Abstract
High-grade gliomas (HGGs) include the most common and the most aggressive primary brain tumor of adults and children. Despite multimodality treatment, most high-grade gliomas eventually recur and are ultimately incurable. Several studies suggest that the initiation, progression, and recurrence of gliomas are driven, at least partly, by cancer stem-like cells. A defining characteristic of these cancer stem-like cells is their capacity to self-renew. We have identified a hypoxia-induced pathway that utilizes the Hypoxia Inducible Factor 1α (HIF-1α) transcription factor and the JAK1/2-STAT3 (Janus Kinase 1/2 - Signal Transducer and Activator of Transcription 3) axis to enhance the self-renewal of glioma stem-like cells. Hypoxia is a commonly found pathologic feature of HGGs. Under hypoxic conditions, HIF-1α levels are greatly increased in glioma stem-like cells. Increased HIF-1α activates the JAK1/2-STAT3 axis and enhances tumor stem-like cell self-renewal. Our data further demonstrate the importance of Vascular Endothelial Growth Factor (VEGF) secretion for this pathway of hypoxia-mediated self-renewal. Brefeldin A and EHT-1864, agents that significantly inhibit VEGF secretion, decreased stem cell self-renewal, inhibited tumor growth, and increased the survival of mice allografted with S100β-v-erbB/p53−/− glioma stem-like cells. These agents also inhibit the expression of a hypoxia gene expression signature that is associated with decreased survival of HGG patients. These findings suggest that targeting the secretion of extracellular, autocrine/paracrine mediators of glioma stem-like cell self-renewal could potentially contribute to the treatment of HGGs.
Similar content being viewed by others
Introduction
The cancer stem cell model proposes that cells within a tumor exhibiting the features of stem cells drive tumor development.1 Cancer cells expressing markers of normal stem cells and having the ability to self-renew have been identified in a variety of human cancers including high-grade gliomas (HGGs).2, 3, 4, 5, 6 Glioma-derived stem-like cells have been demonstrated to have potent tumorigenic capacity4, 5, 6 and display increased resistance to treatments such as radiation and chemotherapy.7, 8, 9 In addition, these stem-like cells have also been implicated in tumor recurrence.10, 11, 12 Successfully targeting this cell population could have significant implications for the future treatment of tumors like HGG, which despite optimal medical treatment, have a poor prognosis.10, 11, 12
Several studies suggest that the tumor microenvironment plays a key role in cancer stem cell biology.12, 13, 14, 15, 16, 17, 18, 19 Hypoxia, which is a defining feature of the HGG microenvironment,20, 21 has been shown to promote self-renewal of glioma stem-like cells,13, 16, 19 but to date little is known about the specific mechanisms driving hypoxia-mediated self-renewal in these tumors. Tumor hypoxia is thought to arise in solid tumors due to rapid tumor growth and aberrant blood vessel formation.22, 23 The presence of hypoxic tumor tissue has been shown to be a prognostic factor associated with advanced stages of malignancy and poor clinical outcome.24, 25, 26, 27 Important molecular and cellular effects of hypoxia are mediated by the hypoxia-inducible factor 1α (HIF-1α) which is a transcription factor that is stabilized in the absence of oxygen.28, 29 High levels of HIF-1α have been observed in a wide variety of human cancers30, 31 and are correlated with poor prognosis in HGG patients.25, 32 Research on HIF-1α activity to date has focused on its role in inducing angiogenesis, metabolic alterations, and other adaptive changes.28, 33, 34
We sought to examine the role of hypoxia in the self-renewal of glioma stem-like cells.13, 16, 19 Using cells from the S100β-v-erbB/p53−/− mouse model of spontaneous HGG,35 we discovered that hypoxia leads to increased HIF-1α expression resulting in enhanced signal transducer and activator of transcription 3 (STAT3)-mediated self-renewal. Janus Kinase (JAK) 1 and 2 were required for STAT3 activation in these glioma stem-like cells, as was Vascular Endothelial Growth Factor (VEGF). Our findings suggest that when glioma stem-like cells respond to hypoxia, HIF-1α enhances expression of secreted factors such as VEGF, which act in a paracrine/autocrine fashion to initiate a signaling pathway leading to the activation of the JAK/STAT axis to promote self-renewal.
Results
The increase in glioma stem cell self-renewal during hypoxia is dependent on HIF-1α and STAT3 phosphorylation
To study the effect of hypoxia on glioma-derived stem-like cells, we derived tumor sphere cultures (TSCs) from spontaneous HGGs arising in the S100β-v-erbB/p53−/−mouse model.35 Tumors arising from S100β-v-erbB/p53−/−animals have been reported to present the histopathologic features classic of HGGs including increased cellularity, endothelial proliferation, necrosis, and increased nuclear atypia35, 36(data not shown). We utilized TSCs for the propagation of glioma stem-like cells, as these conditions have been shown to maintain glioma-derived stem-like cell proliferation4, 5, 37 and to preserve more accurately than conventional serum-based monolayer cultures the phenotype and genotype of the primary glial tumors from which these cultured tumor cells are derived.38, 39 The S100β-v-erbB/p53−/−model is characterized by the spontaneous development of central nervous system (CNS) tumors (Supplementary Figure 1A) as the result of the transgenic expression of two of the most frequent mutations found in HGGs; namely, high-level expression of EGFR and loss of p53.40 Stem-like cells isolated from these CNS tumors grow as spheroids in culture (Supplementary Figure 1B), and they express classic stem cell markers (Supplementary Figure 1C).41 When grown as spheroids they also demonstrate defining functional properties of tumor stem-like cells such as self-renewal (Supplementary Figure 1D) and tumor initiating capacity (Supplementary Figure 1E).
We examined the effect of hypoxia on the self-renewal capacity of S100β-v-erbB/p53−/− glioma stem-like cells, as determined by assaying subsphere formation at limiting dilution and colony formation in soft agar (Figures 1a and b). We found that in cultures derived from two different tumors more spheroids arose when incubated under hypoxic conditions than when incubated under normoxic conditions as observed in two representative cultures, S100β-v-erbB/p53−/− TSC1 (TSC1) and S100β-v-erbB/p53−/− TSC2 (TSC2), in Figure 1a. Consistent with this observation, the number of colonies formed in soft agar when these two cell cultures were incubated under hypoxic conditions was significantly increased compared to the cells cultured in normoxic conditions (Figure 1b). These data provide evidence that hypoxia enhances self-renewal of S100β-v-erbB/p53−/− stem-like cells.
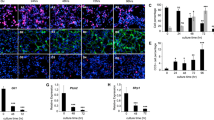
HIF-1α and STAT3 phosphorylation enhances glioma self-renewal during hypoxia. (a) Effect of hypoxia on TSC1 and TSC2 tumor subsphere formation (7 days). Data points represent the percentage of plated cells that grew as spheres in three independent experiments conducted in triplicate and are presented as the mean±s.d. (*P<0.05). (b) Effect of hypoxia on TSC1 (left) and TSC2 (right) colony formation in soft agar (7 days). Data points represent three independent experiments conducted in triplicate and are presented as the mean±s.d. (*P<0.01). (c) Western blot analysis of HIF-1α, phospho-STAT3, and total STAT3 in TSC1 incubated in hypoxia. (d) Western blot analysis of HIF-1α, phospho-STAT3, and total STAT3 in TSC2 incubated in hypoxia. (e) Effect of shRNA HIF-1α expression on TSC1 tumor subsphere formation in hypoxia (7 days). Data points represent the percentage of plated cells that grew as spheres in three independent experiments conducted in triplicate and are presented as the mean±s.d. (*P<0.05). (f) Effect HIF-1α shRNA expression on TSC1 colony formation in soft agar during hypoxia (7 days). Data points represent three independent experiments conducted in triplicate and are presented as the mean±s.d. (*P<0.01). (g) Effect of S3I-201 (100 μM) treatment on TSC1 tumor subsphere formation in hypoxia (7 days). Data points represent the percentage of plated cells that grew as spheres in three independent experiments conducted in triplicate and are presented as the mean±s.d. (*P<0.05). (h) Effect of S3I-201 (100 μM) treatment on TSC1 colony formation in soft agar during hypoxia (7 days). Data points represent three independent experiments conducted in triplicate and are presented as the mean±s.d. (*P<0.01).
We observed that the level of HIF-1α, the main mediator of the cellular response to hypoxia,28 increased during hypoxia (Figures 1c and d). We sought to determine whether this transcription factor regulated the enhanced self-renewal of S100β-v-erbB/p53−/− TSCs that occurs during hypoxia (Figures 1a and b). We utilized shRNAs to decrease HIF-1α expression (Supplementary Figures 2A-B). When HIF-1α expression was inhibited in hypoxia, the proportion of sphere forming cells was significantly reduced in a representative S100β-v-erbB/p53−/− cell line, TSC1 (Figure 1e), abrogating the enhanced subsphere forming capacity observed during hypoxia (Figure 1a). Similarly, inhibition of HIF-1α expression with shRNA eliminated the increase in colony formation associated with hypoxia in S100β-v-erbB/p53−/− TSCs (Figure 1f). These data suggest that HIF-1α contributes to the increase in self-renewal during hypoxia.
