Abstract
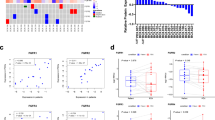
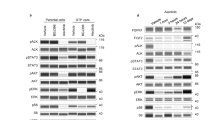
Fibroblast growth factor receptors (FGFRs) can act as driving oncoproteins in certain cancers, making them attractive drug targets. Here we have characterized tumour cell responses to two new inhibitors of FGFR1–3, AZ12908010 and the clinical candidate AZD4547, making comparisons with the well-characterized FGFR inhibitor PD173074. In a panel of 16 human tumour cell lines, the anti-proliferative activity of AZ12908010 or AZD4547 was strongly linked to the presence of deregulated FGFR signalling, indicating that addiction to deregulated FGFRs provides a therapeutic opportunity for selective intervention. Acquired resistance to targeted tyrosine kinase inhibitors is a growing problem in the clinic but has not yet been explored for FGFR inhibitors. To assess how FGFR-dependent tumour cells adapt to long-term FGFR inhibition, we generated a derivative of the KMS-11 myeloma cell line (FGFRY373C) with acquired resistance to AZ12908010 (KMS-11R cells). Basal phosphorylated FGFR and FGFR-dependent downstream signalling were constitutively elevated and refractory to drug in KMS-11R cells. Sequencing of FGFR3 in KMS-11R cells revealed the presence of a heterozygous mutation at the gatekeeper residue, encoding FGFR3V555M; consistent with this, KMS-11R cells were cross-resistant to AZD4547 and PD173074. These results define the selectivity and efficacy of two new FGFR inhibitors and identify a secondary gatekeeper mutation as a mechanism of acquired resistance to FGFR inhibitors that should be anticipated as clinical evaluation proceeds.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 50 print issues and online access
$259.00 per year
only $5.18 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout








Similar content being viewed by others
Accession codes
References
Klint P, Claesson-Welsh L . Signal transduction by fibroblast growth factor receptors. Front Biosci 1999; 4: D165–D177.
Eswarakumar VP, Lax I, Schlessinger J . Cellular signaling by fibroblast growth factor receptors. Cytokine Growth Factor Rev 2005; 16: 139–149.
Ornitz DM, Marie PJ . FGF signaling pathways in endochondral and intramembranous bone development and human genetic disease. Genes Dev 2002; 16: 1446–1465.
Pardo OE, Arcaro A, Salerno G, Raguz S, Downward J, Seckl MJ et al. Fibroblast growth factor-2 induces translational regulation of Bcl-XL and Bcl-2 via a MEK-dependent pathway. J Biol Chem 2002; 277: 12040–12046.
Jeffers M, LaRochelle WJ, Lichenstein HS . Fibroblast growth factors in cancer: therapeutic possibilities. Expert Opin Ther Targets 2002; 6: 469–482.
Knights V, Cook SJ . De-regulated FGF receptors as therapeutic targets in cancer. Pharmacol Ther 2010; 125: 105–117.
Keats JJ, Reiman T, Belch AR, Pilarski LM . Ten years and counting: so what do we know about t(4;14)(p16;q32) multiple myeloma. Leuk Lymphoma 2006; 47: 2289–2300.
Jang JH, Shin KH, Park JG . Mutations in fibroblast growth factor receptor 2 and fibroblast growth factor receptor 3 genes associated with human gastric and colorectal cancers. Cancer Res 2001; 61: 3541–3543.
Ray ME, Yang ZQ, Albertson D, Kleer CG, Washburn JG, Macoska JA et al. Genomic and expression analysis of the 8p11-12 amplicon in human breast cancer cell lines. Cancer Res 2004; 64: 40–47.
Billerey C, Chopin D, Aubriot-Lorton MH, Ricol D, Gil Diez de Medina S, Van Rhijn B et al. Frequent FGFR3 mutations in papillary non-invasive bladder (pTa) tumors. Am J Pathol 2001; 158: 1955–1959.
Reiter A, Sohal J, Kulkarni S, Chase A, Macdonald DH, Aguiar RC et al. Consistent fusion of ZNF198 to the fibroblast growth factor receptor-1 in the t(8;13)(p11;q12) myeloproliferative syndrome. Blood 1998; 92: 1735–1742.
Moffa AB, Tannheimer SL, Ethier SP . Transforming potential of alternatively spliced variants of fibroblast growth factor receptor 2 in human mammary epithelial cells. Mol Cancer Res 2004; 2: 643–652.
Gavine PR, Mooney L, Kilgour E, Thomas AP, Al-Kadhimi K, Beck S et al. AZD4547: an orally bioavailable, potent and selective inhibitor of the fibroblast growth factor receptor tyrosine kinase family. Cancer Res 2012; 72: 2045–2056.
