Abstract
Background:
Lysine-specific demethylase 1 (LSD1 or KDM1A) overexpression correlates with poor survival and castration resistance in prostate cancer. LSD1 is a coregulator of ligand-independent androgen receptor signaling promoting c-MYC expression. We examined the antitumor efficacy of LSD1 inhibition with HCI-2509 in advanced stages of prostate cancer.
Methods:
Cell survival, colony formation, histone methylation, c-MYC level, c-MYC expression, cell cycle changes and in vivo efficacy were studied in castration-resistant prostate cancer cells upon treatment with HCI-2509. In vitro combination studies, using HCI-2509 and docetaxel, were performed to assess the synergy. Cell survival, colony formation, histone methylation and c-myc levels were studied in docetaxel-resistant prostate cancer cells treated with HCI-2509.
Results:
HCI-2509 is cytotoxic and inhibits colony formation in castration-resistant prostate cancer cells. HCI-2509 treatment causes a dose-dependent increase in H3K9me2 (histone H3lysine 9) levels, a decrease in c-MYC protein, inhibition of c-MYC expression and accumulation in the G0/G1 phase of the cell cycle in these cells. PC3 xenografts in mice have a significant reduction in tumor burden upon treatment with HCI-2509 with no associated myelotoxicity or weight loss. More synergy is noted at sub-IC50 (half-maximal inhibitory concentration) doses of docetaxel and HCI-2509 in PC3 cells than in DU145 cells. HCI-2509 has growth-inhibitory efficacy and decreases the c-myc level in docetaxel-resistant prostate cancer cells.
Conclusions:
LSD1 inhibition with HCI-2509 decreases the c-MYC level in poorly differentiated prostate cancer cell lines and has a therapeutic potential in castration- and docetaxel-resistant prostate cancer.
Similar content being viewed by others
Introduction
The efficacy of androgen deprivation therapy in high-risk metastatic prostate cancer is short lived, and tumors evolve during treatment with androgen deprivation therapy to become castration-resistant.1 Treatment with abiraterone, enzalutamide, radium-223, sipuleucel-T and taxane-based chemotherapy provide additional survival benefit;2 however, castration-resistant prostate cancer (CRPC) eventually progresses and remains the second-leading cause of cancer death in men.3
Epigenetic processes have a significant role in embryonic pluripotency, cellular differentiation, cancer progression and epithelial–mesenchymal transition. Lysine-specific demethylase 1 (LSD1), the first histone demethylase enzyme identified, can either function as a corepressor or a coactivator of transcriptional regulation depending on its association with differing binding partners. In association with CoREST4 and NuRD,5 it demethylates mono- and dimethyl histone H3lysine 4 (H3K4me1 and H3K4me2), resulting in transcriptional repression. When LSD1 partners with nuclear androgen or estrogen hormone receptors, it switches its substrate specificity to mono- and dimethyl histone H3lysine 9 (H3K9me1/me2), resulting in transcriptional activation.6 As an androgen receptor (AR) coregulator, LSD1 has a significant role in both hormone-dependent6 and -independent7, 8 prostate cancer.
LSD1 also mediates epithelial–mesenchymal transition and resistance to chemotherapy in mouse hepatocyte cell cultures. Demethylation of H3K9me2 allows derepression of large heterochromatin domains and enables transcription of genes that confer a survival advantage in a challenging cellular environment.9
LSD1 is overexpressed in poorly differentiated prostate cancer and predicts poor survival.10 As prostate cancer progresses through treatment modalities of androgen deprivation and chemotherapy, it acquires a progressively more stem cell-like character.11 AR signaling continues to have a significant role as a driver of CRPC by various mechanisms such as AR gene amplification,12, 13 constitutionally active AR splice variants14, 15 local androgen production and hijacking of AR signaling by other oncogenic pathways and ligands.16, 17 LSD1 colocalizes with the AR in prostate cancer cells and has a critical role in AR signaling, with c-MYC oncogene as a crucial ligand-independent AR target gene.18 c-Myc signaling has a significant role in mediating castration resistance,19 epithelial–mesenchymal transition and docetaxel resistance20 in prostate cancer cells.
Targeting epigenetic mechanisms to treat advanced stages of prostate cancer is an attractive strategy. DNA demethylation,21 bromodomain inhibition18 and HDAC inhibition22 have been studied in this context.
