Abstract
Inhibitors of the mechanistic target of rapamycin (mTOR) are currently used to treat advanced metastatic breast cancer. However, whether an aggressive phenotype is sustained through adaptation or resistance to mTOR inhibition remains unknown. Here, complementary studies in human tumors, cancer models and cell lines reveal transcriptional reprogramming that supports metastasis in response to mTOR inhibition. This cancer feature is driven by EVI1 and SOX9. EVI1 functionally cooperates with and positively regulates SOX9, and promotes the transcriptional upregulation of key mTOR pathway components (REHB and RAPTOR) and of lung metastasis mediators (FSCN1 and SPARC). The expression of EVI1 and SOX9 is associated with stem cell-like and metastasis signatures, and their depletion impairs the metastatic potential of breast cancer cells. These results establish the mechanistic link between resistance to mTOR inhibition and cancer metastatic potential, thus enhancing our understanding of mTOR targeting failure.
Similar content being viewed by others
Introduction
The mechanistic target of rapamycin (mTOR) kinase integrates cues from nutrients and growth factors and is thus a master regulator of cell growth and metabolism.1 As such, mTOR is activated in most cancer types and is frequently associated with poor prognosis.2 Moreover, oncogenic mTOR signaling has a direct role in promoting cancer progression by inducing a pro-invasion translational program.3 This program includes the downregulation of the tuberous sclerosis complex 2 (TSC2) gene, whose product, in a heterodimer with the TSC1 product, serves as a negative regulator of mTOR complex 1 (mTORC1).4 Consequently, loss of Tsc2 in mice promotes breast cancer progression and metastasis.5 Collectively, current knowledge supports the notion that mTOR signaling has a key role in cancer initiation, progression and metastasis.
As mTOR is a key factor in cancer biology, therapies based on its inhibition have been widely studied6 and are central to the treatment of advanced metastatic breast cancer.7 However, the success of monotherapy assays has been limited. Critically, within a relatively short term, allosteric mTOR inhibition concomitantly induces upstream receptor kinase signaling, which mediates therapeutic resistance.8 Thus, therapies that combine allosteric inhibitors (rapamycin (sirolimus) and rapalogs) with inhibitors of growth factor signaling have been extensively evaluated.9 Intriguingly, recent studies have further linked mTOR activity to a stem cell-like cancer phenotype that mediates breast cancer metastasis10, 11 and, using triple-negative (TN) breast cancer cell lines, have described that mTORC1/2 inhibition spares a cell population with stem cell-like properties and enhanced NOTCH activity.12 These results are consistent with previous observations concerning the required activation of mTOR signaling in breast cancer stem-like viability and maintenance,13 the enhancement of NOTCH signaling in poorly differentiated breast tumors14 and the increase of tumor-initiating capacities with mTOR inhibition in liver cancer.15 In this scenario, a fundamental question emerges as to whether relative long-term adaptation or resistance to mTOR inhibition is functionally linked to tumor-initiating properties and, eventually, metastasis.
Here, we explored the hypothesis that mTOR signaling supports metastasis and remains active in therapeutic resistance in metastatic breast cancer. We found that abnormal mTOR signaling enhances tumor-initiating properties and metastatic potential. This activity is dependent on EVI1, which in cooperation with SOX9 sustains a transcriptional reprogramming response.
Results
Active mTORC1 signaling associates with distant metastasis
mTORC1 is the target of one of the latest drugs approved for the treatment of breast cancer in the advanced metastatic setting,7 which suggests that this protein complex has a potential role in supporting metastasis and aggressive features. To study this relationship, a tissue microarray of primary breast tumors was assessed for mTORC1 activity by means of immunohistochemical determination of phospho-Ser235/236-ribosomal protein S6 (pS6), a well-established downstream target of mTORC1.1 An association between pS6 positivity and the basal-like tumor phenotype or CK5 positivity was observed (Figure 1a; Mann–Whitney test P<0.01). Most importantly, an association was also detected between medium-high pS6 positivity and the development of distant metastases (Fisher’s exact test P=0.02; odds ratio (OR)=2.64, 95% confidence interval (CI) 0.95–7.35). Intriguingly, whereas the analyses by tumor subtypes were underpowered, both estrogen receptor (ER)-positive and ER-negative cases suggested a trend toward increased metastatic risk (ORs=4.44 and 1.96, respectively). Thus, enhanced mTOR activity and breast cancer metastatic potential appear to be linked.
mTORC1 activity concurrent with enhanced metastatic potential. (a) Left panels, representative immunohistochemical scores (0, negative, to 3, highest expression) of pS6 staining in the tissue microarray (TMA) of primary breast tumors. Right panel, results for the association between pS6 staining and distant metastasis. (b) Increased expression of mTORC1 pathway components with enhanced metastatic potential of MDA-MB-231 cells. The loading control (α-tubulin, TUBA) is shown. Bottom panel, graph showing quantifications of protein levels relative to parental and TUBA (per sample). (c) Increased pS6 expression in lung metastases developed by LM2 cells. The arrows mark magnified fields. Right panel, box-and-whisker plots for the quantification (pixels/area, p/a) of pS6 intensity; three mice and three similar lung metastases were analyzed in each setting. The P-value of the two-tailed Mann–Whitney test is shown. (d) Left panel, graph showing the in vivo photon flux quantification in mice injected with LM2 and treated with DMSO or everolimus. Representative images from bioluminescence in lungs from DMSO- or everolimus-treated mice are shown. The scale bar depicts the range of photon flux values as a pseudo-color display, with red and blue representing high and low values, respectively. Right top panels, quantification of lung colonization (total metastasis area normalized per total lung area, based on HE). Right bottom panels, representative immunohistochemical results for pS6 and quantification of normalized intensities.
Metastasis dependence on mTORC1 signaling
To test the contribution of mTOR signaling to metastasis, we used the well-defined MDA-MB-231 breast cancer cell line, including its parental poorly metastatic population and the lung metastatic derivatives LM1 and LM2.16 Western blot analyses showed increased levels in LM2 cells of several components of the mTORC1 signaling pathway, and particularly of RAPTOR and RHEB across the sub-populations (Figure 1b). The enhanced signaling in LM2 cells compared with the poorly metastatic parental population was confirmed by quantification of immunohistochemical staining of pS6 in the lung metastases that developed the cells upon tail vein injection (Figure 1c). Expanding on these observations, analysis of TCGA data showed negative correlations between TSC1/2 and an upregulated gene set whose expression was clinically and experimentally associated with breast cancer metastasis to lung (lung metastasis signature (LMS)-up; Pearson’s correlation coefficients (PCCs) <−0.25; P-values <10−8). Notably, this set was derived from the study of LM2 cells.16
Next, we tested the causal role of mTOR activity in the experimental model of lung metastasis. The capacity of LM2 cells to colonize the lung was assessed in the presence or absence of an allosteric mTOR inhibitor. LM2 cells stably expressing green fluorescent protein (GFP) and luciferase were injected into the lateral tail vein of immunocompromised mice, which were then randomly allocated to a group treated with dimethyl sulfoxide (DMSO) or a group treated with everolimus, both for 38 days. A significant reduction of lung colonization (and, as expected, of pS6 intensity) was observed in the latter group, both by measurements of in vivo photon flux and the relative lung metastasis area ex vivo by histology (Figure 1d). Collectively, these data suggest that mTORC1 signaling is associated with breast cancer metastatic potential and that inhibition of mTOR prevents lung metastasis. However, it is unclear whether this association persists in settings of resistance to mTOR inhibitors.
Metastatic resistance to mTOR inhibition
To evaluate the mechanisms responsible for resistance to mTOR inhibitors, we used two independent metastatic tumor models, namely a human TN BRCA1-mutated breast tumor orthotopically engrafted in nude mice (hereafter ortho-xenograft; Supplementary Figure 1) and the TN 4T1 murine breast carcinoma cell line engrafted in syngeneic background mice. Cells from both tumor models showed substantial mTORC1 signaling activity, particularly at the tumor invasive front (Supplementary Figure 1).17 Unexpectedly, although systemic treatment with sirolimus or everolimus blunted primary tumor growth in each model, it did not reduce the number or size of lung metastases (Figure 2a). In addition, and contrary to expectations, the intensity of pS6 staining at the invasive tumor fronts of the primary lesion and in the lung metastases of the sirolimus-treated ortho-xenografts was significantly higher than in the control animals (Figure 2b). Similarly, a key factor in cancer metastasis initially identified in LM2 cells and human data analyses (thus included in LMS-up), FSCN1,16, 18 was found to be significantly overexpressed in both experimental models exposed to mTOR inhibitors (Figure 2c). Subsequent gene expression analysis of the treated tumors revealed coordinated changes concurrent with mTOR inhibition that were associated with LMS activation (Supplementary Figure 2). These changes included overexpression of LMS-up in the sirolimus-treated ortho-xenografts and underexpression of LMS-down in the everolimus-treated 4T1 tumors (as measured by the gene set expression analysis (GSEA), P-values<0.05; Supplementary Figure 2).