Klinowska T, Gavine PR, Mooney L, Parveen N, Al-Kadhimi K, Thomas A et al. The characterization of novel, potent and selective small molecule inhibitors of the FGFR tyrosine kinase in vitro and in vivo. Gordon Research Conference 2–7 March 2008; Lucca, Italy.
Kahan C, Seuwen K, Meloche S, Pouysségur J . Coordinate, biphasic activation of p44 mitogen-activated protein kinase and S6 kinase by growth factors in hamster fibroblasts. Evidence for thrombin-induced signals different from phosphoinositide turnover and adenylylcyclase inhibition. J Biol Chem 1992; 267: 13369–13375.
Weinstein IB, Joe A . Oncogene addiction. Cancer Res 2008; 68: 3077–3080.
Chesi M, Brents LA, Ely SA, Bais C, Robbiani DF, Mesri EA et al. Activated fibroblast growth factor receptor 3 is an oncogene that contributes to tumor progression in multiple myeloma. Blood 2001; 97: 729–736.
Trudel S, Ely S, Farooqi Y, Affer M, Robbiani DF, Chesi M et al. Inhibition of fibroblast growth factor receptor 3 induces differentiation and apoptosis in t(4;14) myeloma. Blood 2004; 103: 3521–3528.
Lombardi L, Poretti G, Mattioli M, Fabris S, Agnelli L, Bicciato S et al. Molecular characterization of human multiple myeloma cell lines by integrative genomics: insights into the biology of the disease. Genes Chromosomes Cancer 2007; 46: 226–238.
Ronchetti D, Greco A, Compasso S, Colombo G, Dell'Era P, Otsuki T et al. Deregulated FGFR3 mutants in multiple myeloma cell lines with t(4;14): comparative analysis of Y373C, K650E and the novel G384D mutations. Oncogene 2001; 20: 3553–3562.
Kimura T, Suzuki H, Ohashi T, Asano K, Kiyota H, Eto Y . The incidence of thanatophoric dysplasia mutations in FGFR3 gene is higher in low-grade or superficial bladder carcinomas. Cancer 2001; 92: 2555–2561.
Tomlinson DC, Baldo O, Harnden P, Knowles MA . FGFR3 protein expression and its relationship to mutation status and prognostic variables in bladder cancer. J Pathol 2007; 213: 91–98.
Al-Ahmadie HA, Iyer G, Janakiraman M, Lin O, Heguy A, Tickoo SK et al. Somatic mutation of fibroblast growth factor receptor-3 (FGFR3) defines a distinct morphological subtype of high-grade urothelial carcinoma. J Pathol 2011; 224: 270–279.
Tomlinson DC, Hurst CD, Knowles MA . Knockdown by shRNA identifies S249C mutant FGFR3 as a potential therapeutic target in bladder cancer. Oncogene 2007; 26: 5889–5899.
Turner N, Lambros MB, Horlings HM, Pearson A, Sharpe R, Natrajan R et al. Integrative molecular profiling of triple negative breast cancers identifies amplicon drivers and potential therapeutic targets. Oncogene 2010; 29: 2013–2023.
Sun S, Jiang Y, Zhang G, Song H, Zhang X, Zhang Y et al. Increased expression of fibroblastic growth factor receptor 2 is correlated with poor prognosis in patients with breast cancer. J Surg Oncol 2011; 105: 773–779.
Tannheimer SL, Rehemtulla A, Ethier SP . Characterization of fibroblast growth factor receptor 2 overexpression in the human breast cancer cell line SUM-52PE. Breast Cancer Res 2000; 2: 311–320.
Hattori Y, Odagiri H, Nakatani H, Miyagawa K, Naito K, Sakamoto H et al. K-sam, an amplified gene in stomach cancer, is a member of the heparin-binding growth factor receptor genes. Proc Natl Acad Sci USA 1990; 87: 5983–5987.
Kuniyasu H, Yasui W, Kitadai Y, Yokozaki H, Ito H, Tahara E . Frequent amplification of the c-met gene in scirrhous type stomach cancer. Biochem Biophys Res Commun 1992; 189: 227–232.
Tsujimoto H, Sugihara H, Hagiwara A, Hattori T . Amplification of growth factor receptor genes and DNA ploidy pattern in the progression of gastric cancer. Virchows Arch 1997; 431: 383–389.
Mor O, Ranzani GN, Ravia Y, Rotman G, Gutman M, Manor A et al. DNA amplification in human gastric carcinomas. Cancer Genet Cytogenet 1993; 65: 111–114.
