Abstract
The mouse haematopoietic stem cell (SC) regulator Latexin (LXN) is the only known homologue of the retinoic acid receptor responder 1 (RARRES1) gene. Both genes lie adjacent on chromosome 3 and differ mostly by the presence of a transmembrane domain in RARRES1. Despite their homology, it is not known whether they possess similar regulatory mechanisms, cellular localization and function. Here, we identified RARRES1 and LXN as highly significantly downregulated genes in human prostate SCs, whose expression was induced by the pro-differentiation agent all-trans retinoic acid (atRA). AtRA induced expression in the most differentiated cells compared with the SC fraction, suggesting that this subpopulation was less responsive to atRA. Small interfering RNA suppression of RARRES1 and LXN enhanced the SC properties of primary prostate cultures, as shown by a significant increase in their colony-forming ability. Expression of both RARRES1 and LXN was co-ordinately repressed by DNA methylation in prostate cancer cell lines and inhibition of RARRES1 and LXN increased the invasive capacity of primary prostate cultures, which also fully rescued an inhibitory effect induced by atRA. Moreover, we showed that RARRES1 and LXN reside within different sub-cellular compartments, providing evidence that RARRES1 is not a plasma membrane protein as previously supposed but is located primarily in the endoplasmic reticulum; whereas LXN was detected in the nucleus of prostate epithelial cells. Thus, LXN and RARRES1 are potential tumour suppressor genes, which are co-ordinately regulated, SC-silenced genes functioning to suppress invasion and colony-forming ability of prostate cancer cells; yet the proteins reside within different sub-cellular compartments.
Similar content being viewed by others
Introduction
Retinoic acid receptor responder 1 (RARRES1) or tazarotene-induced gene 1 was initially identified as the most upregulated gene after retinoic acid receptor (RAR)-β/γ-specific retinoid treatment of skin raft cultures.1 The RARRES1 gene lies adjacent to Latexin (LXN), on chromosome 3q25. LXN was first identified as a marker of neurons in the lateral neocortex of rat brain2, 3 and shares a focal 30% amino-acid sequence homology with RARRES1 (Supplementary Figure 1). The principal differences are in the existence of a putative N-terminal transmembrane domain in RARRES1.
A functional role for suppression of RARRES1 expression in cancer cell lines was confirmed by studies on a metastatic prostate cancer (CaP) cell line (PC3M) where it acted as an invasion suppressor.4 Similar activity has been seen in Epstein–Barr virus-infected nasopharyngeal carcinoma5 and breast carcinoma SUM-159 cells.6 However, no study has determined the effect of RARRES1 on primary cancer cell invasion and more importantly it is not known whether LXN can also suppress invasion. A differentiation-associated function for RARRES1 has been presumed, as its expression is closely associated with differentiation of colorectal adenocarcinoma cells,7 adult adipose-derived mesenchymal stem cells (SCs),8 endometrial tumour cells and colon cancer cell lines,9 although no previous study has directly linked RARRES1 expression with SC differentiation. In contrast, LXN has been identified as a quantitative trait gene responsible for negative regulation of haematopoietic SC (HSC) numbers in mice.10 LXN-deficient HSCs have been shown to possess an enhanced colony-forming ability11 and modulation of LXN expression in gastric carcinoma cell lines affected colony-forming ability in a similar manner.12 However, it is not known whether LXN has a differentiation-associated function similar to RARRES1.
The precise mechanism of action for these two closely related genes has proven elusive. LXN has been described as the only known endogenous carboxypeptidase inhibitor13 and the structure of LXN in complex with carboxypeptidase A4 (CPA4) has been solved.14 As there is primary amino-acid sequence identity in the CPA4-binding site between LXN and RARRES1, both proteins were predicted to function as carboxypeptidase inhibitors.15
Expression of RARRES1 is low in cancer cell lines, but can be induced by the vitamin D3 pro-differentiation agent 1,25-dihydroxyvitamin D in the human colon carcinoma cell line Caco-2.16 RARRES1 expression is repressed by DNA methylation in a number of cancers, including CaP,6, 17, 18, 19, 20, 21, 22, 23, 24 and LXN expression has recently been shown to be controlled by promoter hypermethylation in cancer, including CaP.12, 25, 26, 27
The intracellular localization of RARRES1 has not been defined. Sequence analysis initially predicted RARRES1 to be a transmembrane protein with a small N-terminal intracellular region, a single membrane-spanning hydrophobic region and a long C-terminal extracellular region.4 More recently, RARRES1 was proposed to be a type III transmembrane (plasma membrane) protein based purely on its N-glycosylation status, with its long C-terminal domain now facing the cytoplasm.28 The intracellular localization of LXN in human cells is unknown, but an early study in rat mast cells indicated a cytoplasmic granular distribution that was not associated with lysosomal structures.29
We show here that RARRES1 and LXN are both highly expressed differentiation-associated genes that are not expressed in SC enriched from human primary prostate epithelial cultures. Transcription of both RARRES1 and LXN was co-ordinately regulated by DNA methylation in malignant cell lines but surprisingly not in primary basal cultures. The expression of both RARRES1 and LXN was however further induced by retinoic acid (RA) in these enriched basal prostate cultures. We also demonstrate that both RARRES1 and LXN not only act to suppress invasion in primary epithelial cultures, but also decrease the SC-initiated colony-forming efficiency of primary cultures, in keeping with a SC ‘suppressive’ function. We have also firmly established the intracellular localizations of these two important regulatory proteins: LXN shows a predominantly nuclear expression, whereas RARRES1 is located within the endoplasmic reticular lumen, thus influencing their previously proposed functions.
Results
Expression of RARRES1 and LXN is induced by RA and correlates with DNA methylation in prostate epithelial cell lines
As RARRES1 is known to be upregulated by RA in skin,1 and the homology and adjacent location of RARRES1 and LXN suggest that their expression may be regulated by the same mechanism (Supplementary Figure S1), the regulation of their expression by RA was investigated in prostate epithelial cell lines. Putative binding sites for RA responsive elements (RAREs) are present within 5 kB upstream of the RARRES1 (1 site) and LXN (2 sites) transcription start sites (Figure 1a). Basal (PNT1a, BPH-1, RC165 and Bob) cell lines and the luminal LNCaP cell line were treated with 500 nM atRA over a time course, and mRNA expression of RARRES1 (Figure 1b) and LXN (Figure 1c) quantified. Although three out of four and all four basal cell lines showed a significant increase in RARRES1 and LXN expression respectively after 24 h, the LNCaP cell line showed a modest decrease in expression.