In this study, we investigate the growth-inhibitory effects of HCI-2509 in PC3 and DU145 cell lines. These cell lines have been derived from poorly differentiated metastatic prostate cancer, have low AR gene expression and low levels of AR protein.23 AR signaling continues to have a significant central role in the CRPC regulatory network in both of these cell lines.24
HCI-2509 is a potent, reversible and selective LSD1 inhibitor with demonstrated preclinical efficacy in Ewing’s sarcoma,25 acute myeloid leukemia26 and endometrial cancer.27 HCI-2509 is the lead compound for preclinical studies targeting LSD1 developed from a novel series of LSD1 inhibiting N′-(1-phenylethylidine)-benzo hydrazide compounds.28 Our initial screening studies with several different prostate cancer cell lines revealed cytotoxicity with this class of LSD1 inhibitors. PC3 and DU145 cells represent two distinct groups with regard to clusters of cellular phenotypes in CRPC. These clusters are derived from CRPC gene expression data sets and are based on differences in their active CRPC regulatory pathways.24 For the purpose of our studies, we have chosen these poorly differentiated CRPC cell lines and a docetaxel-resistant prostate cancer cell line as representative of areas of unfulfilled clinical need in this disease.
Materials and methods
Cell culture and cell viabilities
Human prostate cancer cell lines LNCAP, VCAP, 22RV1, PC3 and DU145 were obtained from American Type Culture Collection (ATCC, Manassas, VA, USA) and cultured in Roswell Park Memorial Institute (RPMI) medium supplemented with 10% fetal bovine serum, 100 U ml−1 penicillin and 100 μg ml−1 streptomycin at 37 °C in a humidified atmosphere containing 5% CO2. Cells were seeded in triplicate in 96-well plates at a density of 5000 cells per well, allowed to adhere overnight and then treated with either vehicle (1% dimethyl sulfoxide (DMSO) in RPMI) or increasing concentrations of HCI-2509 (Center for Investigational Therapeutics, Huntsman Cancer Institute, Salt Lake City, UT, USA). After incubation for 72 h, cell viability was assessed using CellTiter-Glo (Promega, Madison, WI, USA).
Further experiments were performed with the CRPC cell lines PC3 and DU145. Cell viabilities of these cell lines were tested using docetaxel (Selleck Pharmaceuticals, Houston, TX, USA).
Colony formation assays
Colony formation assays were performed in duplicate in 6-well plates for each cell line treated with either vehicle (1% DMSO) or serially increasing concentrations of HCI-2509 ranging from 1 nm to 1000 nm at the time of setting up the assay. Using stock solutions of 1.6% agarose (SeaPlaque GTG agarose from Lonza, Allendale, NJ, USA) and 2 × RPMI (RPMI powder; Life Technologies, Grand Island, NY, USA), an underlayer constituted by 0.8% agarose in RPMI was prepared. An overlayer of 5000 treated cells per well suspended in 0.4% agarose in RPMI was prepared. After incubation for 3 weeks, colonies were photographed and counted.
Western blot
Cells were seeded (1 × 106) in 10 cm dishes. Once 50% confluent, these were treated with vehicle (1% DMSO) or HCI-2509 at increasing concentrations of 1, 3 and 10 μm for 24 to 48 h. Treated cells were lysed, their protein extracted and immunoblotted with the following antibodies: β-actin mouse mAb: LI-COR (Lincoln, NE, USA) 926-42212; dimethyl histone H3 (K9) rabbit mAb: Cell Signaling 9753S; c-MYC rabbit mAb: Cell Signaling (Boston, MA, USA) 9402S.
RNA isolation and quantitative PCR
Cells were seeded (1 × 106) in 10 cm dishes. Once 50% confluent, these were treated with 1% DMSO or 1 μm of HCI-2509 for 4 and 24 h. Total RNA was extracted from the cells using an RNeasy Plus Kit (Qiagen, Valencia, CA, USA). cDNA was generated using High Capacity cDNA Reverse Transcription Kit (Applied Biosystems, Boston, MA, USA). This was then amplified, detected and quantified using SYBR green fluorescence (SYBR Green PCR Master Mix; Applied Biosystems) using the ViiA 7 Real-Time PCR System (Boston, MA, USA). The following primers were used: 18S (Thermo Fisher, Boston, MA, USA; Hs03928990_g1) and c-MYC (Thermo Fisher; Hs00153408_m1).
Cell cycle
Cells in active growth phase were treated with vehicle (1% DMSO), HCI-2509 at varying concentrations (1, 3 and 10 μm), docetaxel at 100 nm or a combination of HCI-2509 (10 μm) plus docetaxel (100 nm) for 24 h. Cells were harvested, fixed in ethanol, stained with propidium iodide and then analyzed using a BD FACSCanto Analyzer with Software Diva vs6.1.3 (BD Biosciences, San Jose CA, USA).
