of intron 20, exon 21, and intron 21, respectively, was amplified by PCR from the genomic DNA samples using the following pair of primers: a forward primer on intron 20: 5′-GCAAAATGTAATGTATGCAAAG-3′ and a reverse primer complementary to intron 21: 5′-ATGTTAGTACCTTCTGGATTTCC-3′. The amplified product was directly sequenced (8).
Reverse transcription PCR (RT-PCR) was used to analyze the dystrophin mRNA expressed in lymphocytes or skeletal muscle as described before (11). Full-length dystrophin cDNA was amplified as 10 separate, partially overlapping fragments and directly sequenced (10). To characterize a fragment showing ambiguous sequencing results, a region encompassing exons 19 to 22 was amplified using a forward primer corresponding to a segment of exon 19 (5′-GCCATAGAGCGAGAAAAAGCT-3′) and a reverse primer complementary to a segment of exon 22 (5′- CTGATAGCGCATTGGTGGCAA -3′). The amplified product was purified and subjected to sequencing either directly or after subcloning into T7 blue T vector (24).
Quantification of mRNA.
To quantify each of the three dystrophin mRNA, RT-PCR products labeled with Cy5 were electrophoresed on sequencing gel and its fluorescence was measured. A region encompassing exons 20 to 21 was amplified using a Cy5-labeled forward primer on exon 20 (5′-ATCGCTTTCTATAATCAGCTAC-3′) (Hokkaido System Science Co. Ltd., Sapporo, Japan) and a reverse primer on exon 21 (5′-TTGTCTGTAGCTCTTTCTCTC-3′). The cDNA was amplified by 25 cycles of PCR. The amplified product was separated electrophoretically using a DNA sequencer (Amersham Pharmacia Biotech, Uppsala, Sweden) and each dystrophin mRNA was determined based on the distance it migrated on the gel. Quantification of the respective mRNA was carried out by measuring its peak height.
RESULTS
The biopsied muscle sample showed dystrophic changes on microscopic examination, i.e. size variation in muscle fibers, central nuclear fibers, and degenerative and regenerative fibers. Dystrophin staining showed that the material reactive to the rod-recognizing antibodies was weakly present in the muscle membrane whereas there was no material reactive to the N-terminal or COOH-terminal recognizing antibodies (Fig. 1). Western blot analysis of dystrophin disclosed no significantly visible band (data not shown). A mutation responsible for the intermediate form was investigated in the dystrophin gene.
Immunofluorescent visualization of dystrophin in the patient's biopsied skeletal muscle. Dystrophin staining of biopsied skeletal muscle disclosed that both the N-terminal (upper left panel) and C-terminal (upper right panel) regions were negative, whereas the rod domain was weakly positive (upper middle panel). In normal control, dystrophin is clearly localized to the subsarcoplasmic region (lower panel).
Southern blot analysis failed to disclose any gross gene rearrangement in his genome. Dystrophin mRNA was analyzed as 10 separate fragments by PCR amplification. Direct sequencing of these fragments revealed a completely normal sequence, except one encompassing exons 17 to 25. The ambiguous region was then amplified as a small fragment extending from exons 19 to 22. The amplified product appeared as one band on separation by electrophoresis (Fig. 2a). Direct sequencing showed the ambiguous result again and the amplified products were next sequenced after subcloning. Remarkably, three mRNA with different mutation sequences were found to be present: two had insertions of six (GCACGG) (dys+6) or seven (GGCACGG) (dys+7) nucleotides between exons 20 and 21 (Fig. 2b). Another one had a seven-nucleotide (CATGAAG) deletion of the 5′ end of exon 21 (dys−7) (Fig. 2b). These three mRNA were considered natural products because the exon boundaries were conserved and no other nucleotide change was present in the sequenced exons. However, the mechanism to produce these three mRNA was unclear.