Metastatic resistance to mTOR inhibition. (a) Left panels, graphs showing the average and standard deviation of micro- and macro-metastases observed in the lungs of the DMSO- and sirolimus- or everolimus-treated orthoxenografts and 4T1 tumors, respectively. The results correspond to the last day of treatment, and micro- versus macro-metastases were defined using a 2 mm width threshold, and by examining at least three tissue levels separated by > 20 μM. Right panels, growth rates of the DMSO- and sirolimus- or everolimus-treated tumors. (b) Representative immunohistochemical results for pS6 at the invasive tumor fronts (magnifications; top right panels) and the lung metastases (right panels) of DMSO- or sirolimus/everolimus-treated mice. The middle panels show quantifications, which correspond to three tumors, three equal front areas, and three metastases in each case. (c) Representative immunohistochemical tumor results for FSCN1/Fscn1 in DMSO- or sirolimus/everolimus-treated mice; quantifications are shown in right panels. (d) Recovered pS6 signal with concurrent FSCN1 overexpression through adaptation to everolimus in MCF7 and HCC1937 cells. Days of treatment are shown. (e) Left panel, graph showing the quantification of colonies from untreated and everolimus-adapted cells (12 culture fields were analyzed). The one-tailed t-test P-values are shown. Representative images of cell cultures are shown in right panels. (f) Flow cytometry results showing the cell counts for CD49f/EpCAM and of CD44/CD24 positivity in untreated or everolimus-adapted MCF7 and HCC1937 cells, respectively.
To further study resistance to allosteric mTOR inhibition, we subjected MCF7 ER-positive and HCC1937 TN cells to long-term exposure to 50 and 150 nM of everolimus, respectively. After a period of sensitivity defined by undetectable or very low levels of pS6, both cell lines recovered canonical mTORC1 signaling in 90–120 days (Figure 2d, top panels). Similarly to the in vivo observations, FSCN1 increased concurrently with adaptation to everolimus in both cell settings (Figure 2d, bottom panels). Subsequently, transcriptome analyses showed a significant change of the LMS in both cell lines, and particularly of the LMS-up in HCC1937 cells (Supplementary Figure 3).
Interestingly, both everolimus-adapted cell models showed significantly higher colony-forming capacity, with the higher relative difference found in HCC1937 cells (Figure 2e). Accordingly, fluorescence-activated cell sorting revealed an increase of CD49f+ and of CD44+/CD24− cells in everolimus-adapted MCF7 and HCC1937 cultures, respectively (Figure 2f). Although MCF7 did not show an increase in CD44+/CD24−, CD49f positivity has been linked to cancer stem cell-like properties.19 In addition, quantitative gene expression analysis revealed a significant increase of SOX2 in everolimus-adapted MCF7 cells and, in turn, an increase of NANOG and OCT4 (but not SOX2) in everolimus-adapted HCC1937 cells (Supplementary Figure 4). Notably, an increase in SOX2, but not the two additional stem cell-like markers, has also been described in MCF7 cells resistant to tamoxifen.20 Therefore, by combining in vivo and in vitro models of breast cancer, we reveal that exposure to allosteric mTOR inhibitors consistently promotes metastatic and tumor initiation properties. However, the precise regulators of this aggressive reprogramming remain to be determined.
TSC1/2 expression correlates negatively with tumor-initiating features
Given that differences in colony formation assays and tumor initiation properties were observed in mTOR inhibitor-resistant cell populations, we then explored the association between mTOR signaling and cancer cell initiation features in gene expression profiles from patient samples. To this end, we computed the expression correlations between TSC1 or TSC2 (TSC1/2) and 20 previously defined gene expression signatures using breast cancer data from The Cancer Genome Atlas (TCGA).21 The signatures (full annotation is provided in Supplementary Table 1) include a consensus set derived from the study of embryonic stem cell-like cells (sESCs),22 a consensus set of correlated master regulators of breast cancer stemness-like functions (hereafter sMRS),23 and a MYC-centered regulatory network (sMYC);24 importantly, these signatures were originally associated with poor prognosis and/or metastatic potential of ER-negative breast cancer and other types of cancer.22, 23, 24 The expression profiles of TSC1/2 were found to be negatively correlated (PCCs <−0.10; P-values <0.05) with most of the signatures (Figure 3a). In turn, positive correlations were observed with the downregulated genes that characterize mammary epithelial basal and luminal progenitor cells (Figure 3a).25 Moreover, the expression of TSC1 was positively correlated with a downregulated gene expression signature associated with oncogenic PI3KCA activity in breast cancer.26, 27 Collectively, these results confirmed that mTOR activity is associated with stem cell-like gene expression profiles in breast cancer.
Co-expression analysis and stem cell-like signatures. (a) TCGA network of significant co-expression levels (PCC P-values <0.05) between TSC1 or TSC2 and signatures derived from stem cell-like cell studies (Supplementary Table 1). The nodes represent TSC1/2 and the signatures, and the edges positive (red) or negative (green) correlations. Edge width is proportional to the corresponding PCC value. (b) Clustering correlation of sESC, sMRS and sMYC. The ortho-xenografts are differentially clustered relative to the treatment, and a similar trend is observed for 4T1 tumors. (c) The master regulators of the sMRS (that is, Core-9) are found to be relatively overexpressed upon mTOR inhibition. (d) Significant overexpression of sESC, sMRS and sMYC in regrown ortho-xenografts after sirolimus treatment. The GSEA ESs and the nominal P-values are shown. (e) Most of the regulators of sMRS are relatively overexpressed in everolimus-adapted HCC1937 cells. (f) HMGA1, which is encoded in Core-9, is upregulated upon adaptation to mTOR inhibition, particularly in HCC1937 cells.
Next, we observed that the sESC, sMRS and sMYC largely distinguished the expression profiles of mTOR inhibitor-treated tumors from those treated with DMSO (Figure 3b). The regulators of the sMRS (originally defined as Core-9)23 were commonly overexpressed upon mTOR inhibition (Figure 3c). Furthermore, a strong overexpression of the three signatures was detected when the ortho-xenografts were allowed to re-grow following treatment with sirolimus (relative to the DMSO-treated re-growth, GSEA P-values <0.001; Figure 3d and Supplementary Table 2). Analysis of the three signatures defined above did not reveal significant changes in the cell line models, but most of the regulators of sMRS were found to be overexpressed in HCC1937 cells (Figure 3e). These results were confirmed by western blot analysis of HMGA1 (Figure 3f), which has been associated with poor prognosis and metastatic breast cancer.28 The observed differences between the in vitro and in vivo expression changes may be due to the molecular specificity and/or biological conditions involved in each setting. Globally, however, inhibition of mTOR appears to be coupled to the transcriptional reprogramming that sustains metastatic and tumor initiation features.
EVI1 couples mTOR signaling to metastasis
Feedback activation of known mediators of resistance to rapalogs was not observed in the sirolimus-treated ortho-xenografts, but an increase in phospho-Thr202/Tyr204 ERK (pERK) was detected in 4T1 tumors treated with everolimus (Supplementary Figure 5). Exome sequence comparison between one DMSO- and one sirolimus-treated ortho-xenograft did not identify acquired mutations affecting components of the canonical TSC/mTOR pathway (Supplementary Table 3). In vitro, only modest time-dependent changes of phospho-S473 AKT (pAKT) and phospho-Y703 STAT3 (pSTAT3) were observed (Supplementary Figure 6). Although the lack of detection of known factors of resistance in our in vitro and in vivo analyses may be due to long-term treatments, we next sought to analyze a different mechanism that could be common to all models.
Given the transcriptomic changes observed across the in vivo and in vitro models, the data were analyzed to identify alternative regulators. A significant association (false discovery rate <5%) was observed in the gene expression profiles from ortho-xenograft samples and a target gene set of the ecotropic viral integration site-1 (EVI1) proto-oncogene (Transfac V$EVI1_02; Supplementary Figure 7). Similar associations for predicted EVI1 target sets were observed using data from the 4T1 tumors and MCF7 cells (Supplementary Figure 7). Importantly, expression analysis using TCGA data revealed positive correlations between EVI1 and stem cell-like signatures (sMYC and from mammary stem and progenitor cells),25 in addition to metastatic signatures, including LMS-up,16 a signature from low-burden breast cancer metastatic cells,10 and of breast cancer multipotency promoted by oncogenic PI3KCA26, 27 (Figure 4a). Moreover, a set of 79 commonly overexpressed genes (>0.25 log2; Supplementary Table 4) across the in vivo and in vitro models of mTOR inhibitor resistance showed significant positive co-expression with EVI1 (Figure 4b). Of note, this set included LEF1, which regulates stem cell maintenance in different contexts and is functionally connected to SOXs.29 In addition, this set showed over-representation of gene products involved in actin-cytoskeleton remodeling (Supplementary Table 4).
