Abstract
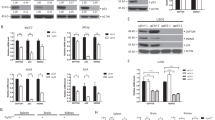
Mutation of p53 is a frequent genetic lesion in pancreatic cancer being an unmet clinical challenge. Mutants of p53 have lost the tumour-suppressive functions of wild type p53. In addition, p53 mutants exert tumour-promoting functions, qualifying them as important therapeutic targets. Here, we show that the class I histone deacetylases HDAC1 and HDAC2 contribute to maintain the expression of p53 mutants in human and genetically defined murine pancreatic cancer cells. Our data reveal that the inhibition of these HDACs with small molecule HDAC inhibitors (HDACi), as well as the specific genetic elimination of HDAC1 and HDAC2, reduce the expression of mutant p53 mRNA and protein levels. We further show that HDAC1, HDAC2 and MYC directly bind to the TP53 gene and that MYC recruitment drops upon HDAC inhibitor treatment. Therefore, our results illustrate a previously unrecognized class I HDAC-dependent control of the TP53 gene and provide evidence for a contribution of MYC. A combined approach targeting HDAC1/HDAC2 and MYC may present a novel and molecularly defined strategy to target mutant p53 in pancreatic cancer.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 50 print issues and online access
$259.00 per year
only $5.18 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
References
Rahib L, Smith BD, Aizenberg R, Rosenzweig AB, Fleshman JM, Matrisian LM . Projecting cancer incidence and deaths to 2030: the unexpected burden of thyroid, liver, and pancreas cancers in the United States. Cancer Res 2014; 74: 2913–2921.
Chang DK, Grimmond SM, Biankin AV . Pancreatic cancer genomics. Curr Opin Genet Dev 2014; 24: 74–81.
Muller PA, Vousden KH . Mutant p53 in cancer: new functions and therapeutic opportunities. Cancer Cell 2014; 25: 304–317.
Oren M, Rotter V . Mutant p53 gain-of-function in cancer. Cold Spring Harb Perspect Biol 2010; 2: a001107.
Freed-Pastor WA, Prives C . Mutant p53: one name, many proteins. Genes Dev 2012; 26: 1268–1286.
Muller PA, Vousden KH . p53 mutations in cancer. Nat Cell Biol 2013; 15: 2–8.
Olive KP, Tuveson DA, Ruhe ZC, Yin B, Willis NA, Bronson RT et al. Mutant p53 gain of function in two mouse models of Li-Fraumeni syndrome. Cell 2004; 119: 847–860.
Doyle B, Morton JP, Delaney DW, Ridgway RA, Wilkins JA, Sansom OJ . p53 mutation and loss have different effects on tumourigenesis in a novel mouse model of pleomorphic rhabdomyosarcoma. J Pathol 2010; 222: 129–137.
Hanel W, Marchenko N, Xu S, Yu SX, Weng W, Moll U . Two hot spot mutant p53 mouse models display differential gain of function in tumorigenesis. Cell Death Differ 2013; 20: 898–909.
Alexandrova EM, Yallowitz AR, Li D, Xu S, Schulz R, Proia DA et al. Improving survival by exploiting tumour dependence on stabilized mutant p53 for treatment. Nature 2015; 523: 352–356.
Schneider G, Henrich A, Greiner G, Wolf V, Lovas A, Wieczorek M et al. Cross talk between stimulated NF-kappaB and the tumor suppressor p53. Oncogene 2010; 29: 2795–2806.
Schneider G, Krämer OH . NFkappaB/p53 crosstalk-a promising new therapeutic target. Biochim Biophys Acta 2011; 1815: 90–103.
Fiorini C, Cordani M, Padroni C, Blandino G, Di Agostino S, Donadelli M . Mutant p53 stimulates chemoresistance of pancreatic adenocarcinoma cells to gemcitabine. Biochim Biophys Acta 2015; 1853: 89–100.
Cooks T, Pateras IS, Tarcic O, Solomon H, Schetter AJ, Wilder S et al. Mutant p53 prolongs NF-kappaB activation and promotes chronic inflammation and inflammation-associated colorectal cancer. Cancer Cell 2013; 23: 634–646.