Mutation analysis. (a) The fragment encompassing exons 19 to 22 was amplified from the patient's cDNA. One clear band was visualized from the index case (p) and was the same size as the control corresponding to 543 bp (c) (lower panel). M refers to a DNA size marker of Hae III-digested φX174 DNA. The amplified region is shown schematically (upper panel). Boxes and the numbers in boxes indicate the exon and exon numbers, respectively. Horizontal arrows under the box indicate location and direction of primers. (b) The sequences of three different clones are shown. Each sequence showed completely normal exons 19 and 20. The sequence of the 3′ end of exon 20 (5′- TGTAAG-3′) is joined to the three different sequences TCAACC (top panel), GCACGG (middle panel), and GGCACG (bottom panel). In the middle and bottom panels, 7- and 6-bp insertions were followed by a completely normal exon 21. In the top panel, seven nucleotides of the 5′ end of exon 21 disappeared. (c) Genomic sequence at the border between intron 20 and exon 21 is shown. The PCR-amplified product from genomic DNA was directly sequenced. The second upstream nucleotide from the 3′ end of intron 20 was G in the index case (G2831−2) (upper panel) whereas it was A in the control (A2831−2) (lower panel). The mutation changed the highly conserved AG dinucleotide to GG. The number 2831 corresponds to the dystrophin cDNA number of the first nucleotide of exon 21. (d) Splicing patterns of intron 20 and candidate splice acceptor sites near the border between intron 20 and exon 21 are shown. Splicing patterns identified in the index case are represented schematically. Three diagonal lines indicate splicing patterns that were activated in the index case. dys+7, dys+6, and dys−7 indicate the activated splice acceptor sites. Boxes and horizontal line indicate the exon and intron 20, respectively. Figure not drawn to scale.
Two inserted sequences shared a common six-nucleotide sequence (GCACGG), but dys+7 had an additional G nucleotide at the 5′ end of the common sequence. A homology search of the common sequences disclosed that five of the six nucleotides were the same as the wild-type sequence of the 3′ end of intron 20 except for one nucleotide. The different nucleotide was an A to G transition at the second-to-last nucleotide of the wild-type intron 20. The possibility that the inserted sequence was from intron 20 was strongly suggested. To examine this possibility, the genomic nucleotides encompassing exon 21 and its flanking introns were sequenced. The 319 nucleotides were completely homologous to the wild-type sequence except for one nucleotide. The same A to G transition at the second-to-last nucleotide of intron 20 (2831st nucleotide of dystrophin cDNA) was identified in the genome (2831–2A>G) (Fig. 2c). Therefore, inserts of six and seven nucleotides, respectively, between exons 20 and 21 of the dystrophin mRNA were concluded to originate from the sequence of the 3′ end of intron 20.
The nucleotide change, 2831–2A>G, disrupted a highly conserved AG dinucleotide sequence at the splice acceptor site. It was reasonable to suspect that a splicing error occurred. However, it was difficult to understand the production of the three dystrophin mRNA with either an insertion or a deletion, especially the two mRNA differing in one additional nucleotide. Remarkably, all three mRNA were found to be produced from the usage of the cryptic splice acceptor sites located nearby the mutated authentic splice acceptor site (Fig. 2d). These sites maintain the high Shapiro's splicing probability score (14) for splice acceptor sites: 0.69, 0.65, and 0.56 at 7 and 6nt upstream and 7nt downstream from the authentic splice site, respectively, whereas the authentic splice acceptor site showed a decrease in its score from 0.83 to 0.62 with the mutation. It was concluded that the three dystrophin mRNA were splicing products obtained via the use of cryptic splice sites in either intron 20 or exon 21.
The ability to produce dystrophin was examined in each dystrophin mRNA. Both dys+7 and dys−7 creating a premature stop codon in exon 20 were identified as nonfunctional. However, dys+6 maintained the translational reading frame and was expected to produce dystrophin. The inserted six nucleotides GCACGG encode alanine and arginine. These two amino acids were inserted between the 874th and 875th amino acid residue located at the 5th repeat of the rod domain of dystrophin. This elongated dystrophin seemed to exert its function because an insertion into the rod domain is not very harmful to dystrophin function (25, 26).