Xenograft studies
All in vivo studies were approved and conducted as per the regulations of the Institutional Animal Care and Use Committee at the University of Utah. Sample size calculation accounted for an attrition rate of 10%. The power calculation was based on a two-tailed test of significance with a type I error of 0.05 and a 0.9 probability of detecting a true difference. This yielded a sample size of 10 animals per arm.
Five million PC3 cells were suspended in 50% media (RPMI) and 50% Matrigel (BD Biosciences), and then implanted in the right hind flank of female nude mice. Animals bearing tumors of adequate size (100 mm3) were randomized based on size into three cohorts of 10 animals each and dosed with either vehicle intraperitoneally (Monday, Wednesday and Friday), 40 mg kg−1 HCI-2509 intraperitoneally (Monday, Wednesday and Friday) or 40 mg kg−1 HCI-2509 orally (per os) (daily, Monday through Friday) for 3 weeks (last dose administered on Day 19). Body weight and tumor volume were measured twice weekly. Blood cell counts (hematocrit, platelet and white blood cell) were measured on days 1 and 24 using a HemaTrue Hematology Analyzer (Heska, Loveland, CO, USA).
Heat maps
PC3 and DU145 cells were seeded in triplicate in 96-well plates at a density of 5000 cells per well and allowed to adhere overnight. These were then treated with vehicle (1% DMSO), increasing doses of HCI-2509 and docetaxel alone and in a 7 × 7 combination at seven different dose levels of each drug clustered around the previously calculated half-maximal inhibitory concentration (IC50s). Cell viability was assessed after 72 h of treatment.
Studies on docetaxel-resistant cells
Docetaxel-resistant PC3D12 (PC3D12) cells29 were obtained, cultured and maintained with 12 nm docetaxel in RPMI growth medium. Cell viabilities, colony formation assays and immunoblotting for c-myc and H3K9me2 upon treatment with HCI-2509 were performed by methods described above.
Results
LSD1 inhibition with HCI-2509 is cytotoxic to prostate cancer cells
A dose-dependent decrease in cell viability is observed after 72 h of treatment with HCI-2509 in all prostate cancer cell lines with IC50s in the high nanomolar to low micromolar range, as shown in Figure 1a.
Effects of HCI-2509 and docetaxel on cell survival in prostate cancer cell lines. The logarithmic drug concentration was plotted versus the percent cell survival and half-maximal inhibitory concentrations (IC50s) were calculated by nonlinear regression analysis using GraphPad Prism (GraphPad Software, San Diego, CA, USA). Dose–response curves showing the percent cell viability at the end of 72 h treatment in (a) VCAP, LNCAP, 22RV1, PC3 and DU145 cells with HCI-2509, and (b) PC3 and DU145 cells with docetaxel.
IC50s for PC3 and DU145 cells using docetaxel are close to 5 nm (Figure 1b).
HCI-2509 is a potent inhibitor of colony formation in CRPC cells
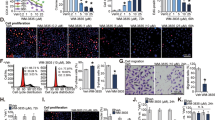
Anchorage-independent growth is inhibited with diminished colony numbers (Figures 2a and b) and size (Figure 2c) in these cells. It is noteworthy that the effect of initial treatment persists for 3 weeks. Anchorage-independent growth is significantly more sensitive to LSD1 inhibition with close to 50% reduction in colony formation at 10 nm concentration. There is complete ablation of colony formation at 1 μm dose of HCI-2509.
Effect of HCI-2509 on colony formation using soft agar assays in castration-resistant prostate cancer cells. A bar graph was plotted to represent the fractional inhibition of colony formation upon treatment with HCI-2509 compared with vehicle using GraphPad Prism in (a) PC3 and (b) DU145 cells Assays were performed in duplicate and error bars indicate the standard deviation. Photographs demonstrate average colony size at × 20 magnification with different doses of HCI-2509 in each cell line (c), with quantification of colony size plotted as a bar graph using GraphPad Prism. Student’s unpaired t-tests, using GraphPad Prism software, were used to calculate two-tailed P-values for each experiment to determine statistical significance between untreated and treated samples. The statistically significant P-values are labeled in each figure with an asterisk.
LSD1 inhibition affects the concentration of transcriptional regulators
Western blot analysis shows that treatment of PC3 and DU145 cells with HCI-2509 results in a dose-dependent increase in H3K9me2 and decrease in c-MYC protein. (Figures 3a and b).
Western blot analysis of H3K9me2 and c-MYC levels upon treatment with increasing concentrations of HCI-2509 as compared with vehicle in (a) PC3 and (b) DU145 cells. The bands of c-MYC, β-actin and H3K9me2 proteins on the immunoblot were cropped using PowerPoint (Windows) to show correlating change in intensities with different levels of treatment with HCI-2509. (Replicated two more times, representative blot shown).