Morton JP, Timpson P, Karim SA, Ridgway RA, Athineos D, Doyle B et al. Mutant p53 drives metastasis and overcomes growth arrest/senescence in pancreatic cancer. Proc Natl Acad Sci USA 2010; 107: 246–251.
Weissmueller S, Manchado E, Saborowski M, Morris JPt, Wagenblast E, Davis CA et al. Mutant p53 drives pancreatic cancer metastasis through cell-autonomous PDGF receptor beta signaling. Cell 2014; 157: 382–394.
Timpson P, McGhee EJ, Morton JP, von Kriegsheim A, Schwarz JP, Karim SA et al. Spatial regulation of RhoA activity during pancreatic cancer cell invasion driven by mutant p53. Cancer Res 2011; 71: 747–757.
Buchwald M, Krämer OH, Heinzel T . HDACi – targets beyond chromatin. Cancer Lett 2009; 280: 160–167.
Fritsche P, Seidler B, Schüler S, Schnieke A, Göttlicher M, Schmid RM et al. HDAC2 mediates therapeutic resistance of pancreatic cancer cells via the BH3-only protein NOXA. Gut 2009; 58: 1399–1409.
Lehmann A, Denkert C, Budczies J, Buckendahl AC, Darb-Esfahani S, Noske A et al. High class I HDAC activity and expression are associated with RelA/p65 activation in pancreatic cancer in vitro and in vivo. BMC Cancer 2009; 9: 395.
Marshall GM, Gherardi S, Xu N, Neiron Z, Trahair T, Scarlett CJ et al. Transcriptional upregulation of histone deacetylase 2 promotes Myc-induced oncogenic effects. Oncogene 2010; 29: 5957–5968.
Ouaissi M, Silvy F, Loncle C, Ferraz da Silva D, Martins Abreu C, Martinez E et al. Further characterization of HDAC and SIRT gene expression patterns in pancreatic cancer and their relation to disease outcome. PLoS One 2014; 9: e108520.
von Burstin J, Eser S, Paul MC, Seidler B, Brandl M, Messer M et al. E-cadherin regulates metastasis of pancreatic cancer in vivo and is suppressed by a SNAIL/HDAC1/HDAC2 repressor complex. Gastroenterology 2009; 137: 361–371 371 e361–365.
Aghdassi A, Sendler M, Guenther A, Mayerle J, Behn CO, Heidecke CD et al. Recruitment of histone deacetylases HDAC1 and HDAC2 by the transcriptional repressor ZEB1 downregulates E-cadherin expression in pancreatic cancer. Gut 2012; 61: 439–448.
Schüler S, Fritsche P, Diersch S, Arlt A, Schmid RM, Saur D et al. HDAC2 attenuates TRAIL-induced apoptosis of pancreatic cancer cells. Mol Cancer 2010; 9: 80.
Peulen O, Gonzalez A, Peixoto P, Turtoi A, Mottet D, Delvenne P et al. The anti-tumor effect of HDAC inhibition in a human pancreas cancer model is significantly improved by the simultaneous inhibition of cyclooxygenase 2. PLoS One 2013; 8: e75102.
Donadelli M, Costanzo C, Beghelli S, Scupoli MT, Dandrea M, Bonora A et al. Synergistic inhibition of pancreatic adenocarcinoma cell growth by trichostatin A and gemcitabine. Biochim Biophys Acta 2007; 1773: 1095–1106.
Piacentini P, Donadelli M, Costanzo C, Moore PS, Palmieri M, Scarpa A . Trichostatin A enhances the response of chemotherapeutic agents in inhibiting pancreatic cancer cell proliferation. Virchows Arch 2006; 448: 797–804.
Yan W, Liu S, Xu E, Zhang J, Zhang Y, Chen X et al. Histone deacetylase inhibitors suppress mutant p53 transcription via histone deacetylase 8. Oncogene 2013; 32: 599–609.
