Abstract
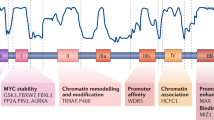
The MYC oncogene is not only deregulated in cancer through abnormally high levels of expression, but also through oncogenic lesions in upstream signalling cascades. Modelling MYC deregulation using signalling mutants is a productive research strategy. For example, the MYC threonine-58 to alanine substitution mutant (T58A) within MYC-homology box 1 is more transforming than wild-type (WT) MYC, because of decreased apoptosis and increased protein stability. Understanding the regulatory mechanisms controlling T58 phosphorylation has led to new approaches for the development of MYC inhibitors. In this manuscript, we have extensively characterized a MYC signalling mutant in which six lysine residues near the highly conserved MYC homology box IV and basic region have been substituted to arginines (6KR). Previous literature suggests these lysines can undergo both ubiquitylation and acetylation. We show MYC 6KR is able to fully rescue the slow growth phenotype of HO15.19 MYC-null fibroblasts, and promote cell cycle entry of serum-starved MCF10A cells. Remarkably, 6KR increased anchorage-independent colony growth compared with WT MYC in both SH-EP and MCF10A cells. Moreover, it was also more potent in promoting xenograft tumour growth of Rat1A and SH-EP cells. Combined, our data identify this region and these six lysines as important residues for the negative regulation of MYC-induced transformation. Mechanistically, we demonstrate that, unlike T58A, the increased transformation is not a result of increased protein stability or a reduced capacity for 6KR to induce apoptosis. Through expression analysis and luciferase reporter assays, we show that 6KR has increased transcriptional activity compared with WT MYC. Combined, through a comprehensive evaluation across multiple cell types, we identify an important regulatory region within MYC. A better understanding of the full scope of signalling through these residues will provide further insights into the mechanisms contributing to MYC-induced tumorigenesis and may unveil novel therapeutic strategies to target Myc in cancer.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 50 print issues and online access
$259.00 per year
only $5.18 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Hann SR . Role of post-translational modifications in regulating c-Myc proteolysis, transcriptional activity and biological function. Semin Cancer Biol 2006; 16: 288–302.
Sears RC . The life cycle of C-myc: from synthesis to degradation. Cell Cycle 2004; 3: 1133–1137.
Popov N, Wanzel M, Madiredjo M, Zhang D, Beijersbergen R, Bernards R et al. The ubiquitin-specific protease USP28 is required for MYC stability. Nat Cell Biol 2007; 9: 765–774.
Bhatia K, Huppi K, Spangler G, Siwarski D, Iyer R, Magrath I . Point mutations in the c-Myc transactivation domain are common in Burkitt's lymphoma and mouse plasmacytomas. Nat Genet 1993; 5: 56–61.
Rabbitts TH, Hamlyn PH, Baer R . Altered nucleotide sequences of a translocated c-myc gene in Burkitt lymphoma. Nature 1984; 306: 760–765.
Chang DW, Claassen GF, Hann SR, Cole MD . The c-Myc transactivation domain is a direct modulator of apoptotic versus proliferative signals. Mol Cell Biol 2000; 20: 4309–4319.
Thibodeaux CA, Liu X, Disbrow GL, Zhang Y, Rone JD, Haddad BR et al. Immortalization and transformation of human mammary epithelial cells by a tumor-derived Myc mutant. Breast Cancer Res Treat 2009; 116: 281–294.
Wang X, Cunningham M, Zhang X, Tokarz S, Laraway B, Troxell M et al. Phosphorylation regulates c-Myc's oncogenic activity in the mammary gland. Cancer Res 2011; 71: 925–936.
Conzen SD, Gottlob K, Kandel ES, Khanduri P, Wagner AJ, O’Leary M et al. Induction of cell cycle progression and acceleration of apoptosis are two separable functions of c-Myc: transrepression correlates with acceleration of apoptosis. Mol Cell Biol 2000; 20: 6008–6018.
Hemann MT, Bric A, Teruya-Feldstein J, Herbst A, Nilsson JA, Cordon-Cardo C et al. Evasion of the p53 tumour surveillance network by tumour-derived MYC mutants. Nature 2005; 436: 807–811.
Adhikary S, Marinoni F, Hock A, Hulleman E, Popov N, Beier R et al. The ubiquitin ligase HectH9 regulates transcriptional activation by Myc and is essential for tumor cell proliferation. Cell 2005; 123: 409–421.
Kim W, Bennett EJ, Huttlin EL, Guo A, Li J, Possemato A et al. Systematic and quantitative assessment of the ubiquitin-modified proteome. Mol Cell 2011; 44: 325–340.
Patel JH, Du Y, Ard PG, Phillips C, Carella B, Chen CJ et al. The c-MYC oncoprotein is a substrate of the acetyltransferases hGCN5/PCAF and TIP60. Mol Cell Biol 2004; 24: 10826–10834.
Zhang K, Faiola F, Martinez E . Six lysine residues on c-Myc are direct substrates for acetylation by p300. Biochem Biophys Res Commun 2005; 336: 274–280.
Wasylishen AR, Stojanova A, Oliveri S, Rust AC, Schimmer AD, Penn LZ . New model systems provide insights into Myc-induced transformation. Oncogene 2011; 30: 3727–3734.