HCI-2509 inhibits c-MYC gene expression in CRPC cells
LSD1 inhibition with HCI-2509 (1 μm) results in inhibition of c-MYC gene transcription in PC3 and DU145 cells at 4 and 24 h treatment time-points (Figures 4a and b).
Quantitative reverse transcription-PCR (RT-PCR) analysis of c-MYC oncogene in (a) PC3 and (b) DU145 cells after treatment with HCI-2509 for 4 and 24 h. Each replicate was normalized to the reference gene (S18) and induction was calculated relative to the vehicle control. The relative expression of c-MYC mRNA in various conditions was plotted in the form of bar graphs and relative inhibition on treatment with HCI-2509 was tested for significance using the unpaired t-test (GraphPad Prism). Assays were performed in quadruplicate and error bars indicate the standard deviation.
LSD1 inhibition by HCI-2509 affects the cell cycle in CRPC cells
Docetaxel causes a G2M arrest in PC3 and DU145 cells. LSD1 inhibition results in the accumulation of cells in the G0/G1 phase of the cell cycle. Treatment with HCI-2509 and docetaxel simultaneously leads to a bimodal peak in G0/G1 and G2/M for both CRPC cell lines (Figures 5a and b).
Cell cycle study of percent cell populations in various phases after treatment with vehicle, HCI-2509 in increasing concentrations and docetaxel alone and in combination with HCI-2509 using (a) PC3 and (b) DU145 cells. Student’s unpaired t-tests, using GraphPad Prism software, were used to calculate two-tailed P-values for each experiment to determine statistical significance between untreated and treated samples. The statistically significant P-values are labeled in each figure with an asterisk. (Replicated two more times, representative graph shown).
PC3 xenografts show in vivo efficacy of HCI-2509
Treatment with HCI-2509 both intraperitoneally and per os causes a significant reduction in tumor burden in PC3 xenografts (Figure 6a) with progressive weight gain in all three arms (Figure 6b). There is no significant change in the platelet count, white blood cell count or hematocrit (Figures 6c–e) in the treatment arms compared with the control arm.
PC3 xenograft study shows efficacy and tolerability of HCI-2509 in vivo. (a) Tumor volume (TV) plotted versus time in different treatment arms—vehicle, intraperitoneal HCI-2509 and per os HCI-2509. The mean TV with error bars to show the standard error of mean (s.e.m.) has been used for the three groups. (b) Relative body weight (BW) of mice in the three treatment arms plotted versus time while on study. Relative mean BW with error bars representing s.e.m. in each group are used. Change in (c) white blood cell count, (d) hematocrit and (e) platelet count in the three treatment arms. Data are represented by creating graphs using the GraphPad Prism software. Statistical analysis of the change in mean tumor volume in each arm with time and treatment was performed using two-way analysis of variance (ANOVA) test. The changes in tumor volume with both forms of treatment as compared with control were found to be significant.
Combination treatment with docetaxel and HCI-2509 can be synergistic
Treatment with docetaxel and HCI-2509 in combination in CRPC cell lines is synergistic, additive or competitive depending on the concentrations used and the cell line studied. The combination is synergistic at sub-IC50 concentrations, with more points of synergy observed in PC3 cells than in DU145 cells. Isobolograms showing the combination points and drug ratios displaying synergy, additivity and competition are presented in Figures 7a–d.
Combination of docetaxel and HCI-2509 in (a) PC3 and (b) DU145 cells. Combination index (CI) values were calculated by Chou–Talalay equation using CalcuSyn (BIOSOFT, Great Shelford, Cambridge, UK). Heat map was created using the CI values at various combination doses to show points of synergy (CI<1). Isobologram of CIs in PC3 and DU145 cells in various combinations (c and d).
HCI-2509 has similar inhibitory effects in docetaxel-resistant prostate cancer cells
IC50 for PC3D12 using docetaxel is >50 nm and the IC50 using HCI-2509 is similar to that for CRPC cells with IC50s in the high nanomolar to low micomolar range. Treatment with HCI-2509 potently inhibits colony formation and diminishes colony size in PC3D12 cells. HCI-2509 treatment in these cells results in a dose-dependent increase in H3K9me2 levels and a decrease in c-myc protein concentration.
Discussion
Our studies demonstrate the efficacy of reversible and specific LSD1 inhibition with HCI-2509 in growth inhibition and c-MYC decrease in poorly differentiated castration- and docetaxel-resistant prostate cancer cell lines with low AR expression.