Wang ZT, Chen ZJ, Jiang GM, Wu YM, Liu T, Yi YM et al. Histone deacetylase inhibitors suppress mutant p53 transcription via HDAC8/YY1 signals in triple negative breast cancer cells. Cell Signal 2016; 28: 506–515.
Li D, Marchenko ND, Moll UM . SAHA shows preferential cytotoxicity in mutant p53 cancer cells by destabilizing mutant p53 through inhibition of the HDAC6-Hsp90 chaperone axis. Cell Death Differ 2011; 18: 1904–1913.
Bradner JE, West N, Grachan ML, Greenberg EF, Haggarty SJ, Warnow T et al. Chemical phylogenetics of histone deacetylases. Nat Chem Biol 2010; 6: 238–243.
Bradner JE, Mak R, Tanguturi SK, Mazitschek R, Haggarty SJ, Ross K et al. Chemical genetic strategy identifies histone deacetylase 1 (HDAC1) and HDAC2 as therapeutic targets in sickle cell disease. Proc Natl Acad Sci USA 2010; 107: 12617–12622.
Bantscheff M, Hopf C, Savitski MM, Dittmann A, Grandi P, Michon AM et al. Chemoproteomics profiling of HDAC inhibitors reveals selective targeting of HDAC complexes. Nat Biotechnol 2011; 29: 255–265.
Lauffer BE, Mintzer R, Fong R, Mukund S, Tam C, Zilberleyb I et al. Histone deacetylase (HDAC) inhibitor kinetic rate constants correlate with cellular histone acetylation but not transcription and cell viability. J Biol Chem 2013; 288: 26926–26943.
Sui X, Shin S, Zhang R, Firozi PF, Yang L, Abbruzzese JL et al. Hdm2 is regulated by K-Ras and mediates p53-independent functions in pancreatic cancer cells. Oncogene 2009; 28: 709–720.
Conradt L, Henrich A, Wirth M, Reichert M, Lesina M, Algul H et al. Mdm2 inhibitors synergize with topoisomerase II inhibitors to induce p53-independent pancreatic cancer cell death. Int J Cancer 2013; 132: 2248–2257.
Azmi AS, Aboukameel A, Banerjee S, Wang Z, Mohammad M, Wu J et al. MDM2 inhibitor MI-319 in combination with cisplatin is an effective treatment for pancreatic cancer independent of p53 function. Eur J Cancer 2010; 46: 1122–1131.
Terzian T, Suh YA, Iwakuma T, Post SM, Neumann M, Lang GA et al. The inherent instability of mutant p53 is alleviated by Mdm2 or p16INK4a loss. Genes Dev 2008; 22: 1337–1344.
Li Y, Guessous F, Kwon S, Kumar M, Ibidapo O, Fuller L et al. PTEN has tumor-promoting properties in the setting of gain-of-function p53 mutations. Cancer Res 2008; 68: 1723–1731.
Lukashchuk N, Vousden KH . Ubiquitination and degradation of mutant p53. Mol Cell Biol 2007; 27: 8284–8295.
Hamilton G, Abraham AG, Morton J, Sampson O, Pefani DE, Khoronenkova S et al. AKT regulates NPM dependent ARF localization and p53mut stability in tumors. Oncotarget 2014; 5: 6142–6167.
Mahboobi S, Sellmer A, Pongratz H, Leonardt M, Krämer O, Böhmer FD et al. Preparation of fused heterocyclic compounds as HDAC6 inhibitors and their uses. PCT Int Appl 2016, WO 2016020369 A1.
Shortt J, Hsu AK, Martin BP, Doggett K, Matthews GM, Doyle MA et al. The drug vehicle and solvent N-methylpyrrolidone is an immunomodulator and antimyeloma compound. Cell Rep 2014; 7: 1009–1019.
Matthews GM, Lefebure M, Doyle MA, Shortt J, Ellul J, Chesi M et al. Preclinical screening of histone deacetylase inhibitors combined with ABT-737, rhTRAIL/MD5-1 or 5-azacytidine using syngeneic Vk*MYC multiple myeloma. Cell Death Dis 2013; 4: e798.