Mateyak MK, Obaya AJ, Adachi S, Sedivy JM . Phenotypes of c-Myc-deficient rat fibroblasts isolated by targeted homologous recombination. Cell Growth Differ 1997; 8: 1039–1048.
Faiola F, Liu X, Lo S, Pan S, Zhang K, Lymar E et al. Dual regulation of c-Myc by p300 via acetylation-dependent control of Myc protein turnover and coactivation of Myc-induced transcription. Mol Cell Biol 2005; 25: 10220–10234.
Berns K, Hijmans EM, Koh E, Daley GQ, Bernards R . A genetic screen to identify genes that rescue the slow growth phenotype of c-myc null fibroblasts. Oncogene 2000; 19: 3330–3334.
Lin CY, Loven J, Rahl PB, Paranal RM, Burge CB, Bradner JE et al. Transcriptional amplification in tumor cells with elevated c-Myc. Cell 2012; 151: 56–67.
Nie Z, Hu G, Wei G, Cui K, Yamane A, Resch W et al. c-Myc is a universal amplifier of expressed genes in lymphocytes and embryonic stem cells. Cell 2012; 151: 68–79.
Callus BA, Ekert PG, Heraud JE, Jabbour AM, Kotevski A, Vince JE et al. Cytoplasmic p53 is not required for PUMA-induced apoptosis. Cell Death Differ 2008; 15: 213–215 author reply 5-6.
Stone J, de Lange T, Ramsay G, Jakobovits E, Bishop JM, Varmus H et al. Definition of regions in human c-myc that are involved in transformation and nuclear localization. Mol Cell Biol 1987; 7: 1697–1709.
Graves JA, Rothermund K, Wang T, Qian W, Van Houten B, Prochownik EV . Point mutations in c-Myc uncouple neoplastic transformation from multiple other phenotypes in rat fibroblasts. PLoS One 5: e13717.
Ngo CV, Gee M, Akhtar N, Yu D, Volpert O, Auerbach R et al. An in vivo function for the transforming Myc protein: elicitation of the angiogenic phenotype. Cell Growth Differ 2000; 11: 201–210.
Gartel AL, Ye X, Goufman E, Shianov P, Hay N, Najmabadi F et al. Myc represses the p21(WAF1/CIP1) promoter and interacts with Sp1/Sp3. Proc Natl Acad Sci USA 2001; 98: 4510–4515.
Marhin WW, Chen S, Facchini LM, Fornace AJ, Penn LZ . Myc represses the growth arrest gene gadd45. Oncogene 1997; 14: 2825–2834.
Oster SK, Mao DY, Kennedy J, Penn LZ . Functional analysis of the N-terminal domain of the Myc oncoprotein. Oncogene 2003; 22: 1998–2010.
Debnath J, Muthuswamy SK, Brugge JS . Morphogenesis and oncogenesis of MCF-10A mammary epithelial acini grown in three-dimensional basement membrane cultures. Methods 2003; 30: 256–268.
Acknowledgements
We thank Dr Martin Eilers for kindly providing the Myc 6KR construct, Dr Garry Nolan for providing retroviral reagents and packaging cell lines. SH-EP Tet21/N-Myc, MCF10A, Rat-1A and HO15.19 cells were kindly provided by Dr Manfred Schwab, Dr Senthil Muthuswamy, Dr Edward Prochownick and Dr John Sedivy, respectively. Additionally we thank Drs Bruno Amati, Chi Dang and Stephen Hann for kindly providing luciferase reporter constructs. We also thank Andrew Rust and Drs Angelina Stojanova and Peter Mullen for technical assistance, and the members of the Penn Lab for helpful discussions and critical review of this manuscript. This research was funded by grants from the Canadian Cancer Society Research Institute and Canadian Institute for Health Research (LZP), Canadian Breast Cancer Foundation Ontario Region Doctoral Fellowships (ARW and AP) and an Ontario Graduate Student Fellowship (MK). BR and LZP hold the Canada Research Chairs in Proteomics & Molecular Medicine and Molecular Oncology, respectively. Additional support was provided by the Ontario Ministry of Health and Long Term Care. The views expressed do not necessarily reflect those of the OMOHLTC.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies the paper on the Oncogene website
Supplementary information
Rights and permissions
About this article
Cite this article
Wasylishen, A., Kalkat, M., Kim, S. et al. MYC activity is negatively regulated by a C-terminal lysine cluster. Oncogene 33, 1066–1072 (2014). https://doi.org/10.1038/onc.2013.36
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/onc.2013.36
Keywords
This article is cited by
-
Multiple direct interactions of TBP with the MYC oncoprotein
Nature Structural & Molecular Biology (2019)
-
Mission Possible: Advances in MYC Therapeutic Targeting in Cancer
BioDrugs (2019)
-
Lysine-52 stabilizes the MYC oncoprotein through an SCFFbxw7-independent mechanism
Oncogene (2017)
-
Inhibiting MYC binding to the E-box DNA motif by ME47 decreases tumour xenograft growth
Oncogene (2017)
-
Selective AKR1C3 inhibitors do not recapitulate the anti-leukaemic activities of the pan-AKR1C inhibitor medroxyprogesterone acetate
British Journal of Cancer (2014)