Abstract
Estrogen receptor α (ERα) is initially expressed in the majority of breast cancers and promotes estrogen-dependent cancer progression by regulating the transcription of genes linked to cell proliferation. ERα status is of clinical importance, as ERα-positive breast cancers can be successfully treated by adjuvant therapy with antiestrogens or aromatase inhibitors. Complications arise from the frequent development of drug resistance that might be caused by multiple alterations, including components of ERα signaling, during tumor progression and metastasis. Therefore, insights into the molecular mechanisms that control ERα expression and stability are of utmost importance to improve breast cancer diagnostics and therapeutics. Here we report that the atypical E3 ubiquitin ligase RNF31 stabilizes ERα and facilitates ERα-stimulated proliferation in breast cancer cell lines. We show that depletion of RNF31 decreases the number of cells in the S phase and reduces the levels of ERα and its downstream target genes, including cyclin D1 and c-myc. Analysis of data from clinical samples confirms correlation between RNF31 expression and the expression of ERα target genes. Immunoprecipitation indicates that RNF31 associates with ERα and increases its stability and mono-ubiquitination, dependent on the ubiquitin ligase activity of RNF31. Our data suggest that association of RNF31 and ERα occurs mainly in the cytosol, consistent with the lack of RNF31 recruitment to ERα-occupied promoters. In conclusion, our study establishes a non-genomic mechanism by which RNF31 via stabilizing ERα levels controls the transcription of estrogen-dependent genes linked to breast cancer cell proliferation.
Similar content being viewed by others
Introduction
The relationship between estrogen signaling and breast cancer was revealed nearly 80 years ago.1 The subsequent discovery and clinical application of antiestrogens and of aromatase inhibitors brought significant survival benefits to breast cancer patients.2, 3 However, up to 40% of patients receiving adjuvant therapy eventually relapse, making resistance to endocrine therapy a significant clinical problem.2, 4 Thus, a detailed understanding of the underlying mechanisms and insight into new facets and components of estrogen signaling is critical in developing novel treatment strategies.
Estrogen is known to exert its function by binding to estrogen receptor (ER) subtypes, ERα and ERβ, which belong to the nuclear receptor family of transcription factors.5 ERα has a main role in breast cancer initiation and proliferation. Overexpression of ERα accelerates the G1-S phase transition, correlating with increased levels of oncogenic proteins, including cyclin D1 and c-myc.6 These proteins promote cell cycle progression by decreasing the level of the cyclin-dependent kinase inhibitors p21Cip/WAF1 and p27Kip1, the activity of which is essential for cell cycle progression.7 Clinically, ERα levels in dysplastic breast epithelial cells correlate with the risk of breast cancer and response to endocrine therapy, and two-thirds of all breast cancers express high levels of ERα.8, 9
There are a number of possible and confirmed mechanisms for the dysregulation of ERα that trigger inappropriate estrogen signaling and drug resistance in breast cancer. Some may relate to the transcriptional and epigenetic control of ERα expression,10, 11 others to the control of ERα activity via ligands, growth factors and post-translational modifications.12, 13, 14 However, our understanding of how ERα protein levels and stability are controlled in breast cancers remains largely unclear. As obvious candidates emerge components of the ubiquitin-proteasome system, including numerous E3 ubiquitin ligases, having been shown to have important roles in breast cancer by modulating estrogen signaling. For example, enzymes such as BRCA1, MDM2 and CHIP have been demonstrated to ubiquitinate ERα, suggesting that they directly trigger proteasomal degradation.11, 13, 15, 16 Despite these seemingly distinct mechanisms, a unifying model emerges wherein ERα turnover by the ubiquitin-proteasome system appears tightly linked to ER-mediated transcription. In fact, E3 ligases appear to be recruited to ER-target genes, suggesting that ER ubiquitination and turnover occurs within short time frames at the chromatin and this tight coupling could be a prerequisite for subsequent rounds of transcription.
That the regulation of ERα stability and turnover is complicated is evident from recent observations that other post-translational modifications such as phosphorylation, SUMOylation and acetylation may crosstalk with ubiquitination.11, 17 Additionally, it is known that diverse ER-associated proteins can modulate ERα stability or even trigger proteasomal degradation and turnover, as demonstrated in the case of the oncogenic coactivator SRC3.18 Interestingly, ERα can be mono-ubiquitinated, which, unlike poly-ubiquitination, does not trigger proteasomal degradation.19 However, the consequences of ERα mono-ubiquitination for its stability and function in breast cancer cells, as well as for the cellular factors that specifically trigger and recognize this type of modification, remain to be characterized.
Here, we identify the atypical E3 ubiquitin ligase RNF31 (alias HOIP and ZIBRA) as one such candidate factor. RNF31 was initially cloned from breast cancer cells based on its elevated mRNA expression,20 but its function in breast tumorigenesis and estrogen signaling has not yet been addressed. Our study characterizes a novel non-genomic mechanism that links RNF31 to the control of ERα ubiquitination and stability and thereby to the transcriptional regulation of estrogen-dependent genes and breast cancer cell proliferation.
Results
RNF31 modulates E2-stimulated proliferation of breast cancer cells
To investigate the role of RNF31 in cell proliferation, we knocked down endogenous RNF31 in the ERα-positive breast cancer cell line MCF-7 using RNA interference (Figure 1a). We observed that RNF31 depletion reduced E2-stimulated cell growth (Figure 1b) and decreased E2-stimulated cell cycle progression (Figure 1c and Supplementary Figure S1A). Importantly, RNF31 depletion reduced the number of cells in the S phase to the same extent as the depletion of ERα in E2-treated cells, which correlated with an increase in the number of cells in the G0-G1 phase. Overexpression of ERα partially reversed the reduced number of cells in the S phase following RNF31 depletion (Figure 1d and Supplementary Figure S1B). The actual level of reversion is difficult to interpret as the conditions for small interfering RNA (siRNA) transfection are different from those of plasmid transfection and it is unlikely that all cells transfected with siRNA are also transfected with plasmid. Together these data suggest that ERα is a key mediator of the proliferative effects of RNF31 in this ER-responsive cellular system. However, also in the absence of E2 treatment (Figure 1c, vehicle), RNF31 depletion reduced the number of cells in the S phase, whereas ERα depletion had no effect under these conditions. Additionally, ERα depletion did not mimic RNF31 depletion with regard to the percentage of cells in the G0-G1 and G2-M phases of the cell cycle.