Nonspecific amine oxidase inhibitors inhibit LSD1 but have off-target side effects.30 Pargyline, an irreversible monoamine oxidase inhibitor, used as an LSD1 inhibitor, does not have any antitumor efficacy in prostate cancer xenograft studies.31 Cryptotanshinone, another LSD1 inhibitor, downregulates AR signaling. However, its action appears to be dependent on the interaction between androgen ligand, AR and LSD1 and it fails to suppress growth in the absence of DHT in LNCAP (a hormone-sensitive prostate cancer cell line) and castration-resistant PC3 cells.32 Reversible specific LSD1 inhibition with namoline, a γ-pyrone, is effective in the androgen-sensitive prostate cancer cell line LNCAP and has in vivo antitumor efficacy, and also causes weight loss in treated mice.33 Our studies demonstrate cytotoxicity of HCI-2509 in PC3 and DU145 cell lines with IC50s in the high nanomolar to low micromolar range (Figure 1a).
LSD1 inhibitors have been shown to inhibit pluripotent cancer cells of germ cell tumor origin without affecting normal somatic cells.34 The remarkable efficacy of HCI-2509 in inhibiting colony formation in our studies (Figures 2a–c) suggests that LSD1 maintains stem cell function in advanced stages of prostate cancer. This finding is reminiscent of the vital role of LSD1 in the maintenance of pluripotency in human embryonic stem cells.35
c-MYC is an important androgen ligand-independent AR target gene that contributes to AR-dependent prostate cancer cell survival in CRPC.18 We demonstrate a dose-dependent decrease in c-MYC protein levels (Figure 3a), with HCI-2509 treatment, correlating with a corresponding increase in H3K9me2 levels (Figure 3b) in castration-resistant cell lines. Quantitative reverse transcription-PCR was performed to evaluate whether the drop in c-MYC protein is due to degradation or a direct effect on gene expression. c-MYC gene expression is significantly reduced at 4 and 24 h after treatment with HCI-2509 (1 μm) (Figure 4).
Downregulation of c-MYC has been shown to rescue tumor cells from chemotherapy and radiation in some studies36 and increase sensitivity to chemotherapy in others.37 The effect of HCI-2509 on cell cycle reflects the effect of c-MYC decrease. Cells accumulate in the G0/G1 phase (Figures 5a and b), correlating with reduction of c-MYC protein. Docetaxel is commonly used in CRPC.38 It causes a G2/M arrest in the cell cycle owing to the stabilization of microtubules of the mitotic spindle.39 The presence of one drug may rescue cytotoxicity by the other or may work synergistically by inhibiting different cell populations. We explored the outcome of combining HCI-2509 with docetaxel in CRPC cell lines at various dose levels, and noted differing levels of synergy for the two cell lines at sub-IC50 doses and competition at higher concentrations (Figures 7a–d), thus reflecting the complexity of the effect of epigenetic manipulation on cells. Synergy was noted more commonly in PC3 cells than in DU145 cells.
Several different mechanisms have been described to explain chemotherapy resistance in cancer including epithelial–mesenchymal transition with TGF-β-induced LSD1 overexpression.9 Previous studies have shown activation of c-MYC expression in docetaxel-resistant prostate cancer cell lines.20 In the castration-resistant setting, use of docetaxel provides an ~3-month average survival advantage38 with eventual development of toxicity or docetaxel resistance.
As an exploratory objective, we studied the effect of HCI-2509 in the docetaxel-resistant cell line, PC3D12. We first performed cell viability studies confirming docetaxel resistance in PC3D12, as demonstrated by an IC50 >50 nm for docetaxel (Figure 8a). There is evidence for the continued efficacy of HCI-2509 in the docetaxel-resistant setting with cytotoxicity (Figure 8b) and potent inhibition of colony formation, as well as a demonstration of increasing H3K9me2 levels and decreasing c-MYC levels with treatment (Figures 8c–e). The efficacy of HCI-2509 in PC3D12 cells suggests that sequencing treatment with LSD1 inhibition after progression on docetaxel in the castration-resistant setting may be a possibility in the future.
Effect of HCI-2509 on docetaxel-resistant PC3D12 cells. Cell viability studies: dose–response curves showing the percent cell viability at the end of 72 h treatment in PC3D12 treated with (a) docetaxel and (b) HCI-2509. Colony formation assays show growth inhibition of colonies (c) and decreased colony size (d) on treatment with HCI-2509. Student’s unpaired t-tests, using GraphPad Prism software, were used to calculate two-tailed P-values for each experiment to determine the statistical significance between untreated and treated samples. The statistically significant P-values are labeled in each figure with an asterisk. (e) Western blot analysis shows increased H3K9me2 and decreased c-MYC levels on treatment with HCI-2509.
