Abstract
Besides its established functions in intermediary metabolism and developmental processes, the nuclear receptor peroxisome proliferator-activated receptor β/δ (PPARβ/δ) has a less defined role in tumorigenesis. In the present study, we have identified a function for PPARβ/δ in cancer cell invasion. We show that two structurally divergent inhibitory ligands for PPARβ/δ, the inverse agonists ST247 and DG172, strongly inhibit the serum- and transforming growth factor β (TGFβ)-induced invasion of MDA-MB-231 human breast cancer cells into a three-dimensional matrigel matrix. To elucidate the molecular basis of this finding, we performed chromatin immunoprecipitation sequencing (ChIP-Seq) and microarray analyses, which identified the gene encoding angiopoietin-like 4 (ANGPTL4) as the major transcriptional PPARβ/δ target in MDA-MB-231 cells, previously implicated in TGFβ-mediated tumor progression and metastatic dissemination. We show that the induction of ANGPTL4 by TGFβ and other oncogenic signals is strongly repressed by ST247 and DG172 in a PPARβ/δ-dependent fashion, resulting in the inhibition of ANGPTL4 secretion. This effect is attributable to these ligands’ ability to induce a dominant transcriptional repressor complex at the site of transcription initiation that blocks preinitiation complex formation through an histone deacetylase-independent, non-canonical mechanism. Repression of ANGPTL4 transcription by inverse PPARβ/δ agonists is functionally linked to the inhibition of cancer cell invasion into a three-dimensional matrix, as (i) invasion of MDA-MB-231 cells is critically dependent on ANGPTL4 expression, (ii) recombinant ANGPTL4 stimulates invasion, and (iii) reverses the inhibitory effect of ST247 and DG172. These findings indicate that a PPARβ/δ–ANGPTL4 pathway is involved in the regulation of tumor cell invasion and that its pharmacological manipulation by inverse PPARβ/δ agonists is feasible.
Similar content being viewed by others
Introduction
Peroxisome proliferator-activated receptor β/δ (PPARβ/δ) is a nuclear receptor whose transcriptional activity is regulated by fatty acid-derived ligands. Although its function in lipid and glucose metabolism, skin repair and macrophage activity is well established, its role in tumorigenesis is unclear and partly controversial.1, 2, 3, 4, 5 PPARβ/δ regulates its target genes through binding to PPAR response elements (PPREs) as heterodimers with a retinoid X receptor (RXR).4 Genome-wide analyses of human myofibroblasts have identified PPRE-mediated repression as a major mechanism of transcriptional regulation by unliganded PPARβ/δ and revealed that only a subset of these repressed genes is activated by an agonist-mediated switch.6
One of the best-established PPARβ/δ target genes is angiopoietin-like 4 (ANGPTL4).7 After secretion, ANGPTL4 is proteolytically cleaved, yielding N-terminal (nANGPTL4) and C-terminal (cANGPTL4) fragments both of which circulate through the blood stream.8 A major function of nANGPTL4 is the inhibition of lipoprotein lipase,9 which is mainly regulated by PPARs,10 while cANGPTL4 appears to have a role in tumor progression and metastasis.11
ANGPTL4 promotes the two-dimensional migration of different cell types in vitro12, 13, 14, 15 and can exert pro-angiogenic effects under certain experimental conditions,16, 17 but the significance of these functions with respect to tumorigenesis is unknown. Furthermore, ANGPTL4 produced by tumor cells is released into the circulation in response to transforming growth factor β (TGFβ), which increases the permeability of lung capillaries and facilitates the extravasation of disseminated cancer cells in a mouse model.15, 18 ANGPTL4 also inhibits anoikis, which is essential for the survival of circulating tumor cells.19 Finally, ANGPTL4 expression is strongly elevated in human clear-cell renal carcinoma,17, 20 correlates with venous invasion in human gastric and colon carcinoma,21, 22 and is part of gene expression signatures associated with distant metastasis and poor outcomes in humans.23, 24 Consistent with these findings, several oncogenic signaling pathways have been shown to converge on the ANGPTL4 gene, including hypoxia-inducible factor-1α,25 AP1 (activator protein 1)26 and SMAD proteins.15, 26 ANGPTL4 transcription is also regulated by the glucocorticoid receptor27 and all members of the PPAR family.9, 26
Previous reports have suggested a function for PPARβ/δ in the two-dimensional migration of different cell types, including keratinocytes28 and vascular smooth muscle cells,29 but its potential significance with respect to cancer cell invasion and metastasis in unknown. In the present study, we have investigated the role of PPARβ/δ-mediated transcriptional repression in cancer cell invasion, with a focus on the PPARβ/δ–ANGPTL4 signaling pathway. Toward this goal, we made use of recently developed subtype-specific PPARβ/δ inhibitors (ST247, DG172; Figure 1a), which downregulate expression of ANGPTL4 by acting as inverse agonists through an unknown mechanism.30, 31, 32 Inverse agonists are defined as ligands that, beyond antagonizing agonist binding, exert an opposite effect as an agonist. Thus, in case of PPARβ/δ, an agonist induces a transcriptional activator complex by facilitating the association of PPARβ/δ with coactivators, whereas an inverse agonist triggers the recruitment of transcriptional corepressors and thereby the formation of a repressor complex.
Invasion of a three-dimensional matrigel matrix by MDA-MB-231 and its inhibition by the inverse PPARβ/δ agonists ST247 and DG172. (a) Chemical structures of ST247 and DG172. (b, c) MDA-MB-231 cells were treated with DMSO or ST247 and analyzed for invasion of a three-dimensional matrigel matrix using serum (10% FCS) as the attractant. Representative images show confocal sections through invaded cells stained for F-actin (red) and DNA (Cytox green) at 20 μm distance to the transwell membrane. Three-dimensional reconstruction shows a side view (z) of the cells’ location relative to the transwell membrane (dashed line). (d) Quantification of invasion assays as in a but including DG172. Horizontal lines indicate the median of biological replicates (N=4 for ST247; N=2 for DG172). ***Significant difference between untreated (solvent only) and treated cells (P<0.001 by t-test).
Results
Invasion of a three-dimensional matrigel matrix by MDA-MB-231 cells is inhibited by inverse PPARβ/δ agonists
The human breast cancer cell line MDA-MB-231 is a well-established model system to study cancer cell invasion. We therefore studied the effect of inverse PPARβ/δ agonists on the serum-induced invasion of MDA-MB-231 cells into a three-dimensional matrigel matrix using an inverse transwell assay (see cartoon in Supplementary Figure S1). Figures 1b–d demonstrates that both inverse PPARβ/δ agonists ST247 and DG172 strongly inhibited invasion. These compounds bear no structural similarities (see Figure 1a), suggesting that off-target effects mediating the observed inhibition are very unlikely. Surprisingly, the activating PPARβ/δ agonists L165,041 and GW501516 did not enhance invasion (not shown), which we attribute to the complexity of the agonist response (see Discussion).
