Abstract
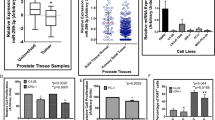
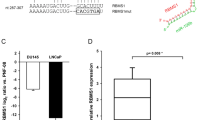
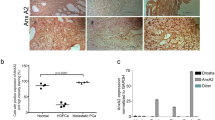
Hormone-sensitive prostate cancer typically progresses to castration resistant prostate cancer (CRPC) after the androgen deprivation therapy. We investigated the impact of microRNAs (miRs) in the transition of prostate cancer to CRPC. MiR-221/-222 was highly expressed in bone metastatic CRPC tumor specimens. We previously demonstrated that transient overexpression of miR-221/-222 in LNCaP promoted the development of the CRPC phenotype. In current study, we show that stably overexpressing miR-221 confers androgen independent (AI) cell growth in LNCaP by rescuing LNCaP cells from growth arrest at G1 phase due to the lack of androgen. Overexpressing of miR-221 in LNCaP reduced the transcription of a subgroup of androgen-responsive genes without affecting the androgen receptor (AR) or AR-androgen integrity. By performing systematic biochemical and bioinformatical analyses, we identified two miR-221 targets, HECTD2 and RAB1A, which could mediate the development of CRPC phenotype in multiple prostate cancer cell lines. Downregulation of HECTD2 significantly affected the androgen-induced and AR-mediated transcription, and downregulation of HECTD2 or RAB1A enhances AI cell growth. As a result of the elevated expression of miR-221, expression of many cell cycle genes was altered and pathways promoting epithelial to mesenchymal transition/tumor metastasis were activated. We hypothesize that a major biological consequence of upregulation of miR-221 is reprogramming of AR signaling, which in turn may mediate the transition to the CRPC phenotype.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 50 print issues and online access
$259.00 per year
only $5.18 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
References
American Cancer Society Cancer Facts and Figs 2012. American Cancer Society, Atlanta, GA, USA, 2012.
Heinlein CA, Chang C . Androgen receptor in prostate cancer. Endocr Rev 2004; 25: 276–308.
Rini BI, Small EJ . Hormone-refractory Prostate Cancer. Curr Treat Options Oncol 2002; 3: 437–446.
Yuan X, Balk SP . Mechanisms mediating androgen receptor reactivation after castration. Urol Oncol 2009; 27: 36–41.
Stanbrough M, Bubley GJ, Ross K, Golub TR, Rubin MA, Penning TM et al. Increased expression of genes converting adrenal androgens to testosterone in androgen-independent prostate cancer. Cancer Res 2006; 66: 2815–2825.
Guo Z, Yang X, Sun F, Jiang R, Linn DE, Chen H et al. A novel androgen receptor splice variant is up regulated during prostate cancer progression and promotes androgen depletion-resistant growth. Cancer Res 2009; 69: 2305–2313.
Sun T, Wang Q, Balk S, Brown M, Lee GS, Kantoff P . The role of microRNA-221 and microRNA-222 in androgen-independent prostate cancer cell lines. Cancer Res 2009; 69: 3356–3363.
Sun T, Yang M, Chen S, Balk S, Pomerantz M, Hsieh CL et al. The altered expression of MiR-221/-222 and MiR-23b/-27b is associated with the development of human castration resistant prostate cancer. Prostate 2012; 72: 1093–1103.
He L, Hannon GJ . MicroRNAs: small RNAs with a big role in gene regulation. Nat Rev Genet 2004; 5: 522–531.
Winter J, Jung S, Keller S, Gregory RI, Diederichs S . Many roads to maturity: microRNA biogenesis pathways and their regulation. Nat Cell Biol 2009; 11: 228–234.
Esquela-Kerscher A, Slack FJ . Oncomirs - microRNAs with a role in cancer. Nat Rev Cancer 2006; 6: 259–269.
Coppola V, De Maria R, Bonci D . MicroRNAs and prostate cancer. Endocr Relat Cancer 2010; 17: F1–17.
Watahiki A, Wang Y, Morris J, Dennis K, O'Dwyer HM, Gleave M et al. MicroRNAs associated with metastatic prostate cancer. PLoS One 2011; 6: e24950.
Fornari F, Gramantieri L, Ferracin M, Veronese A, Sabbioni S, Calin GA et al. MiR-221 controls CDKN1C/p57 and CDKN1B/p27 expression in human hepatocellular carcinoma. Oncogene 2008; 27: 5651–5661.
He H, Jazdzewski K, Li W, Liyanarachchi S, Nagy R, Volinia S et al. The role of microRNA genes in papillary thyroid carcinoma. Proc Natl Acad Sci USA 2005; 102: 19075–19080.