Hingorani SR, Petricoin EF, Maitra A, Rajapakse V, King C, Jacobetz MA et al. Preinvasive and invasive ductal pancreatic cancer and its early detection in the mouse. Cancer Cell 2003; 4: 437–450.
Schönhuber N, Seidler B, Schuck K, Veltkamp C, Schachtler C, Zukowska M et al. A next-generation dual-recombinase system for time- and host-specific targeting of pancreatic cancer. Nat Med 2014; 20: 1340–1347.
Diersch S, Wirth M, Schneeweis C, Jors S, Geisler F, Siveke JT et al. Kras induces EGFR-MYC cross signaling in murine primary pancreatic ductal epithelial cells. Oncogene 2016; 35: 3880–3886.
Saldana-Meyer R, Recillas-Targa F . Transcriptional and epigenetic regulation of the p53 tumor suppressor gene. Epigenetics 2011; 6: 1068–1077.
Kubicek S, Gilbert JC, Fomina-Yadlin D, Gitlin AD, Yuan Y, Wagner FF et al. Chromatin-targeting small molecules cause class-specific transcriptional changes in pancreatic endocrine cells. Proc Natl Acad Sci USA 2012; 109: 5364–5369.
Jamaladdin S, Kelly RD, O'Regan L, Dovey OM, Hodson GE, Millard CJ et al. Histone deacetylase (HDAC) 1 and 2 are essential for accurate cell division and the pluripotency of embryonic stem cells. Proc Natl Acad Sci USA 2014; 111: 9840–9845.
Zeller KI, Jegga AG, Aronow BJ, O'Donnell KA, Dang CV . An integrated database of genes responsive to the Myc oncogenic transcription factor: identification of direct genomic targets. Genome Biol 2003; 4: R69.
Heller G, Schmidt WM, Ziegler B, Holzer S, Mullauer L, Bilban M et al. Genome-wide transcriptional response to 5-aza-2'-deoxycytidine and trichostatin a in multiple myeloma cells. Cancer Res 2008; 68: 44–54.
Yin X, Giap C, Lazo JS, Prochownik EV . Low molecular weight inhibitors of Myc-Max interaction and function. Oncogene 2003; 22: 6151–6159.
Sonnemann J, Marx C, Becker S, Wittig S, Palani CD, Kramer OH et al. p53-dependent and p53-independent anticancer effects of different histone deacetylase inhibitors. Br J Cancer 2014; 110: 656–667.
Peltonen K, Kiviharju TM, Jarvinen PM, Ra R, Laiho M . Melanoma cell lines are susceptible to histone deacetylase inhibitor TSA provoked cell cycle arrest and apoptosis. Pigment Cell Res 2005; 18: 196–202.
Sachweh MC, Drummond CJ, Higgins M, Campbell J, Lain S . Incompatible effects of p53 and HDAC inhibition on p21 expression and cell cycle progression. Cell Death Dis 2013; 4: e533.
Yu X, Guo ZS, Marcu MG, Neckers L, Nguyen DM, Chen GA et al. Modulation of p53, ErbB1, ErbB2, and Raf-1 expression in lung cancer cells by depsipeptide FR901228. J Natl Cancer Inst 2002; 94: 504–513.
Wang Z, Zang C, Cui K, Schones DE, Barski A, Peng W et al. Genome-wide mapping of HATs and HDACs reveals distinct functions in active and inactive genes. Cell 2009; 138: 1019–1031.
Kidder BL, Palmer S . HDAC1 regulates pluripotency and lineage specific transcriptional networks in embryonic and trophoblast stem cells. Nucleic Acids Res 2012; 40: 2925–2939.
Dovey OM, Foster CT, Cowley SM . Emphasizing the positive: A role for histone deacetylases in transcriptional activation. Cell Cycle 2010; 9: 2700–2701.
Kim YJ, Greer CB, Cecchini KR, Harris LN, Tuck DP, Kim TH . HDAC inhibitors induce transcriptional repression of high copy number genes in breast cancer through elongation blockade. Oncogene 2013; 32: 2828–2835.