Genome-wide identification of PPARβ/δ-RXR binding sites in MDA-MB-231 cells
To elucidate the molecular mechanisms underlying the inhibition of tumor cell invasion by ST247 and DG172, we performed chromatin immunoprecipitation sequencing (ChIP-Seq) to identify PPARβ/δ target genes in MDA-MB-231 cells. Deep sequencing of DNA from PPARβ/δ- or RXR-bound chromatin yielded a total of 20 million reads each mappable to unique locations on the human genome. Bioinformatic analysis identified a total of 527 high confidence enrichment peaks (false discovery rate <0.05) for PPARβ/δ (Figure 2a, Supplementary Dataset S1) and 37 415 peaks for RXR (Figure 2a). Peaks for PPARβ/δ and RXR overlapped at 484 genomic regions (Figure 2a; Supplementary Dataset S1), indicating co-occupancy of 92% of PPARβ/δ-binding sites by RXR. Most of the PPARβ/δ–RXR enrichment sites (89%) were found inside or <25 kb upstream of transcribed genomic regions. A de novo motif search (MEME) yielded a 17-bp consensus sequence (AAgTAGGtcAAAGGTcA) that is almost identical to the direct repeat motif (DR-1) previously identified in WPMY-1 cells (AAgTGGGtcAAAGGTcA).6
Genome-wide mapping of chromatin-bound PPARβ/δ-RXR and identification of PPARβ/δ-dependent or ligand-responsive target genes in MDA-MB-231 cells. (a) Left: overlaps between genomic loci with an enrichment of PPARβ/δ or RXR determined by ChIP-Seq; right: overlaps between PPARβ/δ-RXR-bound genes that are regulated by PPARD siRNA, activated by GW501516 or repressed by ST247 (threshold 1.2-fold). (b) Response of individual target genes to PPARD siRNA or GW501516 using the previously established classification6. Different types of responses are shown in different colors. (c) Response of individual target genes to PPARD siRNA or ST247. (d) Validation of PPARβ/δ and RXRα binding to the ANGPTL4 gene by ChIP-qPCR; immunoglobulin G (IgG): negative control antibody; ctrl: unrelated genomic region. (e, f) Validation of expression data by reverse transcriptase–qPCR (averages ±s.d.; N=3). ***, **Significant difference between IgG and PPARβ/δ or RXR in d, between solvent and ligand-treated cells in e, and between si-con and siPPARD treated cells in f (P<0.001, P<0.01 by t-test). Ligand exposure was for 6 h in all cases; none: solvent (DMSO) only.
Identification of PPARβ/δregulated and ligand-responsive target genes
Microarray analysis of MDA-MB-231 cells exposed to (i) control or a previously validated subtype-specific PPARD siRNA, (ii) the agonist GW501516 for 6 h or (iii) the inverse agonist ST247 for 6 h enabled the delineation of subgroups of expression-correlated peaks (Figure 2a; Supplementary Dataset S2). Out of a total of 178 siRNA-responsive genes, 59 were induced by GW501516, 31 were repressed by ST247 and 16 were regulated by both ligands. A correlation analysis of siRNA and agonist responses (Figure 2b) yielded a similar classification scheme as previously described for WPMY-1 cells6, that is, type I (red): agonist-independent repression by PPARβ/δ; type II (blue): agonist-sensitive repression by PPARβ/δ; and type III (green): agonist-independent activation by PPARβ/δ. Interestingly, genes showing a type II response were preferentially repressed by ST247 (Figure 2c), supporting the view that the PPARβ/δ repressor complexes acting on different groups of target genes are not identical. PPARβ/δ-RXR binding to the ANGPTL4 gene and its regulation as a type II response were confirmed by ChIP-qPCR (quantitative PCR; Figure 2d) and by reverse transcriptase–qPCR experiments (Figures 2e and f).
A functional annotation of ST247-repressed genes (Supplementary Dataset S3) identified two gene oncology terms representing metabolic genes with P-values of 0, thus validating the present analysis. Furthermore, the functional annotation identified several groups with a P-value ⩽0.05 that might be interesting in the context of the mechanistically unresolved role of PPARβ/δ in cell proliferation and survival,33 that is, apoptosis, negative regulation of transcription, signal transduction and differentiation. By contrast, gene oncology terms defining genes involved in invasion were not identified.
Inhibition of TGFβ-induced invasion and modulation of the global transcriptional response to TGFβ by the inverse PPARβ/δ agonist ST247
As TGFβ stimulates cancer cell invasion34 and PPARβ/δ and TGFβ signaling pathways are interconnected,26, 35 we investigated how a PPARβ/δ inverse agonist would affect TGFβ-dependent stimulation of cancer cell invasion. As shown in Figure 3a, TGFβ stimulated matrigel invasion by MDA-MB-231 cells, and this effect was clearly reversed by ST247. We therefore analyzed the global transcriptional response to ST247 under these conditions. Microarray analyses identified 107 genes that were induced by a 6-h TGFβ treatment, which was counter-regulated by ST247 for 17 genes (Figure 3b; Supplementary Dataset S4). We also identified 43 genes repressed by TGFβ, which was reversed by ST247 in 13 cases (Figure 3c; Supplementary Dataset S4). Although for most of these genes, ST247 counteracted the induction by TGFβ (colored dots clustering near the abscissa in Figure 3d), ANGPTL4 was repressed even below the basal level expression and clearly showed the strongest regulation (∼40-fold; Figures 3d and e). Finally, we sought to narrow down the number of potential target genes that might be involved in mediating the inhibitory effect of ST247 on invasion. We therefore determined the overlap of genes repressed by ST247 either under basal conditions (fetal calf serum (FCS)) or under TGFβ stimulation and identified a set of 61 genes, again with ANGPTL4 showing the strongest regulation (Figure 3f; Supplementary Dataset S5). This list does not contain other genes with a known role in invasion, although there are few candidates that may have a role in actin-mediated signaling, including genes coding for the ill-defined formins FHOD3 and INF2, or proteins involved in matrix reorganization, such as MMP9. However, because of its exceptionally strong PPARβ/δ responsiveness and its documented role in tumor progression, we subsequently focused on the ANGPTL4 gene.
Inhibition of matrigel invasion by MDA-MB-231 cells and modulation of the global transcriptional response to TGFβ by ST247. (a) Cells were treated with DMSO or ST247 and analyzed for invasion as in Figure 1, except that TGFβ2 (2 ng/ml) was used as a chemoattractant in addition to FCS. Horizontal lines indicate the median of biological replicates (N=3). **Significant differences by t-test (P<0.01). (b) Diagram depicting the set of TGFβ-induced genes (N=107; threshold ⩾3-fold) and counter-regulation by ST247 for a subset of these genes (N=17; threshold ⩾3-fold). Genes upregulated by ST247 alone (threshold ⩾2-fold) were excluded. Cells were treated with TGFβ2 in the presence or absence of ST247 for 6 h, and the transcriptional profiles were determined by microarray analysis. (c) Diagram depicting the set of TGFβ-repressed genes (N=43; threshold ⩾3-fold) and counter-regulation by ST247 for a subset of these genes (N=13; threshold ⩾3-fold). Details as in panel c. (d) Scatter plot showing the response of individual TGFβ target genes to ST247 exposure. Relative expression values determined in the presence of TGFβ and ST247 are plotted against the expression values measured in the presence of TGFβ only. Colored dots represent genes counter-regulated by ST247 by ⩾1.5-fold (e) Effect of ST247 on TGFβ-induced target genes, corresponding to the blue data points in panel d (ST247 effect ⩾1.5-fold; N=17). (f) Venn diagram showing the overlap of genes (N=61) downregulated by ST247 in the absence (left) or presence (right) of TGFβ (threshold ⩾1.5-fold downregulation by ST247).
