Abstract
In acute myeloid leukemia (AML), internal tandem duplication mutations in the FLT3 tyrosine kinase receptor (FLT3-ITD) account for up to 25% of cases and are associated with a poor outcome. In order to better target this AML subtype, a comprehensive view of how FLT3-ITD impacts AML cell biology is required. Here, we found that FLT3-ITD expression increased basal autophagy in AML cells, and that both pharmacological and genetic inhibition of the receptor reduced autophagy in primary AML samples and cell lines. Conditional expression of shRNAs against key autophagy proteins demonstrated that autophagy is required for AML cell proliferation in vitro and for leukemic cells survival in a mouse model of xenograft. Importantly, autophagy inhibition also overcame FLT3 inhibitor resistance both in vitro and in vivo. The transcription factor ATF4 was identified as an essential actor of FLT3-ITD-induced autophagy. Cellular levels of ATF4 were highly dependent on FLT3-ITD activity, and downregulation of ATF4 inhibited autophagy-dependent AML cell proliferation and improved overall mouse survival similarly to autophagy inhibition. These results suggest that targeting autophagy or ATF4 in patients expressing FLT3 mutations may represent a novel promising and innovative therapeutic strategy for AML.
Similar content being viewed by others
Introduction
Acute myeloid leukemia (AML) arise from the malignant clonal expansion of undifferentiated myeloid precursors, resulting in bone marrow failure. Although many AML patients initially respond to conventional induction therapy, relapses are common and carry a very poor prognosis.1 The standard treatments for AML have remained almost unchanged over the past 40 years, demonstrating that despite improvements in our understanding of AML pathogenesis, a more in-depth knowledge of the biology of AML is still required in order to rationally design more efficient therapies. Over the past few decades, extensive molecular characterization studies have highlighted the complex heterogeneity of AML.2 The most common genetic abnormalities, which occur in about 30% of AML patients, lie within the Fms-Like Tyrosine kinase 3 (FLT3) gene which encodes a receptor tyrosine kinase. The most frequent mutations in this gene occur via internal tandem duplication (FLT3-ITD) in the juxta-membrane domain, or through point mutations, usually of the Asp835 residue within the activation loop (FLT3-TKD). Both types of mutations result in constitutive activation of FLT3 and promote leukemic cell proliferation and survival. The poor outcome associated with FLT3-ITD mutations3 has generated great interest in the development of specific tyrosine kinase inhibitors for more than 10 years. Of the new generation of FLT3 inhibitors developed, quizartinib (AC220) is considered the most potent and selective, but resistance still commonly arises due in particular to secondary mutations (for example, FLT3-TKD). Recently, midostaurin (PKC412) has shown efficiency against both FLT3-ITD and FLT3-TKD, and early clinical trials are currently underway (for review, see 4). Although important efforts are aimed at developing more efficient tyrosine kinase inhibitors, novel approaches to eradicate FLT3-mutated leukemic cells must also be developed.
Autophagy is a highly conserved catabolic process used by the cell to degrade and recycle damaged cellular components in response to adverse environmental stimuli. It relies on the formation of the autophagosome, a double-membraned vesicle that sequesters cytoplasmic bulk, proteins and organelles before fusing with lysosomes to degrade the engulfed material. The role of autophagy in cancer is complex since it plays a tumor suppressor role during the early phases of tumor initiation through the prevention of genomic instability yet promotes tumor development in established tumors by promoting cancer cell survival.5 Interestingly, oncogenes such as K-RASG12D, BRAFV600E and the fusion protein BCR-ABL1 were reported to support a high level of basal autophagy in different types of cancers which is required for cell survival and proliferation.6, 7, 8 Moreover, autophagy has also been described to be a mechanism of resistance to different types of drugs or treatments,9, 10 and inhibiting autophagy in this context was shown to enhance treatment efficiency. The publication of these studies has instigated several clinical trials combining the only available in vivo autophagy inhibitor, hydroxychloroquine (HCQ), with conventional treatments in different cancers.11
A few studies have sought to understand the role of autophagy in AML, and suggest that inhibiting autophagy sensitizes particular subgroups of AML cells to chemotherapies12, 13 or to small molecules inhibitors (for example, histone deacetylase inhibitor).14, 15 However, the potential role of autophagy in AML cell biology as a mechanism of progression in FLT3-mutated AML remains to be clarified. Here, we found that FLT3-ITD mutations are able to induce an increase in basal autophagy in leukemic cells, through a previously uncharacterized signaling cascade involving the transcription factor ATF4. Moreover, inhibiting autophagy or ATF4 significantly impaired FLT3-ITD leukemic cell proliferation as well as tumor burden in murine xenograft models. Importantly, autophagy inhibition also overcame FLT3 inhibitor resistance due to FLT3-TKD mutation both in vitro and in vivo. These results suggest that targeting ATF4 or autophagy in AML patients carrying FLT3 mutations may represent a promising alternative therapeutic strategy.
Results
FLT3-ITD increases basal autophagy in AML cells
Little is known of the role of autophagy in AML cell biology, especially in the context of the expression of FLT3-ITD, a mutation frequently found in these pathologies. To evaluate the impact of FLT3-ITD on autophagy levels in AML cells, the effect of FLT3-ITD inhibition was investigated in the two human FLT3-ITD-positive cell lines, MOLM-14 and MV4-11. Pharmacological inhibition of FLT3 activity in these cells by incubation with a specific FLT3 inhibitor for 16 h was followed by a significant decrease in autophagic flux. Indeed, FLT3 inhibition decreased lipidated LC3B (LC3B-II), an autophagosomal membrane marker, in presence of the autophagy inhibitor chloroquine (Figure 1a, Supplementary Figure S1A). Accordingly, the average number of autophagosomes per cell was significantly reduced upon FLT3 inhibition, as evaluated by both immunofluorescence (Figures 1b and c) and cytometry (Supplementary Figure S1B) studies. Similar results were obtained with AC-220 (quizartinib), a FLT3 inhibitor currently being evaluated in phase III clinical trials (Supplementary Figure S1C).
FLT3-ITD supports basal autophagy in leukemic cells. (a) MOLM-14 cells were treated with PBS (control) or FLT3 inhibitor (100 nM) for 16 h in the presence or absence of chloroquine (15 μM), before being processed for western blot analysis of phospho-STAT5 (p-STAT5), STAT5 and LC3B. Actin was used as a loading control. Numbers represent the LC3B-II/actin ratios obtained by densitometric analysis (n=3±s.e.m.). (b, c) MOLM-14 cells were treated with PBS (control) or FLT3 inhibitor (100 nM) for 16 h in the presence or absence of chloroquine (15 μM), then stained for LC3B. Representative confocal pictures are shown in (b), and the graph in (c) represents the number of LC3B dots per cell. Scale bar: 10 μm. (d) MV4-11 cells stably expressing FLT3-specific-inducible shRNA were treated with PBS (control) or doxycycline (2 μg/ml) for 2 days, then bafilomycin (20 nM) was added or not for 2 h before the cells were processed for western blot analysis of FLT3 and LC3B. Actin was used as a loading control. Numbers represent the LC3B-II/actin ratios obtained by densitometric analysis (n=3±s.e.m.). (e, f) MV4-11 cells stably expressing FLT3-specific-inducible shRNA were treated with PBS (control) or doxycycline (2 μg/ml) for 2 days, then chloroquine (15 μM) was added for 16 h before staining for LC3B. Representative confocal pictures are shown in (e), and the graph in (f) represents the number of LC3B dots per cell. Scale bar: 10 μm. (g) Patient samples harboring the FLT3-ITD mutation were treated with PBS (control) or FLT3 inhibitor (100 nM) for 16 h in the presence or absence of chloroquine (10 μM), before being processed for western blot analysis of STAT5, phospho-STAT5 (p-STAT5) and LC3B. Actin was used as a loading control. Numbers represent the LC3B-II/actin ratios obtained by densitometric analysis (n=1). (h) Same experiment as in (g), performed with primary cells from two patients expressing wild-type FLT3. PBS, phosphate buffered saline.