In the index case, it was difficult to decide on a clinical picture on the basis of the reading frame rule because both out-of-frame and in-frame dystrophin mRNA were identified. From the high probability score of the splice acceptor site, it was expected that dys+7 was the most abundant mRNA, thereby resulting in DMD. Clinically, however, the index case was diagnosed as intermediate. To understand its molecular pathogenesis, each of the dystrophin mRNA expressed in lymphocytes was quantitated by RT-PCR using a fluorescent labeled forward primer. From the amplified product, two peaks were visualized at sizes of 300 and 313 bp, corresponding to dys−7 and dys+6, respectively (Fig. 3). Contrary to expectation, a peak at 314 bp corresponding to dys+7 could not be detected. The two peaks were almost equal in height, indicating the parallel use of two acceptor sites. The presence of the in-frame mRNA in half of the total dystrophin mRNA was expected to result in BMD (8). The clinical severity of the index case was not compatible with this finding.
Quantification of each dystrophin mRNA. Dystrophin cDNA encompassing exons 20 to 21 was amplified using a Cy5-labeled forward primer. The amplified product was separated on the sequence gel using an automatic DNA sequencer. Two peaks at 300 bp and was 313 bp long points were visualized from both lymphocytes (lower panel) and skeletal muscle (upper panel). From their sizes, the two products were identified to be dys−7 and dys+6, respectively. A peak between two peaks in the lower panel was considered either nonspecific or a heteroduplex product. However, a peak corresponding to dys+7 could not be identified at 314 bp. The vertical axis indicates fluorescence intensity and the horizontal axis indicates the size of the amplified product as determined using internal marker DNA.
Considering that the mRNA of the dystrophin in muscle reflects the clinical phenotype more accurately than that in lymphocytes, muscle dystrophin mRNA were quantitated. Remarkably, quite different patterns of expression were obtained: one large and one small peak (Fig. 3). From the peak height, it was decided that around 95% of the mRNA was dys−7 and around 5% dys+6. The molecular pathogenesis of intermediate DMD/BMD was considered due to the production of a small fraction of the in-frame dystrophin mRNA.
Here, we describe a case of dystrophinopathy showing intermediate DMD/BMD. Because of a point mutation at the splicing consensus acceptor site, three different dystrophin mRNA were produced. A molecular background of intermediate DMD/BMD was identified because in-frame dystrophin mRNA was produced in a small fraction of muscle mRNA.
DISCUSSION
In this article, we described the molecular analysis of a case of intermediate DMD/BMD. The patient had a novel 2831–2A>G mutation in intron 20 of the dystrophin gene thus disrupting the highly conserved AG dinucleotide at the splice acceptor site (Fig. 2c). Although the mutation was expected to induce skipping of exon 21 by inactivating the splice acceptor site, no skipping was detected by dystrophin mRNA analysis (Fig. 2a). Instead, three aberrant mRNA were identified (Fig. 2b). Among the three, only one was an in-frame mRNA having the ability to produce dystrophin with a two–amino acid insertion. However, this in-frame mRNA (dys+6) accounted for only a small fraction (around 5%) of muscle dystrophin mRNA (Fig. 3) and seemed to be responsible for the clinical phenotype of intermediate DMD/BMD.
The molecular background of intermediate DMD/BMD is not well understood, although the reading frame rule has been documented in DMD and BMD (5, 13). In a case of intermediate DMD/BMD with a nonsense mutation in exon 2 of the dystrophin gene, it was supposed that the in-frame alternative splicing product acted as a modifier of the clinical phenotype of DMD, however, attempts to identify the supposed mRNA in that case failed (27). Our results show the first evidence of production of a case of in-frame mRNA in an intermediate DMD/BMD (Fig. 2b). In the future, extensive analyses of dystrophin mRNA may further clarify the molecular background of intermediate DMD/BMD.