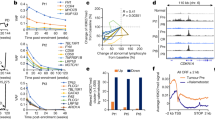
EVI1 couples stemness, metastatic potential and resistance to mTOR inhibition. (a) TCGA network of significant co-expression (PCC P-values <0.05) between EVI1 and signatures derived from stem cell-like cells and/or metastatic settings (Supplementary Table 1). (b) Distributions of PCCs between EVI1 and the commonly overexpressed 79 genes across the studied models or the complete microarray gene list as background control. The P-value of the Mann–Whitney test for the comparison of the distributions is shown. (c) Reduced pS6 levels with EVI1 depletion in cell models. The quantification of pS6/S6 signal ratios is show at the bottom (relative to siControl). (d) Ectopic overexpression of GFP-EVI1 in MCF7 (left panels) and HCC1937 (right panels) cells provides higher viability upon exposure to everolimus, relative to GFP-only overexpression. Also shown are the western blot results for defined markers across the drug-exposed cell cultures. The quantification of pS6/S6 signal ratios is show at the bottom (relative to TUBA per sample). (e) Increased EVI1 binding at predicted target promoters/gene loci with adaptation to everolimus. The fold changes are relative to the immunoglobulin control and the promoter gene targets are shown in the X axis. (f) Relative overexpression of RAPTOR and/or RHEB with adaptation to everolimus in MCF7 and HCC1937 cells. The quantification is show at the bottom (relative to untreated and TUBA per sample). (g) Relative reduction of RAPTOR and RHEB expression following EVI1 depletion, in particular in the everolimus-adapted setting.
EVI1 is essential for hematopoietic stem cell self-renewal30 and its overexpression has been associated with worse recurrence-free, overall and distant metastasis-free survival of ER-negative breast cancer.31 In vitro depletion of EVI1 reduced the levels of pS6, particularly in everolimus-adapted HCC1937 cells (60% reduction, and MCF7 showed a reduction of 10% in any condition; Figure 4c), whereas GFP-EVI1 overexpression conferred higher cellular viability in response to everolimus (Figure 4d). Of note, the levels of pS6 in the GFP-EVI1 overexpression assays were relatively low (Figure 4d), which could be due to the lack of full adaptation and/or the need for EVI1 co-factors.
To further validate the direct transcriptional role of EVI1, chromatin immunoprecipitation (ChIP) assays of predicted transcriptional targets were performed. This analysis revealed increased (one-tailed P-values<0.01) EVI1 binding at the following loci with adaptation to everolimus in at least one cell model: FSCN1 and SPARC (from the LMS-up); SCUBE3 and TCF4 (from V$EVI1_02); and RHEB, RPS6KA1 and RAPTOR (from the mTOR pathway) (Figure 4e). In addition, RAPTOR and RHEB were found to be overexpressed as a function of everolimus adaptation in MCF7 and HCC1937 cells, respectively (Figure 4f; RAPTOR showed transitory underexpression in HCC1937), and EVI1 depletion reduced the expression of both proteins in the everolimus-adapted settings (Figure 4g). However, the expression of FSCN1 was not significantly reduced with EVI1 depletion (Figure 4g) and, in turn, EVI1 was found to be overexpressed with FSCN1 depletion (Supplementary Figure 8). Therefore, these results may reflect the initial response towards mTOR inhibition; in fact, a similar effect was observed for RAPTOR and RHEB when EVI1 was depleted in parental HCC1937 cells (Figure 4g, right panels).
EVI1 cooperates with SOX9
Based on the strong association between EVI1 and stem cell-like/tumor initiation gene expression signatures, we searched for potential EVI1-transcriptional target genes mediating such functions. The V$EVI1_02 gene set and several other stem cell-like cells and/or metastasis-associated gene signatures were positively co-expressed with a key regulator of these functions, SOX9 (Figure 5a).32, 33 Subsequently, whole-genome EVI1 ChIP data corroborated the positive correlation between EVI1 and SOX9 binding sites in HCC1937 cells (Supplementary Figure 9). Interestingly, EVI1 ChIP data also showed a positive correlation with SLUG targets34 in both cell models, and with SNAIL targets34 in HCC1937 (Supplementary Figure 9). Notably, MCF7 differentiation to a basal-like phenotype requires SLUG activity.35
EVI1 cooperates with SOX9 and regulates its expression. (a) TCGA network of significant co-expression (PCC P-values <0.05) between EVI1 or SOX9 and signatures derived from stem cell-like cells and/or metastatic settings (Supplementary Table 1). (b) Increased SOX9 expression in ortho-xenograft tumor fronts of mice treated with sirolimus; the results correspond to at least three ortho-xenografts of each group. (c) Increased SOX9 and ALDH1 expression in everolimus-adapted cells. (d) Graph showing the results from the analysis of the complete drug panel for the correlation between IC50 profiles and the expression of the V$EVI1_02 gene set; drugs are ranked according to PCC log P-values. Negative and positive PCCs are indicated with different colors, and the mTOR inhibitors in the panel are denoted. (e) Left panel, unsupervised clustering and correlation analysis of the difference in EVI1 ChIP results at the SOX9 locus between everolimus-adapted and untreated cells. Right panels, results of ChIP assays targeting a predicted EVI1-binding site in the SOX9 promoter (Supplementary Table 5); the input, control immunoglobulin immunoprecipitation (IP), and EVI1-IP results are shown. The control results for the binding site in FSCN1 are also shown. (f) Depletion of EVI1 leads to a reduction of SOX9 expression in three cell conditions (the results correspond to Figure 4c; the ratios are relative to siControl and TUBA per sample). (g) Depletion of EVI1 leads to a reduction of colony-forming capacity. The results of the one-tailed t-test are shown. (h) Depletion of Evi1 impairs the tumorigenic potential of 4T1 cells. The log-rank P-value is shown for the comparison between the shControl and short hairpin RNA (shRNA)-EVI1 #1; note that transduction with shRNA-EVI1 #2 completely impaired tumor formation so a P-value could not be computed (n.a.).
Expanding on the above results, sirolimus-treated ortho-xenografts and everolimus-adapted cells showed increased SOX9 expression compared with their corresponding controls (Figures 5b and c, respectively; the same antibody did not recognize mouse Sox9). The expression of ALDH1—canonical stem/progenitor marker in normal breast tissue and tumors, and also associated with poor prognosis36—was also detected to be increased with adaptation to everolimus in both MCF7 and HCC1937 cells (Figure 5c). The higher expression at the tumor invasive front is consistent with previous observations of invasive leader cells showing positivity for basal/stem cell-like markers.10, 37
In addition to the results from the models, the analysis of data from hundreds of cell lines38 revealed significant positive correlations between EVI1 and SOX9 expression, and with EVI1 locus copy number (PCCs >0.20, P-values <10−4). Moreover, in this data set, both EVI1 and SOX9 expression correlated positively (that is, linked to resistance) with the half maximal inhibitory concentration (IC50) of temsirolimus (PCCs=0.22 and 0.24, respectively, P-values <10−4). In fact, a ranking-based analysis using the V$EVI1_02 gene set as a surrogate of EVI1 activity showed equivalent results, and the second and fourth most correlated drugs, respectively, were BX-795 (inhibitor of PDK1 activity) and rapamycin (Figure 5d). To further assess these findings, an ER-positive and HER2-positive breast cancer cell model, BT-474, was exposed to 150 nM of everolimus for approximately 100 days and subsequently profiled for gene expression changes; the results showed significant associations with EVI1 and SOX9 targets, and with the commonly overexpressed genes detected across the above in vitro and in vivo models (Supplementary Figure 10).
Further analysis of the predicted functional relationship between EVI1 and SOX9 revealed a positive correlation in the differential (between control and everolimus-adapted) binding of EVI1 at the SOX9 locus (Figure 5e, left panel). Next, targeted ChIP assays confirmed that EVI1 binds at the SOX9 locus in untreated HCC1937 and both everolimus-adapted cell models (Figure 5e, right panel). Thus, depletion of EVI1 reduced the expression of SOX9 in three out of four conditions; however, it remains to be determined which co-factor(s) may maintain SOX9 expression at normal levels in everolimus-adapted HCC1939 cells (Figure 5f). In parallel, depletion of EVI1/Evi1 reduced the colony-forming capacity of both models of everolimus adaptation (Figure 5g) and impaired in vivo tumorigenic potential of 4T1 cells (Figure 5h). Collectively, ChIP, gene/protein expression analyses and in vivo functional assays depict a link between EVI1 and SOX9 in the regulation of stem cell-like and tumor initiation features.