Garofalo M, Di Leva G, Romano G, Nuovo G, Suh SS, Ngankeu A et al. miR-221&222 regulate TRAIL resistance and enhance tumorigenecity through PTEN and TIMP3 down regulation. Cancer Cell 2009; 16: 498–509.
Rao X, Di Leva G, Li M, Fang F, Devlin C, Hartman-Frey C et al. MicroRNA-221/222 confers breast cancer fulvestrant resistance by regulating multiple signaling pathways. Oncogene 2011; 30: 1082–1097.
Zhao JJ, Lin J, Yang H, Kong W, He L, Ma X et al. MicroRNA-221/222 negatively regulates estrogen receptor alpha and is associated with tamoxifen resistance in breast cancer. J Biol Chem 2008; 283: 31079–31086.
Miller TE, Ghoshal K, Ramaswamy B, Roy S, Datta J, Shapiro CL et al. MicroRNA-221/222 confers tamoxifen resistance in breast cancer by targeting p27Kip1. J Biol Chem 2008; 283: 29897–29903.
Terasawa K, Ichimura A, Sato F, Shimizu K, Tsujimoto G . Sustained activation of ERK1/2 by NGF induces microRNA-221 and 222 in PC12 cells. FEBS J 2009; 276: 3269–3276.
Zhang CZ, Zhang JX, Zhang AL, Shi ZD, Han L, Jia ZF et al. MiR-221 and miR-222 target PUMA to induce cell survival in glioblastoma. Mol Cancer 2010; 9: 229.
Wang Q, Li W, Zhang Y, Yuan X, Xu K, Yu J et al. Androgen receptor regulates a distinct transcription program in androgen-independent prostate cancer. Cell 2009; 138: 245–256.
Jiang F, Wang Z . Identification and characterization of PLZF as a prostatic androgen-responsive gene. Prostate 2004; 59: 426–435.
Bohrer LR, Chen S, Hallstrom TC, Huang H . Androgens suppress EZH2 expression via retinoblastoma (RB) and p130-dependent pathways: a potential mechanism of androgen-refractory progression of prostate cancer. Endocrinology 2010; 151: 5136–5145.
Whitfield ML, George LK, Grant GD, Perou CM . Common markers of proliferation. Nat Rev Cancer 2006; 6: 99–106.
Mizuno H, Nakanishi Y, Ishii N, Sarai A, Kitada K . A signature-based method for indexing cell cycle phase distribution from microarray profiles. BMC Genomics 2009; 10: 137.
Taneja SS, Ha S, Garabedian MJ . Androgen stimulated cellular proliferation in the human prostate cancer cell line LNCaP is associated with reduced retinoblastoma protein expression. J Cell Biochem 2001; 84: 188–199.
Love HD, Booton SE, Boone BE, Breyer JP, Koyama T, Revelo MP et al. Androgen regulated genes in human prostate xenografts in mice: relation to BPH and prostate cancer. PLoS One 2009; 4: e8384.
Nelson PS, Clegg N, Arnold H, Ferguson C, Bonham M, White J et al. The program of androgen-responsive genes in neoplastic prostate epithelium. Proc Natl Acad Sci USA 2002; 99: 11890–11895.
Wang Q, Carroll JS, Brown M . Spatial and temporal recruitment of androgen receptor and its coactivators involves chromosomal looping and polymerase tracking. Mol Cell 2005; 19: 631–642.
Jia L, Kim J, Shen H, Clark PE, Tilley WD, Coetzee GA . Androgen receptor activity at the prostate specific antigen locus: steroidal and non-steroidal mechanisms. Mol Cancer Res 2003; 1: 385–392.
Bartel DP . MicroRNAs: target recognition and regulatory functions. Cell 2009; 136: 215–233.
John B, Enright AJ, Aravin A, Tuschl T, Sander C, Marks DS . Human MicroRNA targets. PLoS Biol 2004; 2: e363.
Lewis BP, Burge CB, Bartel DP . Conserved seed pairing, often flanked by adenosines, indicates that thousands of human genes are microRNA targets. Cell 2005; 120: 15–20.
Krek A, Grün D, Poy MN, Wolf R, Rosenberg L, Epstein EJ et al. Combinatorial microRNA target predictions. Nat Genet 2005; 37: 495–500.
Vitelli F, Piccini M, Caroli F, Franco B, Malandrini A, Pober B et al. Identification and characterization of a highly conserved protein absent in the Alport syndrome (A), mental retardation (M), midface hypoplasia (M), and elliptocytosis (E) contiguous gene deletion syndrome (AMME). Genomics 1999; 55: 335–340.
Wang WX, Wilfred BR, Hu Y, Stromberg AJ, Nelson PT . Anti-Argonaute RIP-Chip shows that miRNA transfections alter global patterns of mRNA recruitment to microribonucleoprotein complexes. RNA 2010; 16: 394–404.