Inverse PPARβ/δ agonists inhibit the induction of ANGPTL4 by multiple signaling pathways
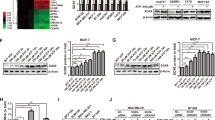
The reported upregulation of ANGPTL4 expression in tumors by different oncogenic signals led us to investigate whether inverse PPARβ/δ agonists might exert dominant-repressive effects. The data in Figure 4a show that upregulation of ANGPTL4 in MDA-MB-231 cells by insulin, 12-O-tetradecanolphorbol myristate acetate, dexamethasone, basic fibroblast growth factor or TGFβ is almost completely inhibited by ST247. By contrast, PAI1, a classical direct TGFβ target gene lacking PPARβ/δ sites26, did not show any detectable response to ST247 (Figure 4b), indicating that TGFβ signaling is intact and does not change under ST247 treatment. Furthermore, the observed inhibition of signal-induced ANGPTL4 expression by inverse PPARβ/δ agonists appears to be of general relevance, as it was also observed in WPMY-1 myofibroblasts (Figure 4c), in Caki-1 renal carcinoma cells (Figure 4d) and upon treatment of WPMY-1 cells with the structurally unrelated inverse PPARβ/δ agonist DG172 (Figure 4c). By contrast, the irreversible PPARβ/δ antagonist GSK378736 failed to show any detectable inhibition (Figure 4c). Finally, the inhibitory effect on both the basal level and TGFβ-induced ANGPTL4 mRNA expression resulted in clearly decreased levels of ANGPTL4 polypeptide(s) in the culture supernatant of WPMY-1 (Figure 4e), Caki-1 cells (Figure 4f) and MDA-MB-231 cells (Figure 4g). By using recombinant full-length and cleaved forms of ANGPTL4, we identified the protein detected by this assay as the C-terminal fragment cANGPTL4 (Supplementary Figure S3).
ST247 and DG172 inhibit the induction of ANGPTL4 by TGFβ and other signaling pathways. (a–d) Reverse transcriptase–qPCR analysis of ANGPTL4 and PAI1 expression in the indicated cell lines treated with TGFβ2 (2 ng/ml), 12-O-tetradecanolphorbol myristate acetate (50 nM), dexamethasone (Dex; 1 μM), insulin (1 μg/ml) or basic fibroblast growth factor (bFGF; 10 ng/ml) in the absence or presence of the different ligands (1 μM) for 6 h. (e–g) Enzyme-linked immunosorbent assay of cANGPTL4 in culture supernatants of the indicated cell lines treated with TGFβ2±ST247 for 24 h. Values are expressed as arbitrary units (a.u.; normalized to 1 for untreated cells) and represent averages ±s.d. (N=3). ***, **, *Significant difference between solvent and ligand-treated cells. n.d., value too low to be determinable. none: as in Figure 1.
ST247 inhibits formation of a transcriptional initiation complex at the ANGPTL4 locus
We next addressed the molecular background of the dominant-repressing effect exerted by inverse PPARβ/δ agonists. ChIP-based analyses using antibodies detecting all forms of the large subunit of RNA polymerase II (RPB1) revealed a decreased accumulation at the transcriptional start site of the ANGPTL4 gene by ST247 in TGFβ-stimulated WPMY-1 cells (61% reduction; Figure 5a). Similar results were obtained with TGFβ-treated MDA-MB-231 cells (Figure 5b) and Caki-1 cells (Figure 5c). These data indicate a strong inhibition by ST247 of TGFβ-induced preinitiation complex formation at the ANGPTL4 gene. ChIP walking of the ANGPTL4 locus showed RPB1 accumulation along the transcribed region upon TGFβ stimulation of Caki-1 cells (Figure 5c), indicating transcriptional elongation. As predicted by the lack of preinitiation complex formation, substantially lower amounts of elongating RNA polymerase II were detected upon ST247 treatment (Figure 5c).
Effect of ST247 on the transcriptional machinery at the ANGPTL4 locus. (a, b) Decreased RNA polymerase II accumulation at the TSS of the ANGPTL4 gene in WPMY-1 (a) or MDA-MB-231 cells (b), determined by ChIP. (c) ChIP walking of RNA polymerase II recruitment to the transcribed region of the ANGPTL4 gene in Caki-1 cells. (d) HDAC3 recruitment to the ANGPTL4 gene in ST247-treated MDA-MB-231 (top) and Caki-1 cells (bottom) in the presence and absence of TGFβ treatment. Cells were treated with 2 ng/ml TGFβ2, 1 μM ST247 or both for 30 min, and ChIP was carried out with a nonspecific immunoglobulin G pool, anti-HDAC3 or anti-RPB1 antibodies. (e) Effect of the HDAC inhibitor trichostatin A TSA (6 h exposure) on basal level and ST247-mediated repression of ANGPTL4 transcription in MDA-MB-231 cells (reverse transcriptase–qPCR analysis). (f) Effects of siRNA-mediated silencing of HDAC3, NCOR1, NCOR2 and SHARP or double knock downs of NCOR1+NCOR2 (siNCOR1+2) or NCOR1+NCOR2+SHARP (siNCOR1+2+SHARP) on basal level and ST247-mediated repression of ANGPTL4 transcription in MDA-MB-231 cells. The efficiency and specificity of the siRNAs used is shown in Supplementary Figure S5. ***, **, *Significant effects of ST247 in a, b, d and e on basal or TGFβ-induced transcription or DNA enrichment, as indicated, and of the indicated siRNAs on basal repression in f (relative to the control siRNA, si-con). none: as in Figure 1.
ST247 triggers the formation of an histone deacetylase (HDAC)-independent repressor complex
The repression of target genes by unliganded PPARβ/δ is mediated by NCoR/SMRT-HDAC3 corepressor complexes, other HDACs and SHARP.37, 38, 39 We therefore analyzed the role of these corepressors in ST247-mediated repression. Surprisingly, ST247 had no effect on the recruitment of HDAC3 to the ANGPTL4 PPREs in MDA-MB-231 cells (Figure 5d; top), although HDAC3 accumulation at the ANGPTL4 gene was seen in Caki-1 cells (Figure 5d; bottom) as previously reported for WPMY-1 cells.30 This finding indicates that HDAC3 does not have a general role in PPARβ/δ-mediated repression by inverse agonists. Consistent with this conclusion, inhibition of all HDAC subtypes by trichostatin A had no effect on the extent of ST247-mediated repression in MDA-MB-231 cells but attenuated basal repression (Figure 5e). However, the siRNA-mediated downregulation of HDAC3 had no significant effect (Figure 5f), suggesting that other HDACs may be involved in basal repression. By contrast, silencing of NCOR1, NCOR2 (SMRT) or SHARP expression by siRNA clearly diminished basal repression but, surprisingly, did not lower the extent of repression by ST247 (Figure 5f; ⩾4.5-fold in all the cases). Furthermore, although the simultaneous silencing of NCOR1, NCOR2 (SMRT) and SHARP completely abolished basal repression, ST247 was still able to repress gene expression (Figure 5f). These data clearly indicate that the ST247-mediated repression of the ANGPTL4 gene is independent of NCoR/SMRT-HDAC complexes and is thus functionally different from the canonical repression mechanism used by PPARβ/δ. PPARβ/δ protein levels were not changed upon treatment with ST247 (Supplementary Figure S2), which is consistent with the conclusion that specific chromatin alterations mediate the inhibitory effect of inverse PPARβ/δ agonists on transcription.
ANGPTL4 induces invasion of MDA-MB-231 cells and counteracts the effect of inverse agonists
Next, we addressed the question as to whether inverse PPARβ/δ agonists target ANGPTL4 to inhibit cancer cell invasion. Toward this goal, we first analyzed the effect of recombinant ANGPTL4 on the invasion of a matrigel matrix by MDA-MB-231 cells. As illustrated in Figures 6a–c, recombinant cANGPTL4 increased the number of invading cells by 80% and was more efficient than the full-length protein at equimolar concentrations, while nANGPTL4 had no detectable effect. Remarkably, recombinant cANGPTL4 also prevented the inhibitory effect of ST247 and DG172 (Figure 6d), indicating that repression of ANGPTL4 transcription indeed mediates the inhibition of invasion by inverse PPARβ/δ agonists, although other PPARβ/δ target genes may be involved as discussed above.
