Abstract
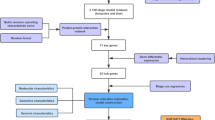
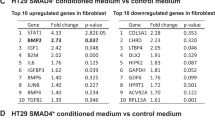
Gene expression-based classification systems have identified an aggressive colon cancer subtype with mesenchymal features, possibly reflecting epithelial-to-mesenchymal transition (EMT) of tumor cells. However, stromal fibroblasts contribute extensively to the mesenchymal phenotype of aggressive colon tumors, challenging the notion of tumor EMT. To separately study the neoplastic and stromal compartments of colon tumors, we have generated a stroma gene filter (SGF). Comparative analysis of stromahigh and stromalow tumors shows that the neoplastic cells in stromahigh tumors express specific EMT drivers (ZEB2, TWIST1, TWIST2) and that 98% of differentially expressed genes are strongly correlated with them. Analysis of differential gene expression between mesenchymal and epithelial cancer cell lines revealed that hepatocyte nuclear factor 4α (HNF4α), a transcriptional activator of intestinal (epithelial) differentiation, and its target genes are highly expressed in epithelial cancer cell lines. However, mesenchymal-type cancer cell lines expressed only part of the mesenchymal genes expressed by tumor-derived neoplastic cells, suggesting that external cues were lacking. We found that collagen-I dominates the extracellular matrix in aggressive colon cancer. Mimicking the tumor microenvironment by replacing laminin-rich Matrigel with collagen-I was sufficient to induce tumor-specific mesenchymal gene expression, suppression of HNF4α and its target genes, and collective tumor cell invasion of patient-derived colon tumor organoids. The data connect collagen-rich stroma to mesenchymal gene expression in neoplastic cells and to collective tumor cell invasion. Targeting the tumor-collagen interface may therefore be explored as a novel strategy in the treatment of aggressive colon cancer.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 50 print issues and online access
$259.00 per year
only $5.18 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Budinska E, Popovici V, Tejpar S, D'Ario G, Lapique N, Sikora KO et al. Gene expression patterns unveil a new level of molecular heterogeneity in colorectal cancer. J Pathol 2013; 231: 63–76.
Guinney J, Dienstmann R, Wang X, de Reynies A, Schlicker A, Soneson C et al. The consensus molecular subtypes of colorectal cancer. Nat Med 2015; 21: 1350–1356.
Marisa L, de RA, Duval A, Selves J, Gaub MP, Vescovo L et al. Gene expression classification of colon cancer into molecular subtypes: characterization, validation, and prognostic value. PLoS Med 2013; 10: e1001453.
De Sousa E Melo F, Wang X, Jansen M, Fessler E, Trinh A, de Rooij LP et al. Poor-prognosis colon cancer is defined by a molecularly distinct subtype and develops from serrated precursor lesions. Nat Med 2013; 19: 614–618.
Perez-Villamil B, Romera-Lopez A, Hernandez-Prieto S, Lopez-Campos G, Calles A, Lopez-Asenjo JA et al. Colon cancer molecular subtypes identified by expression profiling and associated to stroma, mucinous type and different clinical behavior. BMC Cancer 2012; 12: 260.
Roepman P, Schlicker A, Tabernero J, Majewski I, Tian S, Moreno V et al. Colorectal cancer intrinsic subtypes predict chemotherapy benefit, deficient mismatch repair and epithelial-to-mesenchymal transition. Int J Cancer 2013; 134: 552–562.
Sadanandam A, Lyssiotis CA, Homicsko K, Collisson EA, Gibb WJ, Wullschleger S et al. A colorectal cancer classification system that associates cellular phenotype and responses to therapy. Nat Med 2013; 19: 619–625.
Schlicker A, Beran G, Chresta CM, McWalter G, Pritchard A, Weston S et al. Subtypes of primary colorectal tumors correlate with response to targeted treatment in colorectal cell lines. BMC Med Genomics 2012; 5: 66.
Loboda A, Nebozhyn MV, Watters JW, Buser CA, Shaw PM, Huang PS et al. EMT is the dominant program in human colon cancer. BMC Med Genomics 2011; 4: 9.
Sadanandam A, Wang X, de Sousa E Melo F, Gray JW, Vermeulen L, Hanahan D et al. Reconciliation of classification systems defining molecular subtypes of colorectal cancer: interrelationships and clinical implications. Cell Cycle 2014; 13: 353–357.
Hwang WL, Yang MH, Tsai ML, Lan HY, Su SH, Chang SC et al. SNAIL regulates interleukin-8 expression, stem cell-like activity, and tumorigenicity of human colorectal carcinoma cells. Gastroenterology 2011; 141: 291.
Mani SA, Guo W, Liao MJ, Eaton EN, Ayyanan A, Zhou AY et al. The epithelial-mesenchymal transition generates cells with properties of stem cells. Cell 2008; 133: 704–715.
Wellner U, Schubert J, Burk UC, Schmalhofer O, Zhu F, Sonntag A et al. The EMT-activator ZEB1 promotes tumorigenicity by repressing stemness-inhibiting microRNAs. Nat Cell Biol 2009; 11: 1487–1495.