As far as we know, this is the first report to describe the production of three dystrophin mRNA by the usage of different cryptic splice sites. In the literature, four dystrophin mRNA at most are shown to be produced by mechanisms of exon skipping or alternative splicing in mutations at the splice consensus sequence of the dystrophin gene (8, 28, 29), but activation of a cryptic splice site has been limited to one or two sites (30). We do not know why three cryptic splice sites among six sites located near the border between intron 20 and exon 20 were selected. It is clear that Shapiro's probability score is not the sole factor deciding the availability of the cryptic splice acceptor site inasmuch as dys+7 produced from the site with a high Shapiro's score was not quantifiably detected. It is especially difficult to explain why the authentic splice acceptor site containing the mutation was not used, even though it is located at the original position and has a higher Shapiro's splicing probability score (0.62) than the downstream activated site (0.56). Furthermore, the score for the three activated sites is far below the average for all splicing acceptor sites of the dystrophin gene (0.87) (30). These results indicate that unknown factors other than position and the score influence the usage of the splice acceptor site. It seems impossible to say which cryptic acceptor site is used in mutations of the splicing consensus sequence. Therefore, usage patterns of the sites can only be clarified by analyzing mRNA.
The next question is what factor determines the amount of each dystrophin mRNA. The ratio of dys+6 to dys−7 was different between lymphocytes and muscle (Fig. 3). This may be due to either a difference in the use of each cryptic splice site that is regulated by tissue-specific factor(s) or the fate of dystrophin mRNA according to its protein-coding ability (31). More studies are needed to answer this question. Because the amounts of multiple mRNA are different between lymphocyte and muscle (Fig. 3), it is necessary to analyze amounts of each mRNA expressed in muscle for proper analysis of genotype and phenotype correlation in dystrophinopathy.
The large size of both the dystrophin gene (3000 kb) and dystrophin mRNA (14 kb) hampers the identification of mutations of the dystrophin gene. We have used RT-PCR analysis of dystrophin mRNA in lymphocytes for analysis of small mutations and identified several novel mutations (8, 18, 32). In the present case, the ambiguous result of the direct sequencing of a rather large fragment encompassing exons 17 to 25 initiated an extensive study revealing a single nucleotide change in the dystrophin gene. The smaller region encompassing exons 19 to 22 was amplified as a single clear band (Fig. 2a) but was found to be a mixture of three products by sequencing the amplified products using the subcloning sequence method. It is suggested that a subcloning examination of an RT-PCR product be done for a thorough search considering the possibility of a mixture of products differing in only a few nucleotides.
At this moment, there is no effective way to treat DMD. Recently, the conversion of DMD to BMD by changing dystrophin mRNA from out-of-frame to in-frame has attracted much attention as an alternative treatment of DMD (33). The natural modification of the clinical phenotype from DMD to BMD by a mechanism of exon skipping has been reported in BMD cases with nonsense mutations in exons 27, 29, or 72 (8, 9, 34). Therefore, artificial induction of exon skipping is one potential method for alternative gene therapy for DMD. We have shown that skipping of exon 19 of the dystrophin gene can be induced by blocking the function of the splicing enhancer sequence located in exon 19 with an antisense oligonucleotide complementary to the splicing enhancer sequence (35). Skipping of exon 19 in myocytes from a DMD patient with an exon 20 deletion led to the production of an in-frame dystrophin mRNA lacking both the exon 19 and 20 sequence, resulting in dystrophin-positive cells (36). In addition, it was shown that the skipping of exon 46 induced using antisense oligonucleotide complementary to a purine-rich sequence led to the production of dystrophin in DMD-derived muscle cells (37). At the same time, the induction of exon skipping with antisense oligonucleotides complementary to splicing consensus sequences was shown to lead to the production of an internally truncated dystrophin in the mdx mouse, an animal model of DMD (38). These results indicate the potential of these antisense oligonucleotides for treating DMD. In the index case, blocking the activated cryptic splice site within exon 21 with an antisense oligonucleotide may result in a more active use of the upper cryptic splice site that produces in-frame mRNA. In the future, this strategy will be examined for the index case.
We, however, do not know what percentage of dystrophin mRNA should be changed to the in-frame type to obtain a therapeutic effect in alternative gene therapy. It is supposed that the higher the amount of in-frame mRNA present, the milder the muscle weakness. In our previous report, a mild BMD phenotype developed when in-frame dystrophin accounted for nearly 10% of total dystrophin mRNA (8). In this report, a level of around 5% in muscle was not so effective in ameliorating the clinical phenotype. Therefore, at least 10% of dystrophin mRNA should be modified to in-frame mRNA when using alternative gene therapy for DMD.