ANGPTL4-induced invasion by MDA-MB-231 and ANGPTL4-mediated reversal of the inhibitory effect exerted by ST247. (a, b) Effect of recombinant nANGPTL4 (a) and cANGPTL4 (b) on invasion. (c) Quantification of the effects of full-length ANGPTL4 (A4) and the cleaved forms nANGPTL4 (nA4) and cANGPTL4 (cA4) on invasion. (d) Reversal of the inhibitory effect of ST247 or DG172 on invasion by recombinant cANGPTL4. Cells were treated as in Figure 1 and invasion was determined in the presence or absence of cANGPTL4. Horizontal lines indicate the median of biological replicates (N=2 in c; N=3 in d). **, *Significant difference between untreated (solvent only) and treated cells (P<0.01, P<0.05 by t-test). none: as in Figure 1.
Silencing of the ANGPTL4 or PPARD genes confirms their role in invasion
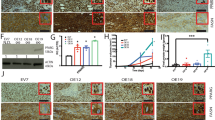
Finally, we sought to obtain additional evidence for the role of PPARβ/δ and ANGPTL4 in invasion of a three-dimensional matrix by MDA-MB-231 cells. As shown in Supplementary Figures S5 and S6, siRNAs against PPARD (si-PPARD) or ANGPTL4 (si-ANGPTL4) clearly reduced the level of PPARβ/δ protein in MDA-MB-231 and the level of cANGPTL4 in culture supernatants. Although si-con (directed at an irrelevant sequence) had no effect on invasion compared with untreated cells (Figures 7a and d), si-ANGPTL4 treatment led to a nearly complete loss of invasion (∼80% reduction; Figures 7b and d), confirming an essential role for ANGPTL4 in this process. By contrast, si-PPARD stimulated invasion (Figures 7c and d), presumably due to derepression of the ANGPTL4 gene (Figure 2f). Importantly, in si-PPARD-treated cells, ST247 had no significant effect on ANGPTL4 expression (Supplementary Figure S8) or invasion (Figure 7e), providing further evidence for the target specificity of this compound.
Oppositional effects of ANGPTL4 or PPARD silencing on matrigel invasion by MDA-MB-231 cells. (a–c) Cells were treated with an irrelevant control siRNA (si-con, panel a), si-ANGPTL4 (panel b) or si-PPARD (panel c) for 48 h and analyzed for invasion as in Figure 1. (d) Quantification of the data of panels a–c. (e) Dependence of the ST247 effect on PPARβ/δ. Cells were treated with si-con or si-PPARD in the presence or absence of ST247 and analyzed for invasion as in Figure 1. Horizontal lines indicate the median of biological replicates (N=3). ***, *Significant difference between untreated (nt) and treated cells (P<0.001, P<0.05 by t-test).
Discussion
A PPARβ/δ–ANGPTL4 pathway involved in cancer cell invasion
The present study provides strong evidence for a PPARβ/δ–ANGPTL4 pathway that modulates cancer cell invasion. This conclusion is based on several key observations as discussed below.
First, PPARβ/δ is linked to invasion as shown by the ability of two structurally divergent inverse PPARβ/δ agonists to inhibit the serum- and TGFβ-triggered invasion into a three-dimensional matrix by MDA-MB-231 cells (Figures 1b–d and 3a). Importantly, this effect is dependent on the presence of PPARβ/δ (Figure 7e), providing strong evidence for the target specificity of these ligands.
Second, our data clearly link PPARβ/δ to the regulation of the ANGPTL4 gene in MDA-MB-231 cells and identify ANGPTL4 as the major transcriptional PPARβ/δ target gene in these cells. This is demonstrated by unique response of ANGPTL4 to the activating agonist GW501516, the repressing inverse agonists ST247 and DG172 and the siRNA-mediated interference with PPARβ/δ expression (Figure 2). Intriguingly, the inverse PPARβ/δ agonists repress ANGPTL4 transcription and ANGPTL4 secretion not only under basal conditions but also under the influence of activating oncogenic signaling pathways, including TGFβ (Figures 3 and 4).
Third, ANGPTL4 is directly linked to invasion. This conclusion is based on the observations that recombinant ANGPTL4 promotes invasion by MDA-MB-231 cells (Figures 6b and c), whereas ANGPTL4 silencing has the opposite effect (Figures 7b and d). This novel role for ANGPTL4 in promoting cancer cell invasion is consistent with clinical observations, suggesting a correlation of ANGPTL4 expression and venous invasion by colon and gastric carcinoma cells.21, 22 Furthermore, in line with our observations, recent reports have described a function for ANGPTL4 in promoting keratinocyte migration and wound healing.12, 40 After secretion, ANGPTL4 is cleaved by proprotein convertases into two biologically active fragments.8, 41 We show that only the C-terminal cleavage product, cANGPTL4, is able to promote invasion when applied as a recombinant protein to MDA-MB-231 cells (Figures 6a–c). Consistent with this finding, we show that all the tested cell lines secrete cANGPTL4 (Figures 4e–f), which is induced by TGFβ and inhibited by inverse PPARβ/δ agonists. This data suggests that the observed effects on invasion are mediated by cANGPTL4.
Finally, our data strongly suggest that ANGPTL4 is a crucial mediator linking PPARβ/δ to invasion. Thus, the inhibition of invasion by ST247 and DG172 is prevented by recombinant cANGPTL4 (Figure 6d), and invasion by MDA-MB-231 cells is highly dependent on ANGPTL4 (Figures 7b and d). Taken together, these findings indicate that repression of the ANGPTL4 gene and inhibition of invasion by inverse PPARβ/δ agonists are functionally connected.
An apparently contradictory finding is the observation that PPARβ/δ agonists do not promote invasion by MDA-MB-231 cells (not shown), although ANGPTL4 transcription is strongly induced (Figure 2e). As an elevation of ANGPTL4 levels should enhance invasion, as seen for the application of recombinant proteins (Figure 6c), we attribute the inefficiency of PPARβ/δ agonists to their pleiotropic effects on the transcriptome. Thus, the agonist GW501516 induces the expression of 234 genes (threshold ⩾1.5-fold; Supplementary Dataset S6) after 48 h of treatment, which corresponds to the conditions of the invasion assay. Among these are several genes coding for proteins with possible direct or indirect functions in actin-dependent and migration-associated signaling pathways, including FHOD3, LCN2, TMPRSS6, ESCSR, CCK, GRP, TGFA, GLI2 and LATS1 (Supplementary Dataset S6; lines 182, 20, 39, 91 101, 117, 166, 212 and 228, respectively). It is conceivable that perturbing the balance of such proteins may interfere with the biological output generated by a single signaling molecule, such as the stimulation of invasion by ANGPTL4. Furthermore, some of the effects of GW501516, including its impact on the cellular ADP/ATP ratio, have been reported to result from PPARβ/δ-independent mechanisms,42 which may also contribute to the observed inhibition of invasion. Similar to our observations with PPARβ/δ agonists, previous reports have described an inhibition rather than stimulation of cancer cell invasion by PPARγ agonists,43, 44, 45, 46 although these ligands induce ANGPTL4 expression.10, 16, 26, 47, 48, 49 It is very likely that this apparent discrepancy is due to the complexity of the transcriptional response to PPARγ agonists, as discussed above for PPARβ/δ ligands.
Interestingly, ST247 also affects several genes with putative functions in the regulation of the actin cytoskeleton and/or migration other than ANGPTL4, such as FHOD3, INF2 and MMP9 (Supplementary Dataset S5; lines 18, 42 and 43, respectively). Even though these genes are repressed to a considerably lesser extent than ANGPTL4 (Supplementary Dataset S5), it is conceivable that other ST247-repressed genes contribute to the inhibition of invasion.