In vivo studies demonstrate the antitumor efficacy of HCI-2509 in PC3 xenografts (Figure 6a). HCI-2509 is well tolerated with progressive weight gain in treated mice (Figure 6b), similar to the control arm. There is no significant myelosuppression (Figures 6c–e). Tolerability and myeloid sparing qualities of therapy are an important consideration in the setting of metastatic CRPC and docetaxel-resistant prostate cancer, due to high frequency of bone metastasis leading to marrow replacement, advanced age, comorbidities and myelotoxicity from previous treatments in this patient population. Our in vivo study supports use of reversible LSD1 inhibition in this clinical setting.
c-MYC is implicated in stem cell maintenance40 and, as an oncogenic transcription factor, has been an elusive target for drug development. BET bromodomain inhibition with JQ1 has been shown to downregulate c-MYC transcription in prostate cancer and multiple myeloma.18, 41 Here we report LSD1 inhibition as yet another potent epigenetic strategy to inhibit c-MYC expression in the setting of advanced prostate cancer.
We conclude that reversible and potent LSD1 inhibition with HCI-2509 has antitumor efficacy in CRPC cell lines, inhibits c-MYC expression, can sensitize prostate cancer cells to docetaxel and inhibits growth in a docetaxel-resistant prostate cancer cell line. This provides a new therapeutic tool, in the space of an unmet clinical need, in patients who have poorly differentiated prostate cancer and are facing limited survival with currently available therapies for CRPC. Neuroendocrine differentiation presents yet another challenge in the treatment of prostate cancer.42 Epigenetic mechanisms such as LSD1-mediated gene regulation may have a role in this phenomenon. This is a possible direction for future studies.
References
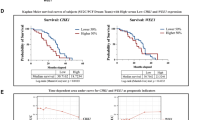
James ND, Spears MR, Clarke NW, Dearnaley DP, De Bono JS, Gale J et al. Survival with newly diagnosed metastatic prostate cancer in the 'Docetaxel Era': data from 917 patients in the control arm of the STAMPEDE Trial (MRC PR08, CRUK/06/019). Eur Urol 2015; 67: 1028–1038.
Flaig TW, Potluri RC, Ng Y, Todd MB, Mehra M . Treatment evolution for metastatic castration-resistant prostate cancer with recent introduction of novel agents: retrospective analysis of real-world data. Cancer Med 2015; 5: 182–191.
Siegel RL, Miller KD, Jemal A . Cancer statistics, 2016. CA Cancer J Clin 2016; 66: 7–30.
Lee MG, Wynder C, Cooch N, Shiekhattar R . An essential role for CoREST in nucleosomal histone 3 lysine 4 demethylation. Nature 2005; 437: 432–435.
Wang Y, Zhang H, Chen Y, Sun Y, Yang F, Yu W et al. LSD1 is a subunit of the NuRD complex and targets the metastasis programs in breast cancer. Cell 2009; 138: 660–672.
Metzger E, Wissmann M, Yin N, Muller JM, Schneider R, Peters AH et al. LSD1 demethylates repressive histone marks to promote androgen-receptor-dependent transcription. Nature 2005; 437: 436–439.
Wang Q, Li W, Zhang Y, Yuan X, Xu K, Yu J et al. Androgen receptor regulates a distinct transcription program in androgen-independent prostate cancer. Cell 2009; 138: 245–256.
Wu X, Deng F, Li Y, Daniels G, Du X, Ren Q et al. ACSL4 promotes prostate cancer growth, invasion and hormonal resistance. OncoTarget 2015; 6: 44849–44863.
McDonald OG, Wu H, Timp W, Doi A, Feinberg AP . Genome-scale epigenetic reprogramming during epithelial-to-mesenchymal transition. Nat Struct Mol Biol 2011; 18: 867–874.
Kahl P, Gullotti L, Heukamp LC, Wolf S, Friedrichs N, Vorreuther R et al. Androgen receptor coactivators lysine-specific histone demethylase 1 and four and a half LIM domain protein 2 predict risk of prostate cancer recurrence. Cancer Res 2006; 66: 11341–11347.
Domingo-Domenech J, Vidal SJ, Rodriguez-Bravo V, Castillo-Martin M, Quinn SA, Rodriguez-Barrueco R et al. Suppression of acquired docetaxel resistance in prostate cancer through depletion of notch- and hedgehog-dependent tumor-initiating cells. Cancer Cell 2012; 22: 373–388.
Koivisto P, Kononen J, Palmberg C, Tammela T, Hyytinen E, Isola J et al. Androgen receptor gene amplification: a possible molecular mechanism for androgen deprivation therapy failure in prostate cancer. Cancer Res 1997; 57: 314–319.