Abbreviations
- DMD:
-
Duchenne muscular dystrophy
- BMD:
-
Becker muscular dystrophy
References
Emery AEH 1993 Duchenne muscular dystrophy. Oxford University Press, Oxford, U.K.
Bushby KM, Gardner-Medwin D 1993 The clinical, genetic and dystrophin characteristics of Becker muscular dystrophy. I. Natural history. J Neurol 240: 98–104
Brooke MH, Fenichel GM, Griggs RC, Mendell JR, Moxley R, Miller JP, Province MA 1983 Clinical investigation in Duchenne dystrophy: 2. Determination of the “power” of therapeutic trials based on the natural history. Muscle Nerve 6: 91–103
Koenig M, Hoffman EP, Bertelson CJ, Monaco AP, Feener C, Kunkel LM 1987 Complete cloning of the Duchenne muscular dystrophy (DMD) cDNA and preliminary genomic organization of the DMD gene in normal and affected individuals. Cell 50: 509–517
Monaco AP, Bertelson CJ, Liechti-Gallati S, Moser H, Kunkel LM 1988 An explanation for the phenotypic differences between patients bearing partial deletions of the DMD locus. Genomics 2: 90–95
Koenig M, Beggs AH, Moyer M, Scherpf S, Heindrich K, Bettecken T, Meng G, Muller CR, Lindlof M, Kaariainen H, de la Chapelle A, Kiuru A, Savontaus M-L, Gilgenkrantz H, Recan D, Chelly J, Kaplan J-C, Covone AE, Archidiacono N, Romeo G, Liechti-Gallati S, Schneider V, Braga S, Moser H, Darras BT, Murphy P, Francke U, Chen JD, Morgan G, Denton M, Greenberg CR, Wrogemann K, Blonden LAJ, van Paassen HMB, van Ommen GJB, Kunkel LM 1989 The molecular basis for Duchenne versus Becker muscular dystrophy: correlation of severity with type of deletion. Am J Hum Genet 45: 498–506
Chelly J, Gilgenkrantz H, Lambert M, Hamard G, Chafey P, Recan D, Katz P, de la Chapelle A, Koenig M, Ginjaar IB, Fardeau M, Tome F, Kahn A, Kaplan J-C 1990 Effect of dystrophin gene deletions on mRNA levels and processing in Duchenne and Becker muscular dystrophies. Cell 63: 1239–1248
Shiga N, Takeshima Y, Sakamoto H, Inoue K, Yokota Y, Yokoyama M, Matsuo M 1997 Disruption of the splicing enhancer sequence within exon 27 of the dystrophin gene by a nonsense mutation induces partial skipping of the exon and is responsible for Becker muscular dystrophy. J Clin Invest 100: 2204–2210
Ginjaar IB, Kneppers AL, v d Meulen JD, Anderson LV, Bremmer-Bout M, van Deutekom JC, Weegenaar J, den Dunnen JT, Bakker E 2000 Dystrophin nonsense mutation induces different levels of exon 29 skipping and leads to variable phenotypes within one BMD family. Eur J Hum Genet 8: 793–796
Roberts RG, Barby TF, Manners E, Bobrow M, Bentley DR 1991 Direct detection of dystrophin gene rearrangements by analysis of dystrophin mRNA in peripheral blood lymphocytes. Am J Hum Genet 49: 298–310
Matsuo M, Masumura T, Nishio H, Nakajima T, Kitoh Y, Takumi T, Koga J, Nakamura H 1991 Exon skipping during splicing of dystrophin mRNA precursor due to an intraexon deletion in the dystrophin gene of Duchenne muscular dystrophy Kobe. J Clin Invest 87: 2127–2131
Tuffery S, Chambert S, Bareil C, Sarda P, Coubes C, Echenne B, Demaille J, Claustres M 1998 Mutation analysis of the dystrophin gene in Southern French DMD or BMD families: from Southern blot to protein truncation test. Hum Genet 102: 334–342
Fajkusova L, Lukas Z, Tvrdikova M, Kuhrova VV, Hajek J, Fajkus J 2001 Novel dystrophin mutations revealed by analysis of dystrophin mRNA: alternative splicing suppresses the phenotypic effect of a nonsense mutation. Neuromuscul Disord 11: 133–138