The expression of RARRES1 and LXN is induced by retinoic acid and correlates with DNA methylation in prostate epithelial cell lines. (a) Depiction of RAREs found in a 5-kB portion of the RARRES1 and LXN promoters using the JASPAR database. (b) qRT-PCR expression data quantifying the expression of RARRES1 and (c) LXN after treatment of basal (PNT2-C2, BPH-1, RC-165n/hTERT and Bob) and luminal (LNCaP) prostate cell lines with 500 nM atRA over a time course. In advance of atRA treatments, cell lines were grown for 24 h in charcoal-stripped serum-supplemented media. Expression relative to an RPLPO control gene; n=3 technical replicates; error bars expressed as range of the mean. (d) The percentage promoter methylation of RARRES1 and LXN in benign and CaP cell lines (Bars=single CpG sites; black line=average of individual CpG sites; error bars expressed as standard deviation of n=3 technical replicates; EpiTect PCR unmethylated and methylated control DNA were used as negative and positive controls, respectively (Qiagen)). (e) mRNA expression of RARRES1 and LXN was analysed by qRT-PCR in benign (PNT2-C2) and CaP (P4E6, PC3 and LNCaP) cell lines after treatment with 1 μM 5-aza-2′-deoxycytidine (AZA) for 96 h. Expression relative to a glyceraldehyde 3-phosphate dehydrogenase (GAPDH) control gene and plotted on a log10 scale; UD: expression undetectable after 40 cycles; error bars expressed as standard deviation of n=2 biological replicates. (f) Dot plot showing the correlation between mRNA expression and DNA methylation of RARRES1 and LXN. Statistical significance values were measured by the Student’s t-test (*P<0.05, **P<0.01, ***P<0.001). DMSO, dimethyl sulfoxide; TSS, transcription start site.
Pyrosequencing methylation analysis performed at the positions indicated in Supplementary Figure S2 showed that the RARRES1 promoter was significantly hypermethylated in the cancer cell lines P4E6 (6%), PC3 (11%) and LNCaP (73%), but not in the non-malignant PNT2-C2 cell line (Figure 1d). LXN showed significant hypermethylation in not only the cancer cell lines P4E6 (82%) and LNCaP (64%), but also at generally lower levels in the benign cell line PNT2-C2 (27%). When prostate cell lines were treated with 1 μM of the demethylating agent 5-Aza-2′-deoxycytidine for 96 h to determine whether methylation was functionally related to the expression of RARRES1 and LXN as measured by quantitative reverse transcription–PCR (qRT–PCR; Figure 1e), RARRES1 expression was significantly induced in cancer cell lines, but not in the benign cell line PNT2-C2. More importantly, LXN expression was also significantly induced in the cancer cell lines P4E6 and LNCaP but not in malignant PC3 cells, which constitutively overexpressed LXN. A direct comparison of the expression of RARRES1 and LXN with the extent of DNA methylation confirmed that hypermethylation of RARRES1 and LXN correlated with a downregulation of expression; cell lines expressing high levels of mRNA had low levels of promoter methylation, but cell lines expressing the lowest levels of mRNA hypermethylated both RARRES1 and LXN (Figure 1f).
RARRES1 and LXN are expressed at low levels in SCs enriched from primary prostate epithelial cultures
Analysis of Affymetrix gene-expression array data30 identified RARRES1 and LXN as two of the most differentially expressed genes in prostate basal cultures, in a comparison between SC and more differentiated committed basal (CB) cells (Supplementary Figure S3). This differential expression was next validated by qRT-PCR in SC, transit amplifying (TA) cells which are the product of asymmetric division upon self renewal of SC and CB cell populations isolated from primary prostate epithelial cultures (see Materials and methods). The primary epithelial cultures described are of an undifferentiated basal phenotype and do not contain a terminally differentiated luminal cell population. RARRES1 showed significantly lower levels of expression in SC from benign prostatic hyperplasia (BPH) samples than more differentiated TA or CB cells. CaP samples showed the same trend in RARRES1 expression; however, the overall expression levels observed in CaP samples were as expected, much lower than in BPH, suggesting that RARRES1 is downregulated in prostate basal cancer cells (Figure 2a). LXN also showed significantly lower levels of expression in SC compared with TA and CB cells (Figure 2b).
RARRES1 and LXN are expressed at low levels in prostate epithelial cultures enriched for stem cells. Absolute gene expression of (a) RARRES1 and (b) LXN in epithelial cell cultures enriched for SC, TA and CB cells derived from BPH (n=5) and CaP (n=6) relative to a standard curve using serial dilutions of RARRES1 (pReceiver-M45) or LXN (pEZ-M06) cDNA expression vectors. Average expression denoted by a horizontal line. Statistical significance values were measured by the Mann–Whitney test (*P<0.05, **P<0.01). (c) Immunofluorescence images of RARRES1 and (d) LXN expression in primary prostate epithelial cultures enriched for SC, TA and CB cells derived from BPH and CaP. Cells were counterstained with 4′-6-diamidino-2-phenylindole to enable nuclear visualization. White scale bar represents 10 μm. (e) Antibody controls used Rabbit IgG instead of primary antibody and secondary antibody only. Filled shapes and symbols in a, b indicate primary samples from different patients.
The mRNA expression patterns were also confirmed by immunofluorescence analysis of RARRES1 and LXN in SC, TA and CB populations from primary prostate epithelial cultures, that is, expression of RARRES1 (Figure 2c) and LXN (Figure 2d) was low in SCs and increased during differentiation to TA and CB cells. In these enriched primary cultures, native RARRES1 expression demonstrated an intense cytoplasmic staining, whereas LXN expression was localized to the nucleus.
RARRES1 and LXN expression is induced by RA in enriched subpopulations from primary basal prostate cultures
In contrast to the data from cell lines, when DNA hypermethylation was analysed in SC, TA and CB subpopulations from primary prostate epithelial cultures derived from BPH or CaP patient samples, no significant hypermethylation was seen in any of the three subpopulations. The average methylation levels of RARRES1 and LXN were less than 10% in all samples tested (Supplementary Figure S4). However, when primary epithelial cultures were treated with various concentrations of atRA over a time course and expression measured by qRT-PCR (Supplementary Figure S5), the expression of RARRES1 and LXN showed dose and time responses to atRA. Expression of both RARRES1 and LXN increased 4-fold after 24 h and up to 500-fold for RARRES1 and 200-fold for LXN after 96 h treatment. As 100 nM atRA for 72 h showed the greatest induction of RARRES1 and LXN expression without phenotypically affecting the cells, primary epithelial cultures derived from BPH and CaP were next treated with this concentration of atRA, and the expression of RARRES1 (Figure 3a) and LXN (Figure 3b) quantified by qRT-PCR in SC, TA and CB populations. After atRA treatment, the expression of RARRES1 and LXN was significantly increased in all cell populations from BPH and CaP. The more differentiated TA and CB cells showed the greatest stimulation (up to 800-fold) compared with the SC population (maximum 20-fold).