RNF31 depletion inhibits cell proliferation and increases G1 arrest in MCF-7 cells. (a) MCF-7 cells were transfected with siRNF31 or siControl and knockdown efficacy was determined by western blot analysis for RNF31, using GAPDH as internal standard for control (Left panel), and qPCR (Right panel). (b) The WST-1 assay was used to determine the cellular metabolic activity at indicated time points after transfection. Cells are treated for indicated times with 10 nM E2 or vehicle. Experiments were done in triplicates. *P<0.05; **P<0.01 for siRNF31 E2 versus siControl E2. (c) RNF31 knockdown induces arrest in the G1 phase and inhibits the E2-mediated progression into the S phase. The effects of RNF31 knockdown were compared with the effects of ERα knockdown in MCF-7 cells. Cells are treated for 24 h with 10 nM E2 or vehicle. The proportion of cells in each phase was measured by fluorescent-activated cell sorting (FACS). Experiments were done in triplicates. *P<0.05; **P<0.01; ***P<0.001 for siRNF31 versus siControl. All values are mean±s.d. (n=3). (d) Overexpression of ERα partially reverses the reduced number of cells in the S phase following RNF31 depletion. MCF-7 cells were cultured in 10% fetal bovine serum (FBS). The proportion of cells in each phase was measured by FACS. Experiments were done in triplicates. *P<0.05; **P<0.01 for siRNF31 versus siControl; siRNF31 versus siRNF31 plus ERα overexpression. All values are mean±s.d. (n=3).
RNF31 depletion decreases ERα levels and modulates expression of ERα target genes
We then addressed the functional consequences of RNF31 depletion on ERα signaling linked to proliferation control. Western blot analysis revealed that RNF31 depletion significantly downregulated protein levels of ERα in MCF-7 cells (Figure 2a). Importantly, ERα downregulation was observed in a similar manner in MDA-MB175 and T47D cells (Supplementary Figure S2A) and could be achieved using two different individual siRNAs of the siRNA pool (Supplementary Figure S2B). To determine whether effects of RNF31 on ERα protein levels were correlated with effects on ERα transcriptional activity, we assayed ERα reporter gene activity following RNF31 depletion or overexpression. Figure 2b shows that RNF31 depletion leads to reduced ERα reporter gene expression, whereas RNF31 overexpression leads to increased reporter gene expression in the presence and absence of estrogen. RNF31 depletion also reduced the expression of endogenous ERα target genes such as ADORA1, pS2 and Cyclin D1 (Figure 2c). Consistent with this, chromatin immunoprecipitation analysis revealed decreased ERα binding to the promoter regions of target genes following RNF31 depletion (Figure 2d). Supplementary Figure 3A shows that inhibition of RNF31 does not affect the endogenous expression of JUND and GAPDH, which were used as negative controls. Furthermore, Supplementary Figure S3B shows that inhibition of RNF31 effects ERα and nuclear factor-κB (NF-κB) signaling but not Liver X Receptor signaling in luciferase assays. Additionally, Supplementary Figure S3C shows that the lack of effect on Liver X Receptor signaling is independent of the presence of ligand. Thus, the effect of RNF31 on cell signaling shows pathway selectivity. The effect on the NF-kB pathway is not surprising considering the established role of RNF31 in modulating this pathway.21, 22, 23 Consistent with the well-known regulation by ERα of its own expression,24 RNF31 depletion downregulated the expression of ERα mRNA (Supplementary Figure S3D) and the binding of ERα to the known ERα-binding site in the ERα promoter (Figure 2d). Global gene expression analysis followed by sub-network enrichment analysis revealed significant regulation of ERα signaling pathways by RNF31 (Table 1). In line with this, RNF31 affects a large number of ERα target genes, both those that have been shown to be upregulated and those that have been shown to be downregulated, in breast cancer cells (Figure 2e). Thus, RNF31 constitutes a regulator of general ERα signaling and its target genes.
RNF31 depletion decreases ERα protein levels and ERα signaling. (a) RNF31 depletion reduces ERα protein levels. MCF-7 cells were transfected with siRNF31 or siControl and treated with 10 nM E2 or vehicle for 72 h. ERα and RNF31 levels were determined by western blot analysis. GAPDH was used as internal control. (b) RNF31 depletion or overexpression affects ERα-dependent expression of an ERE-luciferase reporter gene. MCF-7 cells were transfected with siRNF31 or siControl or with plasmids expressing Myc-tagged RNF31 or Myc-tag vector alone or together with the ERE reporter plasmid. Subsequently, cells were treated with 10 nM E2 or vehicle. Luciferase activity was measured 48 h after transfection. Shown are data from triplicate measurements. ***P<0.001 for siRNF31 versus siControl and RNF31 overexpression versus control. (c) RNF31 depletion reduces the expression of endogenous ERα target genes. MCF-7 cells were transfected with siRNF31 or siControl. 48 h after transfection, cells were treated with 10 nM E2 or vehicle for 6 h. The expression levels of RNF31 and of endogenous ERα target genes (ADORA1, pS2, cyclinD1) were determined by qPCR from triplicate experiments. **P<0.01; ***P<0.001 for siRNF31 versus siControl. (d) RNF31 depletion decreases ERα recruitment to endogenous target gene promoters. MCF-7 cells were transfected with siRNF31 or siControl. Forty-eight hours post-transfection, cells were treated with 10 nM E2 or vehicle for 30 min and chromatin immunoprecipitation (ChIP) assays were performed with ERα antibody or rabbit immunoglobulin G (IgG) and quantified by qPCR. **P<0.01; ***P<0.001 for siRNF31 versus siControl. (e) Heat map of ERα-regulated genes changed by RNF31 depletion in MCF-7 cells. P<0.001 and fold change >2 was set as cutoff to derive regulated genes. All values are mean±s.d. (n=3).
RNF31 is highly expressed and is correlated to ERα target genes in tumor samples
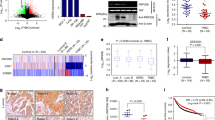
To begin to explore the clinical relevance of the effect of RNF31 on proliferation and estrogen signaling, we analyzed primary breast cancer samples and adjacent tissues for the expression of RNF31 mRNA. We observed high levels of RNF31 expression in tumor tissue compared with adjacent tissues (Figure 3a). Next, we tested known ERα target genes that were also identified as being regulated by RNF31 in MCF-7 cells, for correlation with RNF31 in publically available gene expression profiling data from >2000 breast cancer patients in the TCGA RNA-sequencing25 and KMplot26 databases. Importantly, RNF31 expression correlates with expression of about 70% of the known ERα target genes, in at least one of the clinical gene profiling data sets, identified as being regulated by RNF31 in MCF-7 cells (Table 2). This is exemplified in Figure 3b that shows that the expression of RNF31 positively correlates with the expression of classical ERα target genes, such as TFF1 (pS2) and GREB1 in clinical breast cancer samples. If tumors were separated according to high or low RNF31 expression, a significant correlation to the differential expression of ERα target genes was observed, as further shown in Figure 3c.
RNF31 is highly expressed and is correlated to ERα target genes in tumour samples. (a) RNF31 is highly expressed in breast tumors. The expression of RNF31 was determined for 72 breast tumors and 37 adjacent breast tissues using qPCR. The Student’s t-test was used for comparison. Whiskers, 10th and 90th percentile; box boundaries, 75th and 25th percentile; line within box, median. Dots above the boxes show sample maximum values. asterisk (*) indicates outliers. (b) Correlation between expression of RNF31 and expression of the classical positively regulated ERα target genes GREB1 and pS2 (TFF1). Expression levels are derived from the TCGA mRNA database, test statistics from Pearson product-moment correlation. (c) Heat map showing correlation between RNF31 mRNA levels and mRNA levels of known ERα target genes. Included ERα target genes correspond to genes displaying significant correlation with RNF31 in Supplementary Table S2. RNA expression data are from the TCGA RNA-sequencing database.