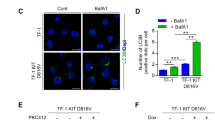
To confirm these results, we genetically downregulated FLT3 in MV4-11 and MOLM-14 cells using either siRNAs or inducible shRNAs against FLT3. Downregulation of the receptor by both of these RNA interference approaches also induced a significant decrease in autophagy in these cells (Figures 1d−f and Supplementary Figure S1D). These data suggest that FLT3-ITD activity increases basal autophagy levels in these leukemic cells. In accordance, as shown by western blot analysis in Supplementary Figure S1E, TF-1 erythroleukemic cells expressing FLT3-ITD displayed increased lipidated LC3B-II in presence of bafilomycin compared to TF-1 cells that did not express the receptor. Moreover, immunofluorescence analysis of LC3B revealed a significantly higher number of autophagosomes in these FLT3-ITD-positive cells (Supplementary Figures S1F and G).
Based on these results, we next asked whether the expression of the oncogenic receptor FLT3-ITD also supports autophagy in AML primary samples. We therefore assessed the level of autophagy upon treatment with FLT3 inhibitors in primary AML samples from patients expressing FLT3-ITD. As shown in Figure 1g, treatment of FLT3-ITD-positive primary cells with FLT3 inhibitor for 16 h decreased the autophagy levels in these cells, whereas it had no effect on primary AML cells expressing wild-type FLT3 (Figure 1h). The efficiency of the FLT3 inhibitor at abrogating FLT3-ITD activity was confirmed in these experiments by monitoring STAT5 phosphorylation status, a marker of FLT3-ITD activity (Figure 1g).
Altogether these complementary biochemical and cellular data collectively demonstrate that FLT3-ITD expression and activity cause an increased level of basal autophagy in AML cells.
The transcription factor ATF4 mediates autophagy downstream of FLT3-ITD
We then investigated which signaling pathways were involved in FLT3-ITD-induced autophagy. Pharmacological inhibition of the JAK2/STAT5, PI3K/AKT or MEK/ERK pathways did not modify the level of autophagy in MOLM-14 cells, showing that none of the major signaling cascades activated by this mutant receptor tyrosine kinase are involved (Supplementary Figures S2A−C).
Since FLT3-ITD was previously reported to localize at the endoplasmic reticulum and to signal from this compartment,16 we wondered whether the eIF2α/ATF4 pathway could be involved. This pathway is induced by different environmental stresses, particularly endoplasmic reticulum stress, and was recently demonstrated to be able to fine tune autophagy.17 As shown in Supplementary Figure S2D, treatment of MOLM-14 cells with a FLT3 inhibitor induced the phosphorylation of eIF2α on Ser51, a marker of activation of the pathway that represses global translation. Surprisingly, eIF2α phosphorylation was accompanied by a dramatic decrease in ATF4 protein levels (Supplementary Figure S2D). This was unexpected since eIF2α phosphorylation generally increases cap-independent translation of ATF4.18 However, these results suggest that ATF4 protein levels are positively controlled by FLT3-ITD activity, independent of their regulation by eIF2α. These results were confirmed by time course experiments in which pharmacological inhibition of FLT3-ITD led to a rapid and complete drop in ATF4 protein levels (Figure 2a). Pharmacological or genetic (RNA interference) inhibition of FLT3 also decreased ATF4 protein levels in MOLM-14 and MV4-11 leukemic cells (Figure 2b and Supplementary Figures S2E and F). In order to precise the mechanisms of ATF4 regulation by FLT3-ITD, we first analyzed ATF4 mRNA levels in response to FLT3 inhibition (Figure 2c) and found that it was not modified in these conditions, suggesting that FLT3-ITD regulates neither transcription nor mRNA stability of ATF4. We then performed time course experiments of cycloheximide in order to estimate the impact of FLT3-ITD on ATF4 turn-over. As shown in Figures 2d and e, FLT3 inhibition does not modify ATF4 turn-over in these experiments. Then, to ask whether FLT3-ITD regulates ATF4 synthesis, cells were first shortly treated with cycloheximide in order to downregulate ATF4 level, and ATF4 re-accumulation was then followed upon removal of cycloheximide, in the presence of MG132 to impair its degradation. Since accumulation of ATF4 in these conditions was significantly decreased upon FLT3 inhibitor treatment, we conclude that FLT3-ITD is important for ATF4 synthesis process rather that for its degradation (Figures 2f and g). Finally, we performed similar experiments in OCI-AML3 cells that express wild-type FLT3, and we found that ATF4 synthesis was not significantly affected by FLT3 inhibitor in these cells (Supplementary Figure S2G).
ATF4 is involved in the FLT3-ITD-dependent signaling required for autophagy. (a) MOLM-14 cells were treated with PBS (control) or FLT3 inhibitor (100 nM) for 30 min, 1, 2 or 16 h and were then processed for western blot analysis of ATF4. Actin was used as a loading control (n=3). (b) MV4-11 cells were treated with FLT3 inhibitor (100 nM) for 2 h, and then processed for western blot analysis of ATF4. Actin was used as a loading control (n=3). (c) MOLM-14 and MV4-11 cells were treated with PBS (control) or FLT3 inhibitor (100 nM) for 6 h then processed for Taqman qPCR analysis of ATF4. GUSB and B2M were used as control genes. Histograms represent the percentage of ATF4 mRNA in cells treated with PBS or FLT3 inhibitor (n=6). (d, e) MOLM-14 cells were treated with cycloheximide (CHX: 50 μg/ml) for 20, 40, 60, 80 or 100 min in the presence (right panel) or absence (left panel) of FLT3 inhibitor (100 nM), before being processed for western blot analysis of ATF4. Actin was used as a loading control. The graph in (e) represents the percentage of ATF4 protein remaining in cells treated with PBS or FLT3 inhibitor in presence of cycloheximide over time (n=3). (f, g). After a pretreatment with cycloheximide (50 μg/ml) of 2 h, MOLM-14 cells were washed to remove cycloheximide, and then treated with MG132 (10 μg/ml), for 1 h, 2 h or 3 h in the presence or absence of FLT3 inhibitor (100 nM), before being processed for western blot analysis of ATF4, eIF2α and P-eIF2α. Actin was used as a loading control. Histogram (g) represents the percentage of ATF4 protein in cells treated with PBS or FLT3 inhibitor in presence of MG132 over time (n=3). (h) MOLM-14 cells were transfected with ATF4-specific siRNA, then 16 h later cells were treated or not with bafilomycin (20 nM) for 2 h, before being processed for western blot analysis of ATF4 and LC3B. Actin was used as a loading control. Numbers represent the LC3B-II/actin ratios obtained by densitometric analysis (n=3±s.e.m.). (i, j) MOLM-14 cells were transfected with ATF4-specific siRNA, then 16 h later cells were treated or not with bafilomycin (20 nM) for 2 h before staining for LC3B. Representative confocal pictures are shown in (i), and the graph in (j) represents the number of LC3B dots per cell. Scale bar: 10 μm. (k) MOLM-14 cells stably expressing ATF4-specific-inducible shRNA were treated with PBS (control) or doxycycline (2 μg/ml) for 2 days. For the final 2 h cells were treated with bafilomycin (20 nM) or not and then processed for western blot analysis of ATF4 and LC3B. Actin was used as a loading control. Numbers represent the LC3B-II/actin ratios obtained by densitometric analysis (n=3±s.e.m.). PBS, phosphate buffered saline.