We found that phosphorylated STAT3, which has been reported to participate in the cellular response to hypoxia,42, 43, 44, 45, 46, 47 also increased in S100β-v-erbB/p53−/− TSCs during hypoxia (Figures 1c and d). We sought to determine whether STAT3 mediated the enhanced self-renewal of S100β-v-erbB/p53−/− TSCs observed during hypoxia (Figures 1a and b). To inhibit STAT3 activity, we utilized the small molecule inhibitor, S3I-201, which selectively binds the SH2 domain of STAT3 to inhibit STAT3 phosphorylation and thus its transcriptional activity (Supplementary Figure 2C–E).48 We observed that following STAT3 inhibition with S3I-201, the subsphere forming potential of S100β-v-erbB/p53−/− glioma stem-like cells was significantly decreased (Figure 1g), abrogating the enhanced subsphere formation associated with hypoxia (Figure 1a). Similarly, S100β-v-erbB/p53−/−colony formation in soft agar was markedly decreased in cells treated with S3I-201 (Figure 1h), and this inhibition was sufficient to eliminate the increase in colony formation associated with hypoxia (Figure 1b). These findings suggest that, during hypoxia, increased STAT3 phosphorylation mediates the increase in self-renewal observed in S100β-v-erbB/p53−/− TSCs.
HIF-1α mediated STAT3 phosphorylation enhances glioma stem cell self-renewal during hypoxia
We found that decreased HIF-1α expression in these TSCs was associated with decreased levels of phosphorylated STAT3 (Figure 2a). Using a STAT3-dependent luciferase reporter assay, we determined that STAT3-dependent transcription, which is typically increased during hypoxia,42, 43, 44, 45, 46, 47 was significantly decreased when HIF-1α expression was decreased in hypoxic S100β-v-erbB/p53−/− cells. (Supplementary Figure 2F). These data suggest that HIF-1α contributes to STAT3 activation that occurs in hypoxic conditions. To evaluate further the role of STAT3 activation during hypoxia, we sought to determine whether STAT3 activation could restore the self-renewal potential that had been reduced by inhibition of HIF-1α during hypoxia (Figures 1e and f). We co-transfected S100β-v-erbB/p53−/− cultures with HIF-1α shRNA and a constitutively activated form of STAT3 (STAT3C)49 (Supplementary Figure 2G). Hypoxic S100β-v-erbB/p53−/− cultures in which HIF-1α expression was decreased formed fewer spheres than control cells transfected with scrambled shRNA (Figure 2b); however, when STAT3C was expressed simultaneously with HIF-1α shRNA, S100β-v-erbB/p53−/− formed subspheres at a frequency indistinguishable from cells transfected with scrambled shRNA alone (Figure 2b). Consistent with this finding, we found that when HIF-1α expression was decreased, expression of STAT3C restored the enhanced hypoxia-induced self-renewal in S100β-v-erbB/p53−/− cultures as determined by colony formation in soft agar (Figure 2c). These data indicated that the role of HIF-1α in the self-renewal of S100β-v-erbB/p53−/− TSCs during hypoxia is at least in part mediated by STAT3 activation.
HIF-1α mediated STAT3 phosphorylation enhances glioma self-renewal. (a) Western blot analysis of HIF-1α, phospho-STAT3, and total STAT3 in TSC1 cells expressing scramble shRNA or HIF-1α shRNA and incubated in normoxia or hypoxia (16 h). (b) Rescue of TSC1 subsphere formation by STAT3C after inhibition of HIF-1α expression. Data points represent the percentage of plated cells that grew as spheres in three independent experiments conducted in triplicate and are presented as the mean±s.d. (*P<0.05). (c) Rescue of TSC1 colony formation in soft agar by STAT3C after inhibition of HIF-1α expression. Data points represent three independent experiments conducted in triplicate and are presented as the mean±s.d. (*P<0.05).
The Janus family of kinases are well-known mediators of the activation of STATs by phosphorylation.50 Given the relatively high expression of JAK1 and JAK2 among the Janus family of kinases in S100β-v-erbB/p53−/− cultures (Supplementary Figure 3A), we sought to confirm the expectation that JAK1 and JAK2 mediated the increase in STAT3 phosphorylation observed during hypoxia and thereby contributed to enhanced TSCs self-renewal. Hence, we treated S100β-v-erbB/p53−/− cultures with Ruxolitinib, a small molecule inhibitor of JAK1 and JAK2 (Supplementary Figure 3B–E). Immunoblotting revealed decreased levels of STAT3 phosphorylation in hypoxic S100β-v-erbB/p53−/− cells treated with increasing, sub-lethal doses of Ruxolitinib (Supplementary Figure 3B and C). Consistent with these observations, Ruxolitinib inhibited the hypoxia-induced increase in TSCs self-renewal as determined by subsphere formation and colony formation in soft agar (Supplementary Figure 3D and E). Similarly, we observed that direct inhibition of JAK1 and JAK2 mRNA expression with shRNA reduced the levels of phosphorylated STAT3 during hypoxia (Supplementary Figure 3F and G). Taken together, these data provide evidence that JAK1 and JAK2 mediate enhanced STAT3 phosphorylation and thus contribute to glioma self-renewal during hypoxia in this mouse model.
Factors released by hypoxic S100β-v-erbB/p53−/− glioma stem-like cells promote STAT3 phosphorylation and enhance self-renewal of glioma stem-like cells
While examining how HIF-1α regulated the activity of the JAK/STAT axis, we observed that incubating TSCs in media conditioned by these same S100β-v-erbB/p53−/− cells cultured under hypoxic conditions (HCM) resulted in a level of STAT3 activation and self-renewal that was much greater than that observed in the same cells cultured in media that had been conditioned by these cells in normoxia (NCM) (Figures 3a–c). This enhanced level of phosphorylated STAT3 following the incubation of S100β-v-erbB/p53−/− cells in HCM was easily detectable by immunoblotting (Figure 3a). Consistent with this observation, TSCs incubated in HCM formed subspheres at a significantly higher frequency than TSCs incubated in NCM (Figure 3b). Also, TSCs incubated in HCM produced a significantly higher number of colonies than cells incubated in NCM (Figure 3c). We used PCR to examine the expression of a number of genes encoding various known regulators of self-renewal in S100β-v-erbB/ p53−/− cultures and found that their expression was markedly increased in cells incubated in HCM when compared to these same cells incubated with fresh media (Figure 3d). Taken together, these results indicate that media conditioned by hypoxic S100β-v-erbB/p53−/− glioma stem-like cells can promote STAT3 phosphorylation and thus enhance glioma self-renewal.
Secreted factors from hypoxic S100β-v-erbB/p53−/− spheroid cells promote STAT3 phosphorylation and enhance self-renewal. (a) Western blot analysis of phospho-STAT3 and total STAT3 in TSC1 cells cultured in NCM, HCM, or growth medium supplemented with OSM (50 ng/ml) (4h). (b) Effect of incubation in NCM or HCM on TSC1 and TSC2 subsphere formation cultured in normoxia (7 days). Data points represent the percentage of plated cells that grew as spheres in three independent experiments conducted in triplicate and are presented as the mean±s.d. (*P<0.01). (c) Effect of incubation in NCM or HCM on TSC1 and TSC2 colony formation in soft agar cultured in normoxia (7 days). Data points represent three independent experiments conducted in triplicate and are presented as the mean±s.d. (*P<0.01). (d) Quantitative RT-PCR analysis of mRNA expression of an array of known regulators of self-renewal in TSC1 incubated in fresh media (M) or HCM (4 h). Data points are from a representative experiment conducted in triplicate and are presented as the mean±s.d. (*P<0.05).
VEGF secreted by hypoxic S100β-v-erbB/p53−/− glioma stem-like cells increases STAT3 phosphorylation and enhances glioma self-renewal
While studying the effect of HCM on the self-renewal of glioma stem-like cells, we discovered that the enhancement of subsphere formation and colony formation in soft agar observed in S100β-v-erbB/p53−/− glioma stem-like cells after incubation in HCM was abrogated by heat inactivation of HCM (Supplementary Figure 4A and B). This finding suggested that a protein may be mediating the increased stem-like cell self-renewal we measured in soft agar during hypoxia. Therefore, we used a multiplexed Luminex-based immunoassay to evaluate NCM and HCM for 32 commonly secreted proteins, many of which are known promoters of STAT3 activation50 (Supplementary Figure 4C). After reviewing the results of this Luminex evaluation, we initially studied IL1α and VEGF because they increased significantly with hypoxia and reagents for their study were readily available (Supplementary Figure 4C).
