Abstract
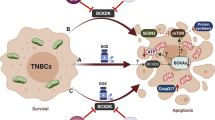
DDX3 is a DEAD box RNA helicase with oncogenic properties. RK-33 is developed as a small-molecule inhibitor of DDX3 and showed potent radiosensitizing activity in preclinical tumor models. This study aimed to assess DDX3 as a target in breast cancer and to elucidate how RK-33 exerts its anti-neoplastic effects. High DDX3 expression was present in 35% of breast cancer patient samples and correlated with markers of aggressiveness and shorter survival. With a quantitative proteomics approach, we identified proteins involved in the mitochondrial translation and respiratory electron transport pathways to be significantly downregulated after RK-33 or DDX3 knockdown. DDX3 localized to the mitochondria and DDX3 inhibition with RK-33 reduced mitochondrial translation. As a consequence, oxygen consumption rates and intracellular ATP concentrations decreased and reactive oxygen species (ROS) increased. RK-33 antagonized the increase in oxygen consumption and ATP production observed after exposure to ionizing radiation and reduced DNA repair. Overall, we conclude that DDX3 inhibition with RK-33 causes radiosensitization in breast cancer through inhibition of mitochondrial translation, which results in reduced oxidative phosphorylation capacity and increased ROS levels, culminating in a bioenergetic catastrophe.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 50 print issues and online access
$259.00 per year
only $5.18 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
References
Tanner NK, Linder P . DExD/H box RNA helicases: from generic motors to specific dissociation functions. Mol Cell 2001; 8: 251–262.
Geissler R, Golbik RP, Behrens SE . The DEAD-box helicase DDX3 supports the assembly of functional 80 S ribosomes. Nucleic Acids Res 2012; 40: 4998–5011.
Botlagunta M, Vesuna F, Mironchik Y, Raman A, Lisok A, Winnard P Jr et al. Oncogenic role of DDX3 in breast cancer biogenesis. Oncogene 2008; 27: 3912–3922.
Bol GM, Vesuna F, Xie M, Zeng J, Aziz K, Gandhi N et al. Targeting DDX3 with a small molecule inhibitor for lung cancer therapy. EMBO Mol Med 2015; 7: 648–669.
Heerma van Voss MR, Vesuna F, Trumpi K, Brilliant J, Berlinicke C, de Leng W et al. Identification of the DEAD box RNA helicase DDX3 as a therapeutic target in colorectal cancer. Oncotarget 2015; 6: 28312–28326.
Wilky BA, Kim C, McCarty G, Montgomery EA, Kammers K, DeVine LR et al. RNA helicase DDX3: a novel therapeutic target in Ewing sarcoma. Oncogene 2016; 35: 2574–2583.
Xie M, Vesuna F, Tantravedi S, Bol GM, Heerma van Voss MR, Nugent K et al. RK-33 radiosensitizes prostate cancer cells by blocking the RNA helicase DDX3. Cancer Res 2016; 76: 6340–6350.
Li Y, Wang H, Wang Z, Makhija S, Buchsbaum D, LoBuglio A et al. Inducible resistance of tumor cells to tumor necrosis factor-related apoptosis-inducing ligand receptor 2-mediated apoptosis by generation of a blockade at the death domain function. Cancer Res 2006; 66: 8520–8528.
Shih JW, Wang WT, Tsai TY, Kuo CY, Li HK, Wu Lee YH . Critical roles of RNA helicase DDX3 and its interactions with eIF4E/PABP1 in stress granule assembly and stress response. Biochem J 2012; 441: 119–129.
Sun M, Song L, Zhou T, Gillespie GY, Jope RS . The role of DDX3 in regulating Snail. Biochim Biophys Acta 2011; 1813: 438–447.
Hagerstrand D, Tong A, Schumacher SE, Ilic N, Shen RR, Cheung HW et al. Systematic interrogation of 3q26 identifies TLOC1 and SKIL as cancer drivers. Cancer Discov 2013; 3: 1044–1057.
Chen HH, Yu HI, Cho WC, Tarn WY . DDX3 modulates cell adhesion and motility and cancer cell metastasis via Rac1-mediated signaling pathway. Oncogene 2015; 34: 2790–2800.
Kondaskar A, Kondaskar S, Kumar R, Fishbein JC, Muvarak N, Lapidus RG et al. Novel, broad spectrum anti-cancer agents containing the tricyclic 5:7:5-fused diimidazodiazepine ring system. ACS Med Chem Lett 2010; 2: 252–256.