Taken together, these data highlight a new eIF2α-independent ATF4 regulation pathway and demonstrate for the first time that FLT3-ITD positively controls the ATF4 protein levels.
Given that ATF4 has been described as a regulator of autophagy in other models, these data also suggested that ATF4 could be a key determinant of the autophagy process downstream of FLT3-ITD. To evaluate this hypothesis, RNA interference (siRNA or inducible shRNA) was used to downregulate ATF4 in MOLM-14 or MV4-11 cells. Strikingly, ATF4 depletion induced a significant inhibition of autophagic flux in all cases, as judged by western blot and immunofluorescence analyses (Figures 2h–k and Supplementary Figure S2H).
Together these novel findings demonstrate that the transcription factor ATF4 is an early target of FLT3-ITD signaling and is involved in FLT3-ITD-mediated autophagy in AML cells.
Autophagy sustains the FLT3-ITD-dependent proliferation of leukemic cells
We next asked whether autophagy could support FLT3-ITD-dependent proliferation since FLT3-ITD-expressing leukemic cells have increased rates of proliferation and autophagy has been shown to support proliferation of various types of cancer cells.19, 20 For this, we first investigated the ability of the VPS34 inhibitor SAR405, a recently described potent inhibitor of autophagy,21 to influence MOLM-14 and MV4-11 cell proliferation. Treatment with SAR405 for 24 to 72 h inhibited autophagy (Figure 3a, Supplementary Figure S3A) and severely impaired proliferation in both cell lines, as judged by trypan blue exclusion and cell counting (Figure 3b, Supplementary Figure S3B). Interestingly, SAR405 did not significantly induce apoptotic cell death in MOLM-14 cells even after 3 days of treatment (Figure 3c), suggesting that the inhibition of proliferation observed in response to this molecule may be due to cytostatic mechanisms.
Targeting autophagy inhibits FLT3-ITD AML cells proliferation. (a) MOLM-14 cells were treated with DMSO (control) or the VPS34 inhibitor SAR-405 (3 μM) for 3 days. For the final 2 h cells were treated or not with bafilomycin (20 nM) and were then processed for western blot analysis of LC3B. Actin was used as a loading control. Numbers represent the LC3B-II/actin ratios obtained by densitometric analysis (n=3±s.e.m.). (b) MOLM-14 cells were treated with DMSO (control) or SAR-405 (3 μM) for 3 days, before the number of cells was assessed by trypan blue exclusion counting. The graph represents the number of cells per day (n=3±s.e.m.). (c) MOLM-14 cells were treated with DMSO (control) or SAR-405 (3 μM) for 3 days before cell death was assessed by annexin V labeling and flow cytometry analysis. The graph represents the percentage of annexin V-positive cells (n=3±s.e.m.). (d) MOLM-14 cells stably expressing an ATG12-specific inducible shRNA were treated with PBS (control) or doxycycline (2 μg/ml) for 3 days. For the final 2 h cells were also treated with bafilomycin (20 nM) or not before being processed for western blot analysis of ATG12 and LC3B. Actin was used as a loading control. Numbers represent the LC3B-II/actin ratios obtained by densitometric analysis (n=3±s.e.m.). (e) MOLM-14 cells stably expressing ATG12-specific inducible shRNA were treated with PBS (control) or doxycycline (2 μg/ml) for 3 days before the number of cells was assessed by trypan blue exclusion counting. The graph represents the number of cells per day (n=3±s.e.m.). (f) MOLM-14 cells stably expressing ATG12-specific inducible shRNA were treated with PBS (control) or doxycycline (2 μg/ml) for 3 days before cell death was assessed by annexin V labeling. The graph represents the percentage of dead cells (n=3±s.e.m.). (g) MOLM-14 cells stably expressing ATF4-specific inducible shRNA were treated with DMSO or doxycycline (2 μg/ml) for 3 days before the number of cells was assessed by trypan blue exclusion counting. The graph represents the number of cells per day (n=3±s.e.m.). (h) MOLM-14 cells stably expressing ATF4-specific inducible shRNA were treated with PBS (control) or doxycycline (2 μg/ml) for 3 days before cell death was assessed by annexin V labeling. The graph represents the percentage of dead cells (n=3±s.e.m.). PBS, phosphate buffered saline.
To investigate more specifically the involvement of autophagy in leukemic cell proliferation, we developed MOLM-14 and MV4-11 cell lines conditionally expressing shRNAs against either ATG12 or ATG5, two key proteins in the autophagy process. The addition of doxycycline efficiently silenced the expression of both ATG12 (Figure 3d, Supplementary Figure S3G) and ATG5 (Supplementary Figures S3C and E) and, as expected, reduced autophagic flux. Consistent with our SAR405 data, downregulation of ATG12 or ATG5 also inhibited MOLM-14 and MV4-11 cell proliferation (Figure 3e, Supplementary Figures S3D, F and H) without inducing significant cell death (Figure 3f). As a control, we verified that doxycycline had no effect on the proliferation of MOLM-14 and MV4-11 cell lines (not shown).
We then asked whether autophagy is as equally important for the proliferation of leukemic cells that do not express FLT3-ITD, using OCI-AML3 cells, which express wild-type FLT3. SAR405 treatment did not affect proliferation or death of OCI-AML3 cells (Supplementary Figures S4A and B). In line with this, ATG12 shRNA-mediated inhibition of autophagy in TF-1 cells did not significantly modify their proliferation rate nor induce apoptosis (Supplementary Figures S4C and D), while similar experiments in TF-1-FLT3-ITD cells resulted in a robust inhibition of proliferation (Supplementary Figure S4E) without inducing cell death (Supplementary Figure S4F). Thus, these data demonstrate that FLT3-ITD expressing cells rely on autophagy for growing.
Since ATF4 was found to be involved in autophagy in FLT3-ITD-positive leukemic cells (Figure 2), we then asked whether this factor was necessary for the proliferation of these cells. ATF4 depletion in MOLM-14 cells using an inducible shRNA against ATF4 induced a dramatic reduction in cell proliferation (Figure 3g), indicating that the FLT3-ITD-dependent autophagy that supports cell proliferation requires the presence of the ATF4 transcription factor. Finally, as previously observed for autophagy inhibition in these cells, ATF4 depletion did not induce significant cell death (Figure 3h).