Greer CB, Tanaka Y, Kim YJ, Xie P, Zhang MQ, Park IH et al. Histone deacetylases positively regulate transcription through the elongation machinery. Cell Rep 2015; 13: 1444–1455.
Reed SM, Quelle DE . p53 acetylation: regulation and consequences. Cancers (Basel) 2014; 7: 30–69.
Wagner T, Brand P, Heinzel T, Krämer OH . Histone deacetylase 2 controls p53 and is a critical factor in tumorigenesis. Biochim Biophys Acta 2014; 1846: 524–538.
Saborowski M, Saborowski A, JPt Morris, Bosbach B, Dow LE, Pelletier J et al. A modular and flexible ESC-based mouse model of pancreatic cancer. Genes Dev 2014; 28: 85–97.
Walz S, Lorenzin F, Morton J, Wiese KE, von Eyss B, Herold S et al. Activation and repression by oncogenic MYC shape tumour-specific gene expression profiles. Nature 2014; 511: 483–487.
Mazur PK, Herner A, Mello SS, Wirth M, Hausmann S, Sanchez-Rivera FJ et al. Combined inhibition of BET family proteins and histone deacetylases as a potential epigenetics-based therapy for pancreatic ductal adenocarcinoma. Nat Med 2015; 21: 1163–1171.
Hessmann E, Schneider G, Ellenrieder V, Siveke JT . MYC in pancreatic cancer: novel mechanistic insights and their translation into therapeutic strategies. Oncogene 2016; 35: 1609–1618.
Wirth M, Schneider G . MYC: a stratification marker for pancreatic cancer therapy. Trends in Cancer 2016; 2: 1–3.
Wirth M, Mahboobi S, Krämer OH, Schneider G . Concepts to target MYC in pancreatic cancer. Mol Cancer Ther 2016; 15: 1792–1798.
Zappasodi R, Cavane A, Iorio MV, Tortoreto M, Guarnotta C, Ruggiero G et al. Pleiotropic antitumor effects of the pan-HDAC inhibitor ITF2357 against c-Myc-overexpressing human B-cell non-Hodgkin lymphomas. Int J Cancer 2014; 135: 2034–2045.
Labisso WL, Wirth M, Stojanovic N, Stauber RH, Schnieke A, Schmid RM et al. MYC directs transcription of MCL1 and eIF4E genes to control sensitivity of gastric cancer cells toward HDAC inhibitors. Cell Cycle 2012; 11: 1593–1602.
Bhadury J, Nilsson LM, Muralidharan SV, Green LC, Li Z, Gesner EM et al. BET and HDAC inhibitors induce similar genes and biological effects and synergize to kill in Myc-induced murine lymphoma. Proc Natl Acad Sci USA 2014; 111: E2721–E2730.
Murakami J, Asaumi J, Kawai N, Tsujigiwa H, Yanagi Y, Nagatsuka H et al. Effects of histone deacetylase inhibitor FR901228 on the expression level of telomerase reverse transcriptase in oral cancer. Cancer Chemother Pharmacol 2005; 56: 22–28.
Kumagai T, Wakimoto N, Yin D, Gery S, Kawamata N, Takai N et al. Histone deacetylase inhibitor, suberoylanilide hydroxamic acid (Vorinostat, SAHA) profoundly inhibits the growth of human pancreatic cancer cells. Int J Cancer 2007; 121: 656–665.
Pei Y, Liu KW, Wang J, Garancher A, Tao R, Esparza LA et al. HDAC and PI3K antagonists cooperate to inhibit growth of MYC-driven medulloblastoma. Cancer Cell 2016; 29: 311–323.
Ronen D, Rotter V, Reisman D . Expression from the murine p53 promoter is mediated by factor binding to a downstream helix-loop-helix recognition motif. Proc Natl Acad Sci USA 1991; 88: 4128–4132.
