Abstract
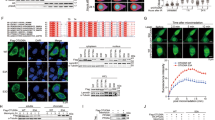
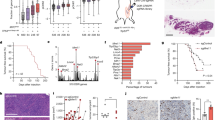
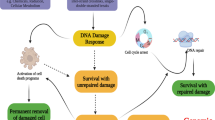
Chromatin DNA damage response (DDR) is orchestrated by the E3 ubiquitin ligase ring finger protein 168 (RNF168), resulting in ubiquitin-dependent recruitment of DDR factors and tumor suppressors breast cancer 1 (BRCA1) and p53 binding protein 1 (53BP1). This ubiquitin signaling regulates pathway choice for repair of DNA double-strand breaks (DSB), toxic lesions whose frequency increases during tumorigenesis. Recruitment of 53BP1 curbs DNA end resection, thereby limiting homologous recombination (HR) and directing DSB repair toward error-prone non-homologous end joining (NHEJ). Under cancer-associated ubiquitin starvation conditions reflecting endogenous or treatment-evoked proteotoxic stress, the ubiquitin-dependent accrual of 53BP1 and BRCA1 at the DNA damage sites is attenuated or lost. Challenging this current paradigm, here we identified diverse human cancer cell lines that display 53BP1 recruitment to DSB sites even under proteasome inhibitor-induced proteotoxic stress, that is, under substantial depletion of free ubiquitin. We show that central to this unexpected phenotype is overabundance of RNF168 that enables more efficient exploitation of the residual-free ubiquitin. Cells with elevated RNF168 are more resistant to combined treatment by ionizing radiation and proteasome inhibition, suggesting that such aberrant RNF168-mediated signaling might reflect adaptation to chronic proteotoxic and genotoxic stresses experienced by tumor cells. Moreover, the overabundant RNF168 and the ensuing unorthodox recruitment patterns of 53BP1, RIF1 and REV7 (monitored on laser micro-irradiation-induced DNA damage) shift the DSB repair balance from HR toward NHEJ, a scenario accompanied by enhanced chromosomal instability/micronuclei formation and sensitivity under replication stress-inducing treatments with camptothecin or poly(ADP-ribose) polymerase (PARP) inhibitor. Overall, our data suggest that the deregulated RNF168/53BP1 pathway could promote tumorigenesis by selecting for a more robust, better stress-adapted cancer cell phenotype, through altered DNA repair, fueling genomic instability and tumor heterogeneity. Apart from providing insights into cancer (patho)biology, the elevated RNF168, documented here also by immunohistochemistry on human clinical tumor specimens, may impact responses to standard-of-care and some emerging targeted cancer therapies.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 50 print issues and online access
$259.00 per year
only $5.18 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout











Similar content being viewed by others
Abbreviations
- 53BP1:
-
p53 binding protein 1
- BRCA1:
-
breast cancer 1
- DDR:
-
DNA damage response
- DSB:
-
double-strand break
- HR:
-
homologous recombination
- IRIF:
-
ionizing radiation induced foci
- NHEJ:
-
non-homologous end joining
- PARP1:
-
poly(ADP-ribose) polymerase 1
- RNF168:
-
ring finger protein 168.
References
Deshaies RJ . Proteotoxic crisis, the ubiquitin-proteasome system, and cancer therapy. BMC Biol 2014; 12: 94.
Halazonetis TD, Gorgoulis VG, Bartek J . An oncogene-induced DNA damage model for cancer development. Science 2008; 319: 1352–1355.
Luo J, Solimini NL, Elledge SJ . Principles of cancer therapy: oncogene and non-oncogene addiction. Cell 2009; 136: 823–837.
Bartek J, Bartkova J, Lukas J . DNA damage signalling guards against activated oncogenes and tumour progression. Oncogene 2007; 26: 7773–7779.
Jackson SP, Bartek J . The DNA-damage response in human biology and disease. Nature 2009; 461: 1071–1078.
Bartkova J, Horejsi Z, Koed K, Kramer A, Tort F, Zieger K et al. DNA damage response as a candidate anti-cancer barrier in early human tumorigenesis. Nature 2005; 434: 864–870.
Bartkova J, Rezaei N, Liontos M, Karakaidos P, Kletsas D, Issaeva N et al. Oncogene-induced senescence is part of the tumorigenesis barrier imposed by DNA damage checkpoints. Nature 2006; 444: 633–637.
