Abstract
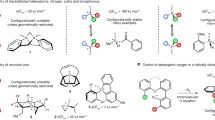
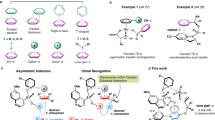
Complex structure and its energy were theoretically predicted between the N-terminal segment of right-handed 310-helical peptide (1) and chiral acid based on various amino acids. Two categories of the chiral acids have been chosen. One is N-carbonyl-blocked amino acid for the three-point coordination to the N-terminal sequence of peptide 1. The other acid for the two-point coordination contains no extra carbonyl groups. Energy minimization from the corresponding initial models was performed by semiempirical molecular orbital calculation. In each amino acid species, the three-point coordination, compared with the two-point type, tends to generate larger difference in energies of D-/L-complexes, which are more stable for L-species bound to right-handed helix. In the three-point binding, N-carbonyl-blocked L-amino acid is prone to adopt negative φ values. Density functional method was also applied to smaller analogs, providing similar tendency in complex structure and energy difference. The predictions obtained here are fully consistent with our previous findings [Y. Inai et al., J. Am. Chem. Soc., 125, 8151–8162 (2003)], in which preferential induction of right-handed helix in peptide 1 occurs with N-carbonyl-protected L-amino acid, but inefficiently with simple carboxylic acid. The energetic advantage for the three-point binding implies the function of 310-helical N-terminus to discriminate the chirality of N-carbonyl-blocked peptide acid molecule.
Similar content being viewed by others
Article PDF
References
J. S. Richardson and D. C. Richardson, Science, 240, 1648 (1988).
R. Aurora, R. Srinivasan, and G. D. Rose, Science, 264, 1126 (1994).
L. Serrano and A. R. Fersht, Nature, 342, 296 (1989).
L. G. Presta and G. D. Rose, Science, 240, 1632 (1988).
R. L. Baldwin and G. D. Rose, Trends Biochem. Sci., 24, 26 (1999).
Y. W. Chen and A. R. Fersht, FEBS Lett., 347, 304 (1994).
M. Petukhov, K. Uegaki, N. Yumoto, S. Yoshikawa, and L. Serrano, Protein Sci., 8, 2144 (1999).
Y. Inai, N. Ousaka, and T. Okabe, J. Am. Chem. Soc., 125, 8151 (2003).
Y. Inai and H. Komori, Biomacromolecules, 5, 1231 (2004).
Y. Inai, K. Tagawa, A. Takasu, T. Hirabayashi, T. Oshikawa, and M. Yamashita, J. Am. Chem. Soc., 122, 11731 (2000).
Y. Inai, Y. Ishida, K. Tagawa, A. Takasu, and T. Hirabayashi, J. Am. Chem. Soc., 124, 2466 (2002).
Y. Inai, H. Komori, A. Takasu, and T. Hirabayashi, Biomacromolecules, 4, 122 (2003).
Y. Inai and T. Hirano, ITE Lett. Batt. New Technol. Med., 4, 485 (2003).
For 310-helix, see: a) C. Toniolo and E. Benedetti, Trends Biochem. Sci., 16, 350 (1991).
D. J. Barlow and J. M. Thornton, J. Mol. Biol., 201, 601 (1988).
For the three-point coordination, see: a) C. E. Dalgliesh, J. Chem. Soc., 3940 (1952).
W. H. Pirkle and D. W. House, J. Org. Chem., 44, 1957 (1979).
Y. Okamoto and E. Yashima, Angew. Chem., Int. Ed., 37, 1020 (1998).
For elegant examples of theoretical study on chiral discriminations, see: a) E. Yashima, T. Nimura, T. Matsushima, and Y. Okamoto, J. Am. Chem. Soc., 118, 9800 (1996).