Targeting autophagy or ATF4 reduces AML tumor burden in mice
We next investigated whether autophagy and ATF4 are required for AML proliferation in vivo. MOLM-14 cells stably expressing conditional shRNA against ATG5 or ATF4, or control shRNA, were injected into the tail vein of NOD scid gamma (NSG) mice to generate a leukemic model22 (Figure 4a). Mice were then treated per os with doxycycline to induce shRNA expression. Strikingly, ATG5 or ATF4 silencing strongly decreased total cell tumor burden, as indicated by a reduced percentage of human leukemic cells (hCD45+/hCD33+) present in the murine bone marrow (Figure 4b) and spleen (Figure 4c). In addition, sternums from mice engrafted with shRNA control cells appeared to have a greater invasion of human cells compared to mice engrafted with cells expressing ATG5 or ATF4 shRNA (Figure 4d). Consistently, mouse survival was significantly prolonged upon ATF4 depletion and even more so with ATG5 depletion (Figure 4e). To further validate these results, mice were also engrafted with MOLM-14 cells silenced for another autophagy gene, ATG12, which also greatly improved mice overall survival (Figure 4f).
Targeting autophagy or ATF4 decreases tumor burden and increases survival of mice xenografted with FLT3-ITD AML cells. (a) NSG mice (n=15) engrafted with MOLM-14 cells stably expressing the indicated inducible shRNAs by intravenous (i.v.) injection were treated with sucrose (10 μg/ml) and doxycycline (200 μg/ml) via their drinking water. Twenty-five days post-graft, six mice per group were killed and the number of human cells (hCD45+ and hCD33+) present in the bone marrow and spleen was analyzed by flow cytometry. The remaining nine mice per group were used for overall survival analysis. (b, c) Graphs representing the number of human cells positive for hCD45 and hCD33 within the murine bone marrow (b) and spleen (c). (d) Histochemical MGG (May–Grünwald–Giemsa) staining of sternums of NSG mice engrafted with MOLM-14 cells stably expressing a control inducible shRNA (n=6), an ATG5-specific inducible shRNA (n=6) or an ATF4-specific inducible shRNA (n=6) by i.v. injection. Scale bar: 200 μm. (e) Graph representing the Kaplan–Meier survival curves of mice engrafted as described in (a). (f) NSG mice were engrafted with MOLM-14 cells stably expressing an ATG12-specific inducible shRNA by i.v. injection, and were then treated with sucrose (10 μg/ml) with or without doxycycline (200 μg/ml) via their drinking water. The graph represents the Kaplan–Meier survival curves of mice treated with sucrose (n=7) or sucrose+doxycycline (n=8).
Taken together these results have identified autophagy and ATF4 as key actors in FLT3-ITD-positive AML in this murine xenograft model.
In order to further precise the functional relationship between FLT3-ITD and autophagy, we performed in vitro and in vivo experiments where we combined FLT3-ITD and autophagy inhibition. For this aim, MOLM-14-shATG12 cells were treated with doxycycline, in the presence of the absence of FLT3 inhibitor. As shown Supplementary Figures S5A and B, inhibiting autophagy, or FLT3, or both, induced a similar reduction in cell proliferation, suggesting that FLT3-ITD signaling and autophagy lie in the same signal transduction pathway. We then performed xenograft experiments with MOLM-14-shATG12 cells, and we additionally treated mice with AC-220. In agreement with the in vitro experiments, mice survival was increased to the same extent by AC-220 and doxycycline-induced autophagy inhibition (Supplementary Figure S5C). However, we noticed that inhibiting both autophagy (doxycycline) and FLT3 (AC-220) slightly increased mice survival compared with each inhibition alone. These data suggest that autophagy represents one of the major mechanisms contributing to FLT3-ITD leukemia in this in vivo model, although we do not exclude involvement of other cellular process.
Inhibiting autophagy overcomes acquired resistance to FLT3 inhibitors
Given that we have established autophagy as a necessary process for AML cell proliferation and tumor burden in vivo, we then wondered whether inhibiting autophagy could represent an alternative method of targeting FLT3-ITD-positive cells that are resistant to FLT3 inhibitors. To test this hypothesis, we used a MOLM-14 cell line expressing an FLT3-D835Y mutant (MOLM-14-TKD), a mutation located within the catalytic domain of FLT3, which is well-known for conferring resistance to AC-220. In consequence these cells express both the FLT3-ITD and FLT3-TKD mutants. When MOLM-14-TKD cells were treated with the VPS34 inhibitor SAR405, autophagy and cell proliferation were robustly inhibited (Figures 5a and b), as previously observed in MOLM-14 cells (Figures 3a and b). To further confirm these results, we transduced MOLM-14-TKD cells with conditional shRNA directed against ATG12. In accordance with the data obtained with SAR405, this also significantly reduced the proliferation of these AC-220 resistant cells (Figures 5c and d). Doxycycline had no effect on the proliferation of MOLM-14-TKD cell line (not shown).
Targeting autophagy or ATF4 overcomes resistance to FLT3 inhibitors. (a) MOLM-14 TKD cells were treated with DMSO (control) or SAR405 (a VPS34 inhibitor, 3 μM), for 3 days. For the final 2 h cells were then treated with bafilomycin (20 nM) or not and then processed for western blot analysis of LC3B. Actin was used as a loading control. Numbers represent the LC3B-II/actin ratios obtained by densitometric analysis (n=3±s.e.m.). (b) MOLM-14 TKD cells were treated with DMSO (control) or SAR405 (3 μM) for 3 days before the number of cells was assessed by trypan blue exclusion counting. The graph represents the number of cells per day (n=3±s.e.m.). (c) MOLM-14 TKD cells stably expressing ATG12-specific inducible shRNA were treated with PBS (control) or doxycycline (2 μg/ml) for 3 days. For the final 2 h cells were then treated with bafilomycin (20 nM) or not and processed for western blot analysis of ATG12 and LC3B. Actin was used as a loading control. Numbers represent the LC3B-II/actin ratios obtained by densitometric analysis (n=3±s.e.m.). (d) MOLM-14 TKD cells stably expressing ATG12-specific inducible shRNA were treated with PBS (control) or doxycycline (2 μg/ml) for 3 days before the number of cells was assessed by trypan blue exclusion counting. The graph represents the number of cells per day (n=3±s.e.m.). (e) NSG mice (n=15) engrafted with MOLM-14 TKD cells stably expressing the ATG12 inducible shRNAs by i.v. injection, were treated with sucrose (10 μg/ml) with or without doxycycline (200 μg/ml) via their drinking water and their overall survival was analyzed. The graph represents the Kaplan–Meier survival curves. PBS, phosphate buffered saline.
We next sought to validate these results in vivo. Therefore, NSG mice were engrafted with MOLM14-TKD cells expressing conditional shRNA against ATG12, as performed with MOLM-14 cells (Figure 4). The overall survival of doxycycline-treated mice was greatly improved (Figure 5e) when compared to untreated mice, indicating that targeting autophagy overcomes acquired resistance to FLT3 inhibitors in vivo.