Shapiro MB, Senapathy P 1987 RNA splice junctions of different classes of eukaryotes: sequence statistics and functional implications in gene expression. Nucleic Acids Res 15: 7155–7174
Nakai K, Sakamoto H 1994 Construction of a novel database containing aberrant splicing mutations of mammalian genes. Gene 141: 171–177
Ahn AH, Kunkel LM 1993 The structural and functional diversity of dystrophin. Nat Genet 3: 283–291
Nishio H, Takeshima Y, Narita N, Yanagawa H, Suzuki Y, Ishikawa Y, Minami R, Nakamura H, Matsuo M 1994 Identification of a novel first exon in the human dystrophin gene and of a new promoter located more than 500 kb upstream of the nearest known promoter. J Clin Invest 94: 1037–1042
Hagiwara Y, Nishio H, Kitoh Y, Takeshima Y, Narita N, Wada H, Yokoyama M, Nakamura H, Matsuo M 1994 A novel point mutation (G −1 to T) in a 5′ splice donor site of intron 13 of the dystrophin gene results in exon skipping and is responsible for Becker muscular dystrophy. Am J Hum Genet 54: 53–61
Ikezawa M, Minami N, Takahashi M, Goto Y, Miike T, Nonaka I 1998 Dystrophin gene analysis on 130 patients with Duchenne muscular dystrophy with a special reference to muscle mRNA analysis. Brain Dev 20: 165–168
Tuffery-Giraud S, Chambert S, Demaille J, Claustres M 1999 Point mutations in the dystrophin gene: evidence for frequent use of cryptic splice sites as a result of splicing defects. Hum Mutat 14: 359–368
Schwarze U, Starman BJ, Byers PH 1999 Redefinition of exon 7 in the COL1A1 gene of type I collagen by an intron 8 splice-donor-site mutation in a form of osteogenesis imperfecta: influence of intron splice order on outcome of splice-site mutation. Am J Hum Genet 65: 336–344
Attanasio C, de Moerloose P, Antonarakis SE, Morris MA, Neerman-Arbez M 2001 Activation of multiple cryptic donor splice sites by the common congenital afibrinogenemia mutation, FGA IVS4 + 1 G–>T. Blood 97: 1879–1881
Arahata K, Ishiura S, Ishiguro T, Tsukahara T, Suhara Y, Eguchi C, Ishihara T, Nonaka I, Ozawa E, Sugita H 1988 Immunostaining of skeletal and cardiac muscle surface membrane with antibody against Duchenne muscular dystrophy peptide. Nature 333: 861–863
Surono A, Takeshima Y, Wibawa T, Ikezawa M, Nonaka I, Matsuo M 1999 Circular dystrophin RNAs consisting of exons that were skipped by alternative splicing. Hum Mol Genet 8: 493–500
Angelini C, Beggs AH, Hoffman EP, Fanin M, Kunkel LM 1990 Enormous dystrophin in a patient with Becker muscular dystrophy. Neurology 40: 808–812
Toscano A, Vitiello L, Comi GP, Galvagni F, Miorin M, Prelle A, Fortunato F, Bardoni A, Mora M, Fiumara A, Falsaperla R, Tomelleri G, Tonin P, Danieli GA, Vita G 1995 Duplication of dystrophin gene and dissimilar clinical phenotype in the same family. Neuromuscul Disord 5: 475–481
Prior TW, Bartolo C, Papp AC, Snyder PJ, Sedra MS, Burghes AH, Mendell JR 1996 Nonsense mutations in a Becker muscular dystrophy and an intermediate patient. Hum Mutat 7: 72–75
Lenk U, Hanke R, Thiele H, Speer A 1993 Point mutations at the carboxy terminus of the human dystrophin gene: implications for an association with mental retardation in DMD patients. Hum Mol Genet 2: 1877–1881
Roberts RG, Gardner RJ, Bobrow M 1994 Searching for the 1 in 2,400,000: a review of dystrophin gene point mutations. Hum Mutat 4: 1–11