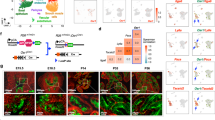
RARRES1 and LXN are induced by retinoic acid in primary prostate cultures. (a) qRT-PCR expression data quantifying the relative expression of RARRES1 and (b) LXN expression after treatment of primary epithelial cell cultures (enriched for SC, TA and CB cells) derived from BPH (n=3) or CaP (n=3) with 100 nM atRA for 72 h. All expression values are relative to an RPLPO endogenous control. Within each subpopulation, expression of all dimethyl sulfoxide (DMSO)-treated and atRA-treated samples was normalized to the DMSO-treated sample showing the lowest expression of RARRES1 or LXN (set at 1). Average expression denoted by a horizontal line. (c) RAR-α, -β and -γ expression was detected by immunofluorescence in primary epithelial cultures derived from CaP, treated with 500 nM atRA for 24 h or a DMSO control. Cells were counterstained with 4′-6-diamidino-2-phenylindole to enable nuclear visualization. White scale bar represents 10 μm. Antibody controls using rabbit or mouse IgG instead of primary antibody and secondary antibody only. (d) Luciferase activity in primary prostate epithelial cell cultures derived from BPH and (e) CaP transfected with a RARE reporter plasmid, and 12 h after transfection, cells were treated with various concentrations of atRA for a further 24 h. Luciferase activity was normalized to the values of the cells transfected with a negative control plasmid (lacked RARE regulatory elements). Statistical significance values were measured by the Student’s t-test (*P<0.05, **P<0.01, ***P<0.001). Filled shapes in a,b indicate primary samples from different patients.
As RA responsiveness in basal prostate cells has not been studied previously, expression of the three receptors and responsiveness to atRA with a synthetic promoter was also measured. All three RAR isoforms were detected; RAR-α showed predominant nuclear expression, RAR-β showed mostly cytoplasmic expression and RAR-γ showed predominant cytoplasmic with some nuclear expression. After atRA treatment, there was no difference in the localization of any of the three RAR isoforms, as RARs are within the nucleus and attached to DNA even in the absence of RA ligand.31 Transfection of primary cells with a reporter plasmid containing a tandem array of RAREs to regulate the expression of the firefly luciferase gene (Figures 3d and e) confirmed that primary cells from both BPH and CaP cells could sustain atRA-dependent gene expression (up to 25- and 15-fold increase, respectively).
RARRES1 is located in the endoplasmic reticulum (ER), and LXN is located in the nucleus of prostate epithelial cells
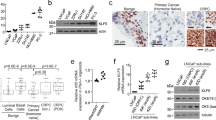
The data presented in Figures 2c and d located native RARRES1 to the cell cytoplasm, and perhaps surprisingly, LXN was predominantly nuclear. To verify these locations, we transfected fusion proteins with haemagglutinin (HA) epitope tags into non-expressing prostate epithelial cells (LNCaP and PC3). As shown in Figure 4a, transfection of a synthetic RARRES1–HA fusion did not result in co-localization with the plasma membrane marker α1-Na/K-ATPase, but did co-stain with the ER marker protein disulphide isomerase (Figures 4b and c), suggesting that RARRES1 was in fact located on the ER lumen membrane. In contrast to RARRES1, LXN-HA was located in the nucleus of LNCaP cells, even in the absence of a canonical nuclear localization signal (Figure 4e). To confirm that RARRES1 was indeed not located in the plasma membrane of cells, cellular fractionation was performed in the PC3 cell line after transfection of the RARRES1-HA construct (Figure 4f). In accordance with the immunofluorescence (IF) data, RARRES1 expression was not detected in the plasma membrane fraction but in the cytoplasmic fraction, which contains ER membranes.
RARRES1 is located in the ER, and LXN is located in the nucleus of prostate epithelial cell lines. Confocal immunofluorescence images depicting the location of HA-tagged RARRES1 in LNCaP (a, b) and PC3 (c) cells and LXN in LNCaP (d) cells 24 h after transfection. Cells we co-stained with anti-HA tag and (a) anti-α1-Na+/K+-ATPase (plasma membrane marker), (b, c) anti-protein disulphide isomerize (PDI; ER marker), (d) anti-LXN antibodies. Cells were counterstained with 4′-6-diamidino-2-phenylindole (DAPI) to enable nuclear visualization. White scale bar represents 10 μm. (e) Antibody controls using rabbit or mouse IgG instead of primary antibody and secondary antibody only. (f) Western blot data showing protein levels of RARRES1-HA (33 kDa) in unbroken cell fraction (UBC), wash fraction (Wash 1) and plasma membrane fraction (PM) of PC3 cells transfected with HA-tagged RARRES1 or reagent-only control (Mock) for 24 h before lysing the cells. Blots were probed with anti-HA (Santa Cruz) primary antibodies and horseradish peroxidase-linked secondary antibodies (Cell Signalling). Antibodies against α1-Na/K-ATPase (AbCam) and β-actin (Sigma) were used as plasma membrane and cytoplasmic internal controls, respectively.
RARRES1 and LXN have opposing effects on the migration of prostate epithelial cell lines
To investigate the functional importance of these homologous genes, wound-healing cell migration assays were initially performed after knockdown or overexpression of RARRES1 in PNT1a cells (Supplementary Figure S6). Compared with the scrambled control, cell migration significantly increased from 56 to 80% with RARRES1 small interfering RNA (siRNA), but was significantly suppressed to 21% after LXN knockdown (Figure 5a). As expected, compared with the vector control, migration decreased after RARRES1 overexpression from 68 to 51% and increased to 88% after LXN overexpression in the same cell line (Figure 5b).
RARRES1 and LXN regulate cell migration, invasion and colony formation in prostate epithelial cell lines and primary cultures. (a) Wound closure assay data showing the motility of PNT1a cells transfected with 10 nM scrambled, RARRES1 or LXN siRNA or (b) empty vector, RARRES1-HA or LXN-HA transfection vectors, 72 h after transfection and 18 h after wounding. (error bars expressed at the standard deviation of n=3 biological replicates). (c) Matrigel invasion assay data showing the relative number of invasive PNT1a cells transfected with 10 nM scrambled (scr), RARRES1 or LXN siRNA (knockdown) or (d) LNCaP cells transfected with vector only, RARRES1-HA or LXN-HA transfection vectors (overexpression). Epithelial cells from the PNT1a (an immortalized cell line from benign prostate; invasion set at 1) and MDA-MB-231 (metastatic breast cancer cell line) cell lines were used as negative and positive controls, respectively (error bars expressed as standard deviation of n=3 technical replicates). (e) Matrigel invasion assay data showing the relative invasion of primary prostate BPH (n=4) and (f) CaP (n=5) epithelial cultures transfected with 50 nM scrambled, RARRES1 or LXN siRNA. All data are 72 h after transfection (least invasive media sample set at 1). (g) Matrigel invasion assay data showing the percentage invasion/migration of a primary prostate BPH epithelial culture initially treated with 100 nM atRA for 18 h, and then transfected with 50 nM scrambled, RARRES1 or LXN siRNA for a further 72 h (PNT1a invasion set at 1). (h) Relative colony-forming assay recovery data after transfection of primary BPH (n=3) and (i) CaP (n=4) epithelial cultures with 50 nM scrambled, RARRES1 or LXN siRNA or untransfected (media; all media samples set at 1) for 96 h. Statistical significance values were measured by the Student’s t-test (*P<0.05, **P<0.01, ***P<0.001). The percentage colony-forming efficiency (CFE) was calculated by dividing the number of colonies by the number of cells plated and relative CFE was calculated by setting the CFE for each media sample at 100%. Filled shapes in e,f,h,i indicate primary samples from different patients.