S. Topiol, M. Sabio, J. Moroz, and W. B. Caldwell, J. Am. Chem. Soc., 110, 8367 (1988).
K. Zborowski and G. Zuchowski, Chirality, 14, 632 (2002).
I. Alkorta, O. Picazo, and J. Elguero, Tetrahedron: Asymmetry, 15, 1391 (2004).
Strictly speaking, we hardly distinguish between the three-point and two-point interactions, because the two-point interactions, prospected for a given chiral additive, should be regarded as three-point or more, if weaker interactions are considered. Here, the number of interaction points is limited to a clear coupling to the three binding N or NH sites of 310-helical N-terminus through hydrogen bonding or ionic interactions (where the ionic species of –COO2− NH3+– is treated as the single-point coordination, whereas hydrogen bonding interaction should be involved. 6a,b) This definition should be reasonable for intuitive understanding of essential binding modes for the present chiral discrimination. For hydrogen-bonding 6a and ionic-binding 6b criteria in protein structures, see: a) I. Y. Torshin, I. T. Weber, and R. W. Harrison, Protein Eng., 15, 359 (2002).
S. Kumar and R. Nussinov, Biophys. J., 83, 1595 (2002).
M. J. S. Dewar, E. G. Zoebisch, E. F. Healy, and J. J. P. Stewart, J. Am. Chem. Soc., 107, 3902 (1985).
For the MOPAC97 program, see: b) J. J. P. Stewart, MOPAC97; Fujitsu Ltd.: Tokyo, Japan, 1998.
For detailed description of MOPAC keywords and their methods, see also: c) J. J. P. Stewart, MOPAC 2000 Manual; Fujitsu Ltd.: Tokyo, Japan, 1999.
J. J. P. Stewart, MOPAC 93 Manual, revision number 2; Fujitsu Ltd.: Tokyo, Japan, 1993.
For the EF-routine, see also: e) J. Baker, J. Comput. Chem., 7, 385 (1986).
Gaussian 03, Revision C.02, M. J. Frisch, G. W. Trucks, H. B. Schlegel, G. E. Scuseria, M. A. Robb, J. R. Cheeseman, J. A. Montgomery, Jr., T. Vreven, K. N. Kudin, J. C. Burant, J. M. Millam, S. S. Iyengar, J. Tomasi, V. Barone, B. Mennucci, M. Cossi, G. Scalmani, N. Rega, G. A. Petersson, H. Nakatsuji, M. Hada, M. Ehara, K. Toyota, R. Fukuda, J. Hasegawa, M. Ishida, T. Nakajima, Y. Honda, O. Kitao, H. Nakai, M. Klene, X. Li, J. E. Knox, H. P. Hratchian, J. B. Cross, V. Bakken, C. Adamo, J. Jaramillo, R. Gomperts, R. E. Stratmann, O. Yazyev, A. J. Austin, R. Cammi, C. Pomelli, J. W. Ochterski, P. Y. Ayala, K. Morokuma, G. A. Voth, P. Salvador, J. J. Dannenberg, V. G. Zakrzewski, S. Dapprich, A. D. Daniels, M. C. Strain, O. Farkas, D. K. Malick, A. D. Rabuck, K. Raghavachari, J. B. Foresman, J. V. Ortiz, Q. Cui, A. G. Baboul, S. Clifford, J. Cioslowski, B. B. Stefanov, G. Liu, A. Liashenko, P. Piskorz, I. Komaromi, R. L. Martin, D. J. Fox, T. Keith, M. A. Al-Laham, C. Y. Peng, A. Nanayakkara, M. Challacombe, P. M. W. Gill, B. Johnson, W. Chen, M. W. Wong, C. Gonzalez, and J. A. Pople, Gaussian, Inc., Wallingford CT, 2004.
P. J. Stephens, F. J. Devlin, C. F. Chabalowski, and M. J. Frisch, J. Phys. Chem., 98, 11623 (1994).
A. D. Becke, J. Chem. Phys., 98, 5648 (1993).
C. Lee, W. Yang, and R. G. Parr, Phys. Rev. B, 37, 785 (1988).
For methods, usages, and full references in Gaussian 03, see: d) Gaussian Online Manual (http://www.gaussian.com).