Collectively, these findings demonstrate that an ATF4/autophagy axis constitutes an important determinant of FLT3-ITD AML proliferation in vivo, and could be considered as a new potential therapeutic target for treating this AML subtype.
Discussion
In this work, we demonstrate that the oncogenic receptor tyrosine kinase FLT3-ITD induces autophagy in AML cells. To date, the role of autophagy in AML development has not been assessed in detail, and the autophagic status of AML cells has also been poorly investigated. Low levels of autophagy have been reported in AML samples in comparison to normal hematopoietic cells23 and, very recently, loss of function mutations of autophagic proteins have been described in AML primary samples.24 Taken together, these two studies suggest that mutation-induced defects in autophagy may participate in AML initiation and progression, in association with other mutational events. In contrast, the key autophagy protein ATG7 has been identified as a resistance factor in AML, with specific importance in response to micro-environmental signals.13 Although highly important, all these studies did not explicitly take into account the heterogeneity of AML subtypes, which may lead to substantial diversity in terms of autophagy status in AML cells. Our work has more specifically addressed this question by focusing on the FLT3-ITD AML subtype, which accounts for up to 25% of AML cases and has been associated with a poor prognosis. This does not imply that high basal autophagy is restricted to FLT3-ITD AML, and it is possible that other AML subtypes may also present high autophagy levels, for example due to the expression of oncogenes other than FLT3-ITD. Interestingly, our results are reminiscent of studies performed in other cancers, which have described the induction of autophagy by oncogenic forms of K-Ras6 or B-Raf,7 or by BCR-ABL.8 In these cases, autophagy was found to support the increased metabolic activity and excessive proliferation rates triggered by these different oncogenes. Our present data are in agreement with this notion, but they also suggest that while sustaining the proliferation of FLT3-ITD leukemic cells, autophagy does not significantly impact on their survival. Indeed, inhibiting autophagy genetically or with pharmacological molecules did not significantly induce cell death in FLT3-ITD AML cells. This is somewhat divergent from the roles of autophagy described downstream of B-Raf or K-Ras, which have been implicated in cell survival and resistance mechanisms to cytotoxic or genotoxic drugs mediated by these oncogenes. Interestingly, in the case of other tyrosine kinase-driven hematological malignancies, such as NPM-ALK-positive anaplastic large cell lymphoma or BCR-ABL-positive chronic myeloid leukemia, inhibiting the oncogenic tyrosine kinase is often found to lead to the induction of a protective autophagy,9, 10 contrasting with our present observations in the FLT3-ITD model. It is also important to note that different studies have described the induction of autophagy-dependent cell death in AML cells in response to different drugs, such as the proteasome inhibitor bortezomib22 or trans-retinoic acid in the case of promyelocytic leukemia.25
These previous studies alongside our present data collectively suggest that the autophagic response to expression or inhibition of oncogenic tyrosine kinases may vary from one pathology to the other and from one oncogene to the other, further stressing the necessity for specific mechanistic studies for each model.
In this study, we have also identified the ATF4 transcription factor as an important modulator of autophagy downstream of FLT3-ITD. To our knowledge this is one of the first reports identifying a function for ATF4 in AML cell biology, and the first evidence of its importance downstream of FLT3-ITD. The induction of the ATF4 protein through the activation of the eIF2α-dependent or -independent integrated stress response has been reported recently in AML in response to different drugs,26, 27 and a recent study implicated ATF4 as a mediator of the unfolded protein response downstream of JUN in AML.28 We found that ATF4 is an early target of FLT3-ITD signaling since its downregulation occurs within 1 h of receptor inhibition in model cell lines. Interestingly, this regulation does not involve eIF2α, the main signaling pathway known to induce ATF4 accumulation, or the canonical signaling networks activated downstream of FLT3-ITD, which include the AKT, ERK or STAT5 pathways. ATF4 regulation has been described at the transcriptional, translational and post-translational level, depending on the model.29 Our results suggest that FLT3-ITD facilitates ATF4 synthesis without affecting stability, so further experiments will be needed to identify the mechanism operating downstream of FLT3-ITD. Interestingly, the signaling pathways leading to the induction of autophagy are different depending on the various oncogenes. An endoplasmic reticulum stress- and eIF2α-dependent process has been implicated downstream of B-Raf,7 MAPK15 involvement was described for BCR-ABL1.8 Different studies have described key autophagic proteins, such as LC3B, Beclin1 or ULK1, as transcriptional targets of ATF4 downstream of eIF2α phosphorylation;30 thus the important question of the mechanism involved in ATF4-dependent regulation of autophagy in FLT3-ITD-positive cells remains opened. Our data also demonstrate that inhibiting autophagy or ATF4 induces a cytostatic decrease in FLT3-ITD leukemic cell proliferation. How autophagy and ATF4 sustain the proliferation of these leukemic cells remains to be clarified. Autophagy has been involved in the regulation of different proliferation and cell cycle events,31, 32, 33 and it was also recently reported to regulate key proliferative signaling molecules such as MEK, ERK34 and MAPK15.8 Additionally, both autophagy and ATF4 are important actors and regulators of energetic metabolism in normal and cancer cells, so it could also impact on the proliferation of leukemic cells through these metabolic functions. Further studies will be needed to identify the molecular and functional links that drive autophagy-dependent proliferation in these leukemic cells.
Much effort has been focused on developing efficient inhibitors against the FLT3-ITD receptor since its expression is associated with a poor outcome and since FLT3-ITD leukemic cells are strongly dependent on its activity for their proliferation and resistance to therapeutic drugs. Clinical trials with quizartinib have yielded encouraging results, but acquired resistance to the inhibitor involving different mechanisms have been described.35 In this context, identifying new Achilles heels of leukemic cells expressing FLT3-ITD appears to be the most important challenge in defining new potential therapeutic strategies. Our data suggest that autophagy or ATF4 could represent such alternatives because certain subsets of AML cells appear to be highly dependent on their activity. Furthermore, we have established that inhibiting autophagy may overcome the resistance to quizartinib induced by point mutations in the catalytic domain of FLT3, a resistance mechanism described in AML patients.36 The only autophagy inhibitor that is approved for in vivo use in humans at the moment is chloroquine, which was tested in association with therapeutic drugs for different types of cancers.11 New efficient autophagy inhibitors have recently been described, including inhibitors of the class III PI3K VPS3437 that can be used in vivo. The results herein suggest that these molecules or their derivatives may constitute efficient tools to inhibit autophagy in vivo in the near future.
As a conclusion, we identified for the first time autophagy as an important effector of FLT3-ITD receptor dependence in AML, and we established the ATF4 transcription factor as a master regulator of this process, underlying the existence of a new targetable signaling pathway in this poor prognosis AML subtype. Our data open the interesting possibility that a similar pathway is operating downstream of mutant tyrosine kinase receptors in other types of cancers.