Our examination of IL1α (Supplementary Figure 4C) revealed that the IL1α/IL1R pathway was not required for the activation of STAT3 during hypoxia (Supplementary Figure 4D–F). We therefore turned our attention to VEGF, which was highly responsive to hypoxic conditions (Figure 4a). VEGF has been previously identified as a canonical target of HIF1α during hypoxia.51 To study the role of VEGF in mediating the self-renewal effects of HCM on S100β-v-erbB/p53−/− cultures, we prepared HCM from which VEGF was depleted, as described in the Materials and Methods section. We confirmed the removal of VEGF from the conditioned media using a Luminex assay (Supplementary Figure 4G). S100β-v-erbB/p53−/− cells cultured in HCM depleted of VEGF revealed decreased levels of phosphorylated STAT3 compared to cells cultured in HCM (Figure 4b). Consistent with this observation, a non-toxic dose of Ki8751 (Supplementary Figure 4H), a small-molecule inhibitor of VEGFR2, readily abrogated the increase in TSCs cultured in HCM of subsphere formation and colony formation in soft agar (Figures 4c and d). Conversely, S100β-v-erbB/p53−/− cells cultured in NCM supplemented with super-physiologic levels of recombinant VEGF protein showed an increase in the level of STAT3 phosphorylation compared to cells cultured in NCM (Figure 4b). While the VEGF levels observed during hypoxia were in the range of 200 pg/ml (Figure 4a), as much as 2.0 × 106 pg/ml of mouse recombinant VEGF was required to produce an increase in STAT3 phosphorylation in S100β-v-erbB/p53−/− cells cultured in NCM (Figure 4b). However, the addition of 200 pg/ml of recombinant VEGF to HCM depleted of endogenous VEGF was sufficient to produce an increase in STAT3 phosphorylation (Supplementary Figure 4I). We interpret these findings to indicate that while VEGF secretion is required for STAT3 activation during hypoxia, VEGF secretion alone may not be sufficient.
VEGF secreted by S100β-v-erbB/p53−/− tumor spheres during hypoxia increases STAT3 phosphorylation and enhances glioma self-renewal. (a) Luminex assay evaluation of VEGF secreted by TSC1 cells cultured under hypoxia (0 h, 8 h, 16 h). Data points are from a representative experiment conducted in triplicate and are presented as the mean±s.d. (*P<0.01). (b) Western blot analysis of phospho-STAT3 and total STAT3 in TSC1 cells cultured in NCM, NCM with bovine serum albumin (BSA) (2 μg/ml), NCM supplemented with recombinant VEGF protein (2 μg/ml), HCM, HCM supplemented with immunoglobulin G (IgG), or HCM depleted of VEGF (4h). (c) Effect of Ki8751 (1 μM) on TSC1 (left) and TSC2 (right) subsphere formation in HCM (7 days). Data points represent the percentage of plated cells that grew as spheres in three independent experiments conducted in triplicate and are presented as the mean±s.d. (*P<0.05). (d) Effect of Ki8751 (1 μM) on TSC1 (left) and TSC2 (right) colony formation in soft agar in HCM (7 days). Data points represent three independent experiments conducted in triplicate and are presented as the mean±s.d. (*P<0.05).
Pharmacological inhibition of secretion eliminates the hypoxia-mediated enhancement of self-renewal in glioma stem-like cells
To examine further the importance of secreted factors in mediating the self-renewal of glioma stem-like cells observed during hypoxia (Figures 1a and b), we examined the effect of Brefeldin A (BFA), a known inhibitor of secretion,52 and EHT-1864, which inhibits Rac1,53, 54 a known mediator of secretion,55, 56, 57, 58, 59, 60, 61, 62, 63, 64 on S100β-v-erbB/p53−/− cells. BFA and EHT-1864, at non-toxic doses that do not affect the proliferation, extent of apoptosis, and viability of these cells (Supplementary Figure 5A–E), markedly decreased VEGF secretion in S100β-v-erbB/p53−/− cells (Figure 5a). To determine if BFA and EHT-1864 would inhibit the increase in self-renewal of S100β-v-erbB/p53−/− cells observed during hypoxia (Figures 1a and b), we examined TSCs treated with BFA and EHT-1864 and found that treatment abrogated the hypoxia-mediated increase in subsphere formation and colony formation in soft agar (Figures 5b and c). Consistent with their function inhibiting secretion, BFA and EHT-1864 had little effect on subsphere formation and colony formation in soft agar when S100β-v-erbB/p53−/− cultures were incubated in HCM (Figures 5d and e).
Pharmacological inhibition of secretion inhibits the increased tumor cell self-renewal induced by hypoxia. (a) Luminex assay evaluation of VEGF secreted by TSC1 cells cultured under hypoxia in the presence of BFA (0.1μM) or EHT-1864 (1 μM) (16h). Data points are from a representative experiment conducted in triplicate and are presented as the mean±s.d. (*P<0.001). (b) Effect of BFA (0.1μM), EHT-1864 (1 μM), BFA (0.1 μM) supplemented with VEGF (400pg) or EHT-1864 (1 μM) supplemented with VEGF (400 pg) on TSC1 subsphere formation in hypoxia (7 days). Data points represent the percentage of plated cells that grew as spheres in a representative experiment conducted in triplicate and are presented as the mean±s.d. (*P<0.01). (c) Effect of BFA (0.1 μM), EHT-1864 (1 μM), BFA (0.1 μM) supplemented with VEGF (400 pg), or EHT-1864 (1 μM) supplemented with VEGF (400pg) on TSC1 colony formation in soft agar during hypoxia (7 days). Data points are from a representative experiment conducted in triplicate and are presented as the mean±s.d. (*P<0.01). (d) Effect of BFA (0.1 μM) or EHT-1864 (1 μM) in the presence of HCM on TSC1 subsphere formation (7 days). Data points represent the percentage of plated cells that grew as spheres in a representative experiment conducted in triplicate and are presented as the mean±s.d. (*P<0.01). (e) Effect of BFA (0.1 μM) or EHT-1864 (1 μM) in the presence of HCM on TSC1 colony formation in soft agar (7 days). Data points are from a representative experiment conducted in triplicate and are presented as the mean±s.d. (*P<0.01).
As shown in Figure 5a, the level of VEGF detected in the media of S100β-v-erbB/p53−/− cultures after incubation in hypoxic conditions for 16 h is in the range of 400 pg/ml. However, the addition of 400 pg/ml of VEGF to the media was not sufficient to rescue the BFA and EHT-1864 induced inhibition of subsphere formation or colony formation in soft-agar (Figures 5b and c). This finding is consistent with our interpretation of the data shown in Figure 4b; namely, VEGF alone, although required (Figures 4b–d), is not sufficient to promote self-renewal and VEGF alone is not sufficient to drive STAT3 phosphorylation (Figure 4b). The addition of VEGF at 2 μg/ml, however, is capable of increasing STAT3 phosphorylation in BFA or EHT-1864 inhibited TSC1 cell cultures (Supplementary Figure 5F). Taken together, these data provide evidence that BFA and EHT-1864 disrupt secretion decreasing the level of secreted VEGF, which is required (Figures 4b and d), though other secreted factors seem to also be required for the increased self-renewal induced by hypoxia in S100gh other sec−/− cells (Figures 5b and c).
BFA and EHT-1864 reduce VEGF secretion in vivo, slow tumor growth, and increase the survival of mice harboring S100β-v-erbB/p53−/− allografts
To test whether the inhibitory effects of BFA and EHT-1864 on the self-renewal of glioma stem-like cells in vitro could be translated to an in vivo model, we treated mice bearing syngeneic S100β-v-erbB/p53−/− tumor allografts with non-toxic doses of either BFA (60 mg/kg), EHT-1864 (80 mg/kg), or 0.9% saline, the drug diluent65, 66, 67 (Supplementary Figures 6A and B).
Once tumors resulting from the injected cells were palpable (approximately 50 mm3), mice were randomly separated into groups (n=7) and treated with one of the following: BFA twice a day for a course of 7 days,65 EHT-1864 once a day for a course of 10 days,66 or 0.9% saline (n=8) twice a day for 10 days (Supplementary Figure 6A). Consistent with our observations in vitro, BFA and EHT-1864 effectively eliminated detectable plasma VEGF after 7 days of treatment (Figure 6a). These treatments did not have a toxic effect on the animals as determined by percent weight change from baseline (Supplementary Figure 6B) and careful observation during these treatment periods did not reveal evidence of pathology or altered behavior. Importantly, we found that treatment with either BFA or EHT-1864 significantly reduced tumor growth (Figures 6b–d). Figure 6b shows tumors that had been removed from drug treated animals which had been sacrificed at day 20 and control animals sacrificed between days 12 and 20 for immunohistochemical evaluation to assess STAT3 phosphorylation (Supplementary Figure 7). Tumors from the control mice treated with saline were clearly larger than tumors isolated from the mice treated with either BFA or EHT-1864 at the end of this experiment on day 20 of treatment (Figure 6b). Three of the five animals in the saline treatment group shown in Figure 6b had to be sacrificed before day 20 due to excessive tumor burden. Tumors from animals treated with either BFA or EHT-1864 were smaller and grew at a significantly slower rate than tumors treated with saline (Figures 6b and d). Moreover, the mice treated with either BFA or EHT-1864 demonstrated significantly higher survival rates than mice treated with saline (Figure 6e). These findings are consistent with our in vitro data indicating that BFA and EHT-1864 eliminate hypoxia-mediated enhancement of self-renewal in glioma stem-like cells (Figures 5a and c).