In vivo evaluation of the EVI1, SOX9 and mTOR relationship
The functional cooperation between EVI1 and SOX9 was evaluated in vivo using LM2 and 4T1 cells transduced with a short-hairpin RNA scrambled control or directed against EVI1, and with or without concomitant overexpression of Sox9 (mouse protein). Thus, EVI1 depletion significantly reduced the capacity to colonize the lungs, and concurrent Sox9 overexpression partially rescued metastatic potential (Figures 6a and b). As shown in everolimus-adapted cell lines, EVI1/Evi1 depletion caused a significant decrease of both SOX9 and pS6 expression in the corresponding metastasis (Supplementary Figure 11). Concurrent Sox9 overexpression recovered pS6 signal in 4T1 but not in LM2 cells (Supplementary Figure 11), which suggest differences in the precise regulation of mTOR activity between the models.
In vivo assessment of the role of EVI1 and SOX9. (a, b) Depletion of EVI1/Evi11 expression (using two different short hairpin RNAs (shRNAs)) in LM2 and 4T1 cells reduced lung colonization, and Sox9 overexpression partially recovered this potential, left panels. (a) Right panels show representative images of lungs and their respective HE staining. (c) Depletion of SOX9 and overexpression of Sox9 reduced and increased, respectively, lung colonization of LM2 cells. Treatment with everolimus of shControl LM2 cells also reduced lung colonization. (d) Depletion of Sox9 or treatment with everolimus of 4T1 cells reduced lung colonization. (e, f) Depletion of FSCN1/Fscn1 expression in LM2 and 4T1 cells reduced lung colonization, and concurrent treatment with everolimus further impaired this potential.
In addition to EVI1, depletion of SOX9 in LM2 cells led to a significant decrease in lung colonization capacity and, conversely, Sox9 overexpression increased this capacity (Figure 6c and Supplementary Figure 12). Depletion of Sox9 in 4T1 cells and everolimus treatment of both cell models also reduced lung colonization (Figures 6c and d). In addition, depletion of FSCN1 in both models also led to a substantial impairment of lung colonization (Figures 6e and f). Moreover, a greater effect was observed when the animals were simultaneously treated with everolimus (Figures 6e and f), which fully suppressed pS6 signal (Supplementary Figure 11). Collectively, these results indicate that transcriptional reprogramming mediated by EVI1-SOX9 is one of the key factors in metastatic resistance to mTOR inhibition.
Discussion
We provide evidence of the association between EVI1-SOX9 function, mTOR inhibition resistance and metastasis in breast cancer. We also show that EVI1-SOX9-mediated transcriptional reprogramming drives the molecular processes that support breast cancer tumor initiation features and metastatic potential in therapeutic resistance (Figure 7). These data are coherent and expand on the concept that cancer stem cell-like cell populations have high tumor-initiating capacity and are frequently the source of therapy resistance and metastasis.39 Data from hundreds of cell lines38 suggest that the proposed mechanism is relevant in settings beyond breast cancer. Importantly, EVI1 maps in a genomic region (including PI3KCA and SOX2) whose amplification is an independent predictor of breast cancer recurrence.40 This region is frequently found to be amplified in basal-like and BRCA1-mutated breast cancer,21 as well as in non-small cell lung and ovarian cancers.41 In addition, EVI1 amplification is independent of PIK3CA mutations,41 which further reinforces the link with basal-like breast cancer and is consistent with a role in resistance to allosteric mTOR inhibition. It remains to be determined whether EVI1 expression is upregulated through genomic amplification and/or whether its function is enhanced by biochemical modifications by casein kinase II42 and/or ERK signaling as seen to be activated in our 4T1 assays.
Proposed mechanistic model. In untreated cancer, low TSC1/2 expression is associated with enhanced mTORC1 activity and, therefore, with a primary metastatic and stemness phenotype. In cancer treated for mTOR inhibition, EVI1-SOX9 become activated (in part by overexpression) and positively sustain the following features: mTOR signaling (through upregulation of RHEB and RAPTOR), metastatic potential (through LMS-up and other signatures) and stemness (through at least SOX9).
Our preclinical findings and mechanistic model are consistent with and expand on recent observations across different neoplastic settings. EVI1 contributes to epithelial-to-mesenchymal transition (EMT) and invasion of acute myeloid leukemia,43 and influences EMT in ovarian cancer cells,44 whereas EMT mediates resistance to rapamycin.45 In the latter study, MCF7 cells transfected with constitute active SNAIL showed increased ERK signaling and decreased sensitivity to rapamycin. Thus, our study provides a mechanistic explanation for these observations. In addition, it has recently been shown that SOX2 and SOX9 mediated the maintenance of latent metastatic stem cell-like cells,46 and it was previously demonstrated that SLUG and SOX9 cooperatively determine mammary stem cell state,32 and that SOX9 function links tumor initiation and invasion.33 Moreover, a stem cell-like cancer phenotype that mediates breast cancer metastasis is predicted to exhibit abnormal mTORC1 signaling10 and, in turn, mTORC1/2 inhibition promotes stem cell-like properties and enhanced NOTCH1 activity in TN breast cancer cell lines.12 Thus, enhanced mTOR signaling impairs cell differentiation by potentiating NOTCH1 activity, and this signaling is found to be increased in poorly differentiated breast tumors.14 Intriguingly, NOTCH may also regulate SOX9 expression,47 which, in turn, is a master regulator of stem and progenitor cells.48 In parallel, allosteric mTOR inhibition increases the number of tumor-initiating cells in a model of liver cancer15 and the metastatic potential in a model of pancreatic neuroendocrine cancer.49 In this scenario, our study proposes that EVI1 and SOX9 functionally cooperate to sustain mTORC1 activity, EMT and metastatic potential, thereby providing new insights into therapeutic resistance.
Our findings—particularly those from MCF7 cell assays—may have clinical implications for the established use of mTOR inhibitors in endocrine-resistant ER-positive advanced metastatic breast cancer,7 in which resistance to treatment is eventually reported. Our results indicate that exposure to allosteric mTOR inhibition selects a stem cell-like cancer cell population with metastatic capacity, which may therefore promote disease progression. Although in vivo assays may be warranted to further assess this observation, and while we cannot rule out that the specific population may arise through the acquisition of new mutations, it is noteworthy that the Breast Cancer Trials of Oral Everolimus-2 (BOLERO-2) study for the efficacy of everolimus plus exemestane in endocrine resistance showed similar benefits for patients with or without visceral metastases.50 Nevertheless, full and durable pathway inhibition—such as obtained that by the next generation of targeted drugs51—may fully impair metastatic resistance.
Materials and methods
Tissue microarray
The tissue microarray included 138 infiltrating ductal breast carcinoma tumors collected at the Department of Pathology of the MD Anderson Cancer Center, Madrid (Spain). The patients underwent surgery between 2003 and 2004, and all tumors were classified as grade 3. According to the TNM system, 45 tumors belonged to stage I, 48 to stage II and 45 to stage III-IV. The linked data included ER (n=104), progesterone receptor (n=127) and epidermal growth factor receptor 2 (HER2; n=125) status, and CK5 expression (n=128), absence/presence of lymph node metastasis (n=124), and absence/presence of distant metastasis (n=127). The tissue microarray contained duplicated cases and normal tissue, and the immunohistochemical results were scored independently and blindly (to molecular and clinical status). Selection of the highest value for a given case, blindly to its status, solved discordant scores. The study was approved by the ethics committee of the MD Anderson Cancer Center and written informed consent was obtained from all patients.