RNF31 associates with ERα and increases its stability
Further support for the functional cooperation of RNF31 with ERα was obtained by co-immunoprecipitation (co-IP) of the overexpressed proteins from HEK-293 cells (Supplementary Figure S4A) and of the endogenous proteins from MCF-7 cells (Figure 4a and Supplementary Figures S4B and C). Additionally, overexpression of RNF31 upregulated ERα protein levels in MCF-7 cells under steady-state conditions (Figure 4b). Upon inhibition of protein synthesis by cycloheximide, RNF31 depletion accelerated ERα degradation and this effect was observed both in the presence and absence of estrogen (Figures 4c and d). It is well established that ERα regulates its own expression in MCF-7 cells,24 making it difficult to distinguish direct effects of RNF31 on ERα protein and mRNA levels in this cell line. We therefore performed assays in HEK-293 cells, in which RNF31 but not ERα is expressed. In the presence of the proteasome inhibitor MG132, ERα was stabilized also in the absence of RNF31 and overexpression of RNF31 did not further increase the stability of ERα (Figure 4e). In summary, these data suggest that RNF31 increases ERα protein stability via inhibition of its proteasome-dependent degradation.
RNF31 associates with ERα and increases its stability. (a) Co-IP assays reveal associations between endogenous RNF31 and ERα in MCF-7 cells. (b) Over-expression of RNF31 increases endogenous ERα protein levels in MCF-7 cells. MCF −7 cells were transfected with plasmids expressing Myc-tagged RNF31 or the Myc-tag alone. Forty-eight hours after transfection, whole-protein extracts were prepared and subjected to western blot analysis. The levels of RNF31, ERα and the internal control GAPDH were determined by western blot analysis. The predicted molecular weights are indicated. (c) Depletion of RNF31 decreases endogenous ERα protein stability in MCF-7 cells. Cells were transfected with siRNF31 or siControl. After 72 h, cells were treated with 100 μmol/l cycloheximide for different times as indicated before whole protein extraction. The levels of ERα and the internal control GAPDH were determined by western blot analysis. The experiments were done in triplicates. Image J was applied to quantify ERα and GAPDH density. (d) Depletion of RNF31 decreases endogenous ERα protein stability in MCF-7 cells in the presence of E2. Cells were transfected with siRNF31 or sicontrol. After 72 h, cells were treated with 10 nM E2 for 24 h. Then cells were treated with 100 μmol/l cycloheximide for different times as indicated before whole protein extraction. The levels of RNF31, ERα and the internal control GAPDH were determined by western blot analysis. The experiments were done with triplicates. Image J was applied to quantify the ERα and GAPDH density. (e) Overexpression of RNF31 does not further increase the stability of ERα in the presence of the proteasome inhibitor MG132. HEK-293 cell were transfected with plasmids expressing Myc-tagged RNF31 or the Myc-tag alone. Forty-eight hours post transfection, cells were treated with 100 nmol/l MG132 or vehicle. Cells were harvested 6 h after MGM132 treatment and whole protein extracts were prepared. The levels of RNF31, ERα and the internal control GAPDH were determined by western blot analysis.
The RBR domain of RNF31 is required for the association with, and stabilizing effect on, ERα
To delineate the domains of RNF31 that are important for the association with ERα, wild-type RNF31 and deletion variants thereof (Figure 5a) were expressed together with ERα in HEK-293 cells. Co-IP revealed a requirement of the C-terminal ring between ring fingers (RBR) domain for association with ERα (Figure 5b). We also overexpressed the same RNF31 variants in MCF-7 cells to determine the effect on endogenous ERα. In line with the notion that the RBR domain is required for RNF31 to associate with ERα, it is also required for the regulation of ERα levels (Figure 5c). In contrast, the ubiquitin-associated domain seems to be dispensable for associations with, and stabilization of, ERα. Finally, estrogen response element (ERE)-luciferase assays show that the RBR region of RNF31 is required to increase transcription-dependent ERα signaling in MCF-7 cells (Figure 5d).
The RNF31 RBR domain is required for association with and stabilization of ERα. (a) RNF31 domain structure and deletion mutants used in this study (full-length, ΔUBA and ΔRBR). ZnF_RBZ, putative zinc-finger ubiquitin-binding domain; UBA, ubiquitin-associated domain, mediates interaction with RBCK1/LUBAC; RING-IBR-RING (RBR), atypical E3 ubiquitin ligase domain. (b) RNF31 association with ERα requires the RBR domain. HEK-293 cells were transfected with ERα together with plasmids expressing Myc-tagged full-length RNF31, variants deleting the UBA or RBR domains, respectively, or the Myc-tag alone. Cells were harvested 24 h after transfection and whole-cell extracts were prepared for co-IP. The predicted molecular weights of the RNF31 derivatives and ERα are indicated. (c) The RBR domain is necessary for the RNF31-mediated increase of endogenous ERα protein levels. MCF-7 cells were transfected with plasmids expressing Myc-tagged RNF31 derivatives or the Myc-tag alone, as indicated. 48 h after transfection, whole-cell extracts were prepared and levels of ERα protein assayed by western blot analysis. The predicted molecular weights of RNF31 variants, ERα and the loading control GAPDH are indicated. (d) The RBR domain of RNF31 is required to increase ERα signaling. MCF-7 cells were transfected with plasmids expressing Myc-tagged RNF31 derivatives, or the Myc-tag alone, as indicated, along with an ERE-luciferase reporter plasmid. Luciferase activity was measured 48 h after transfection and calculated from experiments performed in triplicates. Data are shown as mean±s.d. (n=3). ***P<0.001 for wt-RNF31 versus RNF31 deletion domains/empty vector; NA, P>0.05 for wt-RNF31 versus RNF31 deletion domains.