BFA and EHT-1864 reduce VEGF secretion, slow tumor growth, and increase the survival of mice harboring S100β-v-erbB/p53−/− allografts. (a) Plasma VEGF levels of mice growing glioma tumor allografts treated either with vehicle (n=3), BFA (n=4), or EHT-1864 (n=4) (7 days). Data points are presented as the mean±s.d. (*P<0.001) (b) Gross appearance of tumors extracted 12 to 20 days after treatment initiation with vehicle (n=5), BFA (n=4), or EHT-1864 (n=4). (c) Estimated tumor volumes of S100β-v-erbB/p53−/− allografts treated with vehicle (n=8) or BFA (n=7). Dashed line indicates end of treatment. When comparing all vehicles against all BFA treated allografts, P<0.009. (d) Tumor volume of TSC1 allografts treated with vehicle (n=8) or EHT-1864 (n=7). Dashed line indicates end of treatment. When comparing all vehicles against all EHT-1864 treated allografts, P<0.0004. (e) Kaplan–Meier survival curve of mice that grew glioma tumor allografts and were treated either with vehicle (n=8), BFA (n=7), or EHT-1864 (n=7). Log-rank tests comparing the survival curves for vehicle control against those of BFA and EHT-184 yield P-values of 0.0056 and 0.0017, respectively.
BFA and EHT-1864 disrupt the glioma stem-like cell hypoxia gene expression signature associated with poor survival of glioblastoma patients
In an initial effort to explore the clinical relevance of our current findings, we developed a ‘stem-like cell hypoxia gene expression signature’ comprised of genes whose expression differed most extensively when these cells were incubated in hypoxia rather than normoxia. To develop the signature, we compared RNA expression profiles of S100β-v-erbB/p53−/− TSC1 cultures incubated in hypoxia (n=3) and S100β-v-erbB/p53−/− TSC1 cultures incubated in normoxia (n=3). We selected genes that had at least a 4-fold change in expression and a t-test P-value of less than or equal to 0.005 (Supplementary Table 1). Unexpectedly, no downregulated genes met these strict criteria. Genes whose expression met these criteria were combined to prepare a ‘hypoxia signature’ that was comprised of 19 genes found to be upregulated during hypoxia (Figure 7a). Gene Ontology analysis of these genes in PantherDB68 revealed that the genes belonged to nine different protein classes (Figure 7b) and potentially participate in six different pathways including pathways well-known to be associated with hypoxia such as the ‘Hypoxia response via HIF1a,’ ‘VEGF signaling’ and ‘Angiogenesis’ (Figure 7c).
BFA and EHT disrupt a glioma stem-like cell hypoxia signature associated with poor survival. (a) Heat map representation of the expression levels of the 19 genes conforming the glioma stem-like cell hypoxia signature in TSC1 cells treated with either DMSO, BFA (0.1 μM), or EHT-1864 (1 μM) grown in hypoxia or normoxia (16 h) as indicated in the figure. (b) Pie charts describing the Gene Ontology protein classes of the genes in a. (c) Pie charts describing the Gene Ontology pathways of the genes in a. (d) Kaplan–Meier survival curves comparing GBM patients with high hypoxia signature scores (H) and GBM patients with low hypoxia signature scores (L). Log-rank test comparing these survival curves has a P-value of 0.0109.
We found that this signature was strongly associated with survival in a cohort of 580 GBM patients from the Cancer Genome Atlas (TCGA) database (Glioblastoma Multiforme TCGA, Provisional).69 Using the Cox proportional hazards model, we derived a ‘hypoxia signature score’ for each GBM patient within this cohort based on the mRNA expression data of the genes from our signature.70, 71 Scores of <0 were categorized as ‘Low’ and scores greater than 0 were categorized as ‘High.’ We observed that the population of GBM patients with high signature scores had significantly decreased survival than those with low scores (Log rank P-value=0.0109) (Figure 7d).
Finally, to assess the ability of BFA or EHT-1864 to block the downstream effects of hypoxia, we examined our hypoxia signature in S100β-v-erbB/p53−/− TSC1 treated with either DMSO (n=3), BFA (n=3), or EHT-1864 (n=3) during hypoxia (Figure 7a and Supplementary Tables 2 and 3). We found that glioma stem-like cells treated with BFA or EHT-1864 had markedly decreased expression levels of most genes from this signature (Figure 7a). These findings suggest a potential role for the use of secretion inhibitors like BFA and EHT-1864 in the treatment of GBM to inhibit the downstream effects of hypoxia and improve patient survival.
Discussion
An increasing body of studies describing the importance of factors secreted by cancer stem cells has emerged in recent years.12, 18, 72, 73, 74 These studies suggest that glioma stem-like cells secrete soluble proteins into the tumor microenvironment establishing autocrine/paracrine loops to promote their own maintenance.12, 18, 72, 73, 74 For example, secretion of Gremlin1 by glioma stem cells decreases bone morphogenetic protein (BMP) signaling to inhibit BMP-mediated glioma stem cell differentiation and loss of self-renewal capacity.18 Similarly, secretion of Sema3C by glioma stem cells has been shown to promote stem cell self-renewal through the activation of PlexinA2/PlexinD1 receptors.73 Recent studies have also demonstrated that cytokines, such as interleukin (IL)-6, can establish autocrine loops to influence glioma stem cell self-renewal.74 Soluble factors secreted by glioma stem cells can also indirectly promote their growth and survival. For example, glioma stem cell secreted factors promote activation of myeloid differentiation primary response gene 88-Toll-Like Receptor 4 (MyD88-TLR4).72 MyD88-TLR4 signaling promotes microglial IL-6 secretion,72 which in turn can stimulate glioma stem cell self-renewal.74 Thus, stem cell secretion can indirectly as well as directly through paracrine/autocrine loops promote cancer stem cell maintenance in HGGs.
Even though several studies of cancer stem cells have identified the importance of environmental influences on stem cell maintenance,13, 15, 16, 17, 75, 76 the specific mechanisms regulating stem-cell responses to those influences are still poorly understood. In this study, we identified a pathway induced by tumor hypoxia, a characteristic feature of the microenvironment found in HGG, that promotes self-renewal by the secretion of VEGF and the resulting activation of STAT3 (Figures 1–3). Using tumor stem-like cells derived from the S100β-v-erbB/p53−/− mouse model of spontaneous HGG,35 we discovered that under hypoxic conditions the expression of HIF-1α is greatly increased in HGGs. Increased HIF-1α results in JAK-mediated activation of STAT3, which in turn mediates enhanced glioma stem-like cell self-renewal (Figures 1 and 2, Supplementary Figure 3). Further, hypoxia-mediated VEGF secretion is required for the activation of STAT3 and the increase in self-renewal associated with hypoxia (Figures 4 and 5). While prior studies describing the effects of VEGF secretion in response to hypoxia have focused largely on their roles inducing angiogenesis,77, 78 we have identified VEGF as a key component of a novel pathway regulating STAT3-mediated self-renewal of HGGs during hypoxia (Figures 1–5) in a manner consistent to what has been previously described in other tumor types.14, 79, 80, 81 Previous studies have identified VEGFR to be preferentially expressed in CD133+ cells in comparison to CD133- cells in human GBM tissue, and determined that VEGFR inhibition was associated with decreased CD133+ survival.82 These investigators also determined that GBM cells with higher levels of VEFGR show increased tumor sphere formation capacity in patient-derived GBM cells suggesting that autocrine/paracrine VEGF-VEGFR signaling could play an important role in glioma stem cell biology.82 Our finding that glioma-derived TSCs required VEGF secretion for the increase in self-renewal observed during hypoxia is consistent with this finding (Figures 4 and 5), and may have implications for numerous tumor biologies including tumor initiation.
We have also identified two agents, Brefeldin A and EHT-1864, that can significantly inhibit VEGF secretion, decrease stem cell self-renewal, and inhibit tumor growth (Figures 5 and 6). This inhibition resulted in the increased survival of mice grafted with S100β-v-erbB/p53−/− glioma stem-like cells (Figure 6). Treatment of HGG stem-like cells with these agents inhibited the expression of a hypoxia signature that strongly associated with decreased survival of HGG patients (Figure 7). In vivo, we observed the therapeutic potential of BFA and EHT-1864 for the treatment of GBM. These findings suggest a novel treatment strategy to inhibit hypoxia-mediated self-renewal of glioma stem-like cells in HGG by targeting the secretion of extracellular, autocrine/paracrine mediators of self-renewal. Future studies in models utilizing human HGG tissues to examine the role of secretion in stem-like cell self-renewal will extend our understanding of glioma biology and may lead to the use of agents that could modulate secretion to improve clinical outcomes for glioma patients.