RARRES1 and LXN regulate invasion of primary prostate epithelial cultures
As RARRES1 and LXN are similarly suppressed by DNA methylation in CaP cell lines, and RARRES1 was previously shown to be a metastasis suppressor, the effect of RARRES1 and LXN on invasion was determined. Compared with the scrambled siRNA, the invasion of relatively non-invasive PNT1a cells significantly increased by almost twofold after RARRES1 knockdown and increased threefold with LXN siRNA (Figure 5c). The reciprocal experiment was performed in LNCaP cells, which show the lowest expression of RARRES1 and LXN (Figure 5d). Compared with the vector control, after both RARRES1 and LXN overexpression, invasion was decreased. However, the overall biological effect of LXN was shown to be predominantly due to a difference in the number of motile cells after modulation of its expression (Supplementary Figure S7). To determine whether RARRES1 and LXN also regulate the invasion and motility of primary prostate epithelial cultures, Matrigel invasion assays after siRNA knockdown were next performed (Supplementary Figure S8). Compared with the scrambled control, the average invasion of primary epithelial BPH cultures showed a highly significant increase after siRNA knockdown of RARRES1 and LXN (Figure 5e). Moreover, the invasive capacity of CaP cultures also showed a highly significant increase after RARRES1 and LXN knockdown (Figure 5f). In contrast to cell lines, LXN did not exert its effects on invasion through inhibition of cell motility (Supplementary Figure S9).
RA suppresses invasion of primary prostate epithelial cultures
To assess the effect of RA on invasion, invasion assays were performed after atRA treatment of primary prostate cultures. A concentration of 100 nM atRA treatment dramatically decreased the average invasion of primary prostate cultures compared with the dimethyl sulfoxide vehicle control (Figure 5g). Furthermore, the invasive capacity of primary cells after atRA treatment was fully rescued by the loss of RARRES1 and LXN expressions (Figure 5g).
RARRES1 and LXN regulate colony formation of primary prostate epithelial cultures
To determine whether RARRES1 and LXN knockdown affected SC functions, such as self-renewal of primary prostate epithelial cultures, colony-forming assays were performed. Compared with the scrambled siRNA control, the average of the relative colony-forming efficiency of BPH cultures was significantly increased (>2-fold; Figure 5h), and CaP cultures significantly increased by almost twofold after RARRES1 knockdown (Figure 5i). Similarly, after LXN knockdown, compared with the scrambled control, the relative colony-forming efficiency of BPH cultures also significantly doubled, and CaP cultures significantly increased by almost twofold.
RARRES1 and LXN do not function through CPA4
Sequence alignment analysis showed that the C-terminal inhibitory loop from LXN, which protrudes into the CPA4 active site is fully conserved in RARRES1, suggesting that RARRES1 may also function through CPA4 (Supplementary Figure S1c). To determine whether RARRES1 and LXN were able to bind to CPA4 in the cell, immunoprecipitation analysis was performed on transfected epitope-tagged RARRES1 and LXN (Supplementary Figure S10). The results showed that neither LXN nor RARRES1 was pulled down with CPA4 in LNCaP cells.
Discussion
This is the first study to show that RARRES1 and LXN are two differentiation-associated genes that are highly significantly downregulated in prostate SCs and whose expression increases through differentiation, suggesting that tight regulation of their expression within the prostate hierarchy is required. RA is well established as a differentiation-inducing agent and its action on the expression of both RARRES1 and LXN was tested. Our results clearly demonstrate that RARRES1 is induced by RA in the prostate; moreover, we identify LXN as a novel RA-responsive gene. This indicates that these two genes not only have homologies in their sequences, but also share important regulatory mechanisms.
RA responsiveness was first suggested by the presence of RAREs upstream of the RARRES1 and LXN transcription start sites, and indeed our results show that both RARRES1 and LXN were induced by atRA in basal prostate cells. Moreover, after atRA treatment of primary prostate basal cultures enriched for SC, TA and CB, the expression of both RARRES1 and LXN was significantly induced in each subpopulation. The differentiated TA and CB cells were more responsive to atRA and consequently induced RARRES1 and LXN expression to a greater magnitude than in the SC fraction. These results correlate with previous studies, which showed that RA promoted differentiation of SCs from a range of different tissues, including human HSC,32, 33 mouse embryonic SC,34 rabbit bone marrow-derived mesenchymal SC35 and human breast cancer SC.36 In the converse experiment, inhibition of retinoid signalling pathways has been shown to induce the expansion of human HSC.37 Prostate SCs also demonstrate high aldehyde dehydrogenase (ALDH) activity38 suggesting that they possess the ability to convert vitamin A to RA. However, as our results show lower induction of RARRES1 and LXN in the SC population, we hypothesise that the SC may be less responsive to atRA, and that the RA ligand produced acts in a paracrine signalling fashion by promoting RA-dependent expression in neighbouring differentiated cells more than in the SC itself. Retinoid treatment can also stimulate both prostate epithelial differentiation and growth;39, 40 whereas squamous metaplasia develops in RAR-γ knockout mice41 and preneoplastic lesions develop after RXR-α inactivation in the prostatic epithelium.42 All these phenomena are consistent with an effect of RA on a tumour-initiating cell or cancer SC. Depending on the stage of prostate development, RA can either positively43 or negatively affect prostate formation and gland development.44 Given the inhibitory effects on adult stem and amplifying cells, both RARRES1 and LXN could have a role in embryonic morphogenesis.
Co-ordinate regulation of RARRES1 and LXN is not only carried out by RARs, but also by DNA methylation. Both genes were hypermethylated in CaP cell lines, corroborating previous findings.17, 18, 27 Surprisingly, RA-induced overexpression of RARRES1 and LXN was absent in the LNCaP cell line, which can indeed sustain RA-dependent gene expression,45 suggesting that the much higher levels of DNA methylation of these genes in this cell line could block the RAR activity in this genomic region.46
We provide evidence of a significant decrease in RARRES1 expression in primary CaP basal cultures compared with BPH, but this repression was not due to DNA methylation. RARRES1 expression is consistently repressed by DNA methylation in CaP tissues,18, 22, 27, 47 which are composed of >99% luminal cells.48 This would suggest that gene silencing independent of DNA methylation occurs in basal cancer cells. In the context of a cancer hierarchy, where basal cancer cells generate aberrant luminal cancer cells, this initial silencing could be a pre-requisite to DNA hypermethylation established in luminal cancer cells. As shown in other studies,49, 50, 51, 52 this phenomenon can be driven by Polycomb complexes, which mark genes with repressive histone modifications that are much more likely to be hypermethylated in cancer.