Gene expression analyses
Pre-processed and normalized data of human breast cancer were taken from the corresponding publication16 and from the TCGA repository (http://tcga-data.nci.nih.gov/tcga/tcgaHome2.jsp).52 RNA samples were extracted using TRIzol Reagent (Invitrogen, Karlsruhe, Germany) and RNeasy Kit (Qiagen, Venlo, Netherlands), and quality was evaluated in an Agilent Bioanalyzer (Foster City, CA, USA) 2100. The RNAs were amplified using the Ribo-SPIA system (NuGEN Technologies Inc., San Carlos, CA, USA) and subsequently hybridized on the Human Genome U219 microarray platform (Affymetrix, Santa Clara, CA, USA; IRB Core Facility, Barcelona, Spain). Gene expression data from the ortho-xenograft, and MCF7 and HCC1937 cell lines have been deposited under the GEO reference GSE39694. Gene expression data from the 4T1 tumors and BT-474 cells have been deposited under the GEO references GSE50712 and GSE85801, respectively. The GSEA and DAVID (for functional term analyses) tools were used with standard parameters.53, 54 The signature correlations were computed by selecting genes with s.d. >1.0 and using the average Z-score value per gene set. The quantification of NANOG, OCT4 and SOX2 gene expression was performed as previously described.55
Antibodies
Anti-total and pAKT (#9272 and #9271, respectively, Cell Signaling Technology, Danvers, MA, USA; #4060 for immunohistochemistry assays), anti-ALDH1 (#611194, BD Biosciences, Oxford, UK), anti-4EBP1 (#9452, Cell Signaling Technology), anti-p4EBP1 (#2855 and #9451, Cell Signaling Technology), anti-ER (#IR151, Dako, Glostrup, Denmark), anti-total and phospho-Thr202/Tyr204 ERK (#4695 and #4376, respectively, Cell Signaling Technology), anti-EVI1 (#2265 and #2593, Cell Signaling Technology; and #A301-691A, Bethyl Laboratories, Montgomery, TX, USA), anti-FSCN1 (#SC-56531, Santa Cruz Biotechnology, Dallas, TX, USA), anti-GFP ChIP grade (#ab290, Abcam, Cambridge, UK), anti-GLUT1 (#652, Abcam), anti-HER2 (#790-100, Ventana, Tucson, AZ, USA), anti-HMGA1 (#129153, Abcam), anti-pIGF1R (#39398, Abcam), anti-IRS1 (#2382 and #9451, Cell Signaling Technology), anti-KI67 (#IR626, Dako), anti-CK19 (#IR615, Dako), anti-PR (#IR168, Dako), anti-RAPTOR (#SC-81537, Santa Cruz Biotechnology), anti-RHEB (#SC-6341, Santa Cruz Biotechnology), anti-S6 (#SC-74459, Santa Cruz Biotechnology), anti-pS6 (#4858, Cell Signaling Technology), anti-S6K (#9202, Cell Signaling Technology), anti-pS6K (#9205, Cell Signaling Technology), anti-SOX9 (#5535, Abcam), anti-total and pSTAT3 (#9132 and #9145, respectively, Cell Signaling Technology), anti-TUBA (#44928, Abcam) and anti-VCL (V9131, Sigma-Aldrich, St Louis, MO, USA). The antibodies used for fluorescence-activated cell sorting were anti-CD24-PE, anti-CD44-APC, anti-CD49f-Alexa-647, and anti-EPCAM-FITC (#555428, 559942, 562473, and 347197, respectively; BD Biosciences).
Immunohistochemistry
The assays were performed on serial paraffin sections (3–4 μm thick) using the EnVision (Dako) or Ultraview (Ventana) systems. Antigen retrieval was performed using citrate- or EDTA-based buffers. Endogenous peroxidase was blocked by pre-incubation in a solution of 3% H2O2 and blocking was performed in 1X phosphate-buffered saline with 5% goat serum or 1% bovine serum albumin and 0.1% Tween 20 (Sigma-Aldrich). In all experiments, equivalent sections were processed without incubation with the primary antibody, which did not reveal immunostaining in any case. Sections were hematoxylin and eosin (HE)-counterstained and examined with an Olympus BX51 (Tokyo, Japan) microscope. The immunohistochemistry microscopic images were color deconvoluted and quantitated using the regions of interest methodology in ImageJ (http://rsb.info.nih.gov/ij). Quantification of tumor fronts was based on rectangular areas of 25 μM × 50–300 μM. When quantifying the results from lung metastases, the complete metastatic area was considered because the fronts were often difficult to outline histologically.
Cell culture
The LM2 cell derivative is a lung metastatic sub-line originated from MDA-MB-231 breast cancer cells from the laboratory of Professor Massagué.16 The 4T1 cells derived from a spontaneous BALB/c mouse breast cancer tumor56 and were obtained from the ATCC (Rockville, MD, USA). The LM2 cells were cultured in Dulbecco's modified Eagle's medium (GIBCO, Karlsruhe, Germany) supplemented with 10% fetal bovine serum (GIBCO), 1x l-glutamine (Biowest, Nuaillé, France) and 1% penicillin/streptomycin (Biowest). The 4T1 cells were cultured in Roswell Park Memorial Institute (RPMI)-1640 medium (Sigma-Aldrich) supplemented with 10% fetal bovine serum, 1x l-glutamine and 1% penicillin/streptomycin. All MDA-MB-231 cells/sub-lines were stably transfected with a thymidine kinase and GFP luciferase construct and sorted for GFP expression. The MCF7 and HCC1937 cell lines were obtained from ATCC and cultured in supplemented Dulbecco's modified Eagle's medium and RPMI-1640 medium, respectively. The Matrigel (BD Biosciences) colony formation assays were performed using standard protocols with 5% fetal bovine serum. Everolimus was purchased from Selleck Chemicals (Houston, TX, USA) and LC Laboratories (Woburn, MA, USA). Fluorescence-activated cell sorting was performed using FACS Canto (Becton Dickinson, Franklin Lakes, NJ, USA) and Diva software (Becton Dickinson) package, and antibody-based cell labeling was performed as previously described.55
Western blotting
To analyze extracts, cells were lysed in standard 150 mM NaCl buffer supplemented with protease inhibitor cocktail (Roche Molecular Biochemicals, Mannheim, Germany) and, in some instances, a phosphatase inhibitor was added (1 mM NaF, Sigma-Aldrich). Lysates were clarified twice by centrifugation at 13 000 × g and protein concentration was measured using the Bradford method (Bio-Rad, Solna, Sweden). Lysates were resolved in sodium dodecyl sulfate–polyacrylamide gel electrophoresis gels and transferred to Immobilon-P (Merck Millipore, Billerica, MA, USA) or PVDF membranes (Roche Molecular Biochemicals). Target proteins were identified by detection of horseradish peroxidase-labeled antibody complexes with chemiluminescence using the ECL Western Blotting Detection Kit (GE Healthcare, Amersham, UK).
Lung colonization assays
The Animal Care and Use Committee of IRB Barcelona approved the following animal studies. Female BALB/c nude (MDA-MB- 231 cells) or BALB/c wild-type mice (4T1 cells) were used. For tail vein injections, cells were suspended in 1x phosphate-buffered saline (GIBCO; 200 μl per mouse) and injected into the lateral tail vein of mice using a 26G needle, as previously described.57 Before the injection of cells, mice were anesthetized with ketamine (100 mg/kg body weight) and xylazine (10 mg/kg body weight), and immediately after injection they were imaged for luciferase activity by injecting 50 μl of beetle luciferin potassium salt (Promega, Madison, WI, USA) at 15 mg/ml. To induce the expression of short hairpin RNA in vivo, doxycycline (1 mg/ml, Sigma-Aldrich) was administered ad libitum in drinking water containing 25 mg/ml sucrose (Sigma-Aldrich). When indicated, DMSO solution (at the same concentration as for the compound test, 5%) or everolimus (5 mg/kg; SC-218452, Santa Cruz Biotechnology) was administered daily by intraperitoneal injection. Mice were monitored weekly using IVIS imaging, unless otherwise indicated. Lung tumor development was followed up once a week by bioluminescence imaging of the upper dorsal region that corresponds to lung position. Bioluminescent images were quantified with Living Image 2.60.1 software (Perkin-Elmer, Waltham, MA, USA). All values were normalized to those obtained at day 0. The HE staining of lung sections scored the lung colonization capacity of 4T1 cells 3 weeks post-inoculation. Five sections, separated by 50 μm, per mouse lung were counted. The average of the total metastatic area normalized to total lung area was measured. The average total lung metastasis area for all mice was then plotted. The tissue was dissected, fixed in 10% buffered formalin (Sigma-Aldrich), and embedded in paraffin. Sections (3 μm thick) were stained with HE. To analyze the metastatic area, images were taken with a scanner, and the area of each metastatic lesion was quantified with the ImageJ software. Five images per section/animal were evaluated, and the average area was plotted. The Fiji Trainable Weka segmentation, an ImageJ plugin based on the Weka58 Java machine learning library, was used to classify images on the basis of local colorimetric, textural and structural features in the neighborhood of each pixel. Images were processed with a custom macro created at the Microscopy Core Facility of IRB Barcelona. The two-tailed non-parametric Mann–Whitney test was used to assess significance of the immunohistochemical staining results.