Materials and methods
Reagents
The following reagents were purchased from the companies indicated: S3I-201 (Calbiochem, San Diego, CA, USA), Brefeldin A (BFA) (Selleckchem, Houston, TX, USA), EHT-1864 (Selleckchem, Houston, TX, USA), Ki8751 (Selleckchem, Houston, TX, USA), Ruxolitinib (Selleckchem, Houston, TX, USA), Interleukin-1 receptor-associated kinase (IRAK) inhibitor (Calbiochem), Interleukin-1a receptor antagonist (ProSpec, Rehovot, Israel), Interleukin-1a mouse recombinant protein (ProSpec, Rehovot, Israel), and VEGFA mouse recombinant protein (ProSpec, Rehovot, Israel). These reagents were dissolved in dimethyl sulfoxide (DMSO) (Sigma-Aldrich, St Louis, MO, USA), distributed into small aliquots, and stored at −20 ºC. Drugs for in vivo experiments were dissolved immediately before use in 0.9% saline (Hospira, Lake Forest, IL, USA).
Cell culture
HGG tissue was obtained from CNS tumors arising in S100β-v-erbB/p53−/− mice (3–6 months old) using sterile techniques. Throughout our experiments we evaluated cells from TSCs. These cells invariably were a mixture of tumor stem-like cells and cells that emerged from them during culture. TSCs were established by re-suspending the dissociated tumor cells from independent tumors in Dulbecco’s Modified Eagle’s Medium-F12 (DMEM/F-12; 50:50; Mediatech, Manassas, VA, USA) supplemented with B27 (Invitrogen, Carlsbad, CA, USA), and 1% penicillin (10 000 units/ml)/streptomycin (10 000 μg/ml) as previously described.83 Two of these cultures, TSC1 and TSC2, were chosen for further experimentation. These cells were derived and characterized in the Israel Laboratory. They have been tested for mycoplasma contamination.
Hypoxic conditions were achieved using a hypoxia chamber (MIC-101; Billups-Rothenberg, Del Mar, CA, USA) flushed with a mixture of 94% N2, 5% CO2, and 1% O2. To remove VEGF from tissue culture media, we immunopreciptated VEGF from conditioned media utilizing an anti-VEGFA antibody (R&D Systems, Minneapolis, MN, USA. Catalog #AF-493-NA) at a concentration of 230 ng/ml and protein A/G beads (Pierce, Rockford, IL, USA) following the manufacturer’s protocol. NCM and HCM were prepared by incubating S100β-v-erbB/p53−/− TSCs in normoxic or hypoxic conditions, respectively, for 16 h. No change in the viability of TSCs was observed following 16 h of incubation in either normoxic or hypoxic conditions. Following incubation, the conditioned media was clarified by centrifugation and filtration through a 0.40 μm filter.
Cell growth and viability
TSCs were seeded into six-well plates at 50,000 cells per ml and grown for 7 days. Cell counts and viability were determined in a Nexcelom Auto 2000 Cell Viability Counter (Nexcelom Bioscience, Lawrence, MA, USA) using a Trypan Blue (Gibco, Gaithersburg, MD, USA) exclusion assay. Annexin V/PI staining was performed using the Annexin V/PI Apoptosis Detection Kit II (BD Biosciences, San Jose, CA, USA) following the manufacturer's protocol. The FL1 and FL3 channels were used to detect Annexin V-FITC and PI, respectively. To determine the drug response curves for BFA and EHT-1864, cells were plated in a 96-well plate at a concentration of 5,000 cells per well. They were allowed to grow overnight and then treated with several concentrations of BFA and EHT-1864 for 7 days as indicated in Supplementary Figure 5B and C. 3-(4,5-dimethylthiazol-2-yl)-2,5-diphenyl-2H-tetrazolium hydrobromide (MTT) was then added to each well at a final concentration of 0.5mg/ml, and cells were incubated for four hours. The absorbance of solubilized formazan was measured at an optical density of 570 nm using a microplate reader (Bio-Rad, Hercules, CA, USA).84
Soft-agar colony formation assay
TSCs were incubated in Accutase as obtained from the manufacturer (EMD Millipore, Billerica, MA, USA) for 5 min, dissociated into single cells, and suspended in soft agar as previously described.85 Following incubation under normoxic (21% oxygen) or hypoxic (1% oxygen) conditions for 7 days, colonies were fixed by adding ice-cold methanol to the agar plate for 10 min. These cultured cells were then stained with 0.005% crystal violet in phosphate-buffered saline, and colonies > 0.2 mm in diameter were counted. If a drug was used in these experiments, the soft agar contained these drugs at the stated concentrations, and the cells were fed with 300 μl/well (six-well plate) of fresh medium containing the drug every 3 days.
Subsphere formation assay at limiting dilution
Spheres were incubated with Accutase, dissociated into single cells, and seeded into 96-well plates as previously described at either a limiting dilution of one cell per well or as indicated in the text.86 Following incubation under normoxic (21% oxygen) or hypoxic (1% oxygen) conditions for 7 days, wells that contained at least one sphere of a diameter of more than 0.05mm were scored as positive. The percentage of sphere-forming cells was determined as the ratio of the total number of wells containing at least one sphere of tumor cells to the total number of wells into which cells were plated. If cells were transfected with plasmid DNA, they were dissociated and seeded for this subsphere formation assay 24 h after transfection. If a drug was used in these experiments, cells were fed with 50 μl/well (96-well plate) of fresh medium containing the drug every 3 days.
Luciferase assay
Luciferase activity was measured using the Dual-Luciferase Reporter Assay System (Promega, Madison, WI, USA) and quantified using a LMax II microplate luminometer (Molecular Devices, Sunnyvale, CA, USA) as per the manufacturer’s protocol. TSCs were seeded at 300 000 cells per plate in 6 cm dishes and transfected with a STAT3-dependent Firefly luciferase reporter and a Renilla luciferase reporter (phRL TK-luc) together with other plasmids as indicated in the text. S3I-201 (Calbiochem) was added to cells 16 h before they were harvested for evaluation. Substrates for firefly luciferase and Renilla luciferase were added to cell extracts immediately following their preparation. STAT3-dependent luciferase activity was normalized to Renilla luciferase values.
Plasmids and transfection
HuSHTM shRNA constructs (Origene Technologies, Rockville, MD, USA) were used to reduce gene expression of mouse HIF-1α. MISSION shRNA constructs (Sigma-Aldrich) were used to reduce the expression of JAK1 and JAK2. shRNA sequences are provided in Supplementary Table 4. Control cultures for these experiments were transfected with either the pRS-sh-scrambled control vector (Origene Technologies, Rockville, MD, USA) or the TRC1.5 (Sigma-Aldrich) control vector, respectively. The pRc/CMV-STAT3c plasmid (plasmid 8722, Addgene, Cambridge, MA, USA)49 was used to express a constitutively active form of STAT3, STAT3C. pEGFP (Clontech, Mountain View, CA, USA) was used in our studies as the control for STAT3C transfection experiments. The m67 pTATA TK-luc plasmid (plasmid 8688, Addgene, Cambridge, MA, USA)87 was used as the backbone for constructing a luciferase reporter, pSTAT3 TK-luc, to measure the transcriptional activity of STAT3. Four copies of the STAT-binding site (TTCCCGTAA) were introduced into the AccI and BamHI sites of pTATA-TK-Luc upstream of the minimal promoter. The Renilla luciferase reporter phRL tk-luc (Promega, Madison, WI, USA) was used in the luciferase assay as per the manufacturer’s protocol.
STAT3 DNA binding assay
DNA binding activity of STAT3 was examined using a STAT3 Filter Plate Assay (Signosis, Sunnyvale, CA, USA) following the manufacturer’s instructions. Nuclear or cytoplasmic extracts were prepared as previously described.88 Protein extracts (5μg) were mixed with biotin-labeled STAT3 DNA binding oligonucleotides and incubated at 16 °C for 30 min to allow the formation of STAT3-DNA complexes. Following incubation, DNA-protein complexes were bound to the Signosis filter plate, while unbound DNA probes were removed. The DNA bound to STAT3 on the hybridization plate was then incubated with a streptavidin-HRP conjugate. STAT3 binding was determined using a luminol-based chemiluminescence reagent, West Pico Super Signal (Pierce, Rockford, IL, USA), per the manufacturer’s instructions.
Multiplex luminex cytokine assay
Multiplex Luminex analyses of mouse plasma and tissue culture media supernatants were carried out by the Norris Cotton Cancer Center Immune Monitoring and Flow Cytometry Shared Resource.89
Mouse tumor allografts
To establish tumor allografts, we injected TSC1 (5.0 × 105 cells) subcutaneously into the flank of female, 4 to 6-week-old FVB-NJ mice. Tumor size was measured in three dimensions every other day with calipers once the tumors became palpable, and the tumor volume was calculated assuming the shape was ellipsoid. This study was powered at 80% to see a volume difference of 400 mm3 between treatment groups with a statistical significance of P=0.05. Mice were allocated to treatment groups using a random number generator. Brefeldin A (60 mg/kg) and EHT-1864 (80 mg/kg) were administered to mice intraperitoneally either twice a day for 7 days65 or once a day for 10 days,66 respectively. Mice were sacrificed when tumors reached 2000 mm3 in accordance with the principles and procedures proscribed by the Dartmouth Institutional Animal Care and Use Committee (IACUC). The date of sacrifice defined the duration of survival in our studies. The groups were not blinded from the investigator during monitoring. Blood was collected from the retro-orbital plexus and mixed with heparin (Sagent Pharmaceuticals, Schaumburg, IL, USA) one week after the initiation of treatment, and plasma was collected by centrifugation, 15 min at 1500 g, 4 °C.