RNF31, via its RBR domain, triggers ERα mono-ubiquitination
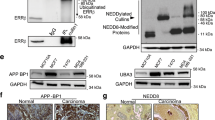
The requirement of the RNF31 RBR domain, implicated in catalyzing atypical ubiquitination of substrates, led us to investigate the consequences of RNF31 depletion or overexpression on the ubiquitination status of ERα in HEK-293 cells and in MCF-7 cells. Western blot analysis of ERα in lysates from HEK-293 cells overexpressing RNF31 and ERα revealed the presence of potentially mono-ubiquitinated ERα species (Figure 6a). Importantly, depletion of endogenous RNF31 lead to reduced levels of mono-ubiquitinated ERα in MCF-7 cells (Figure 6b). The higher-molecular weight ERα species were reduced upon deletion of the RBR domain, whereas no change was seen upon deletion of the ubiquitin-associated domain (Figure 6c). More direct evidence that ERα was specifically subject to modification by ubiquitination was obtained by immunoprecipitation-based ubiquitination assays. Ubiquitinated species were enriched using a ubiquitin antibody from HEK-293 cell extracts overexpressing ERα together with RNF31 variants and analyzed by western blots for the presence of ERα in the precipitates (Figure 6d). The results confirmed that RNF31 overexpression increased the amounts of mono-ubiquitinated ERα. Consistent with the important role of the RBR domain of RNF31 for its association with, and stabilizing effect on, ERα, the presence of mono-ubiquitinated ERα was dependent on the RBR domain of RNF31 (Figure 6e). To further support the functional link between mono-ubiquitination, stability and signaling by ERα, we used the ubiquitin ligase-deficient RNF31 mutant R1/2M, in which cysteine residues required for the transfer of ubiquitin to substrates have been mutated.27 We overexpressed the RNF31 R1/2M in MCF-7 cells to determine the effect on endogenous ERα and ERE-luciferase activity. Figures 7a and b show that R1/2M failed to increase endogenous ERα protein levels and ERE-luciferase activity. If co-transfected with ERα into HEK-293 cells, RNF31 R1/2M failed to induce mono-ubiquitination of ERα (Figures 7c and d). Altogether, these data point at functional links between the RNF31 ubiquitin ligase domain, ERα mono-ubiquitination and ERα stabilization.
RNF31 triggers ERα mono-ubiquitination. (a) Detection of a potentially mono-ubiquitinated form of ERα upon RNF31 overexpression. HEK-293 cells were transfected with ERα together with plasmids expressing Myc-tagged RNF31 or the Myc-tag alone. Forty-eight hours after transfection, whole-cell extracts were prepared and levels of ERα protein assayed by western blot analysis. The predicted molecular weights of RNF31, ERα, mono-ubiquitinated ERα and of the internal control GAPDH are indicated. (b) Detection of endogenous mono-ubiquitinated ERα upon RNF31 depletion. MCF-7 cells were transfected with siRNF31 or siControl. Forty-eight hours after transfection, whole-cell extracts were prepared and levels of ERα protein assayed by western blot analysis. The predicted molecular weights of RNF31, ERα, mono-ubiquitinated ERα and the internal control GAPDH are indicated. (c) Deletion of the RNF31 RBR domain abolishes the potentially mono-ubiquitinated form of ERα. HEK-293 cells were transfected with ERα together with plasmids expressing Myc-tagged full-length RNF31 derivatives or the Myc-tag alone. Forty-eight hours post transfection, cell extracts were prepared and ERα forms were detected by western blot analysis. The predicted molecular weights of RNF31, ERα, mono-ubiquitinated ERα and of the internal control GAPDH are indicated. (d) Direct evidence for ERα mono-ubiquitination. Immunoprecipitation of ubiquitinated proteins from MCF-7 cell extracts upon overexpression of RNF31. Ubiquitinated ERα species were detected by western blots using anti-ERα, identifying a prominent 75 kDa mono-ubiquitinated ER form. (e) ERα mono-ubiquitination requires the RNF31 RBR domain. Plasmids expressing Myc-tagged RNF31 derivatives were transfected into HEK-293 cells together with the ERα expression plasmid. Whole-cell extracts were subjected to immunoprecipitation of ERα and subsequently analyzed for ubiquitinated ERα forms by western blot analysis using anti-ubiquitin. The predicted molecular weight of mono-ubiquitinated ERα is indicated.
The ubiquitin ligase activity of RNF31 is required for ERα stabilization, signaling and mono-ubiquitination. (a) The mono-ubiquitination function of RNF31 is required for the RNF31-mediated increase of endogenous ERα protein levels. MCF-7 cells were transfected with plasmids expressing Myc-tagged RNF31, Myc-tagged RNF31 R1/2M, in which cysteine residues responsible for the transfer of ubiquitin to substrates have been mutated or the Myc-tag alone, as indicated. Forty-eight hours after transfection, whole-cell extracts were prepared and levels of ERα protein assayed by western blot analysis. The predicted molecular weights of RNF31 variants, ERα and the loading control GAPDH are indicated. (b) The mono-ubiquitination function of RNF31 is required to increase ERα signaling. MCF-7 cells were transfected with plasmids expressing Myc-tagged RNF31, Myc-tagged RNF31 R1/2M or the Myc-tag alone, as indicated, along with an ERE-luciferase reporter plasmid. Luciferase activity was measured 48 h after transfection and calculated from experiments performed in triplicates. Data are shown as mean±s.d. (n=3). ***P<0.001 for wt-RNF31 versus RNF31 R1/2M or Myc-tag. RNF31 deletion domains/empty vector; NA, P>0.05 for wt-RNF31 versus RNF31 deletion domains. (c, d) Mutation of the RNF31 ubiquitin ligase domain abolishes mono-ubiquitination of ERα. (c) HEK-293 cells were transfected with ERα together with plasmids expressing Myc-tagged RNF31, RNF31 R1/2M or the Myc-tag alone. Forty-eight hours post transfection, cell extracts were prepared and ERα was detected by western blot analysis. The predicted molecular weights of RNF31, ERα, mono-ubiquitinated ERα and the internal control GAPDH are indicated. (d) HEK-293 cells were transfected with ERα together with plasmids expressing Myc-tagged full-length RNF31, RNF31 R1/2M or the Myc-tag alone. After 48 h, whole-cell extracts were subjected to immunoprecipitation of ERα and subsequently analyzed for ubiquitinated ERα forms by western blot analysis using anti-ubiquitin antibodies. The predicted molecular weights of RNF31, IgG, ERα and mono-ubiquitinated ERα are indicated.
RNF31 and ERα co-occur mainly in the cytosol
RNF31 is known to be involved in diverse ubiquitination pathways both in the cytoplasm and in the nucleus, but its intracellular localization and action in relation to ER signaling in breast cancer cells remained unclear. Immunocytochemistry revealed that endogenous RNF31 is localized to the nuclear and cytosol compartments of MCF-7 cells (Figure 8a, for quantification see Supplementary Figure S5). Furthermore, endogenous ERα and RNF31 were found to be prevalent in the cytosol both in the presence and absence of E2 in MCF-7 cells (Figure 8a), and a similar protein distribution was seen with ectopically expressed proteins in HEK-293 cells (Supplementary Figure S6). In MCF-7 cells, treatment with E2 redistributed a significant fraction of ERα to the nucleus, whereas the RNF31 distribution remained unaffected. Consistent with the immunocytochemistry results, co-IP assays revealed that the association of RNF31 and ERα occurs mainly in the cytosolic fraction (Figure 8b). Furthermore, chromatin immunoprecipitation assays failed to detect the recruitment of RNF31 to ER-responsive genes in MCF-7 cells (Supplementary Figure S7).