Vellinga TT, Borovski T, de Boer VC, Fatrai S, van Schelven S, Trumpi K et al. SIRT1/PGC1alpha-dependent increase in oxidative phosphorylation supports chemotherapy resistance of colon cancer. Clin Cancer Res 2015; 21: 2870–2879.
Viale A, Corti D, Draetta GF . Tumors and mitochondrial respiration: a neglected connection. Cancer Res 2015; 75: 3685–3686.
LeBleu VS, O'Connell JT, Gonzalez Herrera KN, Wikman H, Pantel K, Haigis MC et al. PGC-1alpha mediates mitochondrial biogenesis and oxidative phosphorylation in cancer cells to promote metastasis. Nat Cell Biol 2014; 16: 1001–1015.
Lu CL, Qin L, Liu HC, Candas D, Fan M, Li JJ . Tumor cells switch to mitochondrial oxidative phosphorylation under radiation via mTOR-mediated hexokinase II inhibition—a Warburg-reversing effect. PLoS One 2015; 10: e0121046.
Qin L, Fan M, Candas D, Jiang G, Papadopoulos S, Tian L et al. CDK1 enhances mitochondrial bioenergetics for radiation-induced DNA repair. Cell Rep 2015; 13: 2056–2063.
Croft D, Mundo AF, Haw R, Milacic M, Weiser J, Wu G et al. The Reactome pathway knowledgebase. Nucleic Acids Res 2014; 42: D472–D477.
Franceschini A, Szklarczyk D, Frankild S, Kuhn M, Simonovic M, Roth A et al. STRING v9.1: protein-protein interaction networks, with increased coverage and integration. Nucleic Acids Res 2013; 41: D808–D815.
Li Y, D'Aurelio M, Deng JH, Park JS, Manfredi G, Hu P et al. An assembled complex IV maintains the stability and activity of complex I in mammalian mitochondria. J Biol Chem 2007; 282: 17557–17562.
Acin-Perez R, Bayona-Bafaluy MP, Fernandez-Silva P, Moreno-Loshuertos R, Perez-Martos A, Bruno C et al. Respiratory complex III is required to maintain complex I in mammalian mitochondria. Mol Cell 2004; 13: 805–815.
Zhang X, Fryknas M, Hernlund E, Fayad W, De Milito A, Olofsson MH et al. Induction of mitochondrial dysfunction as a strategy for targeting tumour cells in metabolically compromised microenvironments. Nat Commun 2014; 5: 3295.
Pelicano H, Zhang W, Liu J, Hammoudi N, Dai J, Xu RH et al. Mitochondrial dysfunction in some triple-negative breast cancer cell lines: role of mTOR pathway and therapeutic potential. Breast Cancer Res 2014; 16: 434.
Chen Y, McMillan-Ward E, Kong J, Israels SJ, Gibson SB . Oxidative stress induces autophagic cell death independent of apoptosis in transformed and cancer cells. Cell Death Differ 2008; 15: 171–182.
Youle RJ, Narendra DP . Mechanisms of mitophagy. Nat Rev Mol Cell Biol 2011; 12: 9–14.
Huang Z, Zhou L, Chen Z, Nice EC, Huang C . Stress management by autophagy: implications for chemoresistance. Int J Cancer 2016; 139: 23–32.
Eiermann W, Vallis KA . Locoregional treatments for triple-negative breast cancer. Ann Oncol 2012; 23 (Suppl 6): vi30–vi34.
Oshiumi H, Sakai K, Matsumoto M, Seya T . DEAD/H BOX 3 (DDX3) helicase binds the RIG-I adaptor IPS-1 to up-regulate IFN-beta-inducing potential. Eur J Immunol 2010; 40: 940–948.
Antonicka H, Shoubridge EA . Mitochondrial RNA granules are centers for posttranscriptional RNA processing and ribosome biogenesis. Cell Rep 2015; 10: 920–932.
Tu YT, Barrientos A . The human mitochondrial DEAD-Box protein DDX28 resides in RNA granules and functions in mitoribosome assembly. Cell Rep 2015; 10: 854–864.
Shih JW, Tsai TY, Chao CH, Wu Lee YH . Candidate tumor suppressor DDX3 RNA helicase specifically represses cap-dependent translation by acting as an eIF4E inhibitory protein. Oncogene 2008; 27: 700–714.
Lee CS, Dias AP, Jedrychowski M, Patel AH, Hsu JL, Reed R . Human DDX3 functions in translation and interacts with the translation initiation factor eIF3. Nucleic Acids Res 2008; 36: 4708–4718.