Real-time PCR
RNA was isolated using the RNeasy Kit (Qiagen, Germantown, MD, USA) and reverse transcribed using the iScript cDNA synthesis kit (Bio-Rad, Hercules, CA, USA). To examine mRNA levels, we utilized the iQ SYBR Green Supermix (Bio-Rad) and MyIQ Real-Time PCR system (Bio-Rad). Expression data were expressed relative to untreated cells or to DMSO-treated cells and normalized to 18S Ct values. PCR primer sequences are provided in Supplementary Table 5.
Western blotting
Cellular extracts from cells lysed in RIPA buffer were size fractionated in 6% SDS-PAGE gels and subsequently transferred to PVDF (EMD Millipore, Billerica, MA, USA) or nitrocellulose (GE Healthcare Bio-sciences, Pittsburgh, PA, USA). Antibodies reactive against total STAT3 (Cell Signaling, Danvers, MA, USA. Catalog # 12640), phospho-STAT3 (Cell Signaling, Danvers, MA, USA. Catalog # 9145), and HIF-1α (Novus Biologicals, Littleton, CO, USA. Catalog #NB100-449) were diluted 1:1000 in Tris-Buffered Saline, 0.1% Tween 20 (TBST) containing 5% bovine serum albumin (BSA) for overnight immunoblotting. A horseradish peroxidase (HRP)-conjugated anti-actin antibody (Sigma-Aldrich) was used at 1:50,000 dilution in TBST containing 5% BSA to detect β-actin, a loading control.
Immunohistochemistry
Tumor tissues were fixed in 4% paraformaldehyde in phosphate-buffered saline and then embedded in paraffin. Immunohistochemical reactions were performed in the Norris Cotton Cancer Center Research Pathology Shared Resource (Lebanon, NH, USA) using standard techniques.
Microarray analysis
Mouse RNA samples were labeled using the Affymetrix WTPLus kit according to manufacturer’s guidelines, and probed using the Clariom S Mouse Array. Raw data generated from Clariom S Mouse Arrays were processed using Affymetrix Expression Console Software. CEL files containing feature intensity values were converted into summarized expression values using by Robust Multi-array Average (RMA) which consists of background adjustment, quantile normalization and summarization across all chips. All samples passed QC thresholds for hybridization, labeling and the expression of housekeeping gene controls. The complete normalized data set is available on GEO under Series accession number GSE103022.
Bioinformatics and statistical analysis
To generate a hypoxia signature score for each of the 580 GBM patients from the TCGA database (Glioblastoma Multiforme TCGA, Cell 2013),69 we utilized the Cox proportional hazards model as previously described.70, 71 Scores of less than 0 were categorized as ‘Low’ and scores greater than 0 were categorized as ‘High.’ The code used to generate this is available at GitHub (https://github.com/lharr/gbm_surv_sig). All data are representative of multiple independent experiments as indicated in the figure legends, and they are presented as the mean±s.d. We used the two-tailed, two-sample student’s T-test to evaluate the differences between two groups accounting for variance, and Log-rank (Mantel–Cox) to perform significance analysis of two survival curves. P<0.05 was considered statistically significant. The data met the assumptions of the tests. To compare the effect of BFA, EHT-1864 and vehicle control on tumor size in our animal study, we used a mixed-effects model and multiple chained imputation using Bayesian regression as previously described.90, 91, 92, 93 No samples or animals were excluded from the analysis. Heat maps and statistical analyses were performed with GraphPad Prism (GraphPad Software Inc, La Jolla, CA, USA) and R. Gene Ontology analysis was performed using the Protein Analysis Through Evolutionary Relationships (PANTHER) database.68
Accession codes
References
Rosen JM, Jordan CT . The increasing complexity of the cancer stem cell paradigm. Science 2009; 324: 1670–1673.
Lapidot T, Sirard C, Vormoor J, Murdoch B, Hoang T, Caceres-Cortes J et al. A cell initiating human acute myeloid leukaemia after transplantation into SCID mice. Nature 1994; 367: 645–648.
Al-Hajj M, Wicha MS, Benito-Hernandez A, Morrison SJ, Clarke MF . Prospective identification of tumorigenic breast cancer cells. Proceedings of the National Academy of Sciences of the United States of AmericaProc Natl Acad Sci USA 2003; 100: 3983–3988.
Hemmati HD, Nakano I, Lazareff JA, Masterman-Smith M, Geschwind DH, Bronner-Fraser M et al. Cancerous stem cells can arise from pediatric brain tumors. Proceedings of the National Academy of Sciences of the United States of AmericaProc Natl Acad Sci USA 2003; 100: 15178–15183.
Singh SK, Clarke ID, Terasaki M, Bonn VE, Hawkins C, Squire J et al. Identification of a cancer stem cell in human brain tumors. Cancer ResearchCancer ResCancer Res 2003; 63: 5821.
Singh SK, Clarke ID, Hide T, Dirks PB . Cancer stem cells in nervous system tumors. Oncogene 2004; 23: 7267–7273.
Bao S, Wu Q, McLendon RE, Hao Y, Shi Q, Hjelmeland AB et al. Glioma stem cells promote radioresistance by preferential activation of the DNA damage response. Nature 2006; 444: 756–760.
Liu G, Yuan X, Zeng Z, Tunici P, Ng H, Abdulkadir IR et al. Analysis of gene expression and chemoresistance of CD133(+)cancer stem cells in glioblastoma. Molecular CancerMol Cancer 2006; 5: 67.
Huang Q, Zhang Q-B, Dong J, Wu Y-Y, Shen Y-T, Zhao Y-D et al. Glioma stem cells are more aggressive in recurrent tumors with malignant progression than in the primary tumor, and both can be maintained long-term in vitro. BMC Cancer 2008; 8: 304.
Jackson M, Hassiotou F, Nowak A . Glioblastoma stem-like cells: at the root of tumor recurrence and a therapeutic target. Carcinogenesis 2015; 36: 177–185.
Clevers H . The cancer stem cell: premises, promises and challenges. Nat Med 2011. 313–319.
Lathia JD, Mack SC, Mulkearns-Hubert EE, Valentim CLL, Rich JN . Cancer stem cells in glioblastoma. Genes Dev 2015; 29: 1203–1217.
Soeda A, Park M, Lee D, Mintz A, Androutsellis-Theotokis A, McKay RD et al. Hypoxia promotes expansion of the CD133-positive glioma stem cells through activation of HIF-1α. Oncogene 2009; 28: 3949–3959.
Beck B, Driessens G, Goossens S, Youssef KK, Kuchnio A, Caauwe A et al. A vascular niche and a VEGF-Nrp1 loop regulate the initiation and stemness of skin tumours. Nature 2011; 478: 399–403.
Charles N, Ozawa T, Squatrito M, Bleau A-M, Brennan CW, Hambardzumyan D et al. Perivascular nitric oxide activates notch signaling and promotes stem-like character in PDGF-induced glioma cells. Cell Stem Cell 2010; 6: 141–152.
Heddleston JM, Li Z, Hjelmeland AB, Rich JN . The hypoxic microenvironment maintains glioblastoma stem cells and promotes reprogramming towards a cancer stem cell phenotype. Cell Cycle 2009; 8: 3274–3284.
Hjelmeland AB, Wu Q, Heddleston JM, Choudhary GS, MacSwords J, Lathia J et al. Acidic stress promotes a glioma stem cell phenotype. Cell Death Differ 2011; 18: 829–840.
Yan K, Wu Q, Yan DH, Lee CH, Rahim N, Tritschler I et al. Glioma cancer stem cells secrete Gremlin1 to promote their maintenance within the tumor hierarchy. Genes Dev 2014; 28: 1085–1100.
Yun Z, Lin Q . Hypoxia and regulation of cancer cell stemness. Adv Exp Med Biol 2014; 772: 41–53.
Brat DJ, Castellano-Sanchez AA, Hunter SB, Pecot M, Cohen C, Hammond EH et al. Pseudopalisades in glioblastoma are hypoxic, express extracellular matrix proteases, and are formed by an actively migrating cell population. Cancer ResearchCancer ResCancer Res 2004; 64: 920.
Rong Y, Durden DL, Van Meir EG, Brat DJ . 'Pseudopalisading' necrosis in glioblastoma: a familiar morphologic feature that links vascular pathology, hypoxia, and angiogenesis. J Neuropathol Exp Neurol 2006; 65: 529–539.
Brown JM, Wilson WR . Exploiting tumour hypoxia in cancer treatment. Nat Rev Cancer 2004; 4: 437–447.
Vordermark D, Brown JM . Endogenous markers of tumorhypoxia. Strahlentherapie und OnkologieStrahlenther Onkol 2003; 179: 801–811.
Spence AM, Muzi M, Swanson KR, Sullivan F, Rockhill JK, Rajendran JG et al. Regional hypoxia in glioblastoma multiforme quantified with [18F] fluoromisonidazole positron emission tomography before radiotherapy: correlation with time to progression and survival. Clinical Cancer ResearchCancer ResCancer Res 2008; 14: 2623.