Evangelou K, Bartkova J, Kotsinas A, Pateras IS, Liontos M, Velimezi G et al. The DNA damage checkpoint precedes activation of ARF in response to escalating oncogenic stress during tumorigenesis. Cell Death Differ 2013; 20: 1485–1497.
Zhao Y, Brickner JR, Majid MC, Mosammaparast N . Crosstalk between ubiquitin and other post-translational modifications on chromatin during double-strand break repair. Trends Cell Biol 2014; 24: 426–434.
Lukas J, Lukas C, Bartek J . More than just a focus: the chromatin response to DNA damage and its role in genome integrity maintenance. Nat Cell Biol 2011; 13: 1161–1169.
Reinhardt HC, Yaffe MB . Phospho-Ser/Thr-binding domains: navigating the cell cycle and DNA damage response. Nat Rev Mol Cell Biol 2013; 14: 563–580.
Thorslund T, Ripplinger A, Hoffmann S, Wild T, Uckelmann M, Villumsen B et al. Histone H1 couples initiation and amplification of ubiquitin signalling after DNA damage. Nature 2015; 527: 389–393.
Doil C, Mailand N, Bekker-Jensen S, Menard P, Larsen DH, Pepperkok R et al. RNF168 binds and amplifies ubiquitin conjugates on damaged chromosomes to allow accumulation of repair proteins. Cell 2009; 136: 435–446.
Stewart GS, Panier S, Townsend K, Al-Hakim AK, Kolas NK, Miller ES et al. The RIDDLE syndrome protein mediates a ubiquitin-dependent signaling cascade at sites of DNA damage. Cell 2009; 136: 420–434.
Bunting SF, Callen E, Wong N, Chen HT, Polato F, Gunn A et al. 53BP1 inhibits homologous recombination in Brca1-deficient cells by blocking resection of DNA breaks. Cell 2010; 141: 243–254.
Daley JM, Sung P . 53BP1, BRCA1, and the choice between recombination and end joining at DNA double-strand breaks. Mol Cell Biol 2014; 34: 1380–1388.
Densham RM, Garvin AJ, Stone HR, Strachan J, Baldock RA, Daza-Martin M et al. Human BRCA1-BARD1 ubiquitin ligase activity counteracts chromatin barriers to DNA resection. Nat Struct Mol Biol 2016; 23: 647–655.
Callen E, Di Virgilio M, Kruhlak MJ, Nieto-Soler M, Wong N, Chen HT et al. 53BP1 mediates productive and mutagenic DNA repair through distinct phosphoprotein interactions. Cell 2013; 153: 1266–1280.
Chapman JR, Barral P, Vannier JB, Borel V, Steger M, Tomas-Loba A et al. RIF1 is essential for 53BP1-dependent nonhomologous end joining and suppression of DNA double-strand break resection. Mol Cell 2013; 49: 858–871.
Xu G, Chapman JR, Brandsma I, Yuan J, Mistrik M, Bouwman P et al. REV7 counteracts DNA double-strand break resection and affects PARP inhibition. Nature 2015; 521: 541–544.
Zimmermann M, Lottersberger F, Buonomo SB, Sfeir A, de Lange T . 53BP1 regulates DSB repair using Rif1 to control 5' end resection. Science 2013; 339: 700–704.
Zong D, Callen E, Pegoraro G, Lukas C, Lukas J, Nussenzweig A . Ectopic expression of RNF168 and 53BP1 increases mutagenic but not physiological non-homologous end joining. Nucleic Acids Res 2015; 43: 4950–4961.
Botuyan MV, Lee J, Ward IM, Kim JE, Thompson JR, Chen J et al. Structural basis for the methylation state-specific recognition of histone H4-K20 by 53BP1 and Crb2 in DNA repair. Cell 2006; 127: 1361–1373.
Fradet-Turcotte A, Canny MD, Escribano-Diaz C, Orthwein A, Leung CC, Huang H et al. 53BP1 is a reader of the DNA-damage-induced H2A Lys 15 ubiquitin mark. Nature 2013; 499: 50–54.
Acs K, Luijsterburg MS, Ackermann L, Salomons FA, Hoppe T, Dantuma NP . The AAA-ATPase VCP/p97 promotes 53BP1 recruitment by removing L3MBTL1 from DNA double-strand breaks. Nat Struct Mol Biol 2011; 18: 1345–1350.
Mallette FA, Mattiroli F, Cui G, Young LC, Hendzel MJ, Mer G et al. RNF8- and RNF168-dependent degradation of KDM4A/JMJD2A triggers 53BP1 recruitment to DNA damage sites. EMBO J 2012; 31: 1865–1878.