Edwards J, Krishna NS, Grigor KM, Bartlett JM . Androgen receptor gene amplification and protein expression in hormone refractory prostate cancer. Br J Cancer 2003; 89: 552–556.
Antonarakis ES, Lu C, Wang H, Luber B, Nakazawa M, Roeser JC et al. AR-V7 and resistance to enzalutamide and abiraterone in prostate cancer. N Engl J Med 2014; 371: 1028–1038.
Dehm SM, Schmidt LJ, Heemers HV, Vessella RL, Tindall DJ . Splicing of a novel androgen receptor exon generates a constitutively active androgen receptor that mediates prostate cancer therapy resistance. Cancer Res 2008; 68: 5469–5477.
Abreu-Martin MT, Chari A, Palladino AA, Craft NA, Sawyers CL . Mitogen-activated protein kinase kinase kinase 1 activates androgen receptor-dependent transcription and apoptosis in prostate cancer. Mol Cell Biol 1999; 19: 5143–5154.
Craft N, Shostak Y, Carey M, Sawyers CL . A mechanism for hormone-independent prostate cancer through modulation of androgen receptor signaling by the HER-2/neu tyrosine kinase. Nat Med 1999; 5: 280–285.
Gao L, Schwartzman J, Gibbs A, Lisac R, Kleinschmidt R, Wilmot B et al. Androgen receptor promotes ligand-independent prostate cancer progression through c-Myc upregulation. PLoS One 2013; 8: e63563.
Bernard D, Pourtier-Manzanedo A, Gil J, Beach DH . Myc confers androgen-independent prostate cancer cell growth. J Clin Invest 2003; 112: 1724–1731.
Hatano K, Yamaguchi S, Nimura K, Murakami K, Nagahara A, Fujita K et al. Residual prostate cancer cells after docetaxel therapy increase the tumorigenic potential via constitutive signaling of CXCR4, ERK1/2 and c-Myc. Mol Cancer Res 2013; 11: 1088–1100.
Festuccia C, Gravina GL, D'Alessandro AM, Muzi P, Millimaggi D, Dolo V et al. Azacitidine improves antitumor effects of docetaxel and cisplatin in aggressive prostate cancer models. Endocr-Relat Cancer 2009; 16: 401–413.
Hwang JJ, Kim YS, Kim MJ, Kim DE, Jeong IG, Kim C-S . Histone deacetylase inhibitor potentiates anticancer effect of docetaxel via modulation of Bcl-2 family proteins and tubulin in hormone refractory prostate cancer cells. J Urol 2010; 184: 2557–2564.
Alimirah F, Chen J, Basrawala Z, Xin H, Choubey D . DU-145 and PC-3 human prostate cancer cell lines express androgen receptor: implications for the androgen receptor functions and regulation. FEBS Lett 2006; 580: 2294–2300.
Hu Y, Gu Y, Wang H, Huang Y, Zou YM . Integrated network model provides new insights into castration-resistant prostate cancer. Scientific Rep 2015; 5: 17280.
Sankar S, Theisen ER, Bearss J, Mulvihill T, Hoffman LM, Sorna V et al. Reversible LSD1 inhibition interferes with global EWS/ETS transcriptional activity and impedes Ewing sarcoma tumor growth. Clin Cancer Res 2014; 20: 4584–4597.
Fiskus W, Sharma S, Shah B, Portier BP, Devaraj SGT, Liu K et al. Highly effective combination of LSD1 (KDM1A) antagonist and pan-histone deacetylase inhibitor against human AML cells. Leukemia 2014; 28: 2155–2164.
Theisen ER, Gajiwala S, Bearss J, Sorna V, Sharma S, Janat-Amsbury M . Reversible inhibition of lysine specific demethylase 1 is a novel anti-tumor strategy for poorly differentiated endometrial carcinoma. BMC Cancer 2014; 14: 752.
Sorna V, Theisen ER, Stephens B, Warner SL, Bearss DJ, Vankayalapati H et al. High-throughput virtual screening identifies novel N'-(1-phenylethylidene)-benzohydrazides as potent, specific, and reversible LSD1 inhibitors. J Med Chem 2013; 56: 9496–9508.
O'Neill AJ, Prencipe M, Dowling C, Fan Y, Mulrane L, Gallagher WM et al. Characterisation and manipulation of docetaxel resistant prostate cancer cell lines. Mol Cancer 2011; 10: 126.
Zheng YC, Ma J, Wang Z, Li J, Jiang B, Zhou W et al. A systematic review of histone lysine-specific demethylase 1 and its inhibitors. Med Res Rev 2015; 35: 1032–1071.