Wilson WR, Hay MP . Targeting hypoxia in cancer therapy. Nat Rev Cancer 2011; 11: 393–410.
Vaupel P, Mayer A . Hypoxia in cancer: significance and impact on clinical outcome. Cancer and Metastasis ReviewsCancer Metastat Rev 2007; 26: 225–239.
Nordsmark M, Bentzen SM, Rudat V, Brizel D, Lartigau E, Stadler P et al. Prognostic value of tumor oxygenation in 397 head and neck tumors after primary radiation therapy. An international multi-center study. Radiother Oncol 2005; 77: 18–24.
Semenza GL . Hypoxia-inducible factor 1: oxygen homeostasis and disease pathophysiology. Trends in Molecular MedicineTrends Mol Med 2001; 7: 345–350.
Pouyssegur J, Dayan F, Mazure NM . Hypoxia signalling in cancer and approaches to enforce tumour regression. Nature 2006; 441: 437–443.
Harris AL . Hypoxia [mdash] a key regulatory factor in tumour growth. Nat Rev Cancer 2002; 2: 38–47.
Søndergaard KL, Hilton DA, Penney M, Ollerenshaw M, Demaine AG . Expression of hypoxia-inducible factor 1α in tumours of patients with glioblastoma. Neuropathol Appl Neurobiol 2002; 28: 210–217.
Liu Q, Cao P . Clinical and prognostic significance of HIF-1α in glioma patients: a meta-analysis. Inr J Clin Exp Med 2015; 8: 22073–22083.
Semenza GL . Targeting HIF-1 for cancer therapy. Nat Rev Cancer 2003; 3: 721–732.
Keith B, Simon MC . Hypoxia Inducible Factors, stem cells and cancer. Cell 2007; 129: 465–472.
Weiss WA, Burns MJ, Hackett C, Aldape K, Hill JR, Kuriyama H et al. Genetic determinants of malignancy in a mouse model for oligodendroglioma. Cancer ResearchCancer ResCancer Res 2003; 63: 1589.
Louis DN, Perry A, Reifenberger G, von Deimling A, Figarella-Branger D, Cavenee WK et al. The 2016 World Health Organization classification of tumors of the central nervous system: a summary. Acta Neuropathol 2016; 131: 803–820.
Singh SK, Hawkins C, Clarke ID, Squire JA, Bayani J, Hide T et al. Identification of human brain tumour initiating cells. Nature 2004; 432: 396–401.
De Witt Hamer PC, Van Tilborg AA, Eijk PP, Sminia P, Troost D, Van Noorden CJ et al. The genomic profile of human malignant glioma is altered early in primary cell culture and preserved in spheroids. Oncogene 2007; 27: 2091–2096.
Lee J, Kotliarova S, Kotliarov Y, Li A, Su Q, Donin NM et al. Tumor stem cells derived from glioblastomas cultured in bFGF and EGF more closely mirror the phenotype and genotype of primary tumors than do serum-cultured cell lines. Cancer Cell 2006; 9: 391–403.
Parsons DW, Jones S, Zhang X, JC-H Lin, Leary RJ, Angenendt P et al. An integrated genomic analysis of human glioblastoma multiforme. Science 2008; 321: 1807–1812.
Harris MA, Yang H, Low BE, Mukherje J, Guha A, Bronson RT et al. Cancer stem cells are enriched in the side-population cells in a mouse model of glioma. Cancer Res 2008; 68: 10051–10059.
Lee MY, Joung YH, Lim EJ, Park J-H, Ye S-K, Park T et al. Phosphorylation and activation of STAT proteins by hypoxia in breast cancer cells. Breast 2006; 15: 187–195.
Kang S-H, Yu MO, Park K-J, Chi S-G, Park D-H, Chung Y-G . Activated STAT3 regulates hypoxia-induced angiogenesis and cell migration in human glioblastoma. Neurosurgery 2010; 67: 1386–1395.
Selvendiran K, Bratasz A, Kuppusamy ML, Tazi MF, Rivera BK, Kuppusamy P . Hypoxia induces chemoresistance in ovarian cancer cells by activation of signal transducer and activator of transcription 3. Int J Cancer 2009; 125: 2198–2204.
Gray MJ, Zhang J, Ellis LM, Semenza GL, Evans DB, Watowich SS et al. HIF-1alpha, STAT3, CBP/p300 and Ref-1/APE are components of a transcriptional complex that regulates Src-dependent hypoxia-induced expression of VEGF in pancreatic and prostate carcinomas. Oncogene 2005; 24: 3110–3120.
Jung JE, Lee HG, Cho IH, Chung DH, Yoon SH, Yang YM et al. STAT3 is a potential modulator of HIF-1-mediated VEGF expression in human renal carcinoma cells. FASEB J 2005; 19: 1296–1298.
Noman MZ, Buart S, Van Pelt J, Richon C, Hasmim M, Leleu N et al. The cooperative induction of hypoxia-inducible factor-1 alpha and STAT3 during hypoxia induced an impairment of tumor susceptibility to CTL-mediated cell lysis. J Immunol 2009; 182: 3510–3521.
Siddiquee K, Zhang S, Guida WC, Blaskovich MA, Greedy B, Lawrence HR et al. Selective chemical probe inhibitor of Stat3, identified through structure-based virtual screening, induces antitumor activity. Proc Natl Acad Sci USA 2007; 104: 7391–7396.
Bromberg JF, Wrzeszczynska MH, Devgan G, Zhao Y, Pestell RG, Albanese C et al. Stat3 as an oncogene. Cell 1999; 98: 295–303.
Yu H, Lee H, Herrmann A, Buettner R, Jove R . Revisiting STAT3 signalling in cancer: new and unexpected biological functions. Nat Rev Cancer 2014; 14: 736–746.
Pugh CW, Ratcliffe PJ . Regulation of angiogenesis by hypoxia: role of the HIF system. Nat Med 2003; 9: 677–684.
Misumi Y, Misumi Y, Miki K, Takatsuki A, Tamura G, Ikehara Y . Novel blockade by brefeldin A of intracellular transport of secretory proteins in cultured rat hepatocytes. J Biol Chem 1986; 261: 11398–11403.
Shutes A, Onesto C, Picard V, Leblond B, Schweighoffer F, Der CJ . Specificity and mechanism of action of EHT 1864, a novel small molecule inhibitor of rac family small GTPases. J Biol Chem 2007; 282: 35666–35678.
Onesto C, Shutes A, Picard V, Schweighoffer F, Der CJ . Characterization of EHT 1864, a novel small molecule inhibitor of Rac family small GTPases. Methods in EnzymologyMethods Enzymol 2008; 439: 111–129.
Williams JA, Chen X, Sabbatini ME . Small G proteins as key regulators of pancreatic digestive enzyme secretion. Am J Physiol 2009; 296: E405–E414.
Akbar H, Kim J, Funk K, Cancelas JA, Shang X, Chen L et al. Genetic and pharmacologic evidence that Rac1 GTPase is involved in regulation of platelet secretion and aggregation. Journal of Thrombosis and HaemostasisThromb HaemostasisJ Thromb Haemost 2007; 5: 1747–1755.
Li Q, Ho CS, Marinescu V, Bhatti H, Bokoch GM, Ernst SA et al. Facilitation of Ca(2+)-dependent exocytosis by Rac1-GTPase in bovine chromaffin cells. J Physiol 2003; 550 (Pt 2): 431–445.
Stanley AC, Wong CX, Micaroni M, Venturato J, Khromykh T, Stow JL et al. The Rho GTPase Rac1 is required for recycling endosome-mediated secretion of TNF in macrophages. Immunol Cell Biol 2014; 92: 275–286.
Li J, Luo R, Kowluru A, Li G . Novel regulation by Rac1 of glucose- and forskolin-induced insulin secretion in INS-1 β-cells. Am J Physiol 2004; 286: E818.
Davila J, Laws MJ, Kannan A, Li Q, Taylor RN, Bagchi MK et al. Rac1 regulates endometrial secretory function to control placental development. PLoS GeneticsPLoS Genet 2015; 11: e1005458.
Hwaiz R, Rahman M, Zhang E, Thorlacius H . Platelet secretion of CXCL4 is Rac1‐dependent and regulates neutrophil infiltration and tissue damage in septic lung damage. Br J Pharmacol 2015; 172: 5347–5359.
Hwaiz R, Rahman M, Syk I, Zhang E, Thorlacius H . Rac1-dependent secretion of platelet-derived CCL5 regulates neutrophil recruitment via activation of alveolar macrophages in septic lung injury. Journal of Leukocyte Biology 2015; 97: 975–984.
Goyal P, Brünnert D, Ehrhardt J, Bredow M, Piccenini S, Zygmunt M . Cytokine IL-6 secretion by trophoblasts regulated via sphingosine-1-phosphate receptor 2 involving Rho/Rho-kinase and Rac1 signaling pathways. MHR: Basic science of reproductive medicine 2013; 19: 528–538.