Devgan SS, Sanal O, Doil C, Nakamura K, Nahas SA, Pettijohn K et al. Homozygous deficiency of ubiquitin-ligase ring-finger protein RNF168 mimics the radiosensitivity syndrome of ataxia-telangiectasia. Cell Death Differ 2011; 18: 1500–1506.
Gudjonsson T, Altmeyer M, Savic V, Toledo L, Dinant C, Grofte M et al. TRIP12 and UBR5 suppress spreading of chromatin ubiquitylation at damaged chromosomes. Cell 2012; 150: 697–709.
Dantuma NP, Groothuis TA, Salomons FA, Neefjes J . A dynamic ubiquitin equilibrium couples proteasomal activity to chromatin remodeling. J Cell Biol 2006; 173: 19–26.
Jacquemont C, Taniguchi T . Proteasome function is required for DNA damage response and fanconi anemia pathway activation. Cancer Res 2007; 67: 7395–7405.
Bekker-Jensen S, Rendtlew Danielsen J, Fugger K, Gromova I, Nerstedt A, Lukas C et al. HERC2 coordinates ubiquitin-dependent assembly of DNA repair factors on damaged chromosomes. Nat Cell Biol 2010; 12: 80–86. sup pp 1-12.
Panier S, Boulton SJ . Double-strand break repair: 53BP1 comes into focus. Nat Rev Mol Cell Biol 2014; 15: 7–18.
Chauhan D, Hideshima T, Anderson KC . Proteasome inhibition in multiple myeloma: therapeutic implication. Annu Rev Pharmacol Toxicol 2005; 45: 465–476.
Gething MJ . Role and regulation of the ER chaperone BiP. Semin Cell Dev Biol 1999; 10: 465–472.
Livraghi L, Garber JE . PARP inhibitors in the management of breast cancer: current data and future prospects. BMC Med 2015; 13: 188.
Ricks TK, Chiu HJ, Ison G, Kim G, McKee AE, Kluetz P et al. Successes and challenges of PARP inhibitors in cancer therapy. Front Oncol 2015; 5: 222.
Bouwman P, Aly A, Escandell JM, Pieterse M, Bartkova J, van der Gulden H et al. 53BP1 loss rescues BRCA1 deficiency and is associated with triple-negative and BRCA-mutated breast cancers. Nat Struct Mol Biol 2010, biology 17: 688–695.
Certo MT, Ryu BY, Annis JE, Garibov M, Jarjour J, Rawlings DJ et al. Tracking genome engineering outcome at individual DNA breakpoints. Nat Methods 2011; 8: 671–676.
Rodgers K, McVey M . Error-prone repair of DNA double-strand breaks. J Cell Physiol 2016; 231: 15–24.
Ma T, Chen Y, Zhang F, Yang CY, Wang S, Yu X . RNF111-dependent neddylation activates DNA damage-induced ubiquitination. Mol Cell 2013; 49: 897–907.
Tang J, Cho NW, Cui G, Manion EM, Shanbhag NM, Botuyan MV et al. Acetylation limits 53BP1 association with damaged chromatin to promote homologous recombination. Nat Struct Mol Biol 2013; 20: 317–325.
Hjerpe R, Thomas Y, Chen J, Zemla A, Curran S, Shpiro N et al. Changes in the ratio of free NEDD8 to ubiquitin triggers NEDDylation by ubiquitin enzymes. Biochem J 2012; 441: 927–936.
Watanabe S, Watanabe K, Akimov V, Bartkova J, Blagoev B, Lukas J et al. JMJD1C demethylates MDC1 to regulate the RNF8 and BRCA1-mediated chromatin response to DNA breaks. Nat Struct Mol Biol 2013; 20: 1425–1433.
Kocylowski MK, Rey AJ, Stewart GS, Halazonetis TD . Ubiquitin-H2AX fusions render 53BP1 recruitment to DNA damage sites independent of RNF8 or RNF168. Cell Cycle 2015; 14: 1748–1758.
Mattiroli F, Vissers JH, van Dijk WJ, Ikpa P, Citterio E, Vermeulen W et al. RNF168 ubiquitinates K13-15 on H2A/H2AX to drive DNA damage signaling. Cell 2012; 150: 1182–1195.
Munoz MC, Laulier C, Gunn A, Cheng A, Robbiani DF, Nussenzweig A et al. RING finger nuclear factor RNF168 is important for defects in homologous recombination caused by loss of the breast cancer susceptibility factor BRCA1. J Biol Chem 2012; 287: 40618–40628.