Intracellular localization analysis and model of crosstalk between RNF31 and ERα signaling in breast cancer cells. (a) MCF-7 cells were treated with 10 nM E2 or vehicle for 30 min before fixation. Intracellular localization of RNF31 (red) and ERα (green) was determined by immunofluorescence staining. Nuclei (blue) were stained with 4',6-diamidino-2-phenylindole (DAPI). Shown are representative images. For quantitative analysis see Supplementary Figure S5. (b) Co-IP assay reveals the interaction between RNF31 and ERα in the cytoplasm. The Subcellular Protein Fractionation Kit (Thermo Scientific, 78840) was used for extraction of cytoplasmic and nuclear proteins from MCF-7 cells. Vinculin and Histone 3 were used to identify the quality of cytoplasmic and nuclear fractions, respectively (left panel). Co-IP assays were performed with RNF31 antibody for precipitation and ERα antibody for detection (right panel). (c) Hypothetical model for the functional interplay of RNF31 with ERα signaling in breast cancer cells. RNF31 associates with ERα predominantly in the cytoplasm and promotes mono-ubiquitination of ERα, thereby counteracting proteasome-mediated degradation. The so elevated ERα protein levels cause increased genomic ERα signaling, for example, transcription of E2-target genes linked to the proliferation of breast cancer cells. RNF31, as part of the LUBAC ubiquitin ligase complex, has an established independent function in NFκB signaling. To which extent the two signaling pathways also crosstalk remains to be investigated.
Discussion
Here we report that the atyical ubiquitin ligase RNF31 associates with, and stabilizes, ERα in breast cancer cells, leading to increased estrogen signaling and cellular proliferation. Importantly, we also observe correlations between RNF31 and ERα target gene expression in breast cancer samples. Our study suggests a novel non-genomic mechanism of ER stability control (see model Figure 8c). On the basis of these data we propose that selective modulation of RNF31 expression and/or functional cooperation with ERα could be a strategy to inhibit proliferation in ERα-positive breast cancers.
RNF31—a mediator of crosstalk between proliferative signaling pathways in breast cancer cells
Our analysis of MCF-7 cell proliferation, in conjunction with siRNA-mediated depletion of RNF31 or ERα, revealed a requirement of RNF31 for breast cancer cell growth and specifically for E2-stimulated cell cycle progression. This suggests that ERα is one of the key mediators of the proliferative effects of RNF31 in ERα-positive breast cancer cells. Indeed, unbiased pathway analysis of the RNF31-dependent transcriptome, as determined by microarray expression analysis in MCF-7 cells, confirmed the functional link of RNF31 to the estrogen-signaling pathway. Importantly, a functional link between RNF31 and the estrogen-signaling pathway was also observed in breast cancer samples. In addition, the analysis revealed links to the NFκB pathway, and to the p53 pathway, known for its key role in cell cycle control. In addition, a previous study reported that RNF31 participates in transforming growth factor β and Wnt/β-catenin signaling pathways,28 both of which play fundamental roles in breast cancer. These pathways may contribute to the ERα/E2-independent effects of RNF31 depletion, consistent with the observation that RNF31 is also expressed in ERα-negative cell lines (data not shown).
Irrespective of the precise contribution of these additional RNF31-regulated pathways, it is likely that they crosstalk with the estrogen signaling pathway in ERα-positive cancers, instead of acting independently. One intriguing candidate for such cross talk is the NFκB signaling pathway, implicated in both inflammation and oncogenesis in cancer cells.29, 30 A number of recent studies have demonstrated that RNF31, together with the linear ubiquitin chain assembly complex (LUBAC) complex, is a critical component of cytoplasmic NFκB activation cascades, in other cell types.21, 22, 23 However, a first direct evidence for both positive and negative crosstalk has been provided by studying tumor necrosis factor α-activation of the NFκB pathway, and its interference with ER signaling, in breast cancer cell lines.29 Future studies of the individual facets of this crosstalk with particular emphasis on the RNF31 network should help to re-evaluate newly developed anti-inflammatory ER ligands,29 along with attempts to target components of the RNF31/NF-kB pathway.
The place of RNF31 within the atypical RBR family of E3 ubiquitin ligases
Among the more than 700 putative E3 ubiquitin ligases in humans, members of the RING-in between rings (IBR)-RING (RBR) family have attracted recent attention due to their uncommon ubiquitination mechanism and their regulatory signaling functions.31 Unlike classic E3s, including RING and HECT ligases, RBR ligases seem to preferentially catalyze mono-ubiquitination or linear poly-ubiquitination of substrates, which does not lead to substrate degradation and turnover.32, 33 The very recent clarification of the unique RBR ubiquitination mechanism in part resolves the initial paradox that RBR ligases often stabilize their substrates. Additionally, mechanisms of RBR-mediated ubiquitination are often substrate-dependent.31, 32 RBR ligases appear therefore well suited to fulfill diverse regulatory roles in intracellular signaling and transcription. This has perhaps best been demonstrated in case of the Parkin protein (PARK2), implicated in Parkinson disease, and in the case of PARC (CUL9) which is implicated in p53 tumor suppression.34, 35, 36, 37
There is a growing interest in the linear ubiquitin chain assembly complex referred to as LUBAC, implicated in NFκB signaling, and thus potentially in multiple disorders, such as cancer and chronic inflammatory and metabolic diseases.21, 22 Most notably, RNF31 functions as the main E3 ligase of LUBAC, which contains additional subunits including RBCK1 and Sharpin. Both RNF31 and RBCK1 function independently from each other, and thus without the entire LUBAC complex, to modulate various nuclear receptor pathways. For example, we have previously demonstrated that RBCK1 inhibits cellular proliferation of ERα-positive breast cancer cells via several genomic/transcriptional mechanisms: RBCK1 regulates ERα gene transcription, and it acts as a transcriptional cofactor that co-occupies E2-responsive promoters.38 The combined data of our previous and current studies suggest that RNF31 and RBCK1 affect ERα signaling by very distinct mechanisms. Unlike RBCK1, RNF31 cannot be directly recruited to ERα target promoters in MCF-7 cells, consistent with the dominant cytoplasmic localization of RNF31 that we found.
RNF31 stabilizes ERα via a unique ubiquitination mechanism
In contrast to RBCK1, RNF31 appears specifically linked to the control of ERα protein stability and mono-ubiquitination. This mechanism of action is consistent with a previously reported and similar role of RNF31 in stabilizing the steroidogenic orphan receptor DAX-1.27 The obvious link between RNF31 and mono-ubiquitination explains why RNF31 action leads to substrate stabilization instead of degradation, although some mechanistic details remain unrevealed. On the basis of our experimental data, the following two mechanisms are likely to contribute: (1) RNF31 mono-ubiquitinates ERα, either alone or in complex with yet another E3 ligase. Importantly, RNF31 does not act in complex with RBCK1/LUBAC, as deletion of the RNF31 ubiquitin-associated domain, necessary for RBCK1 interaction, does not abolish mono-ubiquitination and stabilization of ERα (this study), or of DAX-1.27 However, the enzymatically intact RBR-type ubiquitin ligase domain is necessary for these events, and its activity might be further modified in a substrate (that is, ERα)-dependent manner. (2) RNF31 stabilizes ERα by preventing poly-ubiquitination, which may involve direct binding to mono-ubiquitinated ERα. Intriguingly, RNF31 has several putative Zn-finger type ubiquitin-binding domains, whose functionality and specificity, due to the lack of consensus signatures,27 remain to be determined.