Role of PPARβ/δ and ANGPTL4 in migration and invasion in other experimental systems and cell types
The novel role for PPARβ/δ in cancer cell invasion described in the present study extends previous reports suggesting a function for PPARβ/δ in the two-dimensional migration of different cell types, including keratinocytes28 and vascular smooth muscle cells.29 However, the molecular mechanisms underlying PPARβ/δ-modulated cancer cell invasion appear to be different. Thus, keratinocytes with a disrupted Ppard gene have a diminished migratory capacity,28 although ANGPTL4 transcription is likely to be derepressed in the absence of PPARβ/δ as suggested by siRNA experiments with other cell types (for example, Figure 2b). Furthermore, we have observed that the MDA-MB-231 cells used in the present study do not show any significant migration in scratch assays, indicating that the molecular mechanisms associated with three-dimensional invasion and affected by inverse PPARβ/δ agonists are distinct from those regulating two-dimensional migration.
Our observations are consistent with other published reports12, 13, 14, 15 describing a role for ANGPTL4 in stimulation migration. One of the best documented example for a migration-promoting role of ANGPTL4 is wound healing.12 ANGPTL4 specifically interacts with integrins β1 and β5 on keratinocytes and consequently activates numerous integrin-mediated intracellular signaling cascades, including focal adhesion kinase, the RhoGTPase RAC1 and protein kinase C, which may all contribute to the promotion of migration.7 ANGPTL4 also activates 14-3-3σ-dependent signaling pathways, protein kinase C and the transcription factor AP-1, which enhances keratinocyte differentiation.40 In cancer cells, ANGPTL4 has been reported to mediate different pro-metastatic effects, in particular, the promotion of extravasation15 and the inhibition of anoikis in circulating tumor cells.19 Although the former function appears to depend on the extracellular interaction of ANGPTL4 with VE-cadherin and claudin-5,18 resulting in the disintegration of tight junctions, its function in preventing anoikis is mediated by a redox-based signaling mechanism.19 This signaling pathway involves the integrin-mediated activation of NADPH oxidase (NOX1), which generates an elevated oncogenic O2−/H2O2 ratio. This, in turn, triggers activation of the SRC kinase and the downstream PI3K/PKB and ERK (extracellular signal–regulated kinase) signaling cascades, promoting cell survival.19 However, it also has to be mentioned that a negative role for ANGPTL4 has been decribed for endothelial cells, associated with RAF-ERK signaling.50 However, in this system, inhibition of migration appears to be due to a specific biological mechanism, as it appears to be linked to diminished chemotaxis and decreased cell proliferation.50
We currently do not know which of these pathways are relevant for ANGPTL4-mediated invasion and its inhibition by inverse PPARβ/δ agonists. ST247 does not cause any significant change in the surface expression of integrins β1 and β5 (Supplementary Figure S7), supporting the view that its effect is mainly mediated by decreasing the availability of cANGPTL4 and consequently by suppressing cANGPTL4-mediated signaling events. Future studies will have to dissect the complexity of ANGPTL4-triggered signaling cascades in the context of invasion to answer this highly relevant question.
Although our findings regarding the role of ANGPTL4 in invasion are consistent with the published data on its role in migration eluded to above, they are in conflict with another report51 suggesting an inhibitory role for ANGPTL4 in cancer cell invasion and metastasis formation in two mouse tumor cell lines, that is, Lewis lung carcinoma (LLC1) and B16 melanoma. We therefore tested LLC1 cells in our assays. ANGPTL4 was repressed by inverse PPARβ/δ agonists in LLC1 cells (data not shown) similarly to the human cell lines analyzed in Figure 4. Invasion by LLC1 was weak compared with MDA-MD-231 cells but was induced by recombinant cANGPTL4 and inhibited by ST247 (Supplementary Figure S9). Intriguingly, TGFβ did not induce the invasion of LLC1 in the same assay (data not shown). This supports the conclusion that ANGPTL4 has a central role in invasion, as the murine ANGPTL4 gene lacks functional SMAD-binding sites and consequently is not inducible by TGFβ.26
The discrepancy to the published data51 may be due to differences in the experimental approaches, as Galaup et al. used transfected cells overexpressing ANGPTL4. Under these conditions, the protein may be differently localized, processed and/or post-translationally modified compared with the moderately expressed endogenous protein or recombinant ANGPTL4. Furthermore, the effects observed by these authors did not correlate within the level of ANGPTL4 expression by different clones, suggesting that clonal variability may have been involved. The same laboratory also observed an inhibitory effect of extracellular matrix-bound ANGPTL4 on the migration of endothelial cells.52 It is possible that in this case a different, integrin-independent mechanism is involved compared with the effect exerted by soluble cANGPTL4 observed in the present study. The relationship between the processing of ANGPTL4 and its different biological functions is poorly understood and remains a subject for future investigations.7
Induction of a dominant-repressor complex at the ANGPTL4 gene by inverse PPARβ/δ agonists
Intriguingly, ST247 and DG172 inhibit cancer cell invasion triggered by diverse stimuli, that is, serum, where the active component is probably lysophosphatidic acid signaling through the PI3K-PAK1-ERK pathway53 and through TGFβ signaling via SMAD proteins.15, 26 Our ChIP data (Figures 5a–c) show that these inverse agonists prevent preinitiation complex formation, irrespective of the binding of activating transcription factors to the ANGPTL4 gene, such as SMAD3 or AP1 (Supplementary Figure S4), demonstrating the dominant nature of the repressing effect. Our data also indicate that these ligands trigger the recruitment of corepressors to chromatin-bound PPARβ/δ, such as HDAC3 (Figure 5d). However, several lines of evidence suggest that the canonical repressor complex containing NCoR/SMRT and HDACs mediating repression by the unliganded receptor functionally differs from the complex established in response to inverse PPARβ/δ agonists. First, HDAC3 is recruited in Caki-1 cells but not in MDA-MB-231 cells (Figure 5d), although the latter show a stronger response to ST247. Second, trichostatin A alleviates basal repression but has no effect on the repression by ST247 (Figure 5e). Third, the silencing of NCOR1 and/or NCOR2 (SMRT) derepresses ANGPTL4 transcription, but repression by ST247 is not affected (Figure 5f). Likewise, knock down of SHARP partially counteracts ANGPTL4 repression but has no detectable effect of ST247-mediated repression (Figure 5f). These findings suggest that inverse PPARβ/δ agonists trigger the formation of an unusual PPARβ/δ repressor complex exerting dominant effects on other transcription factors. It will be intriguing to identify the interacting proteins responsible for this effect in future studies.
Perspectives
ANGPTL4 has been associated with tumor angiogenesis, metastatic spread and an unfavorable prognosis in human breast cancer.15, 17, 18, 19, 20, 21, 22, 23, 24 One of the compounds showing a strong effect on cancer cell invasion in the present study, DG172, has the required pharmacokinetic properties for in vivo applications in mice.31 Our studies now pave the way to investigate the effect of DG172 in mouse models of tumorigenesis and to assess its potential with respect to a pharmacological interference with ANGPTL4-driven cancer cell invasion.