Munoz MC, Yanez DA, Stark JM . An RNF168 fragment defective for focal accumulation at DNA damage is proficient for inhibition of homologous recombination in BRCA1 deficient cells. Nucleic Acids Res 2014; 42: 7720–7733.
Inbar-Rozensal D, Castiel A, Visochek L, Castel D, Dantzer F, Izraeli S et al. A selective eradication of human nonhereditary breast cancer cells by phenanthridine-derived polyADP-ribose polymerase inhibitors. Breast Cancer Res 2009; 11: R78.
Burrell RA, McClelland SE, Endesfelder D, Groth P, Weller MC, Shaikh N et al. Replication stress links structural and numerical cancer chromosomal instability. Nature 2013; 494: 492–496.
Di Micco R, Fumagalli M, Cicalese A, Piccinin S, Gasparini P, Luise C et al. Oncogene-induced senescence is a DNA damage response triggered by DNA hyper-replication. Nature 2006; 444: 638–642.
Gorgoulis VG, Vassiliou LV, Karakaidos P, Zacharatos P, Kotsinas A, Liloglou T et al. Activation of the DNA damage checkpoint and genomic instability in human precancerous lesions. Nature 2005; 434: 907–913.
Chen L, Madura K . Increased proteasome activity, ubiquitin-conjugating enzymes, and eEF1A translation factor detected in breast cancer tissue. Cancer Res 2005; 65: 5599–5606.
Velimezi G, Liontos M, Vougas K, Roumeliotis T, Bartkova J, Sideridou M et al. Functional interplay between the DNA-damage-response kinase ATM and ARF tumour suppressor protein in human cancer. Nat Cell Biol 2013; 15: 967–977.
Greer EL, Brunet A . FOXO transcription factors at the interface between longevity and tumor suppression. Oncogene 2005; 24: 7410–7425.
Tran H, Brunet A, Grenier JM, Datta SR, Fornace AJ Jr., DiStefano PS et al. DNA repair pathway stimulated by the forkhead transcription factor FOXO3a through the Gadd45 protein. Science 2002; 296: 530–534.
Mistrik M, Vesela E, Furst T, Hanzlikova H, Frydrych I, Gursky J et al. Cells and stripes: a novel quantitative photo-manipulation technique. Sci Rep 2016; 6: 19567.
Wiederschain D, Wee S, Chen L, Loo A, Yang G, Huang A et al. Single-vector inducible lentiviral RNAi system for oncology target validation. Cell Cycle 2009; 8: 498–504.
Wee S, Wiederschain D, Maira SM, Loo A, Miller C, deBeaumont R et al. PTEN-deficient cancers depend on PIK3CB. Proc Natl Acad Sci USA 2008; 105: 13057–13062.
Tiscornia G, Singer O, Verma IM . Production and purification of lentiviral vectors. Nat Protoc 2006; 1: 241–245.
Acknowledgements
We thank Jan Bouchal, Katerina Bouchalova and our colleagues from the Laboratory of Genome Integrity for technical assistance, suggestions and comments on the manuscript. This work was supported by grants from the following foundations: Grant Agency of the Czech Republic 13-17555S, Czech National Program of Sustainability LO1304, the Kellner Family Foundation, the Norwegian Financial Mechanism CZ09 (Project PHOSCAN 7F14061), MEYS CR (LM2015062 Czech-BioImaging), the internal grant IGA-LF-2016-030, the EU operation program CZ.1.07/2.3.00/30.0004, the Danish National Research Foundation (DNRF125, project CARD), Danish Cancer Society, the Swedish Research Council, the Lundbeck Foundation, Cancer Fonden, and the Danish Council for Independent Research.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies this paper on the Oncogene website
Rights and permissions
About this article
Cite this article
Chroma, K., Mistrik, M., Moudry, P. et al. Tumors overexpressing RNF168 show altered DNA repair and responses to genotoxic treatments, genomic instability and resistance to proteotoxic stress. Oncogene 36, 2405–2422 (2017). https://doi.org/10.1038/onc.2016.392
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/onc.2016.392
This article is cited by
-
A drug repurposing strategy for overcoming human multiple myeloma resistance to standard-of-care treatment
Cell Death & Disease (2022)
-
Unravelling roles of error-prone DNA polymerases in shaping cancer genomes
Oncogene (2021)
-
Ubiquitin and the DNA double-strand break repair pathway
Genome Instability & Disease (2020)
-
Rewiring E2F1 with classical NHEJ via APLF suppression promotes bladder cancer invasiveness
Journal of Experimental & Clinical Cancer Research (2019)