Chen Z, Zhang C, Wu D, Chen H, Rorick A, Zhang X et al. Phospho-MED1-enhanced UBE2C locus looping drives castration-resistant prostate cancer growth. EMBO J 2011; 30: 2405–2419.
Bernassola F, Karin M, Ciechanover A, Melino G . The HECT family of E3 ubiquitin ligases: multiple players in cancer development. Cancer Cell 2008; 14: 10–21.
Haas AK, Yoshimura S, Stephens DJ, Preisinger C, Fuchs E, Barr FA . Analysis of GTPase-activating proteins: Rab1 and Rab43 are key Rabs required to maintain a functional Golgi complex in human cells. J Cell Sci 2007; 120: 2997–3010.
Weide T, Teuber J, Bayer M, Barnekow A . MICAL-1 isoforms, novel rab1 interacting proteins. Biochem Biophys Res Commun 2003; 306: 79–86.
Stinson S, Lackner MR, Adai AT, Yu N, Kim HJ, O'Brien C et al. TRPS1 targeting by miR-221/222 promotes the epithelial-to-mesenchymal transition in breast cancer. Sci Signal 2011; 4: ra41.
Wang Z, Li Y, Kong D, Banerjee S, Ahmad A, Azmi AS et al. Acquisition of epithelial-mesenchymal transition phenotype of gemcitabine-resistant pancreatic cancer cells is linked with activation of the notch signaling pathway. Cancer Res 2009; 69: 2400–2407.
Yu J, Cao Q, Mehra R, Laxman B, Yu J, Tomlins SA et al. Integrative genomics analysis reveals silencing of beta-adrenergic signaling by polycomb in prostate cancer. Cancer Cell 2007; 12: 419–431.
Santagata S, Demichelis F, Riva A, Varambally S, Hofer MD, Kutok JL et al. JAGGED1 expression is associated with prostate cancer metastasis and recurrence. Cancer Res 2004; 64: 6854–6857.
Pfeil K, Eder IE, Putz T, Ramoner R, Culig Z, Ueberall F et al. Long-term androgen-ablation causes increased resistance to PI3K/Akt pathway inhibition in prostate cancer cells. Prostate 2004; 58: 259–268.
Klein KA, Reiter RE, Redula J, Moradi H, Zhu XL, Brothman AR et al. Progression of metastatic human prostate cancer to androgen independence in immunodeficient SCID mice. Nat Med 1997; 3: 402–408.
Irizarry RA, Hobbs B, Collin F, Beazer-Barclay YD, Antonellis KJ, Scherf U et al. Exploration, normalization, and summaries of high density oligonucleotide array probe level data. Biostatistics 2003; 4: 249–264.
Saeed AI, Bhagabati NK, Braisted JC, Liang W, Sharov V, Howe EA et al. TM4 microarray software suite. Methods Enzymol 2006; 411: 134–193.
Nelson PT, De Planell-Saguer M, Lamprinaki S, Kiriakidou M, Zhang P, O'Doherty U et al. A novel monoclonal antibody against human Argonaute proteins reveals unexpected characteristics of miRNAs in human blood cells. RNA 2007; 13: 1787–1792.
Acknowledgements
This work was supported by a SPORE in Prostate Cancer 2 P50 CA090381-06 and TS was supported by a Department of Defense (DoD) Prostate Cancer Training Award W81XWH-09-1-0372. We thank Dr Peter T Nelson at University of Kentucky and Dr Zissimos Mourelatos at University of Pennsylvania for kindly providing us the anti-AGO antibody. We also thank Dr Wang-xia Wang for technical suggestions on AGO-RIP-Chip experiments.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies this paper on the Oncogene website
Supplementary information
Rights and permissions
About this article
Cite this article
Sun, T., Wang, X., He, H. et al. MiR-221 promotes the development of androgen independence in prostate cancer cells via downregulation of HECTD2 and RAB1A. Oncogene 33, 2790–2800 (2014). https://doi.org/10.1038/onc.2013.230
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/onc.2013.230
Keywords
This article is cited by
-
E3 ubiquitin ligase HECTD2 mediates melanoma progression and immune evasion
Oncogene (2021)
-
Rab1A promotes cell proliferation and migration by upregulating Gli1 in colorectal cancer
Scientific Reports (2021)
-
Wnt/β-catenin signal transduction pathway in prostate cancer and associated drug resistance
Discover Oncology (2021)
-
Potential microRNA-related targets in clearance pathways of amyloid-β: novel therapeutic approach for the treatment of Alzheimer’s disease
Cell & Bioscience (2019)
-
Expression analysis and implication of Rab1A in gastrointestinal relevant tumor
Scientific Reports (2019)