Boissiere-Michot F, Lazennec G, Frugier H, Jarlier M, Roca L, Duffour J et al. Characterization of an adaptive immune response in microsatellite-instable colorectal cancer. Oncoimmunology 2014; 3: e29256.
Phillips SM, Banerjea A, Feakins R, Li SR, Bustin SA, Dorudi S . Tumour-infiltrating lymphocytes in colorectal cancer with microsatellite instability are activated and cytotoxic. Br J Surg 2004; 91: 469–475.
Calon A, Lonardo E, Berenguer-Llergo A, Espinet E, Hernando-Momblona X, Iglesias M et al. Stromal gene expression defines poor-prognosis subtypes in colorectal cancer. Nat Genet 2015; 47: 320–329.
Isella C, Terrasi A, Bellomo SE, Petti C, Galatola G, Muratore A et al. Stromal contribution to the colorectal cancer transcriptome. Nat Genet 2015; 47: 312–319.
Mesker WE, Junggeburt JM, Szuhai K, de HP, Morreau H, Tanke HJ et al. The carcinoma-stromal ratio of colon carcinoma is an independent factor for survival compared to lymph node status and tumor stage. Cell Oncol 2007; 29: 387–398.
Mesker WE, Liefers GJ, Junggeburt JM, van Pelt GW, Alberici P, Kuppen PJ et al. Presence of a high amount of stroma and downregulation of SMAD4 predict for worse survival for stage I-II colon cancer patients. Cell Oncol 2009; 31: 169–178.
Julien S, Merino-Trigo A, Lacroix L, Pocard M, Goere D, Mariani P et al. Characterization of a large panel of patient-derived tumor xenografts representing the clinical heterogeneity of human colorectal cancer. Clin Cancer Res 2012; 18: 5314–5328.
Calon A, Espinet E, Palomo-Ponce S, Tauriello DV, Iglesias M, Cespedes MV et al. Dependency of colorectal cancer on a TGF-beta-driven program in stromal cells for metastasis initiation. Cancer Cell 2012; 22: 571–584.
Nishida N, Nagahara M, Sato T, Mimori K, Sudo T, Tanaka F et al. Microarray analysis of colorectal cancer stromal tissue reveals upregulation of two oncogenic miRNA clusters. Clin Cancer Res 2012; 18: 3054–3070.
Yoshihara K, Shahmoradgoli M, Martinez E, Vegesna R, Kim H, Torres-Garcia W et al. Inferring tumour purity and stromal and immune cell admixture from expression data. Nat Comm 2013; 4: 2612.
Jorissen RN, Gibbs P, Christie M, Prakash S, Lipton L, Desai J et al. Metastasis-associated gene expression changes predict poor outcomes in patients with Dukes stage B and C colorectal cancer. Clin Cancer Res 2009; 15: 7642–7651.
Smith JJ, Deane NG, Wu F, Merchant NB, Zhang B, Jiang A et al. Experimentally derived metastasis gene expression profile predicts recurrence and death in patients with colon cancer. Gastroenterology 2010; 138: 958–968.
Huang, da W, Sherman BT, Lempicki RA . Systematic and integrative analysis of large gene lists using DAVID bioinformatics resources. Nat Protoc 2009; 4: 44–57.
Huang da W, Sherman BT, Zheng X, Yang J, Imamichi T, Stephens R et al. Extracting biological meaning from large gene lists with DAVID. Curr Protoc Bioinformatics 2009, Chapter 13 Unit 13.11.
Fatrai S, van Schelven SJ, Ubink I, Govaert KM, Raats D, Koster J et al. Maintenance of clonogenic KIT(+) human colon tumor cells requires secretion of stem cell factor by differentiated tumor cells. Gastroenterology 2015; 149: 692–704.
Boyd M, Bressendorff S, Moller J, Olsen J, Troelsen JT . Mapping of HNF4alpha target genes in intestinal epithelial cells. BMC Gastroenterol 2009; 9: 68.
Lussier CR, Babeu JP, Auclair BA, Perreault N, Boudreau F . Hepatocyte nuclear factor-4alpha promotes differentiation of intestinal epithelial cells in a coculture system. Am J Physiol Gastrointest Liver Physiol 2008; 294: G418–G428.
Stegmann A, Hansen M, Wang Y, Larsen JB, Lund LR, Ritie L et al. Metabolome, transcriptome, and bioinformatic cis-element analyses point to HNF-4 as a central regulator of gene expression during enterocyte differentiation. Physiol Genomics 2006; 27: 141–155.
Santangelo L, Marchetti A, Cicchini C, Conigliaro A, Conti B, Mancone C et al. The stable repression of mesenchymal program is required for hepatocyte identity: a novel role for hepatocyte nuclear factor 4alpha. Hepatology 2011; 53: 2063–2074.
Yao HS, Wang J, Zhang XP, Wang LZ, Wang Y, Li XX et al. Hepatocyte nuclear factor 4alpha suppresses the aggravation of colon carcinoma. Mol Carcinog e-pub ahead pf print 25 March 2015.