Ortho-xenograft
The patient was a 33-year-old woman with a pathological germline BRCA1 mutation and diagnosed with breast cancer shortly after pregnancy. At diagnosis she presented a locally advanced TN ductal infiltrating carcinoma of the breast (T4) with involvement of ipsilateral nodes (N2) and lung metastasis. Primary systemic chemotherapy was initiated with TAC regimen for four cycles, followed by mastectomy to prevent local complications because of extensive breast involvement. Following surgery, the patient received further chemotherapy with the same regimen. The patient was diagnosed with brain metastases shortly after and died 8 months post-diagnosis as a result of disease progression. Mutational analysis of BRCA1 was carried out by the Molecular Diagnostics Unit (Catalan Institute of Oncology, Barcelona, Spain) following standards for genetic testing and pathological determination. The patient provided written informed consent, and the study was approved by the IDIBELL Ethics Committee. Female athymic (nu/nu) mice (Harlan, Harlan Laboratories, Barcelona, Spain) between 4 and 6 weeks of age were used for engraftment. The orthotopic model developed histologically detectable lung metastases in a period of approximately 50 days after engraftment. The protocol was reviewed and approved by the IDIBELL Animal Care and Use Committee. A daily oral treatment with sirolimus (Rapamune) or control solution (DMSO, Sigma-Aldrich) was applied.
Exome analysis
The National Centre for Genomic Analysis (CNAG) carried out exome sequencing. Sequence capture and amplification was performed using Agilent (Agilent Technologies, Palo Alto, CA, USA) SureSelect Human All Exon kit (Agilent) according to the manufacturer’s instructions. Paired-end sequencing was performed on a HiSeq2000 instrument (Illumina, San Diego, CA, USA) using 76-base reads. Reads were aligned to the reference genome (GRCh37) and BAM files were generated using SAMtools. Duplicates were removed using SAMtools and custom scripts, and single-nucleotide variant calling was performed using a combination of SAMtools and Sidrón algorithms as described previously.59 The reads were first aligned to mouse genome (mm9), and those read-pairs that did not align to mouse were then aligned to the human genome following the same pipeline as above. Only mismatch variants were taken into account and small insertions and deletions were not counted. Common variants, defined as those present in dbSNP135 with a minor allele frequency > 1%, were filtered out.
4T1 tumors
The animal studies were conducted using protocols that had undergone appropriate review and approval at the New York University School of Medicine. Balb/C mice were injected subcutaneously with 5 × 104 4T1 cells, measured for tumor size at day 10, and randomly organized in two equivalent groups that were treated with DMSO solution (the same concentration as for the compound test) or everolimus (5 mg/kg; SC-218452, Santa Cruz Biotechnology) daily by intraperitoneal injection. Tumors were excised at day 23 and processed. Half of each tissue sample was used for immunohistochemistry and half for gene expression microarray analysis. For the tumorigenicity assays, 250 000 4T1 cells were injected at the orthotopic site, mixed with growth factor-reduced Matrigel (BD Biosciences) before inoculation (1:1). Once palpable, tumors were measured with a digital caliper, and the tumor volume was calculated. The ethics committee of the CIC bioGUNE approved these assays.
ChIP assays
Assays were prepared using 107 cells of each cell line per condition. Chromatin was fragmented by sonication (Bioruptor, Diagenode, Denville, NJ, USA) for 30 min (30-s pulses, 30-s pauses) and assays were carried out following the manufacturer’s protocol (kch-mahigh-A16, HighCell# ChIP Kit, Diagenode), using anti-EVI1 (#2593, Cell Signaling Technology) or an equal amount of IgG isotype as negative control (#2729, Cell Signaling Technology). The amount of DNA was analyzed by real-time polymerase chain reactions using SYBR Green-based assays (Applied Biosystems, Life Technologies, Foster City, CA, USA). The results were calculated using the ΔCt method. A genomic region of the GAPDH gene was used as negative control (Diagenode). The corresponding human genome coordinates, EVI1-binding sites and primers designed for the assays are detailed in Supplementary Table 5. Whole-genome ChIP data were obtained by hybridization to SurePrint G3 Human Promoter 1x1M microarrays (IRB Core Facility) and analyzed by MACS (version 2.0.9).60 The data have been deposited under the GEO reference GSE50905. The complete ranking of differential EVI1 binding between adapted and sensitive MCF7 or HCC1937 cells was used as input for the GSEA of transcription factor targets.
Gene expression alterations
Stable LM2 and 4T1 cell lines expressing short hairpin RNAs were generated as described previously.57 The shFSCN1/Fscn1 (that is, targeting both human and mouse gene expression) #1 and shEVI1/Evi1 #2 were encoded in lentiviral vectors (inducible pTRIPZ lentiviral short hairpin RNAs, GE Dharmacon, Lafayette, CO, USA). The short hairpins were induced by 1 μg/ml doxycycline for 72 h. The shControl and shEVI1/Evi1 #1 were encoded in a retrovirus pGFP-V-RS (OriGene, Rockville, MD, USA). The shFSCN1/Fscn1 #1 was encoded in a pSUPER (Addgene, Cambridge, MA, USA) vector. The shSox9 (against mouse gene sequence) and shSOX9 (human) were obtained from the MISSION library (SHCLND-NM_011448 and SHCLND-NM_000346, respectively; Sigma-Aldrich). An additional shSOX9 was obtained from Addgene, catalog #40644. For Sox9 overexpression, the corresponding coding sequence was cloned into a lentiviral pWXL vector. Stable cell lines expressing the various constructs described above were generated under puromycin selection for 48 h. The siRNA against EVI1 expression was an ON-TARGETplus SMARTpool (L-006530-02-0010, Dharmacon). The following primer sequences were used to assess gene expression changes in real-time (using SYBR Green, Applied Biosystems) polymerase chain reaction assays: EVI1, 5′-CATTGGGAACAGCAACCAT-3′ and 5′-GGTCACCAAAGCCTTTTCAT-3′; Evi1, 5′-CACAGAAAGTCCAAATCACAGG-3′ and 5′-GCCACACGTTGGAGGAAC-3′; Sox9, 5′-GTACCCGCATCTGCACAAC-3′ and 5′-CTCCTCCACGAAGGGTCTCT-3′; ACTB, 5′-GGAGTGGGTGGAGGCAG-3′ and 5′-AACTAAGGTGTGCACTTTTGTTC-3′; and mL32, 5′-GAAACTGGCGGAAACCCA-3′ and 5′-GGATCTGGCCCTTGAACCTT-3′.
Genomics of drug sensitivity data analyses
For the correlation analysis between the basal expression of EVI1, its predicted target genes and the drug responses across cancer cell lines, data were downloaded from the GDS project (web-release April 2012).38 This data set included IC50 values for 131 drugs that were assessed in a panel of 638 human cancer cell lines. The basal gene expression data were downloaded from ArrayExpress reference E-MTAB-783. Non-annotated probes were removed and expression values were averaged when multiple probes mapped to the same gene. Correlation scores and P-values were computed using the PCC. The EVI1 target set included 20 genes that were represented by at least one microarray probe (Supplementary Table 6). The extent of the basal expression of the EVI1 targets was quantified using an enrichment score (ES) computed with a Matlab implementation of the GSEA algorithm. To estimate ES significance, a null model was created by generating 10 000 random gene sets (of the same size as the EVI1 target set) and used to query the data set through GSEA. Next, two inverse Gaussian distributions (for positive and negative ES values) were fitted on the resulting empirical distribution and used to compute P-values. The correlations between IC50 profiles and ESs were computed by considering only cell lines whose basal expression profile yielded a significant ES (P<0.05), according to the null model. The enrichment P-values of mTOR inhibitors among drugs whose IC50 profile was anti-correlated with the EVI1 target ES were computed using Fisher’s exact test and considering the total set of 131 drugs.
References
Zoncu R, Efeyan A, Sabatini DM . mTOR: from growth signal integration to cancer, diabetes and ageing. Nat Rev Mol Cell Biol 2011; 12: 21–35.
Courtney KD, Corcoran RB, Engelman JA . The PI3K pathway as drug target in human cancer. J Clin Oncol 2010; 28: 1075–1083.
Hsieh AC, Liu Y, Edlind MP, Ingolia NT, Janes MR, Sher A et al. The translational landscape of mTOR signalling steers cancer initiation and metastasis. Nature 2012; 485: 55–61.
Crino PB, Nathanson KL, Henske EP . The tuberous sclerosis complex. N Engl J Med 2006; 355: 1345–1356.
Nasr Z, Robert F, Porco Jr JA, Muller WJ, Pelletier J . eIF4F suppression in breast cancer affects maintenance and progression. Oncogene 2013; 32: 861–871.