RNF31—a candidate target for breast cancer diagnosis and therapy?
What evidence supports the clinical relevance of our findings about the role of RNF31 in estrogen signaling? Indeed, RNF31 was originally discovered in screens for genes that were differentially expressed in breast cancer cells compared with normal breast epithelial cells.20 Consistent with this, using a limited number of samples, we found that RNF31 is more highly expressed in breast cancer tissues as compared with the surrounding normal tissues. In addition, consistent with the expression of RNF31 mRNA in ERα-positive breast cancer cell lines, RNF31 mRNA appears highly expressed in ERα-positive breast tumor tissue. Importantly, the notion that RNF31 is expressed in both ERα-positive and negative cancers is not contradictory to our model but rather points at the convergence of distinct signaling pathways toward proliferation and tumorigenesis, as discussed above.
Given our data, it will be interesting but technically challenging to determine whether high RNF31 protein levels in tumor samples also cause a stabilization of ERα protein levels. Optimally, cancer samples containing equal levels of ERα mRNA but different levels of RNF31 (mRNA and protein) should be quantitated for ERα protein. An additional complication arises from the fact that functional associations between RNF31 and ERα occur mainly in the cytoplasm, which will be more complicated to analyze by immunohistochemistry than the nuclear fraction, traditionally used to score for ERα positivity in clinical samples. These concerns apply to monoclonal antibody-based biochemical methods as well, as already noted in 1986.39 An interesting question is therefore whether some tumors that were initially scored as ER-negative could be enriched in cytoplasmic receptors, and whether this enrichment is dependent on cytoplasmic RNF31. Also relevant is the recent observation that clinically used markers including ERα, PR and HER2 protein levels are unstable throughout tumor progression and possibly influenced by adjuvant therapy.9 As alterations of both epigenetic/transcriptional (mRNA) and post-translational (protein) mechanisms are likely to contribute to the loss of ERα, post-translational ER regulators such as RNF31 have to be taken into consideration as well. In light of these concerns, the possible convergence of RNF31 and ER signaling in RNF31/ERα-positive breast cancers should be of outmost interest for the clinical diagnosis and treatment decisions and requires further clarification.
In conclusion, although ERα has already been well documented to have a critical role in the etiology and progression of breast cancer, RNF31 emerges as a new component of ER signaling in ERα-positive breast cancers. To identify novel therapeutic strategies for treating ERα-positive breast cancers, modulation of ERα levels is one feasible approach to inhibit estrogen signaling and subsequently cellular proliferation. As our study identifies RNF31 as a modulator of ERα protein levels in breast cancer cells, RNF31 could constitute a valuable diagnostic tool and/or a drug target for ERα-positive breast cancers.
Materials and methods
Cell culture
MCF-7 and HEK-293 cells were cultured in Dulbecco’s Modified Eagle Medium (DMEM) (Invitrogen, Carlsbad, CA, USA) supplemented with 10% fetal bovine serum and 1% penicillin/streptomycin (Invitrogen) at 37 °C in a humidified atmosphere of 5% CO2 in air. T47D and MDA-MB-175 cells were cultured in RPMI 1640 (Invitrogen) supplemented with 10% fetal bovine serum and 1% penicillin-streptomycin. 17β-estradiol (E2; Sigma-Aldrich, St Louis, MO, USA) was dissolved in ethanol. For proliferation assays, cells were cultured in fetal calf serum treated with dextran-coated charcoal and phenol red-free DMEM.
Plasmids
RNF31 (pcDNA-Myc-RNF31) constructs were described previously.27 The pcDNA3-ERα plasmid, the ERE-TK-Luc reporter and the pRL-TK control have been described previously.40
siRNA transfection
Cells were transfected with 50–100 nM siRNA. RNF31 siRNAs correspond to the siGENOME SMARTpool RNF31 (Dharmacon D-0021419, Thermo Fisher Scientific, Waltham, MA, USA), consisting of four single siRNAs, which were used in previous studies.27 ERα siRNA and control siRNA were Stealth Select siRNA (Invitrogen). INTERFERin transfection reagent (Polyplus Transfection, New York, NY, USA) was used according to the manufacturer’s protocol.
Quantitative PCR
Quantitative PCR was performed as previously described.41 Primer sequences for qPCR are provided in Supplementary Table S1.
Western blotting
Cells were lysed with NP-40 lysis buffer (Roche Diagnostics GmbH, Mannheim, Germany). Antibodies used were anti-GAPDH (ab8425), anti-RNF31 (ab46322) from Abcam, and anti-Myc (9E10), anti-ubiquitin (FL76), anti-HA antibody (F-7), anti-ERα (1D5 and HC-20) from Santa Cruz Biotechnology, Inc., Dallas, TX, USA.
Luciferase reporter assays
Luciferase activity was measured using the Dual-Luciferase Reporter Assay (Promega, Mannheim, Germany).
Quantification of cell viability
The cell number was determined using the WST-1 cell proliferation reagent (Roche) as previously described.38
Fluorescent-activated cell sorting
MCF-7 cells were transfected with siRNF31, siERα or siControl. After 24 h, the medium was changed into phenol red-free DMEM with 2% dextran-coated charcoal-treated fetal calf serum. After an additional 24 h, the cells were fixed in 70% ethanol for 30 min and stained by propidium iodide. The BD LSR II flow cytometer (BD Bioscience, Franklin Lakes, NJ, USA) was used to measure the flow fluorescence intensity. For the rescue experiment, MCF-7 cells were transfected with siRNF31 or siControl. After 24 h, the cells were transfected with 2 μg Flag-ERα vector or empty Flag vector and cultured in DMEM medium with 10% fetal calf serum.
Chromatin immunoprecipitation assays
Chromatin immunoprecipitation assays were performed essentially as previously described.41 For detail see Supplementary Data.
Co-immunoprecipitation
Co-IP was performed essentially as previously described.41 The Subcellular Protein Fractionation Kit (Thermo Scientific, Waltham, MA, USA, 78840) was used for extraction of cytoplasmic and nuclear proteins.
Protein stability assay
MCF-7 cells were transfected with 100 nM siRNF31 or siControl. 24 h post-transfection, cells were treated with E2 or vehicle for 24 h. The cells were then treated with cycloheximide (100 μM) or MG132 (100 μM) for 48 h. Samples were harvested and analyzed by western blotting.