AtRA treatment reduced the invasive capacity of the prostate cultures, which agrees with a number of studies showing that retinoids have a role in suppressing invasion in vitro and metastasis in vivo.53, 54, 55, 56, 57, 58, 59 We propose that this was due to atRA treatment shifting the cultures to a more differentiated, and therefore less invasive phenotype, which expresses high levels of RARRES1 and LXN. Similarly, studies in breast cancer have shown that only the most primitive CD44+CD24− breast cancer SC displayed an enhanced invasive capacity.60 More importantly, a lack of expression of both RARRES1 and LXN increased the invasive capacity of primary prostate cultures, and fully rescued the inhibitory effect of atRA on invasion, and inhibition of RARRES1 and LXN enhanced the SC properties of primary prostate cultures as shown by a significant increase in their colony-forming ability. These novel functional results support a role for both RARRES1 and LXN as newly identified controllers of human SC differentiation.
We also provide evidence that RARRES1 is not a plasma membrane protein, as previously supposed, but is co-localized with an ER lumen marker. This is also the first study to provide a localization for LXN, which conversely resided within the nucleus. This difference in intracellular location of the two homologues suggests that they may have differing functions, despite possessing correlating expression patterns and regulatory mechanisms. A recent study28 showed that RARRES1 was able to interact with the cytosolic carboxypeptidase AGBL2 in the HEK 293 cell line, and an interaction between LXN and CPA4 has been described,14 however, we showed that neither LXN nor RARRES1 interacted with CPA4 in LNCaP cells. Taking the localization data into account, it is possible for RARRES1 but it would be less likely for nuclear LXN, to form a complex with both AGBL2 and CPA4 in prostate cells, which are primarily cytoplasmic and secreted proteins respectively. The differing localization patterns and potentially different interaction partners could account for the contrasting functions of the two homologues. Indeed, this seems to be the case in prostate cell lines, where we showed that both genes had opposing functions: LXN promoted only cell migration, whereas RARRES1 repressed cell migration and invasion. Recently, RARRES1 was identified as the major secreted product, probably lacking the N-terminal membrane anchor, from human plexiform neurofibroma-derived Schwann cells (but not normal Schwann cells).61 This provides further evidence for its primary location in the ER.
We conclude that RARRES1 and LXN are two co-ordinately regulated genes whose expression is repressed in CaP. This data suggests that RA might have an important role in prostate differentiation by inducing the expression of these novel differentiation-associated SC-silenced genes, whereas reducing the invasive capacity of CaP cultures. The differing cellular localizations of both genes could account for their contrasting functions; RARRES1 can probably function to suppress invasion by binding to AGBL2 but the function of LXN may be carried out via interactions with different protein partners. A loss of RARRES1 expression in cancer progression suggests that there would be an increase in the amount of detyrosinated tubulin, whose role in tumour invasion has been proposed recently, along with the suggestion that high RARRES1 levels have a role in SC differentiation and the epithelial-to-mesenchymal transition.62 Elucidating the protein networks that allow these highly similar genes to function in contrasting ways will provide valuable insights into the complex regulation of SC differentiation as well as invasion and metastasis in CaP.
Materials and methods
Cell culture and treatments
A list of cell lines (Supplementary Table 1) and primary prostate tissues (Supplementary Table 2) used are provided as supplementary data. Tissues were disaggregated and epithelial cells isolated as described previously.63, 64, 65 Primary epithelial cultures were enriched for cell subpopulations based on the expression of α2β1-integrin by rapid collagen adhesion and CD133 by MACS selection as described previously:63, 64 (i) an undifferentiated population of cells with SC characteristics (SC; α2β1-integrinhighCD133+), (ii) transit amplifying progeny (TA; α2β1-integrinhighCD133−) and (iii) committed to differentiation committed basal cells (CB; α2β1-integrinlowCD133−). All cells were certified free of mycoplasma and genotyped using the ATCC-approved Powerplex 1.2 system (Promega, Madison, WI, USA) to ensure authenticity.
RNA extraction, cDNA synthesis and qRT-PCR
RNA was extracted from cells using the RNeasy mini kit (Qiagen, Hilden, Germany). Total RNA (50 ng/500 ng) was reverse transcribed using random hexamer primers (Invitrogen, Carlsbad, CA, USA) and Superscript III reverse transcriptase (Invitrogen). Quantitative real-time PCR was performed using TaqMan gene expression assays on the ABI Prism 7000 Sequence Detection System (Applied Biosystems, Foster City, CA, USA) or the SsoFast Probes Supermix on the CFX96 Real-Time PCR Detection System (Bio-Rad Laboratories, Hercules, CA, USA). Analyses were carried out using the delta–delta Ct method for relative quantification66 or standard curve method for absolute quantification.
Pyrosequencing methylation analysis
DNA was extracted from cells and tissues using the DNeasy Blood and Tissue Kit (Qiagen) and the QIAamp DNA micro kit (Qiagen) for small samples. DNA (50 ng/500 ng) was bisulphite converted using the EpiTect Bisulfite Kit (Qiagen). RARRES1 and LXN were amplified by PCR (Platinum TAQ, Invitrogen) with specific primers (see Supplementary Table 3) for regions within their CpG Island, and sequenced using the Pyromark Q24 pyrosequencer (Qiagen).
Immunofluorescence
Antibodies used were: RARRES1 (Santa Cruz Biotechnology, Santa Cruz, CA, USA), LXN (Sigma-Aldrich, St Louis, MO, USA), HA (Santa Cruz), RAR-α, -β, -γ (Santa Cruz), protein disulphide isomerase (AbCam, Cambridge, MA, USA), α1-Na/K-ATPase (AbCam) antibodies or rabbit IgG (Sigma) or mouse IgG (R&D Systems, Minneapolis, MN, USA) isotype negative control antibodies. Alexa Fluor 488 or 568 secondary antibodies (Invitrogen) were also used for detection.
Luciferase assay
Cignal RARE reporter (luc) kit plasmids (SABiosciences, Qiagen) were transfected into primary cultures using TransIT-Prostate transfection kit (Mirus Bio, Madison, WI, USA) as transfection reagent, following manufacturer’s protocol. Luciferase expression was measured using the Dual-Glo system (Promega, Madison, WI, USA) following manufacturer’s protocol using the Polarstar Optima micro-plate reader (BMG Labtech, Offenburg, Germany).