Fruman DA, Rommel C . PI3K and cancer: lessons, challenges and opportunities. Nat Rev Drug Discov 2014; 13: 140–156.
Baselga J, Campone M, Piccart M, Burris HA 3rd, Rugo HS, Sahmoud T et al. Everolimus in postmenopausal hormone-receptor-positive advanced breast cancer. N Engl J Med 2012; 366: 520–529.
Chandarlapaty S . Negative feedback and adaptive resistance to the targeted therapy of cancer. Cancer Discov 2012; 2: 311–319.
Markman B, Dienstmann R, Tabernero J . Targeting the PI3K/Akt/mTOR pathway—beyond rapalogs. Oncotarget 2010; 1: 530–543.
Lawson DA, Bhakta NR, Kessenbrock K, Prummel KD, Yu Y, Takai K et al. Single-cell analysis reveals a stem-cell program in human metastatic breast cancer cells. Nature 2015; 526: 131–135.
Ruiz de Garibay G, Herranz C, Llorente A, Boni J, Serra-Musach J, Mateo F et al. Lymphangioleiomyomatosis biomarkers linked to lung metastatic potential and cell stemness. PLoS One 2015; 10: e0132546.
Bhola NE, Jansen VM, Koch JP, Li H, Formisano L, Williams JA et al. Treatment of triple-negative breast cancer with TORC1/2 inhibitors sustains a drug-resistant and Notch-dependent cancer stem cell population. Cancer Res 2016; 76: 440–452.
Zhou J, Wulfkuhle J, Zhang H, Gu P, Yang Y, Deng J et al. Activation of the PTEN/mTOR/STAT3 pathway in breast cancer stem-like cells is required for viability and maintenance. Proc Natl Acad Sci USA 2007; 104: 16158–16163.
Ma J, Meng Y, Kwiatkowski DJ, Chen X, Peng H, Sun Q et al. Mammalian target of rapamycin regulates murine and human cell differentiation through STAT3/p63/Jagged/Notch cascade. J Clin Invest 2010; 120: 103–114.
Yang Z, Zhang L, Ma A, Liu L, Li J, Gu J et al. Transient mTOR inhibition facilitates continuous growth of liver tumors by modulating the maintenance of CD133+ cell populations. PLoS One 2011; 6: e28405.
Minn AJ, Gupta GP, Siegel PM, Bos PD, Shu W, Giri DD et al. Genes that mediate breast cancer metastasis to lung. Nature 2005; 436: 518–524.
Issa A, Gill JW, Heideman MR, Sahin O, Wiemann S, Dey JH et al. Combinatorial targeting of FGF and ErbB receptors blocks growth and metastatic spread of breast cancer models. Breast Cancer Res 2013; 15: R8.
Chen L, Yang S, Jakoncic J, Zhang JJ, Huang XY . Migrastatin analogues target fascin to block tumour metastasis. Nature 2010; 464: 1062–1066.
Cariati M, Naderi A, Brown JP, Smalley MJ, Pinder SE, Caldas C et al. Alpha-6 integrin is necessary for the tumourigenicity of a stem cell-like subpopulation within the MCF7 breast cancer cell line. Int J Cancer 2008; 122: 298–304.
Piva M, Domenici G, Iriondo O, Rabano M, Simoes BM, Comaills V et al. Sox2 promotes tamoxifen resistance in breast cancer cells. EMBO Mol Med 2014; 6: 66–79.
TCGA. Comprehensive molecular portraits of human breast tumours. Nature 2012; 490: 61–70.
Wong DJ, Liu H, Ridky TW, Cassarino D, Segal E, Chang HY . Module map of stem cell genes guides creation of epithelial cancer stem cells. Cell Stem Cell 2008; 2: 333–344.
Ben-Porath I, Thomson MW, Carey VJ, Ge R, Bell GW, Regev A et al. An embryonic stem cell-like gene expression signature in poorly differentiated aggressive human tumors. Nat Genet 2008; 40: 499–507.
Kim J, Woo AJ, Chu J, Snow JW, Fujiwara Y, Kim CG et al. A Myc network accounts for similarities between embryonic stem and cancer cell transcription programs. Cell 2010; 143: 313–324.
Lim E, Wu D, Pal B, Bouras T, Asselin-Labat ML, Vaillant F et al. Transcriptome analyses of mouse and human mammary cell subpopulations reveal multiple conserved genes and pathways. Breast Cancer Res 2010; 12: R21.
Koren S, Reavie L, Couto JP, De Silva D, Stadler MB, Roloff T et al. PIK3CA(H1047R) induces multipotency and multi-lineage mammary tumours. Nature 2015; 525: 114–118.
Van Keymeulen A, Lee MY, Ousset M, Brohee S, Rorive S, Giraddi RR et al. Reactivation of multipotency by oncogenic PIK3CA induces breast tumour heterogeneity. Nature 2015; 525: 119–123.
Huang R, Huang D, Dai W, Yang F . Overexpression of HMGA1 correlates with the malignant status and prognosis of breast cancer. Mol Cell Biochem 2015; 404: 251–257.
Kormish JD, Sinner D, Zorn AM . Interactions between SOX factors and Wnt/beta-catenin signaling in development and disease. Dev Dyn 2010; 239: 56–68.
Kataoka K, Sato T, Yoshimi A, Goyama S, Tsuruta T, Kobayashi H et al. Evi1 is essential for hematopoietic stem cell self-renewal, and its expression marks hematopoietic cells with long-term multilineage repopulating activity. J Exp Med 2011; 208: 2403–2416.
Patel JB, Appaiah HN, Burnett RM, Bhat-Nakshatri P, Wang G, Mehta R et al. Control of EVI-1 oncogene expression in metastatic breast cancer cells through microRNA miR-22. Oncogene 2011; 30: 1290–1301.
Guo W, Keckesova Z, Donaher JL, Shibue T, Tischler V, Reinhardt F et al. Slug and Sox9 cooperatively determine the mammary stem cell state. Cell 2012; 148: 1015–1028.
Larsimont JC, Youssef KK, Sanchez-Danes A, Sukumaran V, Defrance M, Delatte B et al. Sox9 controls self-renewal of oncogene targeted cells and links tumor initiation and invasion. Cell Stem Cell 2015; 17: 60–73.
Ye X, Tam WL, Shibue T, Kaygusuz Y, Reinhardt F, Ng Eaton E et al. Distinct EMT programs control normal mammary stem cells and tumour-initiating cells. Nature 2015; 525: 256–260.
Sflomos G, Dormoy V, Metsalu T, Jeitziner R, Battista L, Scabia V et al. A preclinical model for ERalpha-positive breast cancer points to the epithelial microenvironment as determinant of luminal phenotype and hormone response. Cancer Cell 2016; 29: 407–422.
Ginestier C, Hur MH, Charafe-Jauffret E, Monville F, Dutcher J, Brown M et al. ALDH1 is a marker of normal and malignant human mammary stem cells and a predictor of poor clinical outcome. Cell Stem Cell 2007; 1: 555–567.
Cheung KJ, Gabrielson E, Werb Z, Ewald AJ . Collective invasion in breast cancer requires a conserved basal epithelial program. Cell 2013; 155: 1639–1651.
Garnett MJ, Edelman EJ, Heidorn SJ, Greenman CD, Dastur A, Lau KW et al. Systematic identification of genomic markers of drug sensitivity in cancer cells. Nature 2012; 483: 570–575.
Clevers H . The cancer stem cell: premises, promises and challenges. Nat Med 2011; 17: 313–319.
Janssen EA, Baak JP, Guervos MA, van Diest PJ, Jiwa M, Hermsen MA . In lymph node-negative invasive breast carcinomas, specific chromosomal aberrations are strongly associated with high mitotic activity and predict outcome more accurately than grade, tumour diameter, and oestrogen receptor. J Pathol 2003; 201: 555–561.
Hagerstrand D, Tong A, Schumacher SE, Ilic N, Shen RR, Cheung HW et al. Systematic interrogation of 3q26 identifies TLOC1 and SKIL as cancer drivers. Cancer Discov 2013; 3: 1044–1057.
Bard-Chapeau EA, Gunaratne J, Kumar P, Chua BQ, Muller J, Bard FA et al. EVI1 oncoprotein interacts with a large and complex network of proteins and integrates signals through protein phosphorylation. Proc Natl Acad Sci USA 2013; 110: E2885–E2894.
Stavropoulou V, Kaspar S, Brault L, Sanders MA, Juge S, Morettini S et al. MLL-AF9 expression in hematopoietic stem cells drives a highly invasive AML expressing EMT-related genes linked to poor outcome. Cancer Cell 2016; 30: 43–58.