Ubiquitination assays
(A) To detect modified ERα from cell extracts, HEK-293 cells were transfected with 4 μg pcDNA3-ERα together with pCDNA3-Myc-RNF31 including deletion variants. Forty-eight hours post-transfection, cells were treated with 10 uM MG132/vehicle for 8 h, directly lysed and modified/unmodified ERα was detected by western blotting. (B) To directly enrich ubiquitinated ERα from cell extracts, HEK-293 cells were transfected with 4 μg pcDNA3-ERα together with 4 ug pCDNA3-Myc-RNF31 or deletion variants. Forty-eight hours post-transfection, total protein was extracted and pre-cleared by 20 μl protein A slurry and 1 μg rabbit immunoglobulin G for 2 h at 4 °C. The supernatant was collected and immunoprecipitated with 20 μl ubiquitin beads (CalBiochem, no. 662200). Western blot with rabbit anti-ERα antibody was performed to detect ubiquitinated ERα.
Immunofluorescence assay
A rabbit anti-RNF31 polyclonal antibody and mouse anti-ERα monoclonal antibodies were used. For detail see Supplementary Data.
Microarray analysis
The Agilent SurePrint 8 × 60K arrays were used for global gene expression analysis at the core facility for bioinformatics and expression analysis (BEA, www.bea.ki.se), Karolinska Institutet, Sweden, according to standard protocols. Data are deposited in the Gene Expression Omnibus (GEO) database (accession number GSE46010). We applied a filter of P<0.001 for significantly modulated gene expression and at least a 2.0-fold change in mean differential expression.
Analysis of human breast tumor samples
Human samples were from the Iranian Tumor Bank, Cancer Institute of Iran. For detail see Supplementary Data.
Analysis of gene expression in publicly available data sets
Pathway analysis and identification of regulated ERα target genes from microarray data were carried out using Pathway Studio (Elsevier). Analyses of gene expression in breast cancer samples from TCGA25 and KMplot.com26 were carried out in the statistical environment R. Presented correlations are Pearson’s product-moment correlations. Samples were separated into high/low expression of RNF31 by setting a cutoff at the median value.
Statistics
Student’s t-test, Pearson correlation coefficient and Cox regression analysis were used for comparisons. A P-value of <0.05 was considered to be significant.
Accession codes
References
Geschickter CF, Lewis D, Hartman CG . Tumors of the breast related to the oestrin hormone. Am J Cancer 1934; 21: 828–859.
Brown RJ, Davidson NE . Adjuvant hormonal therapy for premenopausal women with breast cancer. Sem Oncol 2006; 33: 657–663.
Hicks C, Kumar R, Pannuti A, Miele L . Integrative analysis of response to tamoxifen treatment in ER-positive breast cancer using GWAS information and transcription profiling. Breast Cancer 2012; 6: 47–66.
Musgrove EA, Sutherland RL . Biological determinants of endocrine resistance in breast cancer. Nat Rev Cancer 2009; 9: 631–643.
Thomas C, Gustafsson JA . The different roles of ER subtypes in cancer biology and therapy. Nat RevCancer 2011; 11: 597–608.
Wong SC, Chan JK, Lee KC, Hsiao WL . Differential expression of p16/p21/p27 and cyclin D1/D3, and their relationships to cell proliferation, apoptosis, and tumor progression in invasive ductal carcinoma of the breast. J Pathol 2001; 194: 35–42.
Cariou S, Donovan JC, Flanagan WM, Milic A, Bhattacharya N, Slingerland JM . Down-regulation of p21WAF1/CIP1 or p27Kip1 abrogates antiestrogen-mediated cell cycle arrest in human breast cancer cells. Proc Natl Acad Sci USA 2000; 97: 9042–9046.
Shaaban AM, Sloane JP, West CR, Foster CS . Breast cancer risk in usual ductal hyperplasia is defined by estrogen receptor-alpha and Ki-67 expression. Am J Pathol 2002; 160: 597–604.
Lindstrom LS, Karlsson E, Wilking UM, Johansson U, Hartman J, Lidbrink EK et al. Clinically used breast cancer markers such as estrogen receptor, progesterone receptor, and human epidermal growth factor receptor 2 are unstable throughout tumor progression. J Clin Oncol 2012; 30: 2601–2608.
Duong V, Boulle N, Daujat S, Chauvet J, Bonnet S, Neel H et al. Differential regulation of estrogen receptor alpha turnover and transactivation by Mdm2 and stress-inducing agents. Cancer Res 2007; 67: 5513–5521.
Ma Y, Fan S, Hu C, Meng Q, Fuqua SA, Pestell RG et al. BRCA1 regulates acetylation and ubiquitination of estrogen receptor-alpha. Mol Endocrinol 2010; 24: 76–90.
Reid G, Hubner MR, Metivier R, Brand H, Denger S, Manu D et al. Cyclic, proteasome-mediated turnover of unliganded and liganded ERalpha on responsive promoters is an integral feature of estrogen signaling. Mol Cell 2003; 11: 695–707.
Fan M, Park A, Nephew KP . CHIP (carboxyl terminus of Hsc70-interacting protein) promotes basal and geldanamycin-induced degradation of estrogen receptor-alpha. Mol Endocrinol 2005; 19: 2901–2914.
Le Romancer M, Poulard C, Cohen P, Sentis S, Renoir JM, Corbo L . Cracking the estrogen receptor's posttranslational code in breast tumors. Endocr Rev 2011; 32: 597–622.
Brekman A, Singh KE, Polotskaia A, Kundu N, Bargonetti J . A p53-independent role of Mdm2 in estrogen-mediated activation of breast cancer cell proliferation. Breast Cancer Res 2011; 13: R3.
Zwart W, Theodorou V, Carroll JS . Estrogen receptor-positive breast cancer: a multidisciplinary challenge. Wiley Interdiscip Rev Syst Biol Med 2011; 3: 216–230.
Berry NB, Fan M, Nephew KP . Estrogen receptor-alpha hinge-region lysines 302 and 303 regulate receptor degradation by the proteasome. Mol Endocrinol 2008; 22: 1535–1551.
Lonard DM, O'Malley BW . SRC-3 transcription-coupled activation, degradation, and the ubiquitin clock: is there enough coactivator to go around in cells? Sci Signaling 2008; 1: pe16.
La Rosa P, Pesiri V, Marino M, Acconcia F . 17beta-Estradiol-induced cell proliferation requires estrogen receptor (ER) alpha monoubiquitination. Cell Signalling 2011; 23: 1128–1135.
Thompson HG, Harris JW, Lin L, Brody JP . Identification of the protein Zibra, its genomic organization, regulation, and expression in breast cancer cells. Exp Cell Res 2004; 295: 448–459.
Ikeda F, Deribe YL, Skanland SS, Stieglitz B, Grabbe C, Franz-Wachtel M et al. SHARPIN forms a linear ubiquitin ligase complex regulating NF-kappaB activity and apoptosis. Nature 2011; 471: 637–641.
Tokunaga F, Nakagawa T, Nakahara M, Saeki Y, Taniguchi M, Sakata S et al. SHARPIN is a component of the NF-kappaB-activating linear ubiquitin chain assembly complex. Nature 2011; 471: 633–636.
Tokunaga F, Sakata S, Saeki Y, Satomi Y, Kirisako T, Kamei K et al. Involvement of linear polyubiquitylation of NEMO in NF-kappaB activation. Nat Cell Biolo 2009; 11: 123–132.