Kunii K, Davis L, Gorenstein J, Hatch H, Yashiro M, Di Bacco A et al. FGFR2-amplified gastric cancer cell lines require FGFR2 and Erbb3 signaling for growth and survival. Cancer Res 2008; 68: 2340–2348.
Wheeler DL, Dunn EF, Harari PM . Understanding resistance to EGFR inhibitors-impact on future treatment strategies. Nat Rev Clin Oncol 2010; 7: 493–507.
Milojkovic D, Apperley J . Mechanisms of resistance to imatinib and second- generation tyrosine inhibitors in chronic myeloid leukemia. Clin Cancer Res 2009; 15: 7519–7527.
Hammerman PS, Janne PA, Johnson BE . Resistance to epidermal growth factor receptor tyrosine kinase inhibitors in non-small cell lung cancer. Clin Cancer Res 2009; 15: 7502–7509.
Ley R, Balmanno K, Hadfield K, Weston C, Cook SJ . Activation of the ERK1/2 signaling pathway promotes phosphorylation and proteasome-dependent degradation of the BH3-only protein, Bim. J Biol Chem 2003; 278: 18811–18816.
Ewings KE, Hadfield-Moorhouse K, Wiggins CM, Wickenden JA, Balmanno K, Gilley R et al. ERK1/2-dependent phosphorylation of BimEL promotes its rapid dissociation from Mcl-1 and Bcl-xL. EMBO J 2007; 26: 2856–2867.
Azam M, Seeliger MA, Gray NS, Kuriyan J, Daley GQ . Activation of tyrosine kinases by mutation of the gatekeeper threonine. Nat Struct Mol Biol 2008; 15: 1109–1118.
Daub H, Specht K, Ullrich A . Strategies to overcome resistance to targeted protein kinase inhibitors. Nat Rev Drug Discov 2004; 3: 1001–1010.
Azam M, Daley GQ . Anticipating clinical resistance to target-directed agents: the BCR-ABL paradigm. Mol Diagn Ther 2006; 10: 67–76.
Mohammadi M, Froum S, Hamby JM, Schroeder MC, Panek RL, Lu GH et al. Crystal structure of an angiogenesis inhibitor bound to the FGF receptor tyrosine kinase domain. EMBO J 1998; 17: 5896–5904.
Qing J, Du X, Chen Y, Chan P, Li H, Wu P et al. Antibody-based targeting of FGFR3 in bladder carcinoma and t(4;14)-positive multiple myeloma in mice. J Clin Invest 2009; 119: 1216–1229.
Squires M, Ward G, Saxty G, Berdini V, Cleasby A, King P et al. Potent, selective inhibitors of fibroblast growth factor receptor define fibroblast growth factor dependence in preclinical cancer models. Mol Cancer Ther 2011; 10: 1542–1552.
Trudel S, Li ZH, Wei E, Wiesmann M, Chang H, Chen C et al. CHIR-258, a novel, multitargeted tyrosine kinase inhibitor for the potential treatment of t(4;14) multiple myeloma. Blood 2005; 105: 2941–2948.
Lamont FR, Tomlinson DC, Cooper PA, Shnyder SD, Chester JD, Knowles MA . Small molecule FGF receptor inhibitors block FGFR-dependent urothelial carcinoma growth in vitro and in vivo. Br J Cancer 2011; 104: 75–82.
Jebar AH, Hurst CD, Tomlinson DC, Johnston C, Taylor CF, MA Knowles . FGFR3 and Ras gene mutations are mutually exclusive genetic events in urothelial cell carcinoma. Oncogene 2005; 24: 5218–5225.
Normanno N, Tejpar S, Morgillo F, De Luca A, Van Cutsem E, Ciardiello F . Implications for KRAS status and EGFR-targeted therapies in metastatic CRC. Nat Rev Clin Oncol 2009; 6: 519–527.
Zhou W, Hur W, McDermott U, Dutt A, Xian W, Ficarro SB et al. A structure-guided approach to creating covalent FGFR inhibitors. Chem Biol 2010; 17: 285–295.
Huynh H, Ngo VC, Fargnoli J, Ayers M, Soo KC, Koong HN et al. Brivanib alaninate, a dual inhibitor of vascular endothelial growth factor receptor and fibroblast growth factor receptor tyrosine kinases, induces growth inhibition in mouse models of human hepatocellular carcinoma. Clin Cancer Res 2008; 14: 6146–6153.
Hilberg F, Roth GJ, Krssak M, Kautschitsch S, Sommergruber W, Tontsch-Grunt U et al. BIBF 1120: triple angiokinase inhibitor with sustained receptor blockade and good antitumor efficacy. Cancer Res 2008; 68: 4774–4782.