Materials and methods
Cell lines and AML samples
The human myeloid leukemia cell lines MV4-11, MOLM-14 were purchased from the Leibniz Institute DSMZ-German Collection of Microorganisms and Cell Cultures (Leibniz, Germany). The MOLM-14 TKD, TF1-FLT3-ITD and OCI-AML3 cell lines were kindly provided by Jérôme Tamburini (Institut Cochin, Paris, France), Paulo de Sepulveda (CRCM, Marseille, France) and Pierre Brousset (CRCT, Toulouse, France), respectively. Cells were grown in RPMI 1640 medium with Glutamax (Gibco, Life Technologies, Carlsbad, CA, USA) supplemented with 10% fetal bovine serum (Sigma, St Louis, CA, USA) and 100 units/ml of penicillin and streptomycin (Invitrogen, Life Technologies Corp., Carlsbad, CA, USA). Fresh and thawed samples from AML patients have been obtained after informed consent and stored at the HIMIP collection (BB-0033-00060). According to the French law, HIMIP collections has been declared to the Ministry of Higher Education and Research (DC 2008-307 collection 1) and obtained a transfer agreement (AC 2008-129) after approbation by the ‘Comité de Protection des Personnes Sud-Ouest et Outremer II’ (ethical committee). Clinical and biological annotations of the samples have been declared to the CNIL (Comité National Informatique et Libertés, that is, Data processing and Liberties National Committee) and is supported by CAPTOR (Cancer Pharmacology of Toulouse-Oncopole and Region). AML primary samples were cultured in Iscove’s modified Dulbecco medium containing 20% fetal calf serum.
Antibodies and reagents
The following antibodies from Cell Signaling Technology (Danvers, MA, USA) were used: rabbit antibodies against LC3B (#2775), phospho-FLT3 (Tyr591, #3461), phospho-Stat-5 (Tyr694, #9351), Stat-5 (#9363), phospho-AKT (Ser473, #9277), AKT (#9272), ERK (#9102), phospho-eIF2α (Ser51, #9721), eIF2α (#9722), VPS34 (#4263), ATG5 (#8540), ATG12 (#4180); and mouse antibodies against phospho-ERK (#9106). Rabbit antibodies from Santa Cruz Biotechnology (Santa Cruz, CA, USA) against Flt-3/Flk-2 (C-20, sc479) and ATF4 (C-20, sc200) were also used. Secondary antibodies labeled with horseradish peroxidase were purchased from Promega (Madison, WI, USA). Mouse monoclonal anti-LC3B from nanoTOOLS was used for immunofluorescence analysis. For flow cytometry studies, the antibodies anti-hCD45 (YB5.B8), anti-CD33 (HI30), anti-annexin V and BD Via-Probe (BD Pharmingen, San Diego, CA, USA) were used. The FLT3 inhibitor III was purchased from Calbiochem (LaJolla, CA, USA) and the FLT3 inhibitor quizartinib AC220 was purchased from Selleck Chemicals (Selleckchem, Houston, USA). The VPS34 inhibitor SAR405 was purchased from APExBIO (ApexBio, Houston, USA). The JAK2-STAT5 inhibitor (PD 0325901) were purchased from Selleck Chemicals, the AKT inhibitor (AKT Inhibitor VIII) was purchased from Calbiochem. Bafilomycin was purchased from InvivoGen (San Diego, CA, USA) and chloroquine was obtained from Sigma-Aldrich (St Louis, CA, USA).
Western blot analysis
Proteins were separated using 4–12% gradient polyacrylamide SDS–PAGE gels (Life Technologies) and electrotransferred to 0.2 μM nitrocellulose membranes (GE Healthcare, Velizy-Villacoublay, France). After blocking in Tris-buffered saline with 0.1% Tween and 5% bovine serum albumin, membranes were blotted overnight at 4 °C with the appropriate primary antibodies. Primary antibodies were detected using the appropriate horseradish peroxidase-conjugated secondary antibodies. Immunoreactive bands were visualized by enhanced chemiluminescence (PI32209; Thermo Fisher Scientific, Bremen, Germany) with a Syngene camera. Densitometric analyses of immunoblots were performed using the GeneTools software.
Immunofluorescence analysis
Briefly, cells were seeded onto coverslips coated with 0.01% poly-L-lysine (Sigma), then fixed in 4% formaldehyde for 8 min and subjected to ice-cold methanol treatment for 5 min. After phosphate buffered saline (PBS) washes, cells were incubated in 0.1% Triton X-100 in PBS/5% bovine serum albumin for 15 min and then incubated with anti-LC3B antibodies (2 μg/ml) for 45 min. Cells were then washed again before incubation with an anti-mouse Alexa-488 secondary antibody (Invitrogen, Life Technologies Corp.) for 30 min, followed by washing and mounting in medium containing 4′6-diamidino-2-phenylindole (Invitrogen, Life Technologies Corp.). Images were acquired using a confocal Zeiss LSM 780. For quantification, fields were chosen arbitrarily based on 4′6-diamidino-2-phenylindole staining, and the number of LC3B dots per cell of at least 50 cells was determined with ImageJ software.
Flow cytometry
To determine the number of apoptotic and dead cells, cells were washed then resuspended in annexin V binding buffer (BD Pharmingen). Annexin V-APC (5 μl per 100 000 cells) was added and cells were incubated for 10 min in the dark. Data were acquired on a Macsquant (Miltenyi Biotec, Paris, France) flow cytometer. To analyze autophagosome-associated LC3B, we used the Millipore’s FlowCellect Autophagy LC3 Antibody-based Assay Kit, following the manufacturer’s instructions.
SiRNA transfection
The transfection of small interfering RNA (siRNA) into MOLM-14 and MV4-11 cells was carried out using the Neon transfection system (Life technologies), according to the manufacturer’s recommendations. Briefly, 70 nM of siRNA was used per condition. RNA interference-mediated gene knockdown was achieved using pre-validated Santa Cruz Biotechnology siRNAs for ATF4 (sc-35112), Qiagen small interfering RNAs (siRNAs) for FLT3 (SI0005987) and Dharmacon (Chicago, IL, USA) siGENOME Control Pool Non-Targeting #2 (D-001206-14-20).
Lentiviral infection of MOLM-14 and MOLM-14 TKD cells
Lentiviral particles were generated by calcium phosphate transient transfection into 293T cells. Briefly, 293T cells seeded into a 10-cm dish were transfected with 62.5 μl CaCl2 (2 M), 500 μl HBS 2X, 418 μl H2O, 3.5 μg pVSV-G (env), 6.5 μg p8.1 (tat, pol, rev, gag) and 10 μg of inducible shRNA against ATG5, ATG12 or VPS34. Then, 72 h after cell transfection, 2 ml of the supernatants containing the virus particles were collected and added to 2.106 MOLM-14 or MOLM-14 TKD cells in a six-well plate. Polybrene (8 μg/ml) was added and spinoculation was performed by centrifuging cells for 60 min at 800 g (QIAGEN Inc., Hilden, Germany). Seventy-two hours after transduction, the medium containing the viruses was removed. Then, after an additional 24 h, transduced cells were selected with 1 μg/ml puromycin. All shRNA experiments were performed on the cell bulk, treated or not for 72 h with 1 μg/ml doxycycline for shRNA induction. The following sequences were used 5′>3′:
-
shRNA ATG12: purchased from Dharmacon (Ref RHS4696-20075354) (CCGGGGACTCATTGACTTCATCACTCGAGTGATGAAGTCAATGAGTCC).
-
shRNA CTL: purchased from Sigma (Ref HC016 MISSION pLKO.1-puro Non-Target shRNA Control) (CGGGCGCGATAGCGCTAATAATTTCTCGAGAAATTATTAGCGCTATCGCGCTTTTTG).