Materials and methods
Chemicals
TGFβ2, 12-O-tetradecanolphorbol-13-acetate, insulin and dexamethasone were purchased from Sigma-Aldrich (Steinheim, Germany), basic fibroblast growth factor and trichostatin A from Biomol (Hamburg, Germany), L165,041 from Calbiochem (Darmstadt, Germany) and GW501516 from Axxora (Lörrach, Germany). ST247, DG172 and GSK3787 were synthesized as described.30, 31, 32, 36
Cell culture
MDA-MB-231 cells were purchased from Caliper Life Science (Mainz, Germany) (MDA-MB-231-luc2), WPMY-1 and Caki-1 cells were obtained from the ATCC. Cells were maintained in DMEM (Dulbecco’s modified Eagle’s medium; MDA-MB-231, WPMY-1) or McCoy’s A medium (Caki-1) supplemented with 10% fetal bovine serum, 100 U/ml penicillin and 100 μg/ml streptomycin in a humidified incubator at 37 °C and 5% CO2.
Three-dimensional Matrigel invasion-assays
MDA-MB-231 cells were treated for 48 h with ST247 or DG172 (each at 1 μM) or DMSO (dimethyl sulfoxide; 1:10 000) as solvent control. Transwell inserts (Thincerts, Greiner Bio-One, Solingen, Germany)) were coated with 50 μl growth factor-reduced Matrigel (BD Biosciences; Heidelberg, Germany) at 5 μg/μl and cell invasion was analyzed essentially as described.54 Briefly, cells were harvested and 15 000 cells were seeded on the bottom of the transwell inserts and allowed to adhere for 1 h. Thincerts were inverted and medium, containing ST247, DG172 or DMSO as indicated, was added to the top (containing 10% FCS) and the lower compartment (containing 0.5% FCS). Cells were fixed with 8% formaldehyde after 24 h, stained as indicated and invaded cells were quantified from six randomly chosen sections from each thincert using a confocal microscope (Zeiss LSM 700). ANGPTL4 peptides (Enzo Life Science, Lörrach, Germany) were imbedded at 2 μM in the matrix and TGFβ2 (2 ng/ml) was added to the upper chamber of the thincert as indicated.
Immunoblotting
Immunoblots were performed according to the standard protocols using the following antibodies: α-PPARβ/δ (sc-74517; Santa Cruz, Heidelberg, Germany); α-HDAC3 (sc-11417; Santa Cruz) and α-NCOR1 (No. 5948; Cell Signaling, Frankfurt am Main, Germany).
ANGPTL4 enzyme-linked immunosorbent assay
ANGPTL4 levels in cell culture supernatants were determined by a commercial ELISA kit (RayBio, Norcross, GA, USA or Beneficial Solution, Santa Clara, CA, USA) according to the manufacturer’s instructions. Briefly, MDA-MB-231 cells were grown to a semi-confluent state, the growth medium was replaced with fresh medium containing 10 U/ml heparin, and ligands and/or TGFβ (as indicated) were added after 12 h. Culture supernatant was harvested for analysis 6 h later. WPMY-1 and Caki-1 cells were grown without heparin. Fresh medium and ligands and/or TGFβ were added 2h after seeding, and supernatants were harvested after 24 h of treatment.
siRNA transfections
siRNA transfections were carried out essentially as described6 using pools of 4 siRNAs per gene (Dharmacon, Dreieich, Germany). Cells were seeded at a density of 1 × 106 cells per 6 cm dish in 4 ml DMEM with 10% FCS and cultured for 4 h. In all, 1280 ng siRNA in 100 μl OptiMEM (Invitrogen, Darmstadt, Germany) and 15 μl HiPerfect (Qiagen, Hilden, Germany) or RNAiMAX (Invitrogen, according to the manufacturer’s instructions) were mixed and incubated for 5–10 min at room temperature before transfection. The cells were replated 24 h post-transfection at a density of 1 × 106 cells per 6 cm dish. Transfection was repeated 48 h after start of the experiment, and cells were passaged after another 24 h. Forty-eight hours following the last transfection, cells were stimulated with ligands as indicated and harvested after another 6 h.
Quantitative reverse transcriptase–PCR
cDNA and qPCR were performed as described.30 L27 was used for normalization. Comparative expression analyses were statistically analyzed by Student’s t-test (two-tailed, equal variance). Primer sequences for L27, ANGPTL4 and PAI1 have been published previously.26, 55
Microarrays
Microarray analyses were carried out as published.6 Raw and normalized microarray data were deposited at EBI ArrayExpress. Probes were considered regulated if they had an averaged log intensity ⩾5, a fold change ⩾1.2 and replicates were within 50% of each other. Probes were assigned to genes as described6 using Ensembl release 65. Microarray data were deposited at EBI ArrayExpress (accession numbers E-MTAB-1098 and E-MTAB-1262).
ChIP-qPCR and ChIP-Seq
ChIP-qPCR was performed and evaluated as described21 using the following antibodies: immunoglobulin G pool, I5006 (Sigma-Aldrich); αPPARβ/δ, sc-7197; α-RXR, sc-774; α-HDAC3, sc-11417; α-RNA polymerase II RPB1, sc-899, sc-9001 and sc-56767; (Santa Cruz). Primer sequences were as described previously.26 Comparative binding analyses were statistically analyzed by Student’s t-test (two-tailed, equal variance). For ChIP-Seq, ChIP samples were sequenced on an Illumina IIx Genome Analyzer. Sequencing data were deposited at EBI ArrayExpress (accession number E-MTAB-1097).
Mapping of ChIP-Seq reads and peak calling
ChIP-Seq mapping and peak calling were performed as described6, 56, 57 except for updated versions of Ensembl (v65), Bowtie (0.12.7) and MACS (1.4.0rc2 20110214). The number of usable reads was 19.986.061 (PPARβ/δ), 20.033.004 (RXR) and 18.742.013 (immunoglobulin G control). Peaks were filtered for a MACS false discovery rate ⩽0.05. Peak overlap Venn diagrams were calculated by building the interval union and testing each resulting interval for overlap with the initial peak sets. Genes were associated with peaks using a method based on by GREAT58. Briefly, each gene received a basal region of 5 kb upstream, 1 kb downstream, which was extended to either the basal region of the next gene or a maximum of 1 Mb. A peak could thus be assigned to multiple genes. Only the first annotated TSS for each gene was used.
Databases, correlation of ChIP and microarray data and motif search
All genomic sequence and gene annotation data were retrieved from Ensembl revision 65 (http://dec2011.archive.ensembl.org/index.html). Significance of overlap between microarray and Chip-Seq-derived gene sets with predefined sets was assessed with Fisher’s exact test. Correction for multiple hypothesis testing was done via the Benjamini–Hochberg procedure. De novo motif search was performed using MEME (version 4.3.0).59
Accession codes
References
Desvergne B, Michalik L, Wahli W . Transcriptional regulation of metabolism. Physiol Rev 2006; 86: 465–514.
Van Ginderachter JA, Movahedi K, Van den Bossche J, De Baetselier P . Macrophages, PPARs, and Cancer. PPAR Res 2008; 2008: 169414.
Montagner A, Rando G, Degueurce G, Leuenberger N, Michalik L, Wahli W . New insights into the role of PPARs. Prostaglandins Leukot Essent Fatty Acids 2011; 85: 235–243.
Peters JM, Shah YM, Gonzalez FJ . The role of peroxisome proliferator-activated receptors in carcinogenesis and chemoprevention. Nat Rev Cancer 2012; 12: 181–195.
Müller R, Kömhoff M, Peters JM, Müller-Brüsselbach S . A Role for PPARβ/δ in Tumor Stroma and Tumorigenesis. PPAR Res 2008; 2008: 534294.
Adhikary T, Kaddatz K, Finkernagel F, Schönbauer A, Meissner W, Scharfe M et al. Genomewide analyses define different modes of transcriptional regulation by peroxisome proliferator-activated receptor-beta/delta (PPARbeta/delta). PLoS One 2011; 6: e16344.
Zhu P, Goh YY, Chin HF, Kersten S, Tan NS . Angiopoietin-like 4: a decade of research. Biosci Rep 2012; 32: 211–219.