Cell fractionation and western blotting
Cells were sonicated using the Bioruptor system (Diagenode, Liège, Belgium). Unbroken cells were pelleted by centrifugation. To collect the intracellular membranes, the supernatant was incubated with ice-cold sodium carbonate (0.1 M, pH 11) for 1 h and ultracentrifuged at 100 000 × g for 1 h, two times. To collect the plasma membrane, the pellet was re-suspended in ice-cold Tris buffer and ultracentrifugated at 115 000 × g for 20 min. Western blot analysis was used to detect expression.
SiRNA knockdown
Cells were transfected with Silencer select (Applied Biosystems) siRNAs targeting RARRES1 (siRNA ID: s11812), LXN (siRNA ID: s230651) or negative control #1 using DharmaFECT 2 (Dharmacon, Lafayette, CO, USA) as transfection reagent for cell lines or oligofectamine (Invitrogen) as transfection reagent for primary cultures.
cDNA expression vector transfection
Cells were transfected with endofree (Qiagen) pReceiver-M45 RARRES1, pEZ-M06 LXN or pReceiver-M06 control vector with eGFP cDNA expression plasmids (GeneCopoeia, Rockville, MD, USA) using TransIT-2020 transfection reagent (Mirus Bio).
Cell migration assay
Cells were treated with 10 nM siRNA or cDNA expression vectors 24 h after plating and after incubation at 37 °C for 48 h, a wound was created using a 1-ml pipette tip. The width of the wound at 0 and 18 h was measured using Volocity software (Perkin Elmer, Waltham, MA, USA), the average (of 10 points) taken and the relative percentage wound closure at 18 h with respect to the starting wound size was calculated.
Matrigel invasion assay
The ability of epithelial cells to migrate through Matrigel was determined by the modified Boyden-chamber method.67 Briefly, cultures transfected with siRNA or expression vectors for 24 h, were plated onto Matrigel (BD Biosciences, Bedford, MA, USA) coated 8 μm filters. RPMI medium supplemented with 10% serum was used as a chemo-attractant. Cells invading through Matrigel and filters were counted 48 h after plating.
Clonogenic recovery assay
Primary cultures were treated with 50 nM siRNA for 96 h and plated at 100 cells per well in a 6-well collagen-I-coated plate (BD Biosciences), in triplicate with irradiated STO feeder cells. When colonies greater than 32 cells (five population doublings) started to emerge within 10–14 days the number of colonies were counted visually.
References
Nagpal S, Patel S, Asano AT, Johnson AT, Duvic M, Chandraratna RA . Tazarotene-induced gene 1 (TIG1), a novel retinoic acid receptor-responsive gene in skin. J Invest Dermatol 1996; 106: 269–274.
Hatanaka Y, Uratani Y, Takiguchi-Hayashi K, Omori A, Sato K, Miyamoto M et al. Intracortical regionality represented by specific transcription for a novel protein, latexin. Eur J Neurosci 1994; 6: 973–982.
Arimatsu Y . Latexin: a molecular marker for regional specification in the neocortex. Neurosci Res 1994; 20: 131–135.
Jing C, El-Ghany MA, Beesley C, Foster CS, Rudland PS, Smith P et al. Tazarotene-induced gene 1 (TIG1) expression in prostate carcinomas and its relationship to tumorigenicity. J Natl Cancer Inst 2002; 94: 482–490.
Kwok WK, Pang JC, Lo KW, Ng HK . Role of the RARRES1 gene in nasopharyngeal carcinoma. Cancer Genet Cytogenet 2009; 194: 58–64.
Peng Z, Shen R, Li YW, Teng KY, Shapiro CL, Lin HJ . Epigenetic repression of RARRES1 is mediated by methylation of a proximal promoter and a loss of CTCF binding. PLoS One 2012; 7: e36891.
Wu CC, Shyu RY, Chou JM, Jao SW, Chao PC, Kang JC et al. RARRES1 expression is significantly related to tumour differentiation and staging in colorectal adenocarcinoma. Eur J Cancer 2006; 42: 557–565.
Ohnishi S, Okabe K, Obata H, Otani K, Ishikane S, Ogino H et al. Involvement of tazarotene-induced gene 1 in proliferation and differentiation of human adipose tissue-derived mesenchymal stem cells. Cell Prolif 2009; 42: 309–316.
Takai N, Kawamata N, Walsh CS, Gery S, Desmond JC, Whittaker S et al. Discovery of epigenetically masked tumor suppressor genes in endometrial cancer. Mol Cancer Res 2005; 3: 261–269.
Liang Y, Jansen M, Aronow B, Geiger H, Van Zant G . The quantitative trait gene latexin influences the size of the hematopoietic stem cell population in mice. Nat Genet 2007; 39: 178–188.
Mitsunaga K, Kikuchi J, Wada T, Furukawa Y . Latexin regulates the abundance of multiple cellular proteins in hematopoietic stem cells. J Cell Physiol 2011; 227: 1138–1147.
Li Y, Basang Z, Ding H, Lu Z, Ning T, Wei H et al. Latexin expression is downregulated in human gastric carcinomas and exhibits tumor suppressor potential. BMC Cancer 2011; 11: 121.
Normant E, Martres MP, Schwartz JC, Gros C . Purification, cDNA cloning, functional expression, and characterization of a 26-kDa endogenous mammalian carboxypeptidase inhibitor. Proc Natl Acad Sci USA 1995; 92: 12225–12229.
Pallares I, Bonet R, Garcia-Castellanos R, Ventura S, Aviles FX, Vendrell J et al. Structure of human carboxypeptidase A4 with its endogenous protein inhibitor, latexin. Proc Natl Acad Sci USA 2005; 102: 3978–3983.
Aagaard A, Listwan P, Cowieson N, Huber T, Ravasi T, Wells CA et al. An inflammatory role for the mammalian carboxypeptidase inhibitor latexin: relationship to cystatins and the tumor suppressor TIG1. Structure 2005; 13: 309–317.
Wood RJ, Tchack L, Angelo G, Pratt RE, Sonna LA . DNA microarray analysis of vitamin D-induced gene expression in a human colon carcinoma cell line. Physiol Genomics 2004; 17: 122–129.
Youssef EM, Chen XQ, Higuchi E, Kondo Y, Garcia-Manero G, Lotan R et al. Hypermethylation and silencing of the putative tumor suppressor Tazarotene-induced gene 1 in human cancers. Cancer Res 2004; 64: 2411–2417.
Zhang J, Liu L, Pfeifer GP . Methylation of the retinoid response gene TIG1 in prostate cancer correlates with methylation of the retinoic acid receptor beta gene. Oncogene 2004; 23: 2241–2249.