Abaeva IS, Marintchev A, Pisareva VP, Hellen CU, Pestova TV . Bypassing of stems versus linear base-by-base inspection of mammalian mRNAs during ribosomal scanning. EMBO J 2011; 30: 115–129.
Soto-Rifo R, Rubilar PS, Limousin T, de Breyne S, Decimo D, Ohlmann T . DEAD-box protein DDX3 associates with eIF4F to promote translation of selected mRNAs. EMBO J 2012; 31: 3745–3756.
Fukumura J, Noguchi E, Sekiguchi T, Nishimoto T . A temperature-sensitive mutant of the mammalian RNA helicase, DEAD-BOX X isoform, DBX, defective in the transition from G1 to S phase. J Biochem 2003; 134: 71–82.
Lai MC, Lee YH, Tarn WY . The DEAD-box RNA helicase DDX3 associates with export messenger ribonucleoproteins as well as tip-associated protein and participates in translational control. Mol Biol Cell 2008; 19: 3847–3858.
Lai MC, Chang WC, Shieh SY, Tarn WY . DDX3 regulates cell growth through translational control of cyclin E1. Mol Cell Biol 2010; 30: 5444–5453.
Tarn WY, Chang TH . The current understanding of Ded1p/DDX3 homologs from yeast to human. RNA Biol 2009; 6: 17–20.
Padmanabhan PK, Zghidi-Abouzid O, Samant M, Dumas C, Aguiar BG, Estaquier J et al. DDX3 DEAD-box RNA helicase plays a central role in mitochondrial protein quality control in Leishmania. Cell Death Dis 2016; 7: e2406.
Chen CY, Chan CH, Chen CM, Tsai YS, Tsai TY, Wu Lee YH et al. Targeted inactivation of murine Ddx3x: essential roles of Ddx3x in placentation and embryogenesis. Hum Mol Genet 2016; 25: 2905–2922.
Koppenol WH, Bounds PL, Dang CV . Otto Warburg's contributions to current concepts of cancer metabolism. Nat Rev Cancer 2011; 11: 325–337.
Weinberg SE, Chandel NS . Targeting mitochondria metabolism for cancer therapy. Nat Chem Biol 2015; 11: 9–15.
Sansone P, Ceccarelli C, Berishaj M, Chang Q, Rajasekhar VK, Perna F et al. Self-renewal of CD133(hi) cells by IL6/Notch3 signalling regulates endocrine resistance in metastatic breast cancer. Nat Commun 2016; 7: 10442.
Jones RA, Robinson TJ, Liu JC, Shrestha M, Voisin V, Ju Y et al. RB1 deficiency in triple-negative breast cancer induces mitochondrial protein translation. J Clin Invest 2016; 126: 3739–3757.
Farnie G, Sotgia F, Lisanti MP . High mitochondrial mass identifies a sub-population of stem-like cancer cells that are chemo-resistant. Oncotarget 2015; 6: 30472–30486.
Matassa DS, Amoroso MR, Lu H, Avolio R, Arzeni D, Procaccini C et al. Oxidative metabolism drives inflammation-induced platinum resistance in human ovarian cancer. Cell Death Differ 2016; 23: 1542–1554.
Qin B, Minter-Dykhouse K, Yu J, Zhang J, Liu T, Zhang H et al. DBC1 functions as a tumor suppressor by regulating p53 stability. Cell Rep 2015; 10: 1324–1334.
Wardman P, Anderson RF, Hodgkiss RJ, Parrick J, Smithen CE, Wallace RG et al. Radiosensitization by non-nitro compounds. Int J Radiat Oncol Biol Phys 1982; 8: 399–401.
Zhang X, Zhou X, Chen R, Zhang H . Radiosensitization by inhibiting complex I activity in human hepatoma HepG2 cells to X-ray radiation. J Radiat Res 2012; 53: 257–263.
Skrtic M, Sriskanthadevan S, Jhas B, Gebbia M, Wang X, Wang Z et al. Inhibition of mitochondrial translation as a therapeutic strategy for human acute myeloid leukemia. Cancer Cell 2011; 20: 674–688.
Moelans CB, de Weger RA, van Blokland MT, Ezendam C, Elshof S, Tilanus MG et al. HER-2/neu amplification testing in breast cancer by multiplex ligation-dependent probe amplification in comparison with immunohistochemistry and in situ hybridization. Cell Oncol 2009; 31: 1–10.