Reisman D, Elkind NB, Roy B, Beamon J, Rotter V . c-Myc trans-activates the p53 promoter through a required downstream CACGTG motif. Cell Growth Differ 1993; 4: 57–65.
Roy B, Beamon J, Balint E, Reisman D . Transactivation of the human p53 tumor suppressor gene by c-Myc/Max contributes to elevated mutant p53 expression in some tumors. Mol Cell Biol 1994; 14: 7805–7815.
Gui CY, Ngo L, Xu WS, Richon VM, Marks PA . Histone deacetylase (HDAC) inhibitor activation of p21WAF1 involves changes in promoter-associated proteins, including HDAC1. Proc Natl Acad Sci USA 2004; 101: 1241–1246.
Scholz C, Weinert BT, Wagner SA, Beli P, Miyake Y, Qi J et al. Acetylation site specificities of lysine deacetylase inhibitors in human cells. Nat Biotechnol 2015; 33: 415–423.
Geismann C, Grohmann F, Sebens S, Wirths G, Dreher A, Hasler R et al. c-Rel is a critical mediator of NF-kappaB-dependent TRAIL resistance of pancreatic cancer cells. Cell Death Dis 2014; 5: e1455.
Ossewaarde JM, de Vries A, Bestebroer T, Angulo AF . Application of a mycoplasma group-specific PCR for monitoring decontamination of mycoplasma-infected Chlamydia sp. strains. Appl Environ Microbiol 1996; 62: 328–331.
Wirth M, Fritsche P, Stojanovic N, Brandl M, Jaeckel S, Schmid RM et al. A simple and cost-effective method to transfect small interfering RNAs into pancreatic cancer cell lines using polyethylenimine. Pancreas 2011; 40: 144–150.
Wirth M, Stojanovic N, Christian J, Paul MC, Stauber RH, Schmid RM et al. MYC and EGR1 synergize to trigger tumor cell death by controlling NOXA and BIM transcription upon treatment with the proteasome inhibitor bortezomib. Nucleic Acids Res 2014; 42: 10433–10447.
Acknowledgements
We thank Dr E. Olson, Dr T. Jacks, Dr A. Lowy, Dr P. Soriano, and Dr D. Tuveson for providing mouse lines. We thank Dr A. Bradley for support and help during the transfer of mouse lines. This work was supported by: Deutsche Krebshilfe [110908 to G.S., 110909 to O.H.K., and 111273 to M.R.], Wilhelm-Sander Stiftung [2016.004.1 to S.M. and G.S., 2010.078.1 to O.H.K], Else Kröner-Fresenius-Stiftung (2016_A43 to M.W.), Deutsche Forschungsgemeinschaft (DFG) [SCHN 959/2-1 to G.S.; SFB824/C9 to G.S. and D.S.; KR 2291/4-1/MA; 2183/1-1 to O.H.K. and S.M., and KR 2291/5-1 to O.H.K.], and DKTK Joint Funding [to R.R., D.S., and G.S.].
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies this paper on the Oncogene website
Rights and permissions
About this article
Cite this article
Stojanovic, N., Hassan, Z., Wirth, M. et al. HDAC1 and HDAC2 integrate the expression of p53 mutants in pancreatic cancer. Oncogene 36, 1804–1815 (2017). https://doi.org/10.1038/onc.2016.344
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/onc.2016.344
This article is cited by
-
Targeted therapy using nanocomposite delivery systems in cancer treatment: highlighting miR34a regulation for clinical applications
Cancer Cell International (2023)
-
Update on histone deacetylase inhibitors in peripheral T-cell lymphoma (PTCL)
Clinical Epigenetics (2023)
-
Inhibition of STAT3Y705 phosphorylation by Stattic suppresses proliferation and induces mitochondrial-dependent apoptosis in pancreatic cancer cells
Cell Death Discovery (2022)
-
Protein acylation: mechanisms, biological functions and therapeutic targets
Signal Transduction and Targeted Therapy (2022)
-
Targeting the MYC interaction network in B-cell lymphoma via histone deacetylase 6 inhibition
Oncogene (2022)