Mizuiri H, Yoshida K, Toge T, Oue N, Aung PP, Noguchi T et al. DNA methylation of genes linked to retinoid signaling in squamous cell carcinoma of the esophagus: DNA methylation of CRBP1 and TIG1 is associated with tumor stage. Cancer Sci 2005; 96: 571–577.
Kwong J, Lo KW, Chow LS, Chan FL, To KF, Huang DP . Silencing of the retinoid response gene TIG1 by promoter hypermethylation in nasopharyngeal carcinoma. Int J Cancer 2005; 113: 386–392.
Yanatatsaneejit P, Chalermchai T, Kerekhanjanarong V, Shotelersuk K, Supiyaphun P, Mutirangura A et al. Promoter hypermethylation of CCNA1, RARRES1, and HRASLS3 in nasopharyngeal carcinoma. Oral Oncol 2008; 44: 400–406.
Ellinger J, Bastian PJ, Jurgan T, Biermann K, Kahl P, Heukamp LC et al. CpG island hypermethylation at multiple gene sites in diagnosis and prognosis of prostate cancer. Urology 2008; 71: 161–167.
Tamura G, So K, Miyoshi H, Honda T, Nishizuka S, Motoyama T . Quantitative assessment of gene methylation in neoplastic and non-neoplastic gastric epithelia using methylation-specific DNA microarray. Pathol Int 2009; 59: 895–899.
Son MS, Kang MJ, Park HC, Chi SG, Kim YH . Expression and mutation analysis of TIG1 (tazarotene-induced gene 1) in human gastric cancer. Oncol Res 2009; 17: 571–580.
Anderton JA, Lindsey JC, Lusher ME, Gilbertson RJ, Bailey S, Ellison DW et al. Global analysis of the medulloblastoma epigenome identifies disease-subgroup-specific inactivation of COL1A2. Neuro Oncol 2008; 10: 981–994.
Muthusamy V, Duraisamy S, Bradbury CM, Hobbs C, Curley DP, Nelson B et al. Epigenetic silencing of novel tumor suppressors in malignant melanoma. Cancer Res 2006; 66: 11187–11193.
Kloth M, Goering W, Ribarska T, Arsov C, Sorensen KD, Schulz WA . The SNP rs6441224 influences transcriptional activity and prognostically relevant hypermethylation of RARRES1 in prostate cancer. Int J Cancer 2012; 131: E897–E904.
Sahab ZJ, Hall MD, Me Sung Y, Dakshanamurthy S, Ji Y, Kumar D et al. Tumor suppressor RARRES1 interacts with cytoplasmic carboxypeptidase AGBL2 to regulate the alpha-tubulin tyrosination cycle. Cancer Res 2011; 71: 1219–1228.
Uratani Y, Takiguchi-Hayashi K, Miyasaka N, Sato M, Jin M, Arimatsu Y . Latexin, a carboxypeptidase A inhibitor, is expressed in rat peritoneal mast cells and is associated with granular structures distinct from secretory granules and lysosomes. Biochem J. 2000; 346 (Pt 3): 817–826.
Birnie R, Bryce SD, Roome C, Dussupt V, Droop A, Lang SH et al. Gene expression profiling of human prostate cancer stem cells reveals a pro-inflammatory phenotype and the importance of extracellular matrix interactions. Genome Biol 2008; 9: R83.
Bastien J, Rochette-Egly C . Nuclear retinoid receptors and the transcription of retinoid-target genes. Gene 2004; 328: 1–16.
Sammons J, Ahmed N, Khokher MA, Hassan HT . Mechanisms mediating the inhibitory effect of all-trans retinoic acid on primitive hematopoietic stem cells in human long-term bone marrow culture. Stem Cells 2000; 18: 214–219.
Luo P, Wang A, Payne KJ, Peng H, Wang JG, Parrish YK et al. Intrinsic retinoic acid receptor alpha-cyclin-dependent kinase-activating kinase signaling involves coordination of the restricted proliferation and granulocytic differentiation of human hematopoietic stem cells. Stem Cells 2007; 25: 2628–2637.
Simandi Z, Balint BL, Poliska S, Ruhl R, Nagy L . Activation of retinoic acid receptor signaling coordinates lineage commitment of spontaneously differentiating mouse embryonic stem cells in embryoid bodies. FEBS Lett 2010; 584: 3123–3130.
Su ZY, Li Y, Zhao XL, Zhang M . All-trans retinoic acid promotes smooth muscle cell differentiation of rabbit bone marrow-derived mesenchymal stem cells. J Zhejiang Univ Sci B. 2010; 11: 489–496.
Ginestier C, Wicinski J, Cervera N, Monville F, Finetti P, Bertucci F et al. Retinoid signaling regulates breast cancer stem cell differentiation. Cell Cycle 2009; 8: 3297–3302.
Chute JP, Muramoto GG, Whitesides J, Colvin M, Safi R, Chao NJ et al. Inhibition of aldehyde dehydrogenase and retinoid signaling induces the expansion of human hematopoietic stem cells. Proc Natl Acad Sci USA 2006; 103: 11707–11712.
van den Hoogen C, van der Horst G, Cheung H, Buijs JT, Lippitt JM, Guzman-Ramirez N et al. High aldehyde dehydrogenase activity identifies tumor-initiating and metastasis-initiating cells in human prostate cancer. Cancer Res 2010; 70: 5163–5173.
Peehl DM, Wong ST, Stamey TA . Vitamin A regulates proliferation and differentiation of human prostatic epithelial cells. Prostate 1993; 23: 69–78.
Seo R, McGuire M, Chung M, Bushman W . Inhibition of prostate ductal morphogenesis by retinoic acid. J Urol 1997; 158 (3 Pt 1): 931–935.
Lohnes D, Mark M, Mendelsohn C, Dolle P, Decimo D, LeMeur M et al. Developmental roles of the retinoic acid receptors. J Steroid Biochem Mol Biol 1995; 53: 475–486.
Huang J, Powell WC, Khodavirdi AC, Wu J, Makita T, Cardiff RD et al. Prostatic intraepithelial neoplasia in mice with conditional disruption of the retinoid X receptor alpha allele in the prostate epithelium. Cancer Res 2002; 62: 4812–4819.
Vezina CM, Allgeier SH, Fritz WA, Moore RW, Strerath M, Bushman W et al. Retinoic acid induces prostatic bud formation. Dev Dyn 2008; 237: 1321–1333.
Aboseif SR, Dahiya R, Narayan P, Cunha GR . Effect of retinoic acid on prostatic development. Prostate 1997; 31: 161–167.
Rivera-Gonzalez GC, Droop AP, Rippon HJ, Tiemann K, Pellacani D, Georgopoulos LJ et al. Retinoic acid and androgen receptors combine to achieve tissue specific control of human prostatic transglutaminase expression: a novel regulatory network with broader significance. Nucleic Acids Res 2012; 40: 4825–4840.