E. Yashima, T. Matsushima, and Y. Okamoto, J. Am. Chem. Soc., 119, 6345 (1997).
E. Yashima, K. Maeda, and Y. Okamoto, Nature, 399, 449 (1999).
T. Ishi-i, M. Crego-Calama, P. Timmerman, D. N. Reinhoudt, and S. Shinkai, Angew. Chem., Int. Ed., 41, 1924 (2002).
Y. Mizuno, T. Aida, and K. Yamaguchi, J. Am. Chem. Soc., 122, 5278 (2000).
E. Yashima, K. Maeda, and T. Nishimura, Chem. Eur. J., 10, 42 (2004).
D. S. Schlitzer and B. M. Novak, J. Am. Chem. Soc., 120, 2196 (1998).
For instance, see: a) S. S. Zimmerman, M. S. Pottle, G. Némethy, and H. A. Scheraga, Macromolecules, 10, 1 (1977).
For elegant example of systematic search in peptide conformation, see: M. Oka, Y. Baba, A. Kagemoto, and A. Nakajima, Polym. Bull., 25, 87 (1991).
For the general definition of torsion angle for peptide conformation, see: IUPAC-IUB Commission on Biochemical Nomenclature, Biochemistry, 9, 3471 (1970).
D. Ajò, M. Casarin, and G. Granozzi, J. Mol. Struct. (THEOCHEM), 86, 297 (1982).
Y. Inai, T. Oshikawa, M. Yamashita, K. Tagawa, and T. Hirabayashi, Biopolymers, 70, 310 (2003).
G. Alagona, C. Ghio, and C. Pratesi, J. Comput. Chem., 12, 934 (1991).
M. A. Broda, D. Siodlak, and B. Rzeszotarska, J. Pep. Sci., 11, 235 (2005).
M. J. Citra, M. G. Paterlini, T. B. Freedman, A. Fissi, and O. Pieroni, Proc. SPIE-Int. Soc. Opt. Eng., 2089, 478 (1993); SciFinder Scholar, American Chemical Society, CAN 121:109655.
C. Alemán and J. J. Perez, Biopolymers, 33, 1811 (1993).
D. Siodlak, B. Rzeszotarska, M. A. Brodal, A. E. Koziol, and E. Kolodziejczyk, Acta Biochim. Pol., 51, 145 (2004).
F. S. Nandel and B. Khare, Biopolymers, 77, 63 (2005).
M. A. Thompson, ArgusLab 4.0.1, Planaria Software LLC, Seattle, WA, 2004 (http://www.arguslab.com).
The ΔED–L value (1.89) and complex structure (Figure 1) for Boc-Leu-OH are updated from those in ref 2a. Several conformers for the Leu side chain are considered in the present initial modeling, as mentioned in the experimental section.
While the energy bias was interpreted from steric repulsion between a helical chain and a chiral molecule, 2a,b the present explanation for 2–5 species of Leu, Ala, Phe, and Val should be more comprehensive.
B. Pengo, F. Formaggio, M. Crisma, C. Toniolo, G. M. Bonora, Q. B. Broxterman, J. Kamphuis, M. Saviano, R. Iacovino, F. Rossi, and E. Benedetti, J. Chem. Soc., Perkin Trans. 2, 1651 (1998).
For the definition of torsion angles in β-amino acid, see:a) I. L. Karle, A. Pramanik, A. Banerjee, S. Bhattacharjya, and P. Balaram, J. Am. Chem. Soc., 119, 9087 (1997).
R. P. Cheng, S. H. Gellman, and W. F. DeGrado, Chem. Rev., 101, 3219 (2001). For the general definition, see ref 12.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Inai, Y., Ousaka, N. & Miwa, Y. Theoretical Comparison between Three-Point and Two-Point Binding Modes for Chiral Discrimination upon the N-Terminal Sequence of 310-Helix. Polym J 38, 432–441 (2006). https://doi.org/10.1295/polymj.38.432
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1295/polymj.38.432