Sironi M, Pozzoli U, Cagliani R, Comi GP, Bardoni A, Bresolin N 2001 Analysis of splicing parameters in the dystrophin gene: relevance for physiological and pathogenetic splicing mechanisms. Hum Genet 109: 73–84
Hentze MW, Kulozik AE 1999 A perfect message: RNA surveillance and nonsense-mediated decay. Cell 96: 307–310
Wibawa T, Takeshima Y, Mitsuyoshi I, Wada H, Surono A, Nakamura H, Matsuo M 2000 Complete skipping of exon 66 due to novel mutations of the dystrophin gene was identified in two Japanese families of Duchenne muscular dystrophy with severe mental retardation. Brain Dev 22: 107–112
Matsuo M 2002 Duchenne and Becker muscular dystrophy: from gene diagnosis to molecular therapy. IUBMB Life 53: 1–6
Melis MA, Muntoni F, Cau M, Loi D, Puddu A, Boccone L, Mateddu A, Cianchetti C, Cao A 1998 Novel nonsense mutation (C–>A nt 10512) in exon 72 of dystrophin gene leading to exon skipping in a patient with a mild dystrophinopathy. Hum Mutat ( suppl 1): S137–S138
Pramono ZA, Takeshima Y, Alimsardjono H, Ishii A, Takeda S, Matsuo M 1996 Induction of exon skipping of the dystrophin transcript in lymphoblast-oid cells by transfecting an antisense oligodeoxynucleotide complementary to an exon recognition sequence. Biochem Biophys Res Commun 226: 445–449
Takeshima Y, Yagi M, Ishikawa Y, Ishikawa Y, Minami R, Nakamura H, Matsuo M 2001 Oligonucleotides against a splicing enhancer sequence led to dystrophin production in muscle cells from a Duchenne muscular dystrophy patient. Brain Dev 23: 788–798
van Deutekom JCT, Bremmer-Bout M, Janson AAM, Ginjaar IB, Baas F, den Dunnen JT, van Ommen GJ 2001 Antisense-induced exon skipping restores dystrophin expression in DMD patient derived muscle cells. Hum Mol Genet 10: 1547–1554
Mann CJ, Honeyman K, Cheng AJ, Ly T, Lloyd F, Fletcher S, Morgan JE, Partridge TA, Wilton SD 2001 Antisense-induced exon skipping and synthesis of dystrophin in the mdx mouse. Proc Natl Acad Sci U S A 98: 42–47
Acknowledgements
The authors thank N. Kageyama for her help in preparing the manuscript.
Author information
Authors and Affiliations
Corresponding author
Additional information
Supported by a grant from the Ministry of Education, Science, Sports and Culture of Japan and a research grant (11B-1) from the Ministry of Health, Labor and Welfare, Japan.
Rights and permissions
About this article
Cite this article
Adachi, K., Takeshima, Y., Wada, H. et al. Heterogous Dystrophin mRNA Produced by a Novel Splice Acceptor Site Mutation in Intermediate Dystrophinopathy. Pediatr Res 53, 125–131 (2003). https://doi.org/10.1203/00006450-200301000-00021
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1203/00006450-200301000-00021
This article is cited by
-
Faulty RNA splicing: consequences and therapeutic opportunities in brain and muscle disorders
Human Genetics (2017)
-
A novel splicing silencer generated by DMD exon 45 deletion junction could explain upstream exon 44 skipping that modifies dystrophinopathy
Journal of Human Genetics (2014)
-
Mutation spectrum of the dystrophin gene in 442 Duchenne/Becker muscular dystrophy cases from one Japanese referral center
Journal of Human Genetics (2010)
-
Identification of seven novel cryptic exons embedded in the dystrophin gene and characterization of 14 cryptic dystrophin exons
Journal of Human Genetics (2007)
-
A nonsense mutation-created intraexonic splice site is active in the lymphocytes, but not in the skeletal muscle of a DMD patient
Human Genetics (2006)