Wang M, Liu X, Guo J, Weng X, Jiang G, Wang Z et al. Inhibition of LSD1 by pargyline inhibited process of EMT and delayed progression of prostate cancer in vivo. Biochem Biophys Res Commun 2015; 467: 310–315.
Wu CY, Hsieh CY, Huang KE, Chang C, Kang HY . Cryptotanshinone down-regulates androgen receptor signaling by modulating lysine-specific demethylase 1 function. Int J Cancer 2012; 131: 1423–1434.
Willmann D, Lim S, Wetzel S, Metzger E, Jandausch A, Wilk W et al. Impairment of prostate cancer cell growth by a selective and reversible lysine-specific demethylase 1 inhibitor. Int J Cancer 2012; 131: 2704–2709.
Wang J, Lu F, Ren Q, Sun H, Xu Z, Lan R et al. Novel histone demethylase LSD1 inhibitors selectively target cancer cells with pluripotent stem cell properties. Cancer Res 2011; 71: 7238–7249.
Adamo A, Sese B, Boue S, Castano J, Paramonov I, Barrero MJ et al. LSD1 regulates the balance between self-renewal and differentiation in human embryonic stem cells. Nat Cell Biol 2011; 13: 652–659.
von Bueren AO, Oehler C, Shalaby T, von Hoff K, Pruschy M, Seifert B et al. c-MYC expression sensitizes medulloblastoma cells to radio- and chemotherapy and has no impact on response in medulloblastoma patients. BMC Cancer 2011; 11: 74.
Leonetti C, Biroccio A, Candiloro A, Citro G, Fornari C, Mottolese M et al. Increase of cisplatin sensitivity by c-myc antisense oligodeoxynucleotides in a human metastatic melanoma inherently resistant to cisplatin. Clin Cancer Res 1999; 5: 2588–2595.
Berthold DR, Pond GR, Soban F, de Wit R, Eisenberger M, Tannock IF . Docetaxel plus prednisone or mitoxantrone plus prednisone for advanced prostate cancer: updated survival in the TAX 327 study. J Clin Oncol 2008; 26: 242–245.
Herbst RS, Khuri FR . Mode of action of docetaxel—a basis for combination with novel anticancer agents. Cancer Treat Rev 2003; 29: 407–415.
Araki R, Hoki Y, Uda M, Nakamura M, Jincho Y, Tamura C et al. Crucial role of c-Myc in the generation of induced pluripotent stem cells. Stem Cells 2011; 29: 1362–1370.
Delmore Jake E, Issa Ghayas C, Lemieux Madeleine E, Rahl Peter B, Shi J, Jacobs Hannah M et al. BET bromodomain inhibition as a therapeutic strategy to target c-Myc. Cell 2011; 146: 904–917.
Sun Y, Niu J, Huang J . Neuroendocrine differentiation in prostate cancer. Am J Transl Res 2009; 1: 148–162.
Acknowledgements
We thank Dr William Watson and Dr Amanda O’Neill, University College Dublin, Ireland, for providing docetaxel-resistant PC3 cells. This project received financial support from Huntsman Cancer Institute and Huntsman Cancer Foundation. This work was supported by the University of Utah Flow Cytometry Facility in addition to the National Cancer Institute through Award Number 5P30CA042014-24. We are grateful to Ellen Wilson, PhD, for her discussion and critical feedback.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
SS is a founder and equity holder of Salarius Pharmaceuticals. JB is an equity holder of Salarius Pharmaceuticals. SG’s spouse is an equity holder of Salarius Pharmaceuticals. The remaining authors declare no conflicts of interest.
Rights and permissions
This work is licensed under a Creative Commons Attribution-NonCommercial-NoDerivs 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-nd/4.0/
About this article
Cite this article
Gupta, S., Weston, A., Bearrs, J. et al. Reversible lysine-specific demethylase 1 antagonist HCI-2509 inhibits growth and decreases c-MYC in castration- and docetaxel-resistant prostate cancer cells. Prostate Cancer Prostatic Dis 19, 349–357 (2016). https://doi.org/10.1038/pcan.2016.21
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/pcan.2016.21
This article is cited by
-
The emerging roles of histone demethylases in cancers
Cancer and Metastasis Reviews (2024)
-
Repression of LSD1 potentiates homologous recombination-proficient ovarian cancer to PARP inhibitors through down-regulation of BRCA1/2 and RAD51
Nature Communications (2023)
-
Epigenetic regulation in human cancer: the potential role of epi-drug in cancer therapy
Molecular Cancer (2020)