Stevens TA, Meech R . BARX2 and estrogen receptor-alpha (ESR1) coordinately regulate the production of alternatively spliced ESR1 isoforms and control breast cancer cell growth and invasion. Oncogene 2006; 25: 5426–5435.
TCGA, Comprehensive molecular portraits of human breast tumours. Nature 2012; 490: 61–70.
Gyorffy B, Lanczky A, Eklund AC, Denkert C, Budczies J, Li Q et al. An online survival analysis tool to rapidly assess the effect of 22 277 genes on breast cancer prognosis using microarray data of 1809 patients. Breast Cancer Res Treat 2010; 123: 725–731.
Ehrlund A, Anthonisen EH, Gustafsson N, Venteclef N, Robertson Remen K, Damdimopoulos AE et al. E3 ubiquitin ligase RNF31 cooperates with DAX-1 in transcriptional repression of steroidogenesis. Mol Cell Biol 2009; 29: 2230–2242.
Ehrlund A, Jonsson P, Vedin LL, Williams C, Gustafsson JA, Treuter E . Knockdown of SF-1 and RNF31 affects components of steroidogenesis, TGFbeta, and Wnt/beta-catenin signaling in adrenocortical carcinoma cells. PLoS One 2012; 7: e32080.
Frasor J, Weaver A, Pradhan M, Dai Y, Miller LD, Lin CY et al. Positive cross-talk between estrogen receptor and NF-kappaB in breast cancer. Cancer Res 2009; 69: 8918–8925.
Nettles KW, Bruning JB, Gil G, Nowak J, Sharma SK, Hahm JB et al. NFkappaB selectivity of estrogen receptor ligands revealed by comparative crystallographic analyses. Nat Chem Biol 2008; 4: 241–247.
Eisenhaber B, Chumak N, Eisenhaber F, Hauser MT . The ring between ring fingers (RBR) protein family. Genome Biol 2007; 8: 209.
Smit JJ, Monteferrario D, Noordermeer SM, van Dijk WJ, van der Reijden BA, Sixma TK . The E3 ligase HOIP specifies linear ubiquitin chain assembly through its RING-IBR-RING domain and the unique LDD extension. EMBO J 2012; 31: 3833–3844.
Kirisako T, Kamei K, Murata S, Kato M, Fukumoto H, Kanie M et al. A ubiquitin ligase complex assembles linear polyubiquitin chains. EMBO J 2006; 25: 4877–4887.
Woo MG, Xue K, Liu J, McBride H, Tsang BK . Calpain-mediated processing of p53-associated parkin-like cytoplasmic protein (PARC) affects chemosensitivity of human ovarian cancer cells by promoting p53 subcellular trafficking. J Biol Chem 2012; 287: 3963–3975.
Mulhall JP, Barnas J, Kobylarz K, Mueller A . p53-Associated Parkin-like cytoplasmic protein (Parc) short-interfering RNA (siRNA) alters p53 location and biology of Peyronie's disease fibroblasts. BJU Int 2010; 106: 1706–1713.
da Costa CA, Sunyach C, Giaime E, West A, Corti O, Brice A et al. Transcriptional repression of p53 by parkin and impairment by mutations associated with autosomal recessive juvenile Parkinson's disease. Nature Cell Biol 2009; 11: 1370–1375.
Kastan MB, Zambetti GP . Parc-ing p53 in the cytoplasm. Cell 2003; 112: 1–2.
Gustafsson N, Zhao C, Gustafsson JA, Dahlman-Wright K . RBCK1 drives breast cancer cell proliferation by promoting transcription of estrogen receptor alpha and cyclin B1. Cancer Res 2010; 70: 1265–1274.
Leclercq G, Bojar H, Goussard J, Nicholson RI, Pichon MF, Piffanelli A et al. Abbott monoclonal enzyme immunoassay measurement of estrogen receptors in human breast cancer: a European multicenter study. Cancer Res 1986; 46: 4233s–4236s.
Putnik M, Zhao C, Gustafsson JA, Dahlman-Wright K . Global identification of genes regulated by estrogen signaling and demethylation in MCF-7 breast cancer cells. Biochem Biophys Res Commun 2012; 426: 26–32.
Zhao C, Matthews J, Tujague M, Wan J, Strom A, Toresson G et al. Estrogen receptor beta2 negatively regulates the transactivation of estrogen receptor alpha in human breast cancer cells. Cancer Res 2007; 67: 3955–3962.
Acknowledgements
We are grateful to the BEA core facility for performing microarray analysis. We thank Indranil Sinha for uploading the microarray data. This work was supported by scholarship from the China Scholarship Council and by a KID faculty grant from the Karolinska Institutet (Jian Zhu), funding from the Center for Biosciences, the Swedish Cancer Society and the Karolinska Institutet (Karin Dahlman-Wright), and grants from the Center for Biosciences, the Swedish Cancer Society and the Swedish Research Council (Eckardt Treuter). Human samples were provided by the Iran National Tumor Bank, which is funded by the Cancer Institute of Tehran University. Imaging, performed at the live cell imaging unit at the Department of Biosciences and Nutrition, was supported by the Knut & Alice Wallenberg foundation, the Swedish Research Council and the Center for Biosciences. This work was supported by scholarship from the China Scholarship Council and by a KID faculty grant from the Karolinska Institutet, funding from the Center for Biosciences, the Swedish Cancer Society, the Swedish Research Council and the Karolinska Institutet. Human samples were provided by the IRAN NATIONAL TUMOR BANK, which is funded by the Cancer Institute of Tehran University. Imaging was performed at the live cell imaging unit at the Department of Biosciences and Nutrition, supported by the Knut & Alice Wallenberg foundation, the Swedish Research Council and the Center for Biosciences.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies this paper on the Oncogene website
Supplementary information
Rights and permissions
This work is licensed under a Creative Commons Attribution-NonCommercial-NoDerivs 3.0 Unported License. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-nd/3.0/
About this article
Cite this article
Zhu, J., Zhao, C., Kharman-Biz, A. et al. The atypical ubiquitin ligase RNF31 stabilizes estrogen receptor α and modulates estrogen-stimulated breast cancer cell proliferation. Oncogene 33, 4340–4351 (2014). https://doi.org/10.1038/onc.2013.573
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/onc.2013.573
Keywords
This article is cited by
-
RNF31 promotes proliferation and invasion of hepatocellular carcinoma via nuclear factor kappaB activation
Scientific Reports (2024)
-
High expression of RNF31 is associated with tumor immune cell infiltration and leads to poor prognosis in liver hepatocellular carcinoma
Scientific Reports (2023)
-
Resonance assignments of the PUB domain of the RNF31 protein
Biomolecular NMR Assignments (2023)
-
RBCK1 is an endogenous inhibitor for triple negative breast cancer via hippo/YAP axis
Cell Communication and Signaling (2022)
-
RNF31 represses cell progression and immune evasion via YAP/PD-L1 suppression in triple negative breast Cancer
Journal of Experimental & Clinical Cancer Research (2022)