Ge H, Yang G, Yu X, Pourbahrami T, Li C . Oligomerization state-dependent hyperlipidemic effect of angiopoietin-like protein 4. J Lipid Res 2004; 45: 2071–2079.
Mandard S, Zandbergen F, van Straten E, Wahli W, Kuipers F, Muller M et al. The fasting-induced adipose factor/angiopoietin-like protein 4 is physically associated with lipoproteins and governs plasma lipid levels and adiposity. J Biol Chem 2006; 281: 934–944.
Mandard S, Zandbergen F, Tan NS, Escher P, Patsouris D, Koenig W et al. The direct peroxisome proliferator-activated receptor target fasting-induced adipose factor (FIAF/PGAR/ANGPTL4) is present in blood plasma as a truncated protein that is increased by fenofibrate treatment. J Biol Chem 2004; 279: 34411–34420.
Tan MJ, Teo Z, Sng MK, Zhu P, Tan NS . Emerging roles of angiopoietin-like 4 in human cancer. Mol Cancer Res 2012; 10: 677–688.
Goh YY, Pal M, Chong HC, Zhu P, Tan MJ, Punugu L et al. Angiopoietin-like 4 interacts with integrins beta1 and beta5 to modulate keratinocyte migration. Am J Pathol 2010; 177: 2791–2803.
Huang XF, Han J, Hu XT, He C . Mechanisms involved in biological behavior changes associated with Angptl4 expression in colon cancer cell lines. Oncol Rep 2012; 27: 1541–1547.
Li H, Ge C, Zhao F, Yan M, Hu C, Jia D et al. HIF-1alpha-activated ANGPTL4 contributes to tumor metastasis via VCAM-1/integrin beta1 signaling in human hepatocellular carcinoma. Hepatology 2011; 54: 910–919.
Padua D, Zhang XH, Wang Q, Nadal C, Gerald WL, Gomis RR et al. TGFbeta primes breast tumors for lung metastasis seeding through angiopoietin-like 4. Cell 2008; 133: 66–77.
Gealekman O, Burkart A, Chouinard M, Nicoloro SM, Straubhaar J, Corvera S . Enhanced angiogenesis in obesity and in response to PPARgamma activators through adipocyte VEGF and ANGPTL4 production. Am J Physiol Endocrinol Metab 2008; 295: E1056–E1064.
Le Jan S, Amy C, Cazes A, Monnot C, Lamande N, Favier J et al. Angiopoietin-like 4 is a proangiogenic factor produced during ischemia and in conventional renal cell carcinoma. Am J Pathol 2003; 162: 1521–1528.
Huang RL, Teo Z, Chong HC, Zhu P, Tan MJ, Tan CK et al. ANGPTL4 modulates vascular junction integrity by integrin signaling and disruption of intercellular VE-cadherin and claudin-5 clusters. Blood 2011; 118: 3990–4002.
Zhu P, Tan MJ, Huang RL, Tan CK, Chong HC, Pal M et al. Angiopoietin-like 4 protein elevates the prosurvival intracellular O(2)(-):H(2)O(2) ratio and confers anoikis resistance to tumors. Cancer Cell 2011; 19: 401–415.
Verine J, Lehmann-Che J, Soliman H, Feugeas JP, Vidal JS, Mongiat-Artus P et al. Determination of angptl4 mRNA as a diagnostic marker of primary and metastatic clear cell renal-cell carcinoma. PLoS One 2010; 5: e10421.
Nakayama T, Hirakawa H, Shibata K, Abe K, Nagayasu T, Taguchi T . Expression of angiopoietin-like 4 in human gastric cancer: ANGPTL4 promotes venous invasion. Oncol Rep 2010; 24: 599–606.
Nakayama T, Hirakawa H, Shibata K, Nazneen A, Abe K, Nagayasu T et al. Expression of angiopoietin-like 4 (ANGPTL4) in human colorectal cancer: ANGPTL4 promotes venous invasion and distant metastasis. Oncol Rep 2011; 25: 929–935.
Hu Z, Fan C, Livasy C, He X, Oh DS, Ewend MG et al. A compact VEGF signature associated with distant metastases and poor outcomes. BMC Med 2009; 7: 9.
Mannelqvist M, Stefansson IM, Bredholt G, Hellem Bo T, Oyan AM, Jonassen I et al. Gene expression patterns related to vascular invasion and aggressive features in endometrial cancer. Am J Pathol 2011; 178: 861–871.
Belanger AJ, Lu H, Date T, Liu LX, Vincent KA, Akita GY et al. Hypoxia up-regulates expression of peroxisome proliferator-activated receptor gamma angiopoietin-related gene (PGAR) in cardiomyocytes: role of hypoxia inducible factor 1alpha. J Mol Cell Cardiol 2002; 34: 765–774.
Kaddatz K, Adhikary T, Finkernagel F, Meissner W, Müller-Brüsselbach S, Müller R . Transcriptional profiling identifies functional interactions of TGFβ and PPARβ/δ signaling: synergistic induction of ANGPTL4 transcription. J Biol Chem 2010; 285: 29469–29479.
Koliwad SK, Kuo T, Shipp LE, Gray NE, Backhed F, So AY et al. Angiopoietin-like 4 (ANGPTL4, fasting-induced adipose factor) is a direct glucocorticoid receptor target and participates in glucocorticoid-regulated triglyceride metabolism. J Biol Chem 2009; 284: 25593–25601.
Tan NS, Icre G, Montagner A, Bordier-ten-Heggeler B, Wahli W, Michalik L . The nuclear hormone receptor peroxisome proliferator-activated receptor beta/delta potentiates cell chemotactism, polarization, and migration. Mol Cell Biol 2007; 27: 7161–7175.
Lim HJ, Lee S, Park JH, Lee KS, Choi HE, Chung KS et al. PPARdelta agonist L-165041 inhibits rat vascular smooth muscle cell proliferation and migration via inhibition of cell cycle. Atherosclerosis 2009; 202: 446–454.
Naruhn S, Toth PM, Adhikary T, Kaddatz K, Pape V, Dörr S et al. High-affinity peroxisome proliferator-activated receptor beta/delta-specific ligands with pure antagonistic or inverse agonistic properties. Mol Pharmacol 2011; 80: 828–838.
Lieber S, Scheer F, Meissner W, Naruhn S, Adhikary T, Müller-Brüsselbach S et al. (Z)-2-(2-bromophenyl)-3-{[4-(1-methyl-piperazine)amino]phenyl}acrylonitrile (DG172): an orally bioavailable PPARbeta/delta-selective ligand with inverse agonistic properties. J Med Chem 2012; 55: 2858–2868.
Toth PM, Naruhn S, Pape VF, Dörr SM, Klebe G, Müller R et al. Development of Improved PPARbeta/delta Inhibitors. ChemMedChem 2012; 7: 159–170.
Peters JM, Gonzalez FJ . Sorting out the functional role(s) of peroxisome proliferator-activated receptor-beta/delta (PPARbeta/delta) in cell proliferation and cancer. Biochim Biophys Acta 2009; 1796: 230–241.
Bierie B, Moses HL . Tumour microenvironment: TGFbeta: the molecular Jekyll and Hyde of cancer. Nat Rev Cancer 2006; 6: 506–520.
Stockert J, Adhikary T, Kaddatz K, Finkernagel F, Meissner W, Müller-Brüsselbach S et al. Reverse crosstalk of TGFβ and PPARβ/δ signaling identified by transcriptional profiling. Nucleic Acids Res 2011; 39: 119–131.
