Abstract
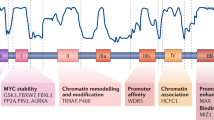
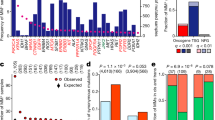
The relevance of changes to the coding sequence of the c-MYC oncogene to malignancy is controversial. Overexpression of a pristine form of MYC is observed in many cancers and is sufficient to drive tumorigenesis in most contexts. Yet missense changes to MYC are found in ~50% of Burkitt’s lymphomas, aggregate within an amino-terminal degron important for proteasomal destruction of MYC, and where examined profoundly enhance the tumorigenic properties of MYC in vitro and in vivo. Much of the controversy surrounding these mutants stems from the limited number of mutations that have been evaluated and their clustering within a single region of the MYC protein; the highly-conserved Myc box I (MbI) element. Here, by analysis of extant genomic data sets, we identify a previously unrecognized hotspot for tumor-associated MYC mutations, located in a conserved central portion of the protein. We show that, despite their distal location in MYC, mutations in this region precisely phenocopy those in MbI in terms of stability, in vitro transformation, growth-promoting properties, in vivo tumorigenesis and ability to escape p53-dependent tumor surveillance mechanisms. The striking parallels between the behavior of tumor-derived mutations in disparate regions of the MYC protein reveals that a common molecular process is disrupted by these mutations, implying an active role for these mutations in tumorigenesis and suggesting that different therapeutic strategies may be needed for treatment of lymphomas expressing wild type versus mutant forms of MYC protein.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 50 print issues and online access
$259.00 per year
only $5.18 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Rabbitts TH, Hamlyn PH, Baer R . Altered nucleotide sequences of a translocated c-myc gene in Burkitt lymphoma. Nature 1983; 306: 760–765.
Westaway D, Payne G, Varmus HE . Proviral deletions and oncogene base-substitutions in insertionally mutagenized c-myc alleles may contribute to the progression of avian bursal tumors. Proc Natl Acad Sci USA 1984; 81: 843–847.
Showe LC, Ballantine M, Nishikura K, Erikson J, Kaji H, Croce CM . Cloning and sequencing of a c-myc oncogene in a Burkitt's lymphoma cell line that is translocated to a germ line alpha switch region. Mol Cell Biol 1985; 5: 501–509.
Murphy W, Sarid J, Taub R, Vasicek T, Battey J, Lenoir G et al. A translocated human c-myc oncogene is altered in a conserved coding sequence. Proc Natl Acad Sci USA 1986; 83: 2939–2943.
Bhatia K, Huppi K, Spangler G, Siwarski D, Iyer R, Magrath I . Point mutations in the c-Myc transactivation domain are common in Burkitt's lymphoma and mouse plasmacytomas. Nat Genet 1993; 5: 56–61.
Bhatia K, Spangler G, Gaidano G, Hamdy N, Dalla-Favera R, Magrath I . Mutations in the coding region of c-myc occur frequently in acquired immunodeficiency syndrome-associated lymphomas. Blood 1994; 84: 883–888.
Clark HM, Yano T, Otsuki T, Jaffe ES, Shibata D, Raffeld M . Mutations in the coding region of c-MYC in AIDS-associated and other aggressive lymphomas. Cancer Res 1994; 54: 3383–3386.
Axelson H, Henriksson M, Wang Y, Magnusson KP, Klein G . The amino-terminal phosphorylation sites of C-MYC are frequently mutated in Burkitt's lymphoma lines but not in mouse plasmacytomas and rat immunocytomas. Eur J Cancer 1995; 31A: 2099–2104.
Hemann MT, Bric A, Teruya-Feldstein J, Herbst A, Nilsson JA, Cordon-Cardo C et al. Evasion of the p53 tumour surveillance network by tumour-derived MYC mutants. Nature 2005; 436: 807–811.
Schmitz R, Young RM, Ceribelli M, Jhavar S, Xiao W, Zhang M et al. Burkitt lymphoma pathogenesis and therapeutic targets from structural and functional genomics. Nature 2012; 490: 116–120.
Giulino-Roth L, Wang K, MacDonald TY, Mathew S, Tam Y, Cronin MT et al. Targeted genomic sequencing of pediatric Burkitt lymphoma identifies recurrent alterations in antiapoptotic and chromatin-remodeling genes. Blood 2012; 120: 5181–5184.