Dutta P, Bui T, Bauckman KA, Keyomarsi K, Mills GB, Nanjundan M . EVI1 splice variants modulate functional responses in ovarian cancer cells. Mol Oncol 2013; 7: 647–668.
Holder AM, Akcakanat A, Adkins F, Evans K, Chen H, Wei C et al. Epithelial to mesenchymal transition is associated with rapamycin resistance. Oncotarget 2015; 6: 19500–19513.
Malladi S, Macalinao DG, Jin X, He L, Basnet H, Zou Y et al. Metastatic latency and immune evasion through autocrine inhibition of WNT. Cell 2016; 165: 45–60.
Chao CH, Chang CC, Wu MJ, Ko HW, Wang D, Hung MC et al. MicroRNA-205 signaling regulates mammary stem cell fate and tumorigenesis. J Clin Invest 2014; 124: 3093–3106.
Malhotra GK, Zhao X, Edwards E, Kopp JL, Naramura M, Sander M et al. The role of Sox9 in mouse mammary gland development and maintenance of mammary stem and luminal progenitor cells. BMC Dev Biol 2014; 14: 47.
Pool SE, Bison S, Koelewijn SJ, van der Graaf LM, Melis M, Krenning EP et al. mTOR inhibitor RAD001 promotes metastasis in a rat model of pancreatic neuroendocrine cancer. Cancer Res 2013; 73: 12–18.
Campone M, Bachelot T, Gnant M, Deleu I, Rugo HS, Pistilli B et al. Effect of visceral metastases on the efficacy and safety of everolimus in postmenopausal women with advanced breast cancer: subgroup analysis from the BOLERO-2 study. Eur J Cancer 2013; 49: 2621–2632.
Rodrik-Outmezguine VS, Okaniwa M, Yao Z, Novotny CJ, McWhirter C, Banaji A et al. Overcoming mTOR resistance mutations with a new-generation mTOR inhibitor. Nature 2016; 534: 272–276.
Koboldt DC, Fulton RS, McLellan MD, Schmidt H, Kalicki-Veizer J, McMichael JF et al. Comprehensive molecular portraits of human breast tumours. Nature 2012; 490: 61–70.
Subramanian A, Tamayo P, Mootha VK, Mukherjee S, Ebert BL, Gillette MA et al. Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc Natl Acad Sci USA 2005; 102: 15545–15550.
Huang DW, Sherman BT, Tan Q, Collins JR, Alvord WG, Roayaei J et al. The DAVID gene functional classification tool: a novel biological module-centric algorithm to functionally analyze large gene lists. Genome Biol 2007; 8: R183.
Palafox M, Ferrer I, Pellegrini P, Vila S, Hernandez-Ortega S, Urruticoechea A et al. RANK induces epithelial-mesenchymal transition and stemness in human mammary epithelial cells and promotes tumorigenesis and metastasis. Cancer Res 2012; 72: 2879–2888.
Aslakson CJ, Miller FR . Selective events in the metastatic process defined by analysis of the sequential dissemination of subpopulations of a mouse mammary tumor. Cancer Res 1992; 52: 1399–1405.
Morales M, Arenas EJ, Urosevic J, Guiu M, Fernandez E, Planet E et al. RARRES3 suppresses breast cancer lung metastasis by regulating adhesion and differentiation. EMBO Mol Med 2014; 6: 865–881.
Witten IH, Frank E, Trigg L, Hall M, Holmes G, Cunningham SJ . Weka: practical machine learning tools and techniques with Java implementations. Proceedings of the ICONIP/ANZIIS/ANNES’99 Workshop on Emerging Knowledge Engineering and Connectionist-Based Information Systems 1999, pp 192–196.
Puente XS, Pinyol M, Quesada V, Conde L, Ordonez GR, Villamor N et al. Whole-genome sequencing identifies recurrent mutations in chronic lymphocytic leukaemia. Nature 2011; 475: 101–105.
Zhang Y, Liu T, Meyer CA, Eeckhoute J, Johnson DS, Bernstein BE et al. Model-based analysis of ChIP-Seq (MACS). Genome Biol 2008; 9: R137.
Acknowledgements
We thank Ander Urruticoechea, Gabriel Capellá and George Thomas for helpful discussion; Ana Isabel Extremera, Antoni Xaubet and Julio Ancochea for administrative support; and Sebastien Tosi from the Advanced Digital Microscopy Facility at IRB Barcelona for image analysis. We also thank Xiyun Deng, Yanna Cao, Tien C Ko, Yi Zhang (The University of Texas Health Science Centre at Houston) and Adrian W Moore (RIKEN Brain Science Institute) for their support in evaluating antibodies against EVI1. This study was supported by the following bodies and grants: the Scientific Foundation ‘Asociación Española Contra el Cáncer’ (AECC, Stable Coordinated Group, Hereditary Cancer); the BBVA Foundation; the Eugenio Rodríguez Pascual Foundation grant 2012; Generalitat de Catalunya AGAUR SGR 2012 grants 283, 290 and 312, and SGR 2014 grants 364, 530, and 535; Spanish Ministry of Health ISCIII FIS grants PI10/00057, PI10/00222, PI10/01422, PI12/01528, PI13/00132, and PI14/00336. ISCIII RTICC grants RD06/0020/1051, RD12/0036/0007, RD12/0036/0008 and RD12/0036/0063; Spanish Ministry of Science and Innovation, ‘Fondo Europeo de Desarrollo Regional (FEDER), una manera de hacer Europa’, MINECO grants SAF2010-20203 and SAF2013-46196; and the Telemaraton 2014 ‘Todos Somos Raros, Todos Somos Únicos’ grant P35. EJA was supported by ‘la Caixa’ PhD fellowship program, F Iorio was supported by a fellowship from the EMBL-EBI and the Wellcome Trust Sanger Institute Postdoctoral (ESPOD) program, and NL-B, ME and RRG were supported by ICREA. No funding bodies had any role in study design, data collection and analysis, decision to publish, or preparation of the manuscript, either in the submission form or the text of the manuscript.
Author contributions
RRG and MAP: performed the design and interpretation of experiments, and wrote the manuscript; FM, EJA, HA, GRG, J Boni, XP, CH, AL, HS, MS, LG-B, XS, J Cerón and JG-M: molecular and cell biology experiments; FM, EJA, HA, SD, MHB-H and A Villanueva: in vivo studies; FM, HA, MM and MDO: chromatin immunoprecipitation assays; JS-M, FI, AI, DC, NB, LP, AG, NL-B, LG-A and JS-R: bioinformatic analyses; FM, HA, J Boni, S Puertas, NG, VH, MM-I and AF: immunohistochemistry experiments; AR-S, A Martínez, MPB, M Gil, CF, AF, IM, S Pernas, MJP, XA, MAS, RB, EC, SM, JV, A Velasco, XM-G, MAQ, AS and GM-B: tissue microarray studies; RV-M and XSP: exome analyses; A Petit, A Vidal, IC, TS and GV: pathological evaluations; AMS, VW, MPL, M Nellist, JVS-M, ME, MJ, LB and JWMM: genetic analyses; FM, HA, J Boni, EG-S and A Cordero: three-dimensional cell culture experiments; JH-L, SRC and J Cortés: breast cancer tumor analyses; JL, FC, IB, A Perkins, J Brunet, FV, OC, M Graupera, NM-M, A Matheu, A Carracedo, TFG, EEWC, MS-C, M Nanjundan and CL: experimental design and interpretation.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies this paper on the Oncogene website
Supplementary information
Rights and permissions
This work is licensed under a Creative Commons Attribution-NonCommercial-NoDerivs 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-nd/4.0/
About this article
Cite this article
Mateo, F., Arenas, E., Aguilar, H. et al. Stem cell-like transcriptional reprogramming mediates metastatic resistance to mTOR inhibition. Oncogene 36, 2737–2749 (2017). https://doi.org/10.1038/onc.2016.427
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/onc.2016.427
This article is cited by
-
RPTOR blockade suppresses brain metastases of NSCLC by interfering the ceramide metabolism via hijacking YY1 binding
Journal of Experimental & Clinical Cancer Research (2024)
-
MECOM permits pancreatic acinar cell dedifferentiation avoiding cell death under stress conditions
Cell Death & Differentiation (2021)
-
Aspirin enhances cisplatin sensitivity of resistant non-small cell lung carcinoma stem-like cells by targeting mTOR-Akt axis to repress migration
Scientific Reports (2019)
-
Emerging Role of SOX Proteins in Breast Cancer Development and Maintenance
Journal of Mammary Gland Biology and Neoplasia (2019)