Radomska HS, Bassères DS, Zheng R, Zhang P, Dayaram T, Yamamoto Y et al. Block of C/EBP alpha function by phosphorylation in acute myeloid leukemia with FLT3 activating mutations. J Exp Med 2006; 203: 371–381.
Pabst T, Mueller BU, Harakawa N, Schoch C, Haferlach T, Behre G et al. AML1-ETO downregulates the granulocytic differentiation factor C/EBPalpha in t(8;21) myeloid leukemia. Nat Med 2001; 7: 444–451.
Kawano M, Hirano T, Matsuda T, Taga T, Horii Y, Iwato K et al. Autocrine generation and requirement of BSF-2/IL-6 for human multiple myelomas. Nature 1988; 332: 83–85.
Binato R, Mencalha A, Pizzatti L, Scholl V, Zalcberg I, Abdelhay E . RUNX1T1 is overexpressed in imatinib mesylate-resistant cells. Mol Med Rep 2009; 2: 657–661.
Todd DE, Densham RM, Molton SA, Balmanno K, Newson C, Weston CR et al. ERK1/2 and p38 cooperate to induce a p21CIP1-dependent G1 cell cycle arrest. Oncogene 2004; 23: 3284–3295.
Davies BR, Logie A, McKay JS, Martin P, Steele S, Jenkins R et al. AZD6244 (ARRY-142886), a potent inhibitor of mitogen-activated protein kinase/extracellular signal-regulated kinase kinase 1/2 kinases: mechanism of action in vivo, pharmacokinetic/pharmacodynamic relationship, and potential for combination in preclinical models. Mol Cancer Ther 2007; 6: 2209–2219.
Waterhouse AM, Procter JB, Martin DMA, Clamp M, Barton GJ . Jalview Version 2 a multiple sequence alignment editor and analysis workbench. Bioinformatics 2009; 25: 1189–1191.
Emsley P, Lohkamp B, Scott WG, Cowtan K . Features and development of Coot. Acta Crystallogr D Biol Crystallogr 2010; 66: 486–501.
Benjamini Y, Hochberg Y . Controlling the false discovery rate: a practical and powerful approach to multiple testing. J Roy Statist Soc Ser B (Methodological) 1995; 57: 289–300.
Vandesompele J, De Preter K, Pattyn F, Poppe B, Van Roy N, De Paepe A et al. Accurate normalization of real-time quantitative RT-PCR data by geometric averaging of multiple internal control genes. Genome Biol 2002; 3: : RESEARCH0034.
Acknowledgements
We would like to thank Anne Segonds-Pichon (Babraham Bioinformatics Group) for statistical analysis of qRT–PCR data and Jonathan Keats, Janet Nutt, Margaret Knowles, Matthias Ebert and Andrew Garner for provision of some of the cell lines used in this study. We would especially like to thank Andrew Garner for initiating this project, and for discussions and encouragement in its early stages. We are grateful to Teresa Klinowska, Nigel Brooks, Elaine Kilgour and Paul Smith for many useful discussions and suggestions throughout. This work was supported by a CASE PhD studentship (VC, née Victoria Knights) funded by the Biotechnology and Biological Sciences Research Council (BBSRC) and AstraZeneca, and a sponsored research agreement between AstraZeneca and the Babraham Institute.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
Laura Blockley, Mark Hampson and Paul Gavine are paid employees of AstraZeneca. All other authors declare no conflict of interest.
Additional information
Supplementary Information accompanies the paper on the Oncogene website
Supplementary information
Rights and permissions
About this article
Cite this article
Chell, V., Balmanno, K., Little, A. et al. Tumour cell responses to new fibroblast growth factor receptor tyrosine kinase inhibitors and identification of a gatekeeper mutation in FGFR3 as a mechanism of acquired resistance. Oncogene 32, 3059–3070 (2013). https://doi.org/10.1038/onc.2012.319
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/onc.2012.319
Keywords
This article is cited by
-
Resistance of HNSCC cell models to pan-FGFR inhibition depends on the EMT phenotype associating with clinical outcome
Molecular Cancer (2024)
-
The FGF/FGFR system in the microglial neuroinflammation with Borrelia burgdorferi: likely intersectionality with other neurological conditions
Journal of Neuroinflammation (2023)
-
Elucidating the potential effects of point mutations on FGFR3 inhibitor resistance via combined molecular dynamics simulation and community network analysis
Journal of Computer-Aided Molecular Design (2023)
-
Discovery of a small molecule ligand of FRS2 that inhibits invasion and tumor growth
Cellular Oncology (2023)
-
FGFR-TKI resistance in cancer: current status and perspectives
Journal of Hematology & Oncology (2021)