-
shRNA ATG5: purchased from Sigma (custom shRNA) (CCGGCCTGAACAGAATCATCCTTAACTCGAGTTAAGGATGATTCTGTTCAGGTTTTTTG).
-
shRNA ATF4: purchased from Sigma (custom shRNA) (CCGGGCCTAGGTCTCTTAGATGATTCTCGAGAATCATCTAAGAGACCTAGGCTTTTT).
Tumor xenografts into NOD SCID gamma (NSG) mice
NSG mice were bred at the UMS006 in Toulouse (France) using breeders obtained from Charles River. All animal experimental protocols were approved by the institutional Animal Care and Use Ethical Committee of the UMS006 and the Région Midi-Pyrénées (approval #13-U1037-JES08). NSG mice were treated by an i.p. injection of busulfan (20 mg/kg) on the day before the experiment. Mice were engrafted by injection of 2.106 MOLM-14 shCTL, MOLM-14 shATG5 or MOLM-14 shATF4 cells into the tail vein. After injection, all mice were treated by adding doxycycline (200 μg/ml) and sucrose (10 μg/ml) to the drinking water. After 25 days, mice were killed for human cell engraftment analysis, and the other ones were kept for overall survival evaluation. For establishing human cell engraftment, either mixed bone marrow from tibias and femurs or spleens were crushed into Hanks' Balanced Salt Solution with 1% fetal bovine serum, and then washed with PBS. The number of human cells (hCD33+/hCD45+ cells) in the bone marrow and spleen was then evaluated by flow cytometry (LSRII Fortessa). For MOLM-14 shATG12 and MOLM-14/TKD shATG12 in vivo experiments, 2.106 cells were injected into the tail vein of the NSG mice. After injection, mice were randomly split into two groups: one group was treated with 200 μg/ml doxycycline and 10 μg/ml sucrose in the drinking water (n=8) and the other only with sucrose 10 μg/ml (n=7). Overall mouse survival was established under these conditions.
For the other MOLM-14 shATG12 in vivo experiments, 2.106 cells were injected into the tail vein of the NSG mice. After injection, mice were randomly split into four groups: one group was treated with 200 μg/ml doxycycline plus 10 μg/ml sucrose in the drinking water and treated with vehicle (5% Cyclodextrin in distilled water) via oral gavage (n=7), one group was treated only with sucrose 10 μg/ml and treated with vehicle (5% Cyclodextrin in distilled water) via oral gavage (n=7), one group was treated with 200 μg/ml doxycycline plus 10 μg/ml sucrose in the drinking water and treated with vehicle (5% Cyclodextrin plus AC220 5 mg/kg in distilled water) via oral gavage (n=7), one group was treated only with sucrose 10 μg/ml and treated with vehicle (5% Cyclodextrin plus AC220 5 mg/kg in distilled water) via oral gavage (n=7). Overall mouse survival was established under these conditions.
Statistical analysis
Data from three independent experiments are reported as means±standard error of the mean. Statistical analyses were performed using unpaired two-tailed Student t-tests with Prism 5 software (GraphPad Software Inc). Differences were considered as significant for P-values <0.05. *P<0.05; **P<0.01; ***P<0.001. The Kaplan–Meier method was used to estimate leukemia-free survival in xenografted mice. Log-rank P-values were used for comparisons of leukemia-free survival among the three subgroups.
References
Döhner H, Weisdorf DJ, Bloomfield CD . Acute myeloid leukemia. N Engl J Med 2015; 373: 1136–1152.
Papaemmanuil E, Gerstung M, Bullinger L, Gaidzik VI, Paschka P, Roberts ND et al. Genomic classification and prognosis in acute myeloid leukemia. N Engl J Med 2016; 374: 2209–2221.
Kayser S, Schlenk RF, Londono MC, Breitenbuecher F, Wittke K, Du J et al. Insertion of FLT3 internal tandem duplication in the tyrosine kinase domain-1 is associated with resistance to chemotherapy and inferior outcome. Blood 2009; 114: 2386–2392.
Gallogly MM, Lazarus HM . Midostaurin: an emerging treatment for acute myeloid leukemia patients. J Blood Med 2016; 7: 73–83.
White E . The role for autophagy in cancer. J Clin Invest 2015; 125: 42–46.
Guo JY, Chen HY, Mathew R, Fan J, Strohecker AM, Karsli-Uzunbas G et al. Activated Ras requires autophagy to maintain oxidative metabolism and tumorigenesis. Genes Dev 2011; 25: 460–470.
Corazzari M, Rapino F, Ciccosanti F, Giglio P, Antonioli M, Conti B et al. Oncogenic BRAF induces chronic ER stress condition resulting in increased basal autophagy and apoptotic resistance of cutaneous melanoma. Cell Death Differ 2015; 22: 946–958.
Colecchia D, Rossi M, Sasdelli F, Sanzone S, Strambi A, Chiariello M . MAPK15 mediates BCR-ABL1-induced autophagy and regulates oncogene-dependent cell proliferation and tumor formation. Autophagy 2015; 11: 1790–1802.
Bellodi C, Lidonnici MR, Hamilton A, Helgason GV, Soliera AR, Ronchetti M et al. Targeting autophagy potentiates tyrosine kinase inhibitor-induced cell death in Philadelphia chromosome-positive cells, including primary CML stem cells. J Clin Invest 2009; 119: 1109–1123.
Mitou G, Frentzel J, Desquesnes A, Le Gonidec S, AlSaati T, Beau I et al. Targeting autophagy enhances the anti-tumoral action of crizotinib in ALK-positive anaplastic large cell lymphoma. Oncotarget 2015; 6: 30149–30164.
Rebecca VW, Amaravadi RK . Emerging strategies to effectively target autophagy in cancer. Oncogene 2016; 35: 1–11.
Bosnjak M, Ristic B, Arsikin K, Mircic A, Suzin-Zivkovic V, Perovic V et al. Inhibition of mTOR-dependent autophagy sensitizes leukemic cells to cytarabine-induced apoptotic death. PLoS One 2014; 9: e94374.
Piya S, Kornblau SM, Ruvolo VR, Mu H, Ruvolo PP, McQueen T et al. Atg7 suppression enhances chemotherapeutic agent sensitivity and overcomes stroma-mediated chemoresistance in acute myeloid leukemia. Blood 2016; 128: 1260–1269.
Gammoh N, Lam D, Puente C, Ganley I, Marks PA, Jiang X . Role of autophagy in histone deacetylase inhibitor-induced apoptotic and nonapoptotic cell death. Proc Natl Acad Sci USA 2012; 109: 6561–6565.
Torgersen ML, Engedal N, Bøe SO, Hokland P, Simonsen A . Targeting autophagy potentiates the apoptotic effect of histone deacetylase inhibitors in t(8;21) AML cells. Blood 2013; 122: 2467–2476.
Choudhary C, Olsen JV, Brandts C, Cox J, Reddy PN, Böhmer FD et al. Mislocalized activation of oncogenic RTKs switches downstream signaling outcomes. Mol Cell 2009; 36: 326–339.