Love C, Sun Z, Jima D, Li G, Zhang J, Miles R et al. The genetic landscape of mutations in Burkitt lymphoma. Nat Genet 2012; 44: 1321–1325.
Salghetti SE, Kim SY, Tansey WP . Destruction of Myc by ubiquitin-mediated proteolysis: cancer-associated and transforming mutations stabilize Myc. Embo J 1999; 18: 717–726.
Welcker M, Orian A, Jin J, Grim JA, Harper JW, Eisenman RN et al. The Fbw7 tumor suppressor regulates glycogen synthase kinase 3 phosphorylation-dependent c-Myc protein degradation. Proc Natl Acad Sci USA 2004; 101: 9085–9090.
Conzen SD, Gottlob K, Kandel ES, Khanduri P, Wagner AJ, O'Leary M et al. Induction of cell cycle progression and acceleration of apoptosis are two separable functions of c-Myc: transrepression correlates with acceleration of apoptosis. Mol Cell Biol 2000; 20: 6008–6018.
Wang X, Cunningham M, Zhang X, Tokarz S, Laraway B, Troxell M et al. Phosphorylation regulates c-Myc's oncogenic activity in the mammary gland. Cancer Res 2011; 71: 925–936.
Egle A, Harris AW, Bouillet P, Cory S . Bim is a suppressor of Myc-induced mouse B cell leukemia. Proc Natl Acad Sci USA 2004; 101: 6164–6169.
McLaughlin J, Chianese E, Witte ON . In vitro transformation of immature hematopoietic cells by the P210 BCR/ABL oncogene product of the Philadelphia chromosome. Proc Natl Acad Sci USA 1987; 84: 6558–6562.
Murphy DJ, Junttila MR, Pouyet L, Karnezis A, Shchors K, Bui DA et al. Distinct thresholds govern Myc's biological output in vivo. Cancer Cell 2008; 14: 447–457.
Faber AC, Corcoran RB, Ebi H, Sequist LV, Waltman BA, Chung E et al. BIM expression in treatment-naive cancers predicts responsiveness to kinase inhibitors. Cancer Discov 2011; 1: 352–365.
Oltersdorf T, Elmore SW, Shoemaker AR, Armstrong RC, Augeri DJ, Belli BA et al. An inhibitor of Bcl-2 family proteins induces regression of solid tumours. Nature 2005; 435: 677–681.
Tworkowski KA, Chakraborty AA, Samuelson AV, Seger YR, Narita M, Hannon GJ et al. Adenovirus E1A targets p400 to induce the cellular oncoprotein Myc. Proc Natl Acad Sci USA 2008; 105: 6103–6108.
Herbst A, Hemann MT, Tworkowski KA, Salghetti SE, Lowe SW, Tansey WP . A conserved element in Myc that negatively regulates its proapoptotic activity. EMBO Rep 2005; 6: 177–183.
Acknowledgements
For reagents, we thank the CSHL Antibody Shared Resource and the Vanderbilt Antibody and Protein Shared Resource. We thank Simone Salghetti for technical assistance. This work was supported by the CSHL Cancer Center Support Grant CA45508, the Vanderbilt Ingram Cancer Center Support grant CA68485 and by US Public Health Service grant CA-13106 from the NCI (WPT/SWL) and Howard Hughes Medical Institute (SWL). SWL is an investigator in the Howard Hughes Medical Institute.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies this paper on the Oncogene website
Supplementary information
Rights and permissions
About this article
Cite this article
Chakraborty, A., Scuoppo, C., Dey, S. et al. A common functional consequence of tumor-derived mutations within c-MYC. Oncogene 34, 2406–2409 (2015). https://doi.org/10.1038/onc.2014.186
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/onc.2014.186
This article is cited by
-
MYC promotes global transcription in part by controlling P-TEFb complex formation via DNA-binding independent inhibition of CDK9 SUMOylation
Science China Life Sciences (2023)
-
Distinct genetic changes reveal evolutionary history and heterogeneous molecular grade of DLBCL with MYC/BCL2 double-hit
Leukemia (2020)
-
MYC dephosphorylation by the PP1/PNUTS phosphatase complex regulates chromatin binding and protein stability
Nature Communications (2018)
-
Glucose induced activation of canonical Wnt signaling pathway in hepatocellular carcinoma is regulated by DKK4
Scientific Reports (2016)