Asahara S, Shibutani Y, Teruyama K, Inoue HY, Kawada Y, Etoh H et al. Ras-related C3 botulinum toxin substrate 1 (RAC1) regulates glucose-stimulated insulin secretion via modulation of F-actin. Diabetologia 2013; 56: 1088–1097.
Sausville EA, Duncan KL, Senderowicz A, Plowman J, Randazzo PA, Kahn R et al. Antiproliferative effect in vitro and antitumor activity in vivo of brefeldin A. Cancer J Sci Am 1996; 2: 52–58.
Desire L, Bourdin J, Loiseau N, Peillon H, Picard V, De Oliveira C et al. RAC1 inhibition targets amyloid precursor protein processing by gamma-secretase and decreases Abeta production in vitro and in vivo. J Biol Chem 2005; 280: 37516–37525.
Dwivedi S, Pandey D, Khandoga AL, Brandl R, Siess W . Rac1-mediated signaling plays a central role in secretion-dependent platelet aggregation in human blood stimulated by atherosclerotic plaque. Journal of Translational MedicineJ Transl Med 2010; 8: 128.
Mi H, Huang X, Muruganujan A, Tang H, Mills C, Kang D et al. PANTHER version 11: expanded annotation data from Gene Ontology and Reactome pathways, and data analysis tool enhancements. Nucleic Acids ResearchNucleic Acids Res 2017; 45: D183–D189.
Brennan CW, Verhaak RGW, McKenna A, Campos B, Noushmehr H, Salama SR et al. The somatic genomic landscape of glioblastoma. Cell 2013; 155: 462–477.
Li H, Gui J . Partial Cox regression analysis for high-dimensional microarray gene expression data. Bioinformatics 2004; 20: i208–i215.
Lu Y, Lemon W, Liu P-Y, Yi Y, Morrison C, Yang P et al. A gene expression signature predicts survival of patients with stage i non-small cell lung cancer. PLoS MedicinePLoS Med 2006; 3: e467.
a Dzaye OD, Hu F, Derkow K, Haage V, Euskirchen P, Harms C et al. Glioma stem cells but not bulk glioma cells upregulate IL-6 secretion in microglia/brain macrophages via toll-like receptor 4 signaling. J Neuropathol Exp Neurol 2016; 75: 429–440.
Man J, Shoemake J, Zhou W, Fang X, Wu Q, Rizzo A et al. Sema3C Promotes The Survival and Tumorigenicity of Glioma Stem Cells Through Rac1 Activation. Cell reports 2014; 9: 1812–1826.
Wang H, Lathia JD, Wu Q, Wang J, Li Z, Heddleston JM et al. Targeting interleukin 6 signaling suppresses glioma stem cell survival and tumor growth. Stem Cells 2009; 27: 2393–2404.
Qiang L, Wu T, Zhang Hw, Lu N, Hu R, Wang Yj et al. HIF-1alpha is critical for hypoxia-mediated maintenance of glioblastoma stem cells by activating Notch signaling pathway. Cell Death Differ 2012; 19: 285–294.
Flavahan WA, Wu Q, Hitomi M, Rahim N, Kim Y, Sloan AE et al. Brain tumor initiating cells adapt to restricted nutrition through preferential glucose uptake. Nat Neurosci 2013; 16: 1373–1382.
Ferrara N . VEGF and the quest for tumour angiogenesis factors. Nat Rev Cancer 2002; 2: 795–803.
Ferrara N, Gerber H-P, LeCouter J . The biology of VEGF and its receptors. Nat Med 2003; 9: 669–676.
Cao Y, E G, Wang E, Pal K, Dutta SK, Bar-Sagi D et al. VEGF exerts an angiogenesis-independent function in cancer cells to promote their malignant progression. Cancer Res 2012; 72: 3912–3918.
Zhao D, Pan C, Sun J, Gilbert C, Drews-Elger K, Azzam DJ et al. VEGF drives cancer-initiating stem cells through VEGFR-2/Stat3 signaling to upregulate Myc and Sox2. Oncogene 2015; 34: 3107–3119.
Goel HL, Mercurio AM . VEGF targets the tumour cell. Nat Rev Cancer 2013; 13: 871–882.
Hamerlik P, Lathia JD, Rasmussen R, Wu Q, Bartkova J, Lee M et al. Autocrine VEGF–VEGFR2–Neuropilin-1 signaling promotes glioma stem-like cell viability and tumor growth. J Exp Med 2012; 209: 507–520.
Almiron Bonnin DA, Ran C, Havrda MC, Liu H, Hitoshi Y, Zhang Z et al. Insulin-mediated signaling facilitates resistance to PDGFR inhibition in proneural hPDGFB-driven gliomas. Mol Cancer Ther 2017; 16: 705–716.
van Meerloo J, Kaspers GJL, Cloos J Cell Sensitivity Assays: The MTT Assay In: Cree IA (ed) Cancer Cell Culture: Methods and Protocols. Humana Press: Totowa, NJ, 2011, p 237–245.
Borowicz S, Van Scoyk M, Avasarala S, Karuppusamy Rathinam MK, Tauler J, Bikkavilli RK et al. The soft agar colony formation assay. J Vis Exp 2014; 92: 51998.
Agro L, O’Brien CA . In vitro and in vivo limiting dilution assay for colorectal cancer. Bio-protocol 2015; 5: e1659.
Besser D, Bromberg JF, Darnell JE, Hanafusa H . A single amino acid substitution in the v-Eyk intracellular domain results in activation of stat3 and enhances cellular transformation. Molecular and Cellular BiologyMol Cell Biol 1999; 19: 1401–1409.
Havrda MC, Johnson MJ, O’Neill CF, Liaw L . A novel mechanism of transcriptional repression of p27kip1 through Notch/HRT2 signaling in vascular smooth muscle cells. Thromb Haemostasis 2006; 96: 361–370.
Lab D . Multiplexed cytokines. Available at http://www.dartmouth.edu/~dartlab/?page=multiplexed-cytokines.
van Buuren S, Groothuis-Oudshoorn K . mice: multivariate imputation by chained equations in R. J Stat Softw 2011; 1: 2011.
Finucane MM, Samet JH, Horton NJ . Translational methods in biostatistics: linear mixed effect regression models of alcohol consumption and HIV disease progression over time. Epidemoil Perspect Innov 2007; 4: 8.
Garabed RB, Johnson WO, Gill J, Perez AM, Thurmond MC . Exploration of associations between governance and economics and country level foot-and-mouth disease status by using Bayesian model averaging. J R Stat Soc A 2008; 171: 699–722.
Laird NM, Ware JH . Random-effects models for longitudinal data. Biometrics 1982; 38: 963–974.
Acknowledgements
We thank Dr Tor Tosteson and Dr Jiang Gui from the Biostatistics Shared Resource at Norris Cotton Cancer Center at Dartmouth for their consultation regarding statistical analysis, and Tabatha Richardson for her administrative support. Microarray processing and data normalization was carried out at Norris Cotton Cancer Center in the Genomics Shared Resource, which was established by equipment grants from the NIH and NSF and is supported in part by a Cancer Center Core Grant (P30CA023108) from the National Cancer Institute. Multiplex Luminex Cytokine Assay was performed in DartLab, the Immune Monitoring and Flow Cytometry Shared Resource at Norris Cotton Cancer Center, supported by a Cancer Center Core Grant (P30CA023108) from the National Cancer Institute and an Immunology COBRE Grant (P30GM10315-15) from the National Institute of General Medical Sciences. Financial support was generously provided by the Theodora B. Betz Foundation, the Jordan and Kyra Memorial Foundation and the Andrea Clark Nelson Medical Research Endowment. Support was generously provided by the Theodora B. Betz Foundation (M.A. Israel), the Jordan and Kyra Memorial Foundation (M.A. Israel), and the Andrea Clark Nelson Medical Research Endowment (M.A. Israel)..
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies this paper on the Oncogene website
Rights and permissions
This work is licensed under a Creative Commons Attribution-NonCommercial-ShareAlike 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-sa/4.0/
About this article
Cite this article
Almiron Bonnin, D., Havrda, M., Lee, M. et al. Secretion-mediated STAT3 activation promotes self-renewal of glioma stem-like cells during hypoxia. Oncogene 37, 1107–1118 (2018). https://doi.org/10.1038/onc.2017.404
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/onc.2017.404
This article is cited by
-
Modulation of hypoxia-inducible factor-1 signaling pathways in cancer angiogenesis, invasion, and metastasis by natural compounds: a comprehensive and critical review
Cancer and Metastasis Reviews (2024)
-
STAT3 and HIF1α cooperatively mediate the transcriptional and physiological responses to hypoxia
Cell Death Discovery (2023)
-
GRB10 is a novel oncogene associated with cell proliferation and prognosis in glioma
Cancer Cell International (2022)
-
Direct and indirect effects of IFN-α2b in malignancy treatment: not only an archer but also an arrow
Biomarker Research (2022)
-
Alpha-enolase (ENO1), identified as an antigen to monoclonal antibody 12C7, promotes the self-renewal and malignant phenotype of lung cancer stem cells by AMPK/mTOR pathway
Stem Cell Research & Therapy (2021)