B'chir W, Maurin AC, Carraro V, Averous J, Jousse C, Muranishi Y et al. The eIF2α/ATF4 pathway is essential for stress-induced autophagy gene expression. Nucleic Acids Res 2013; 41: 7683–7699.
Ameri K, Harris AL . Activating transcription factor 4. Int J Biochem Cell Biol 2008; 40: 14–21.
Lefort S, Joffre C, Kieffer Y, Givel AM, Bourachot B, Zago G et al. Inhibition of autophagy as a new means of improving chemotherapy efficiency in high-LC3B triple-negative breast cancers. Autophagy 2014; 10: 2122–2142.
Yang S, Wang X, Contino G, Liesa M, Sahin E, Ying H et al. Pancreatic cancers require autophagy for tumor growth. Genes Dev 2011; 25: 717–729.
Ronan B, Flamand O, Vescovi L, Dureuil C, Durand L, Fassy F et al. A highly potent and selective Vps34 inhibitor alters vesicle trafficking and autophagy. Nat Chem Biol 2014; 10: 1013–1019.
Larrue C, Saland E, Boutzen H, Vergez F, David M, Joffre C et al. Proteasome inhibitors induce FLT3-ITD degradation through autophagy in AML cells. Blood 2016; 127: 882–892.
Watson AS, Riffelmacher T, Stranks A, Williams O, De Boer J, Cain K et al. Autophagy limits proliferation and glycolytic metabolism in acute myeloid leukemia. Cell Death Discov 2015; 1: 15008.
Visconte V, Przychodzen B, Han Y, Nawrocki ST, Thota S, Kelly KR et al. Complete mutational spectrum of the autophagy interactome: A novel class of tumor suppressor genes in myeloid neoplasms. Leukemia 2016; 31: 505–510.
Isakson P, Bjørås M, Bøe SO, Simonsen A . Autophagy contributes to therapy-induced degradation of the PML/RARA oncoprotein. Blood 2010; 116: 2324–2331.
Sujobert P, Poulain L, Paubelle E, Zylbersztejn F, Grenier A, Lambert M et al. Co-activation of AMPK and mTORC1 induces cytotoxicity in acute myeloid leukemia. Cell Rep 2015; 11: 1446–1457.
Ishizawa J, Kojima K, Chachad D, Ruvolo P, Ruvolo V, Jacamo RO et al. ATF4 induction through an atypical integrated stress response to ONC201 triggers p53-independent apoptosis in hematological malignancies. Sci Signal 2016; 9: ra17.
Zhou C, Martinez E, Di Marcantonio D, Patel-Solanki N, Aghayev T, Peri S et al. JUN is a key transcriptional regulator of the unfolded protein response in acute myeloid leukemia. Leukemia 2017; 31: 1196–1205.
Singleton DC, Harris AL . Targeting the ATF4 pathway in cancer therapy. Expert Opin Ther Targets 2012; 16: 1189–1202.
Pakos-Zebrucka K, Koryga I, Mnich K, Ljujic M, Samali A, Gorman AM . The integrated stress response. EMBO Rep 2016; 17: 1374–1395.
Doménech E, Maestre C, Esteban-Martínez L, Partida D, Pascual R, Fernández-Miranda G et al. AMPK and PFKFB3 mediate glycolysis and survival in response to mitophagy during mitotic arrest. Nat Cell Biol 2015; 17: 1304–1316.
Lee IH, Kawai Y, Fergusson MM, Rovira II, Bishop AJ, Motoyama N et al. Atg7 modulates p53 activity to regulate cell cycle and survival during metabolic stress. Science 2012; 336: 225–228.
Jia W, He MX, McLeod IX, Guo J, Ji D, He YW . Autophagy regulates T lymphocyte proliferation through selective degradation of the cell-cycle inhibitor CDKN1B/p27Kip1. Autophagy 2015; 11: 2335–2345.
Martinez-Lopez N, Athonvarangkul D, Mishall P, Sahu S, Singh R . Autophagy proteins regulate ERK phosphorylation. Nat Commun 2013; 4: 2799.
Kiyoi H . Flt3 inhibitors: recent advances and problems for clinical application. Nagoya J Med Sci 2015; 77: 7–17.
Alvarado Y, Kantarjian HM, Luthra R, Ravandi F, Borthakur G, Garcia-Manero G et al. Treatment with FLT3 inhibitor in patients with FLT3-mutated acute myeloid leukemia is associated with development of secondary FLT3-tyrosine kinase domain mutations. Cancer 2014; 120: 2142–2149.
Pasquier B, El-Ahmad Y, Filoche-Rommé B, Dureuil C, Fassy F, Abecassis PY et al. Discovery of (2S)-8-[(3R)-3-methylmorpholin-4-yl]-1-(3-methyl-2-oxobutyl)-2-(trifluoromethyl)-3,4-dihydro-2H-pyrimido[1,2-a]pyrimidin-6-one: a novel potent and selective inhibitor of Vps34 for the treatment of solid tumors. J Med Chem 2015; 58: 376–400.
Acknowledgements
We are very grateful to Jérôme Tamburini (Institut Cochin, Paris, France), Paulo De Sepulveda (CRCM, Marseille, France) and Pierre Brousset (CRCT, Toulouse, France) for providing us with specific leukemic cell lines (MOLM-14-TKD and MV4-11 shFLT3, TF-1-FLT3-ITD, and OCI-AML3 respectively). We thank Florence Capilla for performing and managing histochemistry experiments, and we acknowledge excellent technical assistance of Mathilde Gotanegre. We are grateful to Manon Farcé and Laetitia Ligat for assistance with flow cytometry and microscopy, respectively. We thank HIMIP collection for patients’ samples. This work was supported by the ANR LABEX TOUCAN, the Cancéropole GSO, la Ligue régionale contre le Cancer and la Ligue nationale contre le Cancer. Quentin Heydt is a recipient of the Ligue Nationale contre le Cancer.
Author contributions
VM-DM, CJ, SM and J-ES conceived and designed the experiments, QH, CL, ES and CJ performed the experiments, VM-DM, CJ and SM wrote the manuscript.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies this paper on the Oncogene website
Rights and permissions
This work is licensed under a Creative Commons Attribution-NonCommercial-NoDerivs 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-nd/4.0/
About this article
Cite this article
Heydt, Q., Larrue, C., Saland, E. et al. Oncogenic FLT3-ITD supports autophagy via ATF4 in acute myeloid leukemia. Oncogene 37, 787–797 (2018). https://doi.org/10.1038/onc.2017.376
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/onc.2017.376
This article is cited by
-
Crosstalk between autophagy and metabolism: implications for cell survival in acute myeloid leukemia
Cell Death Discovery (2024)
-
Structural and non-structural proteins in SARS-CoV-2: potential aspects to COVID-19 treatment or prevention of progression of related diseases
Cell Communication and Signaling (2023)
-
Targeting PP2A-dependent autophagy enhances sensitivity to ruxolitinib in JAK2V617F myeloproliferative neoplasms
Blood Cancer Journal (2023)
-
Targeting the integrated stress response in hematologic malignancies
Experimental Hematology & Oncology (2022)
-
Potential role of autophagy induced by FLT3-ITD and acid ceramidase in acute myeloid leukemia chemo-resistance: new insights
Cell Communication and Signaling (2022)