Shearer BG, Wiethe RW, Ashe A, Billin AN, Way JM, Stanley TB et al. Identification and characterization of 4-chloro-N-(2-{[5-trifluoromethyl)-2-pyridyl]sulfonyl}ethyl)benzamide (GSK3787), a selective and irreversible peroxisome proliferator-activated receptor delta (PPARdelta) antagonist. J Med Chem 2010; 53: 1857–1861.
Dowell P, Ishmael JE, Avram D, Peterson VJ, Nevrivy DJ, Leid M . Identification of nuclear receptor corepressor as a peroxisome proliferator-activated receptor alpha interacting protein. J Biol Chem 1999; 274: 15901–15907.
Shi Y, Hon M, Evans RM . The peroxisome proliferator-activated receptor delta, an integrator of transcriptional repression and nuclear receptor signaling. Proc Natl Acad Sci USA 2002; 99: 2613–2618.
Ricote M, Glass CK . PPARs and molecular mechanisms of transrepression. Biochim Biophys Acta 2007; 1771: 926–935.
Goh YY, Pal M, Chong HC, Zhu P, Tan MJ, Punugu L et al. Angiopoietin-like 4 interacts with matrix proteins to modulate wound healing. J Biol Chem 2010; 285: 32999–33009.
Lei X, Shi F, Basu D, Huq A, Routhier S, Day R et al. Proteolytic processing of angiopoietin-like protein 4 by proprotein convertases modulates its inhibitory effects on lipoprotein lipase activity. J Biol Chem 2011; 286: 15747–15756.
Krämer DK, Al-Khalili L, Guigas B, Leng Y, Garcia-Roves PM, Krook A . Role of AMP kinase and PPARdelta in the regulation of lipid and glucose metabolism in human skeletal muscle. J Biol Chem 2007; 282: 19313–19320.
Farrow B, O’Connor KL, Hashimoto K, Iwamura T, Evers BM . Selective activation of PPARgamma inhibits pancreatic cancer invasion and decreases expression of tissue plasminogen activator. Surgery 2003; 134: 206–212.
Liu H, Zang C, Fenner MH, Possinger K, Elstner E . PPARgamma ligands and ATRA inhibit the invasion of human breast cancer cells in vitro. Breast Cancer Res Treat 2003; 79: 63–74.
Papi A, Rocchi P, Ferreri AM, Guerra F, Orlandi M . Enhanced effects of PPARgamma ligands and RXR selective retinoids in combination to inhibit migration and invasiveness in cancer cells. Oncol Rep 2009; 21: 1083–1089.
Zaytseva YY, Wallis NK, Southard RC, Kilgore MW . The PPARgamma antagonist T0070907 suppresses breast cancer cell proliferation and motility via both PPARgamma-dependent and -independent mechanisms. Anticancer Res 2011; 31: 813–823.
Borland MG, Khozoie C, Albrecht PP, Zhu B, Lee C, Lahoti TS et al. Stable over-expression of PPARbeta/delta and PPARgamma to examine receptor signaling in human HaCaT keratinocytes. Cell Signal 2011; 23: 2039–2050.
Tian L, Zhou J, Casimiro MC, Liang B, Ojeifo JO, Wang M et al. Activating peroxisome proliferator-activated receptor gamma mutant promotes tumor growth in vivo by enhancing angiogenesis. Cancer Res 2009; 69: 9236–9244.
Yoon JC, Chickering TW, Rosen ED, Dussault B, Qin Y, Soukas A et al. Peroxisome proliferator-activated receptor gamma target gene encoding a novel angiopoietin-related protein associated with adipose differentiation. Mol Cell Biol 2000; 20: 5343–5349.
Yang YH, Wang Y, Lam KS, Yau MH, Cheng KK, Zhang J et al. Suppression of the Raf/MEK/ERK signaling cascade and inhibition of angiogenesis by the carboxyl terminus of angiopoietin-like protein 4. Arterioscler Thromb Vasc Biol 2008; 28: 835–840.
Galaup A, Cazes A, Le Jan S, Philippe J, Connault E, Le Coz E et al. Angiopoietin-like 4 prevents metastasis through inhibition of vascular permeability and tumor cell motility and invasiveness. Proc Natl Acad Sci USA 2006; 103: 18721–18726.
Cazes A, Galaup A, Chomel C, Bignon M, Brechot N, Le Jan S et al. Extracellular matrix-bound angiopoietin-like 4 inhibits endothelial cell adhesion, migration, and sprouting and alters actin cytoskeleton. Circ Res 2006; 99: 1207–1215.
Du J, Sun C, Hu Z, Yang Y, Zhu Y, Zheng D et al. Lysophosphatidic acid induces MDA-MB-231 breast cancer cells migration through activation of PI3K/PAK1/ERK signaling. PLoS One 2010; 5: e15940.
Kitzing TM, Sahadevan AS, Brandt DT, Knieling H, Hannemann S, Fackler OT et al. Positive feedback between Dia1, LARG, and RhoA regulates cell morphology and invasion. Genes Dev 2007; 21: 1478–1483.
Rieck M, Meissner W, Ries S, Müller-Brüsselbach S, Müller R . Ligand-mediated regulation of peroxisome proliferator-activated receptor (PPAR) beta/delta: a comparative analysis of PPAR-selective agonists and all-trans retinoic acid. Mol Pharmacol 2008; 74: 1269–1277.
Zhang Z, Cao L, Li J, Liang X, Liu Y, Liu H et al. Acquisition of anoikis resistance reveals a synoikis-like survival style in BEL7402 hepatoma cells. Cancer Lett 2008; 267: 106–115.
Langmead B, Trapnell C, Pop M, Salzberg SL . Ultrafast and memory-efficient alignment of short DNA sequences to the human genome. Genome Biol 2009; 10: R25.
McLean CY, Bristor D, Hiller M, Clarke SL, Schaar BT, Lowe CB et al. GREAT improves functional interpretation of cis-regulatory regions. Nat Biotechnol 2010; 28: 495–501.
Bailey TL, Elkan C . Fitting a mixture model by expectation maximization to discover motifs in biopolymers. Proc Int Conf Intell Syst Mol Biol 1994; 2: 28–36.
Acknowledgements
This work was supported by research grants from the Deutsche Forschungsgemeinschaft to RM (SFB/TRR17-A3 and MU601/13) and by the LOEWE research cluster ‘Tumor and Inflammation’.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies the paper on the Oncogene website
Rights and permissions
This work is licensed under the Creative Commons Attribution-NonCommercial-No Derivative Works 3.0 Unported License. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-nd/3.0/
About this article
Cite this article
Adhikary, T., Brandt, D., Kaddatz, K. et al. Inverse PPARβ/δ agonists suppress oncogenic signaling to the ANGPTL4 gene and inhibit cancer cell invasion. Oncogene 32, 5241–5252 (2013). https://doi.org/10.1038/onc.2012.549
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/onc.2012.549
Keywords
This article is cited by
-
Rapid acceleration of KRAS-mutant pancreatic carcinogenesis via remodeling of tumor immune microenvironment by PPARδ
Nature Communications (2022)
-
Angiopoietin-Like Protein 4 Is Involved in Manganese Superoxide Dismutase-Mediated Suppression of Breast Cancer Cell Growth
Bulletin of Experimental Biology and Medicine (2022)
-
Targeting TRAF3IP2, Compared to Rab27, is More Effective in Suppressing the Development and Metastasis of Breast Cancer
Scientific Reports (2020)
-
Angiopoietin-like protein 1 inhibits epithelial to mesenchymal transition in colorectal cancer cells via suppress Slug expression
Cytotechnology (2019)
-
PPAR-delta promotes survival of breast cancer cells in harsh metabolic conditions
Oncogenesis (2016)