Zhang B, Wang J, Wang X, Zhu J, Liu Q, Shi Z et al. Proteogenomic characterization of human colon and rectal cancer. Nature 2014; 513: 382–387.
Emmink BL, Laoukili J, Kipp AP, Koster J, Govaert KM, Fatrai S et al. GPx2 suppression of H2O2 stress links the formation of differentiated tumor mass to metastatic capacity in colorectal cancer. Cancer Res 2014; 74: 6717–6730.
Belt EJ, Fijneman RJ, van den Berg EG, Bril H, Delis-van Diemen PM, Tijssen M et al. Loss of lamin A/C expression in stage II and III colon cancer is associated with disease recurrence. Eur J Cancer 2011; 47: 1837–1845.
Thomson S, Petti F, Sujka-Kwok I, Mercado P, Bean J, Monaghan M et al. A systems view of epithelial-mesenchymal transition signaling states. Clin Exp Metastasis 2011; 28: 137–155.
Sato T, Stange DE, Ferrante M, Vries RG, van Es JH, Van den BS et al. Long-term expansion of epithelial organoids from human colon, adenoma, adenocarcinoma, and Barrett's epithelium. Gastroenterology 2011; 141: 1762–1772.
Celesti G, Di Caro G, Bianchi P, Grizzi F, Basso G, Marchesi F et al. Presence of Twist1-positive neoplastic cells in the stroma of chromosome-unstable colorectal tumors. Gastroenterology 2013; 145: e615.
Chanrion M, Kuperstein I, Barriere C, El Marjou F, Cohen D, Vignjevic D et al. Concomitant Notch activation and p53 deletion trigger epithelial-to-mesenchymal transition and metastasis in mouse gut. Nat Commun 2014; 5: 5005.
Spaderna S, Schmalhofer O, Hlubek F, Berx G, Eger A, Merkel S et al. A transient, EMT-linked loss of basement membranes indicates metastasis and poor survival in colorectal cancer. Gastroenterology 2006; 131: 830–840.
Tabibzadeh S . Homeostasis of extracellular matrix by TGF-beta and lefty. Front Biosci 2002; 7: d1231–d1246.
Gujral TS, Chan M, Peshkin L, Sorger PK, Kirschner MW, MacBeath G . A noncanonical Frizzled2 pathway regulates epithelial-mesenchymal transition and metastasis. Cell 2014; 159: 844–856.
Caiado F, Carvalho T, Rosa I, Remedio L, Costa A, Matos J et al. Bone marrow-derived CD11b+Jagged2+ cells promote epithelial-to-mesenchymal transition and metastasization in colorectal cancer. Cancer Res 2013; 73: 4233–4246.
Hongo K, Tsuno NH, Kawai K, Sasaki K, Kaneko M, Hiyoshi M et al. Hypoxia enhances colon cancer migration and invasion through promotion of epithelial-mesenchymal transition. J Surg Res 2013; 182: 75–84.
Lu J, Ye X, Fan F, Xia L, Bhattacharya R, Bellister S et al. Endothelial cells promote the colorectal cancer stem cell phenotype through a soluble form of Jagged-1. Cancer Cell 2013; 23: 171–185.
Smith K, Bui TD, Poulsom R, Kaklamanis L, Williams G, Harris AL . Up-regulation of macrophage wnt gene expression in adenoma-carcinoma progression of human colorectal cancer. Br J Cancer 1999; 81: 496–502.
Wei SC, Fattet L, Tsai JH, Guo Y, Pai VH, Majeski HE et al. Matrix stiffness drives epithelial-mesenchymal transition and tumour metastasis through a TWIST1-G3BP2 mechanotransduction pathway. Nat Cell Biol 2015; 17: 678–688.
Miles FL, Sikes RA . Insidious changes in stromal matrix fuel cancer progression. Mol Cancer Res 2014; 12: 297–312.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies this paper on the Oncogene website
Rights and permissions
About this article
Cite this article
Vellinga, T., den Uil, S., Rinkes, I. et al. Collagen-rich stroma in aggressive colon tumors induces mesenchymal gene expression and tumor cell invasion. Oncogene 35, 5263–5271 (2016). https://doi.org/10.1038/onc.2016.60
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/onc.2016.60
This article is cited by
-
KCNQ1 and lymphovascular invasion are key features in a prognostic classifier for stage II and III colon cancer
BMC Cancer (2022)
-
Correlation between diffusion kurtosis and intravoxel incoherent motion derived (IVIM) parameters and tumor tissue composition in rectal cancer: a pilot study
Abdominal Radiology (2022)
-
αSMA+ fibroblasts suppress Lgr5+ cancer stem cells and restrain colorectal cancer progression
Oncogene (2021)
-
IFITM1 expression determines extracellular vesicle uptake in colorectal cancer
Cellular and Molecular Life Sciences (2021)
-
Bile and urine peptide marker profiles: access keys to molecular pathways and biological processes in cholangiocarcinoma
Journal of Biomedical Science (2020)