Vaissiere T, Sawan C, Herceg Z . Epigenetic interplay between histone modifications and DNA methylation in gene silencing. Mutat Res 2008; 659: 40–48.
Tokumaru Y, Harden SV, Sun DI, Yamashita K, Epstein JI, Sidransky D . Optimal use of a panel of methylation markers with GSTP1 hypermethylation in the diagnosis of prostate adenocarcinoma. Clin Cancer Res 2004; 10: 5518–5522.
Grisanzio C, Signoretti S . p63 in prostate biology and pathology. J Cell Biochem 2008; 103: 1354–1368.
Gal-Yam EN, Egger G, Iniguez L, Holster H, Einarsson S, Zhang X et al. Frequent switching of Polycomb repressive marks and DNA hypermethylation in the PC3 prostate cancer cell line. Proc Natl Acad Sci USA 2008; 105: 12979–12984.
Ohm JE, McGarvey KM, Yu X, Cheng L, Schuebel KE, Cope L et al. A stem cell-like chromatin pattern may predispose tumor suppressor genes to DNA hypermethylation and heritable silencing. Nat Genet 2007; 39: 237–242.
Schlesinger Y, Straussman R, Keshet I, Farkash S, Hecht M, Zimmerman J et al. Polycomb-mediated methylation on Lys27 of histone H3 pre-marks genes for de novo methylation in cancer. Nat Genet 2007; 39: 232–236.
Widschwendter M, Fiegl H, Egle D, Mueller-Holzner E, Spizzo G, Marth C et al. Epigenetic stem cell signature in cancer. Nat Genet 2007; 39: 157–158.
Nwankwo JO . Anti-metastatic activities of all-trans retinoic acid, indole-3-carbinol and (+)-catechin in Dunning rat invasive prostate adenocarcinoma cells. Anticancer Res 2002; 22: 4129–4135.
Dahiya R, Park HD, Cusick J, Vessella RL, Fournier G, Narayan P . Inhibition of tumorigenic potential and prostate-specific antigen expression in LNCaP human prostate cancer cell line by 13-cis-retinoic acid. Int J Cancer 1994; 59: 126–132.
Webber MM, Waghray A . Urokinase-mediated extracellular matrix degradation by human prostatic carcinoma cells and its inhibition by retinoic acid. Clin Cancer Res 1995; 1: 755–761.
Lan L, Cui D, Luo Y, Shi BY, Deng LL, Zhang GY et al. Inhibitory effects of retinoic acid on invasiveness of human thyroid carcinoma cell lines in vitro. J Endocrinol Invest 2009; 32: 731–738.
Lotan R . Retinoids as modulators of tumor cells invasion and metastasis. Semin Cancer Biol 1991; 2: 197–208.
Messi E, Florian MC, Caccia C, Zanisi M, Maggi R . Retinoic acid reduces human neuroblastoma cell migration and invasiveness: effects on DCX, LIS1, neurofilaments-68 and vimentin expression. BMC Cancer 2008; 8: 30.
Nakajima M, Lotan D, Baig MM, Carralero RM, Wood WR, Hendrix MJ et al. Inhibition by retinoic acid of type IV collagenolysis and invasion through reconstituted basement membrane by metastatic rat mammary adenocarcinoma cells. Cancer Res 1989; 49: 1698–1706.
Sheridan C, Kishimoto H, Fuchs RK, Mehrotra S, Bhat-Nakshatri P, Turner CH et al. CD44+/CD24- breast cancer cells exhibit enhanced invasive properties: an early step necessary for metastasis. Breast Cancer Res 2006; 8: R59.
Chen HL, Seol H, Brown KJ, Gordish-Dressman H, Hill A, Gallo V et al. Secretome Survey of Human Plexiform Neurofibroma Derived Schwann Cells Reveals a Secreted form of the RARRES1 Protein. Int J Mol Sci 2012; 13: 9380–9399.
Whipple RA, Matrone MA, Cho EH, Balzer EM, Vitolo MI, Yoon JR et al. Epithelial-to-mesenchymal transition promotes tubulin detyrosination and microtentacles that enhance endothelial engagement. Cancer Res 2010; 70: 8127–8137.
Collins AT, Habib FK, Maitland NJ, Neal DE . Identification and isolation of human prostate epithelial stem cells based on alpha(2)beta(1)-integrin expression. J Cell Sci 2001; 114 (Pt 21): 3865–3872.
Richardson GD, Robson CN, Lang SH, Neal DE, Maitland NJ, Collins AT . CD133, a novel marker for human prostatic epithelial stem cells. J Cell Sci 2004; 117 (Pt 16): 3539–3545.
Collins AT, Berry PA, Hyde C, Stower MJ, Maitland NJ . Prospective identification of tumorigenic prostate cancer stem cells. Cancer Res 2005; 65: 10946–10951.
Schmittgen TD, Livak KJ . Analyzing real-time PCR data by the comparative C(T) method. Nat Protoc 2008; 3: 1101–1108.
Albini A, Iwamoto Y, Kleinman HK, Martin GR, Aaronson SA, Kozlowski JM et al. A rapid in vitro assay for quantitating the invasive potential of tumor cells. Cancer Res 1987; 47: 3239–3245.
Acknowledgements
We thank Dr Simon Hayward for provision of BPH-1 cells, Dr Johng Rhim for provision of RC-165N/hTERT cells and Dr David Hudson for provision of Bob cells. This research was funded by a programme grant from Yorkshire Cancer Research (Y256) and project funding from The Freemasons’ Grand Charity (Y257PA).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies this paper on the Oncogenesis website .
Supplementary information
Rights and permissions
This work is licensed under a Creative Commons Attribution-NonCommercial-NoDerivs 3.0 Unported License. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-nd/3.0/
About this article
Cite this article
Oldridge, E., Walker, H., Stower, M. et al. Retinoic acid represses invasion and stem cell phenotype by induction of the metastasis suppressors RARRES1 and LXN. Oncogenesis 2, e45 (2013). https://doi.org/10.1038/oncsis.2013.6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/oncsis.2013.6
Keywords
This article is cited by
-
Latexin deficiency attenuates adipocyte differentiation and protects mice against obesity and metabolic disorders induced by high-fat diet
Cell Death & Disease (2022)
-
Bromodomain-containing protein 4 (BRD4) as an epigenetic regulator of fatty acid metabolism genes and ferroptosis
Cell Death & Disease (2022)
-
Latexin deficiency in mice up-regulates inflammation and aggravates colitis through HECTD1/Rps3/NF-κB pathway
Scientific Reports (2020)
-
Expression of RARRES1 and AGBL2 and progression of conventional renal cell carcinoma
British Journal of Cancer (2020)
-
Cathepsin V Mediates the Tazarotene-induced Gene 1-induced Reduction in Invasion in Colorectal Cancer Cells
Cell Biochemistry and Biophysics (2020)