Carey LA, Perou CM, Livasy CA, Dressler LG, Cowan D, Conway K et al. Race, breast cancer subtypes, and survival in the Carolina Breast Cancer Study. JAMA 2006; 295: 2492–2502.
Vermeulen JF, van de Ven RA, Ercan C, van der Groep P, van der Wall E, Bult P et al. Nuclear Kaiso expression is associated with high grade and triple-negative invasive breast cancer. PLoS One 2012; 7: e37864.
The Medical Research Involving Human Subjects Act [In Dutch: Wet medisch-wetenschappelijk onderzoek met mensen, WMO]. Burgerlijk Wetboek, Staatsblad van het Koninkrijk der Nederlanden, 1998. Available at: http://wetten.overheid.nl/BWBR0009408/2017-03-01.
Bol GM, Raman V, van der Groep P, Vermeulen JF, Patel AH, van der Wall E et al. Expression of the RNA helicase DDX3 and the hypoxia response in breast cancer. PLoS One 2013; 8: e63548.
Angus AG, Dalrymple D, Boulant S, McGivern DR, Clayton RF, Scott MJ et al. Requirement of cellular DDX3 for hepatitis C virus replication is unrelated to its interaction with the viral core protein. J Gen Virol 2010; 91: 122–132.
Chambers AF . MDA-MB-435 and M14 cell lines: identical but not M14 melanoma? Cancer Res 2009; 69: 5292–5293.
Kammers K, Cole RN, Tiengwe C, Ruczinski I . Detecting significant changes in protein abundance. EuPA Open Proteom 2015; 7: 11–19.
Smyth GK . Linear models and empirical bayes methods for assessing differential expression in microarray experiments. Stat Appl Genet Mol Biol 2004; 3: Article 3.
Chen EY, Tan CM, Kou Y, Duan Q, Wang Z, Meirelles GV et al. Enrichr: interactive and collaborative HTML5 gene list enrichment analysis tool. BMC Bioinformatics 2013; 14: 128.
Leary SC, Sasarman F . Oxidative phosphorylation: synthesis of mitochondrially encoded proteins and assembly of individual structural subunits into functional holoenzyme complexes. Methods Mol Biol 2009; 554: 143–162.
Rasband WS . ImageJ 2015. U.S. National Institutes of Health: Bethesda, MD, USA.
Acknowledgements
We thank Bob Cole, Tatiana Boronina and Bob O’ Meally of the Johns Hopkins Mass Spectrometry and Proteomics core facility for their help with the proteomics experiments; Tri Nguyen for his help with interpretation of the electron microscopy images; the Dawson Laboratory at the Johns Hopkins School of Medicine for their help with the mitochondrial translation assay; and Beth Rodgers, who kindly provided us with S35-methionine. This work was financially supported by Utrecht University Alexandre Suerman Stipend (MRHvV), the Dutch Cancer Foundation (UU2013-5851; MRHvV), NIH (R01CA166348 to PTT, R01CA193895 to AL, and R01CA140226 and R01CA131250 to VR), FAMRI (VR) and Safeway (VR).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
VR and PTT have received a patent for the use of RK-33 as a radiosensitizer (US 8,518,901). VR, GMB and PJvD have received a patent for the use of DDX3 as a cancer biomarker (US 9,322,831). PJvD, PTT and VR are on the advisory board of Natsar Pharmaceuticals.
Additional information
Supplementary Information accompanies this paper on the Oncogene website
Supplementary information
Rights and permissions
About this article
Cite this article
Heerma van Voss, M., Vesuna, F., Bol, G. et al. Targeting mitochondrial translation by inhibiting DDX3: a novel radiosensitization strategy for cancer treatment. Oncogene 37, 63–74 (2018). https://doi.org/10.1038/onc.2017.308
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/onc.2017.308
This article is cited by
-
Molecular docking, synthesis, and biological evaluation of 7-azaindole-derivative (7AID) as novel anti-cancer agent and potent DDX3 inhibitor:—an in silico and in vitro approach
Medical Oncology (2022)
-
DDX3X: structure, physiologic functions and cancer
Molecular Cancer (2021)
-
Selective cell death in HIV-1-infected cells by DDX3 inhibitors leads to depletion of the inducible reservoir
Nature Communications (2021)
-
Cytoplasmic DDX3 as prognosticator in male breast cancer
Virchows Archiv (2021)
-
The DEAD-box protein family of RNA helicases: sentinels for a myriad of cellular functions with emerging roles in tumorigenesis
International Journal of Clinical Oncology (2021)